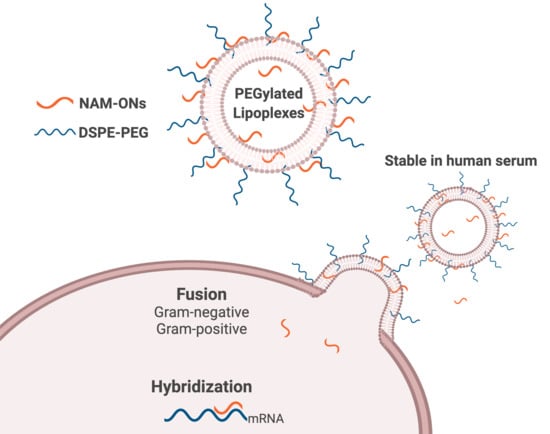
Lipoplexes to Deliver Oligonucleotides in Gram-Positive and Gram-Negative Bacteria: Towards Treatment of Blood Infections
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Design of NAM-ONs
2.3. Preparation of Liposomes and Lipoplexes
2.4. Collection of Human Serum
2.5. Determination of Complexation Stability of Lipoplexes in Buffer and Human Serum by Fluorescence Correlation Spectroscopy (FCS)
2.6. Evaluation of Liposomes’ Interaction with Bacteria by Epifluorescence Microscopy
2.7. Quantification of Liposomes’ Fusion with Different Bacterial Envelopes Using Lipid-Mixing Assay
2.8. Assessment of NAM-ONs Internalization by Confocal Microscopy
2.9. Determination of Cellular Localization of NAM-ONs by Bacterial Fractionation
2.10. Statistical Analysis
3. Results and Discussion
3.1. Characterization of the Liposomes and Lipoplexes
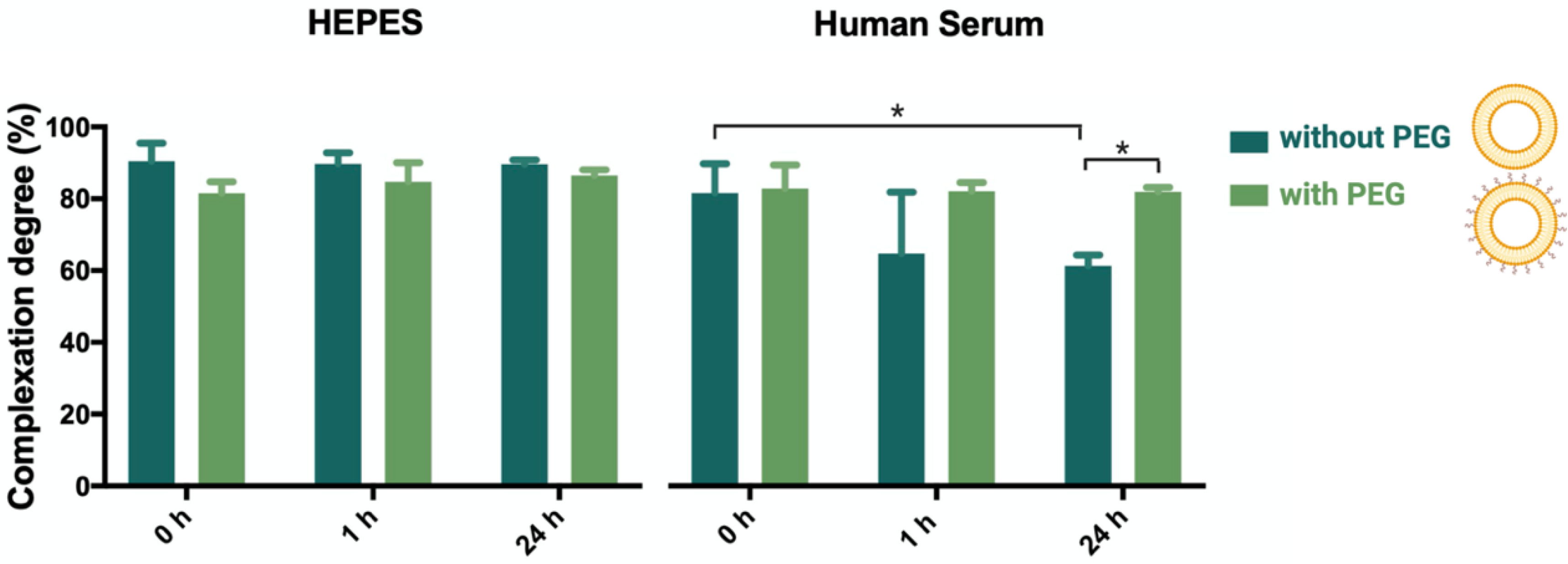
3.2. Complexation Stability of the Lipoplexes in Human Serum
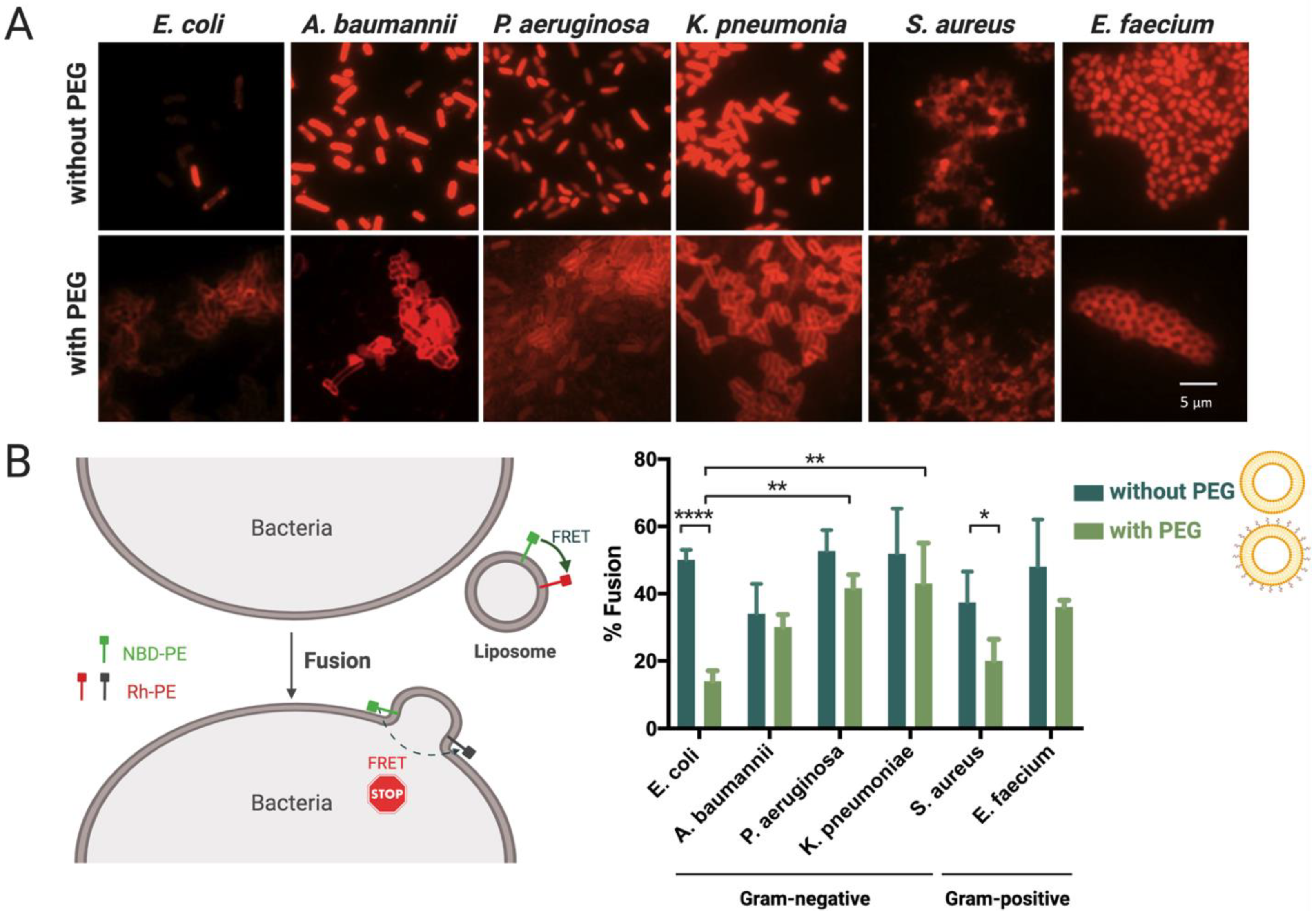
3.3. Interaction and Fusion of Liposomes with Different Bacterial Envelopes
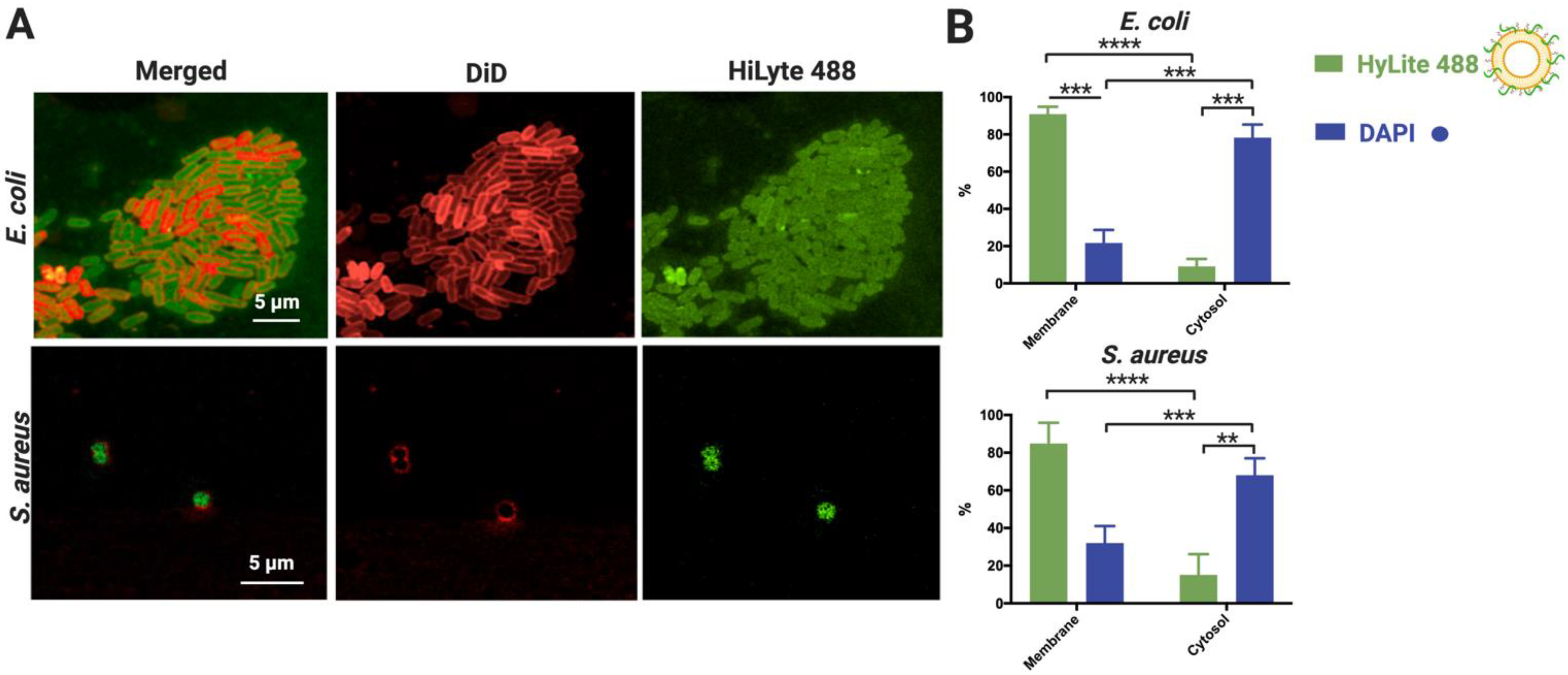
3.4. Characterization of NAM-ONs Internalization and Cellular Localization
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Freire-Moran, L.; Aronsson, B.; Manz, C.; Gyssens, I.C.; So, A.D.; Monnet, D.L.; Cars, O.; The ECDC-EMA Working Group. Critical shortage of new antibiotics in development against multidrug-resistant bacteria—Time to react is now. Drug Resist. Updates 2011, 14, 118–124. [Google Scholar] [CrossRef]
- Zhu, X.; Radovic-Moreno, A.F.; Wu, J.; Langer, R.; Shi, J. Nanomedicine in the management of microbial infection—Overview and perspectives. Nano Today 2014, 9, 478–498. [Google Scholar] [CrossRef] [Green Version]
- Goldstein, E.; MacFadden, D.R.; Karaca, Z.; Steiner, C.A.; Viboud, C.; Lipsitch, M. Antimicrobial resistance prevalence, rates of hospitalization with septicemia and rates of mortality with sepsis in adults in different US states. Int. J. Antimicrob. Agents 2019, 54, 23–34. [Google Scholar] [CrossRef]
- Piddock, L.J. The crisis of no new antibiotics—What is the way forward? Lancet Infect. Dis. 2012, 12, 249–253. [Google Scholar] [CrossRef]
