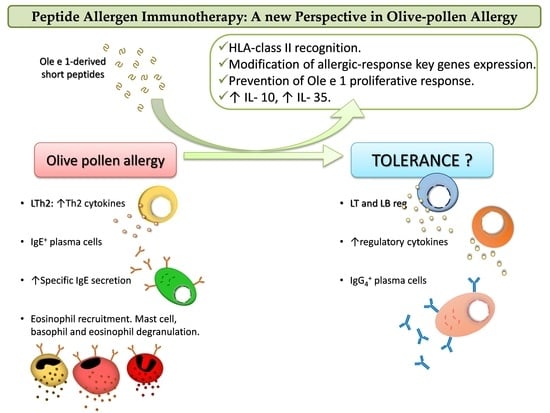
Peptide Allergen Immunotherapy: A New Perspective in Olive-Pollen Allergy
Abstract
1. Allergy and Tolerance
2. Allergen-Specific Immunotherapy (AIT)
3. Peptide Allergen Immunotherapy: State of the Art
Peptide Allergen Immunotherapy as a New Approach in AIT
4. Olive Pollen Allergy
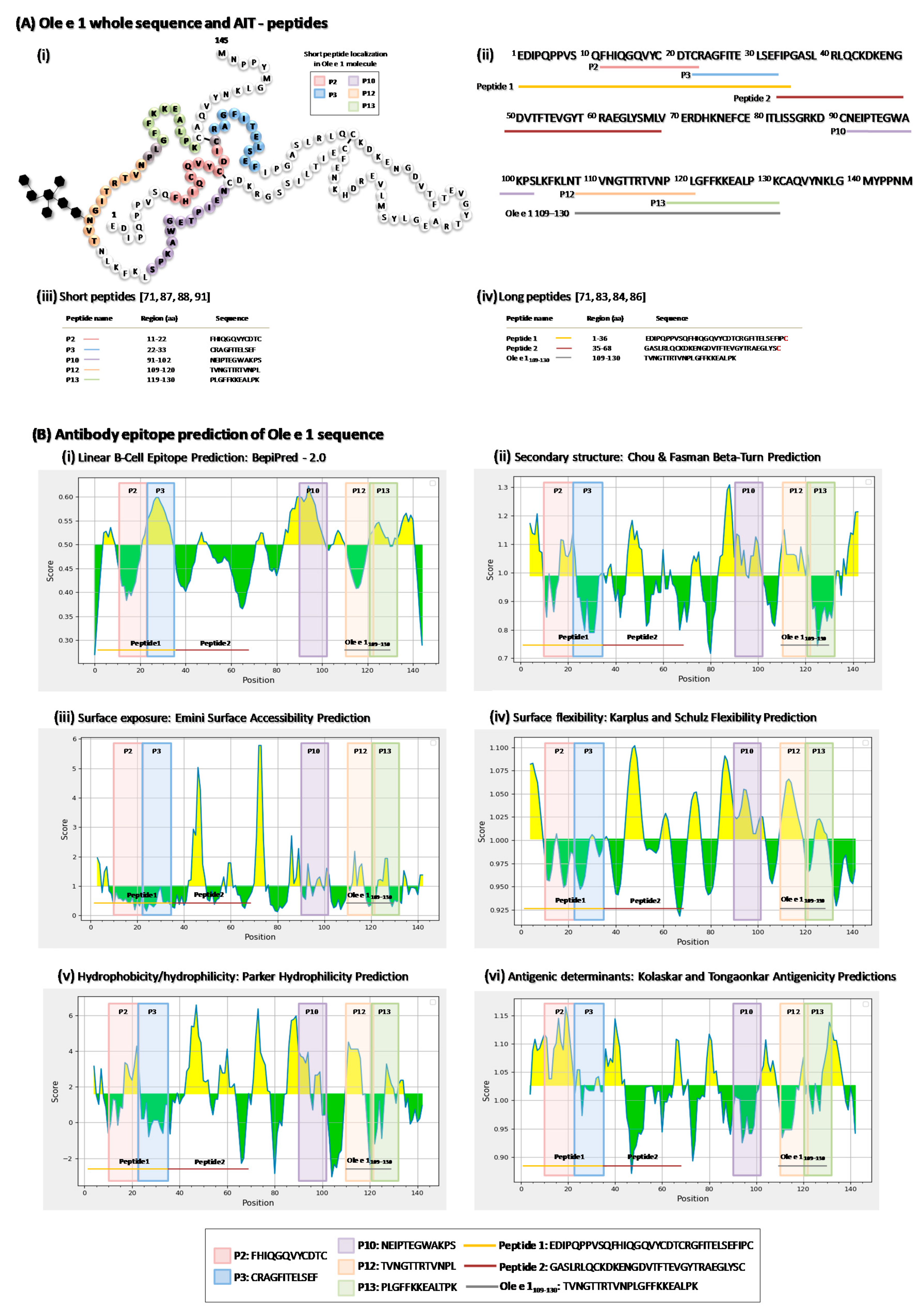
5. Ole e 1-Peptides
5.1. Characterization of Ole e 1-Derived Peptides Immunoregulation
5.1.1. Long Ole e 1-Derived Peptides
5.1.2. Short Ole e 1-Derived Peptides
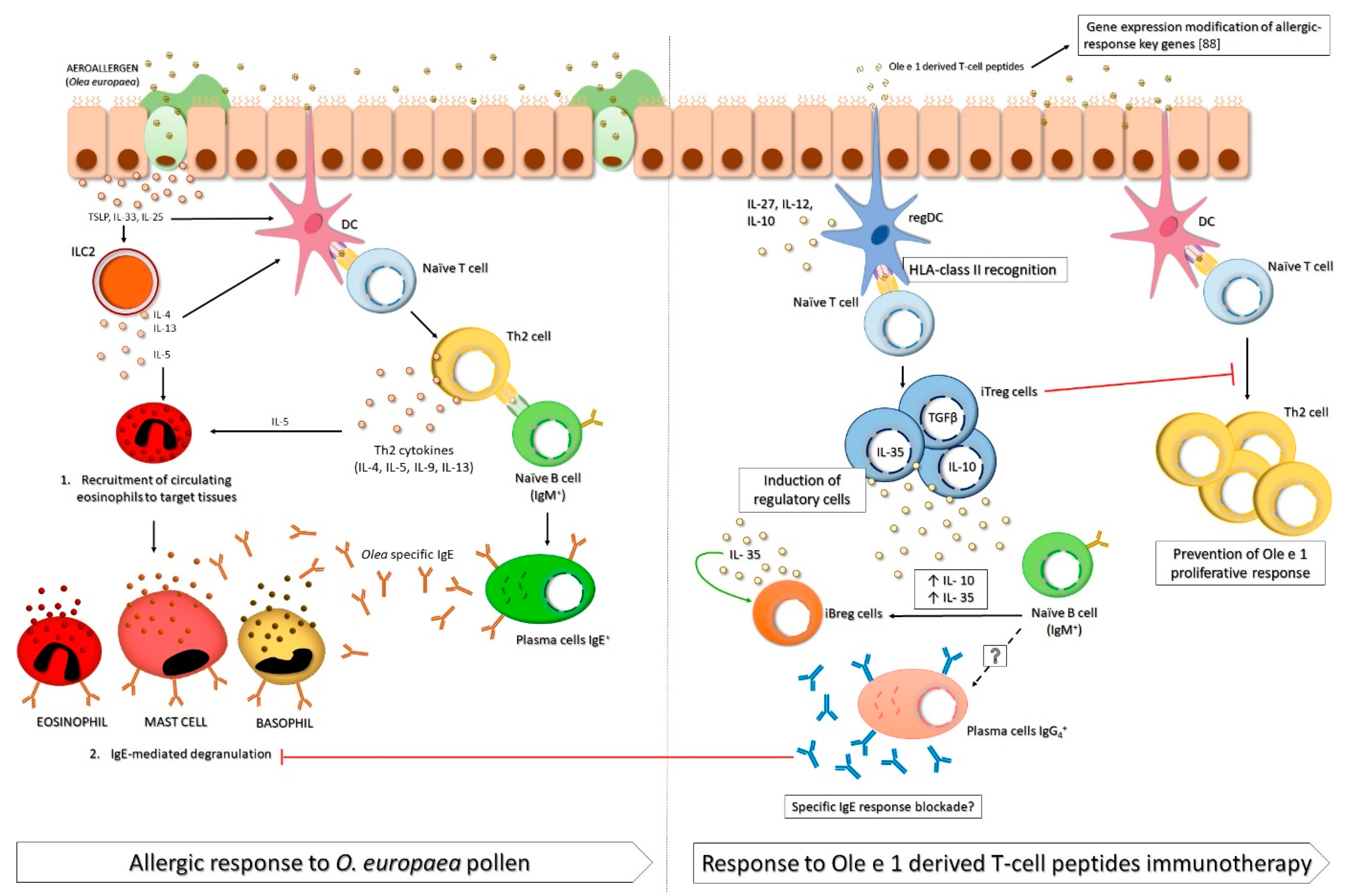
5.2. Ole e 1 T Cell Peptides: Mechanisms of Action
5.2.1. The Use of Peptides in the Era of Precision Medicine
5.2.2. Cellular Response: Treg vs. Th2 Response
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Marfo, K.; Lu, A.; Ling, M.; Akalin, E. Desensitization protocols and their outcome. Clin. J. Am. Soc. Nephrol. 2011, 6, 922–936. [Google Scholar] [CrossRef] [PubMed]
- Kannan, J.A.; Epstein, T.G. Immunotherapy safety: What have we learned from surveillance surveys? Curr. Allergy Asthma. Rep. 2013, 13, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Shamji, M.H.; Durham, S.R. Mechanisms of allergen immunotherapy for inhaled allergens and predictive biomarkers. J. Allergy Clin. Immunol. 2017, 140, 1485–1498. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Krempski, J.W.; Nadeau, K. Advances and novel developments in mechanisms of allergic inflammation. Allergy 2020, 75, 3100–3111. [Google Scholar] [CrossRef]
- Gunaguardana, N.C.; Durham, S.R. New approach to allergen immunotherapy. Ann. Allergy Asthma Inmmunol. 2018, 121, 293–305. [Google Scholar] [CrossRef]
- Zhao, H.; Liao, X.; Kang, Y. Tregs: Where we are and what comes next? Front. Immunol. 2017, 8, 1578. [Google Scholar] [CrossRef]
- Sözener, Z.C.; Mungan, D.; Cevhertas, L.; Ogulur, I.; Akdis, M.; Akdis, C. Tolerance mechanisms in allergen immnunotherapy. Curr. Opin. Allergy Clin. Immunol. 2020, 20, 591–601. [Google Scholar] [CrossRef] [PubMed]
- Akdis, C.A.; Arkwright, P.D.; Brüggen, M.C.; Busse, W.; Gadina, M.; Guttman-Yassky, E.; Kabashima, K.; Mitamura, Y.; Vian, L.; Wu, J.; et al. Type 2 immunity in the skin and lungs. Allergy 2020, 75, 1582–1605. [Google Scholar] [CrossRef]
- Komatsu, N.; Mariotti-Ferrandiz, M.E.; Wang, Y.; Malissen, B.; Waldmann, H.; Hori, S. Heterogeneity of natural Foxp3+ T cells: A committed regulatory T-cell lineage and an uncommitted minor population retaining plasticity. Proc. Natl. Acad. Sci. USA 2009, 106, 1903–1908. [Google Scholar] [CrossRef]