- Bennett, C.F.; Swayze, E. RNA Targeting Therapeutics: Molecular Mechanisms of Antisense Oligonucleotides as a Therapeutic Platform. Annu. Rev. Pharmacol. Toxicol. 2010, 50, 259–293. [Google Scholar] [CrossRef]
- McClorey, G.; Wood, M.J. An overview of the clinical application of antisense oligonucleotides for RNA-targeting therapies. Curr. Opin. Pharmacol. 2015, 24, 52–58. [Google Scholar] [CrossRef]
- Lima, J.F.; Carvalho, J.; Pinto-Ribeiro, I.; Almeida, C.; Wengel, J.; Cerqueira, L.; Figueiredo, C.; Oliveira, C.; Azevedo, N.F. Targeting miR-9 in gastric cancer cells using locked nucleic acid oligonucleotides. BMC Mol. Biol. 2018, 19, 6. [Google Scholar] [CrossRef] [Green Version]
- Bai, H.; You, Y.; Yan, H.; Meng, J.; Xue, X.; Hou, Z.; Zhou, Y.; Ma, X.; Sang, G.; Luo, X. Antisense inhibition of gene expression and growth in gram-negative bacteria by cell-penetrating peptide conjugates of peptide nucleic acids targeted to rpoD gene. Biomaterials 2012, 33, 659–667. [Google Scholar] [CrossRef]
- Xue, X.-Y.; Mao, X.-G.; Zhou, Y.; Chen, Z.; Hu, Y.; Hou, Z.; Li, M.-K.; Meng, J.-R.; Luo, X.-X. Advances in the delivery of antisense oligonucleotides for combating bacterial infectious diseases. Nanomed. Nanotechnol. Biol. Med. 2018, 14, 745–758. [Google Scholar] [CrossRef]
- Mondhe, M.; Chessher, A.; Goh, S.; Good, L.; Stach, J.E.M. Species-Selective Killing of Bacteria by Antimicrobial Peptide-PNAs. PLoS ONE 2014, 9, e89082. [Google Scholar] [CrossRef]
- Santos, R.S.; Figueiredo, C.; Azevedo, N.; Braeckmans, K.; De Smedt, S.C. Nanomaterials and molecular transporters to overcome the bacterial envelope barrier: Towards advanced delivery of antibiotics. Adv. Drug Deliv. Rev. 2018, 136–137, 28–48. [Google Scholar] [CrossRef] [Green Version]
- Hegarty, J.P.; Krzeminski, J.; Sharma, A.K.; Guzman-Villanueva, D.; Weissig, V.; Stewart, D.B., Sr. Bolaamphiphile-based nanocomplex delivery of phosphorothioate gapmer antisense oligonucleotides as a treatment for Clostridium difficile. Int. J. Nanomed. 2016, 11, 3607–3619. [Google Scholar] [CrossRef] [Green Version]
- Hegarty, J.; Stewart, D.B. Advances in therapeutic bacterial antisense biotechnology. Appl. Microbiol. Biotechnol. 2018, 102, 1055–1065. [Google Scholar] [CrossRef]
- Wagner, V.; Dullaart, A.; Bock, A.-K.; Zweck, A. The emerging nanomedicine landscape. Nat. Biotechnol. 2006, 24, 1211–1217. [Google Scholar] [CrossRef] [PubMed]
- Meunier, F.; Prentice, H.G.; Ringdén, O. Liposomal amphotericin B (AmBisome): Safety data from a phase II/III clinical trial. J. Antimicrob. Chemother. 1991, 28, 83–91. [Google Scholar] [CrossRef]
- Rigacci, L.; Mappa, S.; Nassi, L.; Alterini, R.; Carrai, V.; Bernardi, F.; Bosi, A. Liposome-encapsulated doxorubicin in combination with cyclophosphamide, vincristine, prednisone and rituximab in patients with lymphoma and concurrent cardiac diseases or pre-treated with anthracyclines. Hematol. Oncol. 2007, 25, 198–203. [Google Scholar] [CrossRef]
- Schiffelers, R.; Storm, G.; Bakker-Woudenberg, I. Liposome-encapsulated aminoglycosides in pre-clinical and clinical studies. J. Antimicrob. Chemother. 2001, 48, 333–344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Salem, I.I.; Düzgünes, N. Efficacies of cyclodextrin-complexed and liposome-encapsulated clarithromycin against Mycobacterium avium complex infection in human macrophages. Int. J. Pharm. 2003, 250, 403–414. [Google Scholar] [CrossRef]
- Santos, R.S.; Dakwar, G.R.; Zagato, E.; Brans, T.; Figueiredo, C.; Raemdonck, K.; Azevedo, N.F.; De Smedt, S.C.; Braeckmans, K. Intracellular delivery of oligonucleotides in Helicobacter pylori by fusogenic liposomes in the presence of gastric mucus. Biomaterials 2017, 138, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Meng, J.; He, G.; Wang, H.; Jia, M.; Ma, X.; Da, F.; Wang, N.; Hou, Z.; Xue, X.; Li, M.; et al. Reversion of antibiotic resistance by inhibiting mecA in clinical methicillin-resistant Staphylococci by antisense phosphorothioate oligonucleotide. J. Antibiot. 2015, 68, 158–164. [Google Scholar] [CrossRef]
- Wang, H.; Meng, J.; Jia, M.; Ma, X.; He, G.; Yu, J.; Wang, R.; Bai, H.; Hou, Z.; Luo, X. oprM as a new target for reversion of multidrug resistance in Pseudomonas aeruginosa by antisense phosphorothioate oligodeoxynucleotides. FEMS Immunol. Med. Microbiol. 2010, 60, 275–282. [Google Scholar] [CrossRef] [Green Version]
- Xia, X.-R.; Monteiro-Riviere, N.A.; Riviere, J.E. An index for characterization of nanomaterials in biological systems. Nat. Nanotechnol. 2010, 5, 671–675. [Google Scholar] [CrossRef]