- Zhao, S.T.; Wang, C.Z. Regulatory T cells and asthma. J. Zhejiang Univ. Sci. B 2018, 19, 663–673. [Google Scholar] [CrossRef]
- Głobińska, A.; Boonpiyathad, T.; Satitsuksanoa, P.; Kleuskens, M.; van de Veen, W.; Sokolowska, M.; Akdis, M. Mechanisms of allergen-specific immunotherapy: Diverse mechanisms of immune tolerance to allergens. Ann. Allergy Asthma. Immunol. 2018, 121, 306–312. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, K.; Kitani, A.; Strober, W. Cell contact–dependent immunosuppression by CD4(+)CD25(+) regulatory T cells is mediated by cell surface–bound transforming growth factor beta. J. Exp. Med. 2001, 194, 629–644. [Google Scholar] [CrossRef] [PubMed]
- Jordan, M.S.; Boesteanu, A.; Reed, A.J.; Petrone, A.L.; Holenbeck, A.E.; Lerman, M.A.; Naji, A.; Caton, A.J. Thymic selection of CD4+CD25+regulatory T cells induced by an agonist self-peptide. Nat. Immunol. 2001, 2, 301–306. [Google Scholar] [CrossRef] [PubMed]
- Boonpiyathad, T.; Satitsuksanoa, P.; Akdis, M.; Akdis, C.A. IL-10 producing T and B cells in allergy. Semin. Immunol. 2019, 44, 101326. [Google Scholar] [CrossRef]
- Komlósi, Z.I.; Kovács, N.; Sokolowska, M.; van de Veen, W.; Akdis, M.; Akdis, C.A. Mechanisms of Subcutaneous and Sublingual Aeroallergen Immunotherapy: What Is New? Immunol. Allergy Clin. 2020, 40, 1–14. [Google Scholar] [CrossRef]
- Zemmour, D.; Zilionis, R.; Kiner, E.; Klein, A.M.; Mathis, D.; Benoist, C. Single-cell gene expression reveals a landscape of regulatory T cell phenotypes shaped by the TCR. Nat. Immunol. 2018, 19, 291–301. [Google Scholar] [CrossRef]
- Collison, L.W.; Chaturvedi, V.; Henderson, A.L.; Giacomin, P.R.; Guy, C.; Bankoti, J.; Finkelstein, D.; Forbes, K.; Workman, C.K.; Brown, S.A.; et al. IL-35-mediated induction of a potent regulatory T cell population. Nat. Immunol. 2010, 11, 1093–1101. [Google Scholar] [CrossRef]
- Sokolowska, M.; Boonpiyathad, T.; Escribese, M.M.; Barber, D. Allergen-specific immunotherapy: Power of adjuvants and novel predictive biomarkers. Allergy 2019, 74, 2061–2063. [Google Scholar] [CrossRef]
- Palomares, O.; Akdis, M.; Martín-Fontecha, M.; Akdis, C.A. Mechanisms of immune regulation in allergic diseases: The role of regulatory T and B cells. Immunol. Rev. 2017, 278, 219–236. [Google Scholar] [CrossRef]
- Calzada, D.; Baos, S.; Cremades, L.; Cárdaba, B. New Treatments for Allergy: Advances in Peptide Immunotherapy. Curr. Med. Chem. 2018, 25, 2215–2232. [Google Scholar] [CrossRef]
- Platts-Mills, T.; Vaughan, J.; Squillace, S.; Woodfolk, J.; Sporik, R. Sensitisation, asthma, and a modified Th2 response in children exposed to cat allergen: A population-based cross-sectional study. Lancet 2001, 357, 752–756. [Google Scholar] [CrossRef]