- Lundqvist, M.; Stigler, J.; Elia, G.; Lynch, I.; Cedervall, T.; Dawson, K.A. Nanoparticle size and surface properties determine the protein corona with possible implications for biological impacts. Proc. Natl. Acad. Sci. USA 2008, 105, 14265–14270. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Cheng, Q.; Jiang, Q.; Huang, Y.; Liu, H.; Zhao, Y.; Cao, W.; Ma, G.; Dai, F.; Liang, X.; et al. Enhanced endosomal/lysosomal escape by distearoyl phosphoethanolamine-polycarboxybetaine lipid for systemic delivery of siRNA. J. Control. Release 2014, 176, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.; Webster, P.; Davis, M.E. PEGylation significantly affects cellular uptake and intracellular trafficking of non-viral gene delivery particles. Eur. J. Cell Biol. 2004, 83, 97–111. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, Z.; Lin, W.; Chen, S. Gene transfection in complex media using PCBMAEE-PCBMA copolymer with both hydrolytic and zwitterionic blocks. Biomaterials 2014, 35, 7909–7918. [Google Scholar] [CrossRef]
- Deshpande, M.C.; Davies, M.C.; Garnett, M.C.; Williams, P.M.; Armitage, D.; Bailey, L.; Vamvakaki, M.; Armes, S.P.; Stolnik, S. The effect of poly (ethylene glycol) molecular architecture on cellular interaction and uptake of DNA complexes. J. Control. Release 2004, 97, 143–156. [Google Scholar] [CrossRef] [PubMed]
- Nicolosi, D.; Scalia, M.; Nicolosi, V.M.; Pignatello, R. Encapsulation in fusogenic liposomes broadens the spectrum of action of vancomycin against Gram-negative bacteria. Int. J. Antimicrob. Agents 2010, 35, 553–558. [Google Scholar] [CrossRef] [Green Version]
- Remaut, K.; Lucas, B.; Braeckmans, K.; Demeester, J.; De Smedt, S. Pegylation of liposomes favours the endosomal degradation of the delivered phosphodiester oligonucleotides. J. Control. Release 2007, 117, 256–266. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; De Smedt, S.C.; Remaut, K. Fluorescence Correlation Spectroscopy to find the critical balance between extracellular association and intracellular dissociation of mRNA complexes. Acta Biomater. 2018, 75, 358–370. [Google Scholar] [CrossRef] [Green Version]
- Remaut, K.; Lucas, B.; Raemdonck, K.; Braeckmans, K.; Demeester, J.; De Smedt, S. Can we better understand the intracellular behavior of DNA nanoparticles by fluorescence correlation spectroscopy? J. Control. Release 2007, 121, 49–63. [Google Scholar] [CrossRef]