- Meiler, F.; Zumkehr, J.; Klunker, S.; Rückert, B.; Akdis, C.A.; Akdis, M. In vivo switch to IL-10-secreting T regulatory cells in high dose allergen exposure. J. Exp. Med. 2008, 205, 2887–2898. [Google Scholar] [CrossRef]
- Santos, M.C.P.; Serra-Caetano, A.; Pedro, E.; Melo, A.; Caramalho, I.; Barbosa, M.P.; Victorino, R.M.M.; Sousa, A.E. Expansion of FOXP3+ regulatory CD4 T cells upon exposure to hymenoptera venom during the beekeeping season. Allergy 2019, 74, 1182–1184. [Google Scholar] [CrossRef]
- Aguerri, M.; Calzada, D.; Martín, E.; Florido, F.; Quiralte, J.; Delgado, J.; Miranda, A.; López-Cacho, J.M.; Gallardo, S.; Lahoz, C.; et al. FOXP3 and TGF-β: Differential regulatory molecules between sensitization and tolerance to olive pollen. Eur. J. Inflamm. 2012, 10, 193–202. [Google Scholar] [CrossRef]
- Wraith, D.C.; Krishna, M.T. Peptide allergen-specific immunotherapy for allergic airway diseases-State of the art. Clin. Exp. Allergy 2021, 51, 751–769. [Google Scholar] [CrossRef] [PubMed]
- Passalacqua, G.; Bagnasco, D.; Ferrando, M.; Heffler, E.; Puggioni, F.; Canonica, G.W. Current insights in allergen immunotherapy. Ann. Allergy Asthma. Immunol. 2018, 120, 152–154. [Google Scholar] [CrossRef] [PubMed]
- Eckl-Dorna, J.; Villazala-Merino, S.; Linhart, B.; Karaulov, A.V.; Zhernov, Y.; Khaitov, M.; Niederberger-Leppin, V.; Valenta, R. Allergen-specific antibodies regulate secondary allergen-specific immune responses. Front. Immunol. 2019, 9, 3131. [Google Scholar] [CrossRef]
- Komlósi, Z.I.; Kovács, N.; Sokolowska, M.; van de Veen, W.; Akdis, M.; Akdis, C.A. Highlights of Novel Vaccination Strategies in Allergen Immunotherapy. Immunol. Allergy Clin. N. Am. 2020, 40, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Jacquet, A. Perspectives in Allergen-Specific Immunotherapy: Molecular Evolution of Peptide- and Protein-Based Strategies. Curr. Protein. Pept. Sci. 2020, 21, 203–223. [Google Scholar] [CrossRef]
- Dorofeeva, Y.; Shilovskiy, I.; Tulaeva, I.; Focke-Tejkl, M.; Flicker, S.; Kudlay, D.; Khaitov, M.; Karsonova, A.; Riabova, K.; Karaulov, A.; et al. Past, present, and future of allergen immunotherapy vaccines. Allergy 2021, 76, 131–149. [Google Scholar] [CrossRef]
- Marth, K.; Focke-Tejkl, M.; Lupinek, C.; Valenta, R.; Niederberger, V. Allergen Peptides, Recombinant Allergens and Hypoallergens for Allergen-Specific Immunotherapy. Curr. Treat. Options Allergy 2014, 26, 91–106. [Google Scholar] [CrossRef] [PubMed]
- Valenta, R.; Campana, R.; Focke-Tejkl, M.; Niederberger, V. Vaccine development for allergen-specific immunotherapy based on recombinant allergens and synthetic allergen peptides: Lessons from the past and novel mechanisms of action for the future. J. Allergy Clin. Immunol. 2016, 137, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Klimek, L.; Pfaar, O.; Worm, M. New opportunities for allergen immunotherapy using synthetic peptide immune-regulatory epitopes (SPIREs). Expert Rev. Clin. Immunol. 2016, 12, 1123–1135. [Google Scholar] [CrossRef] [PubMed]
- Incorvaia, C.; Montagni, M.; Ridolo, E. The efficiency of peptide immunotherapy for respiratory allergy. Expert Rev. Clin. Pharmacol. 2016, 9, 831–837. [Google Scholar] [CrossRef] [PubMed]
- Moldaver, D.; Larché, M. Immunotherapy with peptides. Allergy 2011, 66, 784–791. [Google Scholar] [CrossRef]
- Alexander, C.; Ying, S.; Kay, A.B.; Larché, M. Fel d 1-derived T cell peptide therapy induces recruitment of CD4+CD25+; CD4+ interferon-gamma+ T helper type 1 cells to sites of allergen-induced late-phase skin reactions in cat-allergic subjects. Clin. Exp. Allergy 2005, 35, 52–58. [Google Scholar] [CrossRef]
- Alexander, C.; Tarzi, M.; Larché, M.; Kay, A.B. The effect of Fel d 1-derived T-cell peptides on upper and lower airway outcome measurements in cat-allergic subjects. Allergy 2005, 60, 1269–1274. [Google Scholar] [CrossRef]
- Verhoef, A.; Alexander, C.; Kay, A.B.; Larché, M. T cell epitope immunotherapy induces a CD4+ T cell population with regulatory activity. PLoS Med. 2005, 2, e78. [Google Scholar] [CrossRef]
- Worm, M.; Lee, H.H.; Kleine-Tebbe, J.; Hafner, R.P.; Laidler, P.; Healey, D.; Buhot, C.; Verhoef, A.; Maillère, B.; Kay, A.B.; et al. Development and preliminary clinical evaluation of a peptide immunotherapy vaccine for cat allergy. J. Allergy Clin. Immunol. 2011, 127, 89–97. [Google Scholar] [CrossRef]
- Larché, M.; Patel, D.; Patel, P.; Salapatek, A.M.; Laidler, P.; Hafner, R. Safety and efficacy of Fel d 1 derived peptide immunotherapy in a double-blind, placebo-controlled environmental exposure chamber (EEC) study. Allergy 2012, 67, 1–97. [Google Scholar]
- Hafner, R.P.; Patel, P.; Salapatek, A.M.; Laider, P.; Larché, M.; Patel, D. Fel d 1 peptide antigen desensitization safety and efficacy in a double-blind, placebo-controlled environmental exposure chamber study. World Allergy Organ. J. 2013, 6 (Suppl. S1), P150. [Google Scholar] [CrossRef]
- Patel, D.; Couroux, P.; Hickey, P.; Salapatek, A.M.; Laider, P.; Larché, M.; Hafner, R.P. Fel d 1-derived peptide antigen desensitization shows a persistent treatment effect 1 year after the start of dosing: A randomized, placebo-controlled study. J. Allergy Clin. Immunol. 2013, 131, 103–109. [Google Scholar] [CrossRef]