- Zhang, H.; Rombouts, K.; Raes, L.; Xiong, R.; De Smedt, S.C.; Braeckmans, K.; Remaut, K. Fluorescence-Based Quantification of Messenger RNA and Plasmid DNA Decay Kinetics in Extracellular Biological Fluids and Cell Extracts. Adv. Biosyst. 2020, 4, e2000057. [Google Scholar] [CrossRef]
- Struck, D.K.; Hoekstra, D.; Pagano, R.E. Use of resonance energy transfer to monitor membrane fusion. Biochemistry 1981, 20, 4093–4099. [Google Scholar] [CrossRef] [PubMed]
- Mugabe, C.; Halwani, M.; Azghani, A.O.; Lafrenie, R.M.; Omri, A. Mechanism of Enhanced Activity of Liposome-Entrapped Aminoglycosides against Resistant Strains of Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2006, 50, 2016–2022. [Google Scholar] [CrossRef] [Green Version]
- Santos, R.S.; Guimarães, N.; Madureira, P.; Azevedo, N.F. Optimization of a peptide nucleic acid fluorescence in situ hybridization (PNA-FISH) method for the detection of bacteria and disclosure of a formamide effect. J. Biotechnol. 2014, 187, 16–24. [Google Scholar] [CrossRef] [Green Version]
- Rocha, R.; Santos, R.S.; Madureira, P.; Almeida, C.; Azevedo, N. Optimization of peptide nucleic acid fluorescence in situ hybridization (PNA-FISH) for the detection of bacteria: The effect of pH, dextran sulfate and probe concentration. J. Biotechnol. 2016, 226, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Azevedo, A.S.; Sousa, I.M.; Fernandes, R.M.; Azevedo, N.; Almeida, C. Optimizing locked nucleic acid/2′-O-methyl-RNA fluorescence in situ hybridization (LNA/2′OMe-FISH) procedure for bacterial detection. PLoS ONE 2019, 14, e0217689. [Google Scholar] [CrossRef]
- Banbula, A.; Bugno, M.; Goldstein, J.; Yen, J.; Nelson, D.; Travis, J.; Potempa, J. Emerging Family of Proline-Specific Peptidases of Porphyromonas gingivalis: Purification and Characterization of Serine Dipeptidyl Peptidase, a Structural and Functional Homologue of Mammalian Prolyl Dipeptidyl Peptidase IV. Infect. Immun. 2000, 68, 1176–1182. [Google Scholar] [CrossRef] [Green Version]