- Hafner, R.P.; Couroux, P.; Armstrong, K.; Patel, D.; Larché, M. Two Year Persistent Treatment Effect Achieved After 4 Doses of Cat-Peptide Antigen Desensitization (Cat- PAD) in an Environmental Exposure Chamber (EEC) Model of Cat Allergy. J. Allergy Clin. Immunol. 2013, 131, AB147. [Google Scholar] [CrossRef]
- Hafner, R.P.; Couroux, P.; Armstrong, K.; Patel, D.; Larché, M.; Haumann, B. Total Nasal Symptom Scores are reduced in an EEC model of cat allergy two years after administration of 4 doses of Cat-PAD. Allergy Eur. J. Allergy Clin. Immunol. 2013, 68, 647. [Google Scholar]
- Pfaar, O.; Klimek, L.; Varga, E.M. Fel d 1 synthetic peptides (Cat-PAD)—Good news for cat owners with children? Pediatr. Allergy Immunol. 2016, 27, 666–670. [Google Scholar] [CrossRef]
- Kündig, T.M.; Senti, G.; Schnetzler, G.; Wolf, C.; Vavricka, B.M.P.; Fulurija, A.; Hennecke, F.; Sladko, K.; Jennings, G.T.; Bachmann, M.F. Der p 1 peptide on virus-like particles is safe and highly immunogenic in healthy adults. J. Allergy Clin. Immunol. 2006, 117, 1470–1476. [Google Scholar] [CrossRef] [PubMed]
- Larché, M.; Hickey, P.; Hebert, J.; Hafner, R.P. Safety and Tolerability of Escalating Doses of House Dust Mite- Peptide Antigen Desensitization (HDM-PAD). J. Allergy Clin. Immunol. 2013, 131, AB37. [Google Scholar] [CrossRef]
- Hafner, R.P.; Couroux, P.; Armstrong, K.; Salapatek, A.M.; Patel, D.; Larché, M. Persistent treatment effect achieved at one year after four doses of Der p derived synthetic peptide immuno-regulatory epitopes in an exposure chamber model of House Dust Mite allergy. J. Allergy Clin. Immunol. 2014, 133, AB289. [Google Scholar] [CrossRef]
- Hafner, R.P.; Salapatek, A.M.; Larché, M.; Ahenkorah, B.; Patel, P.; Pawsey, S. Comparison of the treatment effect of house dust mite synthetic peptides immune-regulatory epitopes in the environmental exposure chamber and field setting two years after a short course of treatment. Allergy 2015, 70, 28. [Google Scholar]
- Spertini, F.; Perrin, Y.; Audran, R.; Pellaton, C.; Boudousquie, C.; Barbier, N.; Thierry, A.C.; Charlon, V.; Reymond, C. Safety and immunogenicity of immunotherapy with Bet v 1-derived contiguous overlapping peptides. J. Allergy Clin. Immunol. 2014, 134, 239–240. [Google Scholar] [CrossRef]
- Spertini, F.; DellaCorte, G.; Kettner, A.; de Blay, F.; Jacobsen, L.; Jutel, M.; Worm, M.; Charlon, V.; Reymond, C. Efficacy of 2 months of allergen-specific immunotherapy with Bet v 1-derived contiguous overlapping peptides in patients with allergic rhinoconjunctivitis: Results of a phase IIb study. J. Allergy Clin. Immunol. 2016, 138, 162–168. [Google Scholar] [CrossRef] [PubMed]
- Kettner, A.; DellaCorte, G.; de Blay, F.; Jacobsen, L.; Jutel, M.; Worm, M.; Charlon, V.; Simonsen, K.; Reymond, C.; Spertini, F. Benefit of Bet v 1 contiguous overlapping peptide immunotherapy persists during first follow-up season. J. Allergy Clin. Immunol. 2018, 142, 678–680 e7. [Google Scholar] [CrossRef] [PubMed]
- Ellis, A.K.; Frankish, C.W.; Armstrong, K.; Steacy, L.; Tenn, M.W.; Pawsey, S.; Hafner, R.P. Persistence of the clinical effect of grass allergen peptide immunotherapy after the second and third grass pollen seasons. J. Allergy Clin. Immunol. 2020, 145, 610–618. [Google Scholar] [CrossRef] [PubMed]
- Zhernov, Y.; Curin, M.; Khaitov, M.; Karaulov, A.; Valenta, R. Recombinant allergens for immunotherapy: State of the art. Curr. Opin. Allergy Clin. Immunol. 2019, 19, 402–414. [Google Scholar] [CrossRef] [PubMed]
- Agache, I. Peptide allergen immunotherapy-unraveling new pathways. J. Allergy Clin. Immunol. 2019, 144, 658–660. [Google Scholar] [CrossRef]
- Rudulier, C.D.; Tonti, E.; James, E.; Kwok, W.W.; Larché, M. Modulation of CRTh2 expression on allergen-specific T cells following peptide immunotherapy. Allergy 2019, 74, 2157–2166. [Google Scholar] [CrossRef]
- Liccardi, G.; D’Amato, M.; D’Amato, G. Oleaceae pollinosis: A review. Int. Arch. Allergy Immunol. 1996, 111, 210–217. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.; Pierantozzi, P.; Searles, P.; Rousseaux, M.C.; García-Inza, G.; Miserere, A.; Bodoira, R.; Contreras, C.; Maestri, D. Olive Cultivation in the Southern Hemisphere: Flowering, Water Requirements and Oil Quality Responses to New Crop Environments. Front. Plant Sci. 2017, 27, 1830. [Google Scholar] [CrossRef] [PubMed]
- Hamman-Khalifa, A.M.; Castro, A.J.; Jiménez-López, J.C.; Rodríguez-García, M.I.; Alché, J.D. Olive cultivar origin is a major cause of polymorphism for Ole e 1 pollen allergen. BMC Plant Biol. 2008, 25, 10. [Google Scholar] [CrossRef]
- Castro, A.J.; Bednarczyk, A.; Schaeffer-Reiss, C.; Rodríguez-García, M.I.; Van Dorsselaer, A.; Alché, J.D. Screening of Ole e 1 polymorphism among olive cultivars by peptide mapping and N-glycopeptide analysis. Proteomics 2010, 10, 953–962. [Google Scholar] [CrossRef]
- Lombardero, M.; Obispo, T.; Calabozo, B.; Lezaún, A.; Polo, F.; Barber, D. Cross-reactivity between olive and other species. Role of Ole e 1-related proteins. Allergy 2002, 57 (Suppl. S71), 29–34. [Google Scholar] [CrossRef]