- Dakwar, G.R.; Braeckmans, K.; Ceelen, W.; De Smedt, S.C.; Remaut, K. Exploring the HYDRAtion method for loading siRNA on liposomes: The interplay between stability and biological activity in human undiluted ascites fluid. Drug Deliv. Transl. Res. 2017, 7, 241–251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alfredsson, V. Cryo-TEM studies of DNA and DNA–lipid structures. Curr. Opin. Colloid Interface Sci. 2005, 10, 269–273. [Google Scholar] [CrossRef]
- Zolnik, B.S.; González-Fernández, A.; Sadrieh, N.; Dobrovolskaia, M.A. Minireview: Nanoparticles and the Immune System. Endocrinology 2010, 151, 458–465. [Google Scholar] [CrossRef]
- Dakwar, G.R.; Zagato, E.; Delanghe, J.; Hobel, S.; Aigner, A.; Denys, H.; Braeckmans, K.; Ceelen, W.; De Smedt, S.; Remaut, K. Colloidal stability of nano-sized particles in the peritoneal fluid: Towards optimizing drug delivery systems for intraperitoneal therapy. Acta Biomater. 2014, 10, 2965–2975. [Google Scholar] [CrossRef] [Green Version]
- Braeckmans, K.; Buyens, K.; Bouquet-Geerardyn, W.; Vervaet, C.; Joye, P.; De Vos, F.; Plawinski, L.; Doeuvre, L.; Angles-Cano, E.; Sanders, N.N.; et al. Sizing Nanomatter in Biological Fluids by Fluorescence Single Particle Tracking. Nano Lett. 2010, 10, 4435–4442. [Google Scholar] [CrossRef] [Green Version]
- Dan, N. Effect of liposome charge and PEG polymer layer thickness on cell-liposome electrostatic interactions. Biochim. Biophys. Acta Biomembr. 2002, 1564, 343–348. [Google Scholar] [CrossRef] [Green Version]
- Xing, H.; Li, J.; Xu, W.; Hwang, K.; Wu, P.; Yin, Q.; Li, Z.; Cheng, J.; Lu, Y. The Effects of Spacer Length and Composition on Aptamer-Mediated Cell-Specific Targeting with Nanoscale PEGylated Liposomal Doxorubicin. ChemBioChem 2016, 17, 1111–1117. [Google Scholar] [CrossRef]
- Düzgüneş, N.; Faneca, H.; de Lima, M.C.P. Methods to monitor liposome fusion, permeability, and interaction with cells. In Liposomes; Springer: Berlin/Heidelberg, Germany, 2010; pp. 209–232. [Google Scholar]
- Chen, T.; Choi, L.S.; Einstein, S.; Klippenstein, M.A.; Scherrer, P.; Cuhis, P.R. Proton-induced permeability and fusion of large unilamellar vesicles by covalently conjugated poly(2-ethylacrylic acid). J. Liposome Res. 1999, 9, 387–405. [Google Scholar] [CrossRef]