- Palomares, O.; Swoboda, I.; Villalba, M.; Balic, N.; Spitzauer, S.; Rodríguez, R.; Valenta, R. The major allergen of olive pollen Ole e 1 is a diagnostic marker for sensitization to Oleaceae. Int. Arch. Allergy Immunol. 2006, 141, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Villalba, M.; Rodríguez, R.; Batanero, E. The spectrum of olive pollen allergens. From structures to diagnosis and treatment. Methods 2014, 66, 44–54. [Google Scholar] [CrossRef]
- Oeo-Santos, C.; Mas, S.; Quiralte, J.; Colás, C.; Blanca, M.; Fernández, J.; Feo Brito, F.; Villalba, M.; Barderas, R. A Hypoallergenic Polygalacturonase Isoform from Olive Pollen Is Implicated in Pollen-Pollen Cross-Reactivity. Int. Arch. Allergy Immunol. 2018, 177, 290–301. [Google Scholar] [CrossRef] [PubMed]
- Jimenez-Lopez, J.C.; Robles-Bolivar, P.; Lopez-Valverde, F.J.; Lima-Cabello, E.; Kotchoni, S.O.; Alché, J.D. Ole e 13 is the unique food allergen in olive: Structure-functional, substrates docking, and molecular allergenicity comparative analysis. J. Mol. Graph. Model. 2016, 66, 26–40. [Google Scholar] [CrossRef] [PubMed]
- Quiralte, J.; Llanes, E.; Barral, P.; Arias de Saavedra, J.M.; Sáenz de San Pedro, B.; Villalba, M.; Florido, J.F.; Rodríguez, R.; Lahoz, C.; Cárdaba, B. Ole e 2 and Ole e 10: New clinical aspects and genetic restrictions in olive pollen allergy. Allergy 2005, 60, 360–365. [Google Scholar] [CrossRef]
- Barber, D.; de la Torre, F.; Feo, F.; Florido, F.; Guardia, P.; Moreno, C.; Quiralte, J.; Lombardero, M.; Villalba, M.; Salcedo, G.; et al. Understanding patient sensitization profiles in complex pollen areas: A molecular epidemiological study. Allergy 2008, 63, 1550–1558. [Google Scholar] [CrossRef]
- Villalba, M.; Batanero, E.; López-Otín, C.; Sánchez, L.M.; Monsalve, R.I.; González de la Peña, M.A.; Lahoz, C.; Rodríguez, R. The amino acid sequence of Ole e I, the major allergen from olive tree (Olea europaea) pollen. Eur. J. Biochem. 1993, 216, 863–869. [Google Scholar] [CrossRef] [PubMed]
- Batanero, E.; Villalba, M.; Rodríguez, R. Glycosylation site of the major allergen from olive tree pollen. Allergenic implications of the carbohydrate moiety. Mol. Immunol. 1994, 31, 31–37. [Google Scholar] [CrossRef]
- Martín-Orozco, E.; Cárdaba, B.; del Pozo, V.; de Andrés, B.; Villalba, M.; Gallardo, S.; Rodriguez-García, M.I.; Fernández, M.C.; Alché, J.D.; Rodriguez, R. Ole e I: Epitope mapping, cross-reactivity with other Oleaceae pollens and ultrastructural localization. Int. Arch. Allergy Immunol. 1994, 104, 160–170. [Google Scholar] [CrossRef] [PubMed]