- Ma, Y.; Wang, Z.; Khalil, H.; Wang, R.; Lu, T.; Zhao, W.; Zhang, Y.; Chen, T.; Chen, J. Fusion between fluid liposomes and intact bacteria: Study of driving parameters and in vitro bactericidal efficacy. Int. J. Nanomed. 2016, 11, 4025–4036. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scriboni, A.B.; Couto, V.M.; Ribeiro, L.N.D.M.; Freires, I.A.; Groppo, F.C.; De Paula, E.; Franz-Montan, M.; Cogo-Müller, K. Fusogenic Liposomes Increase the Antimicrobial Activity of Vancomycin against Staphylococcus aureus Biofilm. Front. Pharmacol. 2019, 10, 1401. [Google Scholar] [CrossRef]
- Nicolosi, D.; Cupri, S.; Genovese, C.; Tempera, G.; Mattina, R.; Pignatello, R. Nanotechnology approaches for antibacterial drug delivery: Preparation and microbiological evaluation of fusogenic liposomes carrying fusidic acid. Int. J. Antimicrob. Agents 2015, 45, 622–626. [Google Scholar] [CrossRef]
- Shi, J.; Mao, N.-F.; Wang, L.; Zhang, H.-B.; Chen, Q.; Liu, H.; Tang, X.; Jin, T.; Zhu, C.-T.; Li, F.-B.; et al. Efficacy of Combined Vancomycin and Fosfomycin against Methicillin-Resistant Staphylococcus aureus in Biofilms in vivo. PLoS ONE 2014, 9, e113133. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, Y.; Hou, X.; Lei, J.; Chen, M.; Cong, S.; Zhang, Y.; Ding, W.; Li, G.; Li, X. Multi-functional Liposomes Enhancing Target and Antibacterial Immunity for Antimicrobial and Anti-Biofilm against Methicillin-Resistant Staphylococcus aureus. Pharm. Res. 2016, 33, 763–775. [Google Scholar] [CrossRef]
- Song, L.; Ahkong, Q.; Rong, Q.; Wang, Z.; Ansell, S.; Hope, M.; Mui, B. Characterization of the inhibitory effect of PEG-lipid conjugates on the intracellular delivery of plasmid and antisense DNA mediated by cationic lipid liposomes. Biochim. Biophys. Acta Biomembr. 2002, 1558, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Verhoef, J.J.F.; Anchordoquy, T.J. Questioning the use of PEGylation for drug delivery. Drug Deliv. Transl. Res. 2013, 3, 499–503. [Google Scholar] [CrossRef]
- Che, J.; Okeke, C.I.; Hu, Z.-B.; Xu, J. DSPE-PEG: A distinctive component in drug delivery system. Curr. Pharm. Des. 2015, 21, 1598–1605. [Google Scholar] [CrossRef]
- Nosova, A.S.; Koloskova, O.; Nikonova, A.A.; Simonova, V.A.; Smirnov, V.; Kudlay, D.; Khaitov, M. Diversity of PEGylation methods of liposomes and their influence on RNA delivery. MedChemComm 2019, 10, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Zweifel, U.L.; Hagstrom, A. Total counts of marine bacteria include a large fraction of non-nucleoid-containing bacteria (ghosts). Appl. Environ. Microbiol. 1995, 61, 2180–2185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stoddard, S.F.; Smith, B.J.; Hein, R.; Roller, B.R.; Schmidt, T.M. rrnDB: Improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res. 2015, 43, D593–D598. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Remaut, K.; Lucas, B.; Braeckmans, K.; Sanders, N.N.; Demeester, J.; De Smedt, S.C. Delivery of Phosphodiester Oligonucleotides: Can DOTAP/DOPE Liposomes Do the Trick? Biochemistry 2006, 45, 1755–1764. [Google Scholar] [CrossRef] [PubMed]
- Peeters, L.; Sanders, N.; Jones, A.; Demeester, J.; De Smedt, S. Post-pegylated lipoplexes are promising vehicles for gene delivery in RPE cells. J. Control. Release 2007, 121, 208–217. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pereira, S.; Santos, R.S.; Moreira, L.; Guimarães, N.; Gomes, M.; Zhang, H.; Remaut, K.; Braeckmans, K.; De Smedt, S.; Azevedo, N.F. Lipoplexes to Deliver Oligonucleotides in Gram-Positive and Gram-Negative Bacteria: Towards Treatment of Blood Infections. Pharmaceutics 2021, 13, 989. https://doi.org/10.3390/pharmaceutics13070989
Pereira S, Santos RS, Moreira L, Guimarães N, Gomes M, Zhang H, Remaut K, Braeckmans K, De Smedt S, Azevedo NF. Lipoplexes to Deliver Oligonucleotides in Gram-Positive and Gram-Negative Bacteria: Towards Treatment of Blood Infections. Pharmaceutics. 2021; 13(7):989. https://doi.org/10.3390/pharmaceutics13070989
Chicago/Turabian StylePereira, Sara, Rita Sobral Santos, Luís Moreira, Nuno Guimarães, Mariana Gomes, Heyang Zhang, Katrien Remaut, Kevin Braeckmans, Stefaan De Smedt, and Nuno Filipe Azevedo. 2021. "Lipoplexes to Deliver Oligonucleotides in Gram-Positive and Gram-Negative Bacteria: Towards Treatment of Blood Infections" Pharmaceutics 13, no. 7: 989. https://doi.org/10.3390/pharmaceutics13070989
APA StylePereira, S., Santos, R. S., Moreira, L., Guimarães, N., Gomes, M., Zhang, H., Remaut, K., Braeckmans, K., De Smedt, S., & Azevedo, N. F. (2021). Lipoplexes to Deliver Oligonucleotides in Gram-Positive and Gram-Negative Bacteria: Towards Treatment of Blood Infections. Pharmaceutics, 13(7), 989. https://doi.org/10.3390/pharmaceutics13070989