- Cárdaba, B.; Del Pozo, V.; Jurado, A.; Gallardo, S.; Cortegano, I.; Arrieta, I.; Del Amo, A.; Tramón, P.; Florido, F.; Sastre, J.; et al. Olive pollen allergy: Searching for immunodominant T-cell epitopes on the Ole e 1 molecule. Clin. Exp. Allergy 1998, 28, 413–422. [Google Scholar] [CrossRef] [PubMed]
- Jespersen, M.C.; Peters, B.; Nielsen, M.; Marcatili, P. BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 2017, 3, W24–W29. [Google Scholar] [CrossRef]
- Chou, P.Y.; Fasman, G.D. Prediction of the Secondary Structure of Proteins from their Amino Acid Sequence. Adv. Enzymol. Relat Areas Mol. Biol. 1978, 47, 45–148. [Google Scholar] [CrossRef] [PubMed]
- Emini, E.A.; Hughes, J.V.; Perlow, D.S.; Boger, J. Induction of hepatitis A virus-neutralizing antibody by a virus-specific synthetic peptide. J. Virol. 1985, 55, 836–839. [Google Scholar] [CrossRef] [PubMed]
- Karplus, P.A.; Schultz, G. Prediction of chain flexibility in proteins. Naturwissenschaften 1985, 72, 212–213. [Google Scholar] [CrossRef]
- Parker, J.M.; Guo, D.; Hodges, R.S. New hydrophilicity scale derived from high-performance liquid chromatography peptide retention data: Correlation of predicted surface residues with antigenicity and X-ray-derived accessible sites. Biochemistry 1986, 23, 5425–5432. [Google Scholar] [CrossRef]
- Kolaskar, A.S.; Tongaonkar, P.C. A semi-empirical method for prediction of antigenic determinants on protein antigens. FEBS Lett. 1990, 276, 172–174. [Google Scholar] [CrossRef]
- Sanchez-Trincado, J.L.; Gomez-Perosanz, M.; Reche, P.A. Fundamentals and Methods for T- and B-Cell Epitope Prediction. J. Immunol. Res. 2017, 201, 2680160. [Google Scholar] [CrossRef]
- Ahmed, R.K.; Maeurer, M.J. T-cell epitope mapping. Methods Mol. Biol. 2009, 524, 427–438. [Google Scholar] [CrossRef]
- Lafuente, E.M.; Reche, P.A. Prediction of MHC-peptide binding: A systematic and comprehensive overview. Curr. Pharm. Des. 2009, 15, 3209–3220. [Google Scholar] [CrossRef]
- Jensen, P.E. Recent advances in antigen processing and presentation. Nat. Immunol. 2007, 8, 1041–1048. [Google Scholar] [CrossRef]
- Dhanda, S.K.; Karosiene, E.; Edwards, L.; Grifoni, A.; Paul, S.; Andreatta, M.; Weiskopf, D.; Sidney, J.; Nielsen, M.; Peters, B.; et al. Predicting HLA CD4 Immunogenicity in Human Populations. Front. Immunol. 2018, 14, 1369. [Google Scholar] [CrossRef]
- Marazuela, E.G.; Rodríguez, R.; Fernández-García, H.; García, M.S.; Villalba, M.; Batanero, E. Intranasal immunization with a dominant T-cell epitope peptide of a major allergen of olive pollen prevents mice from sensitization to the whole allergen. Mol. Immunol. 2008, 45, 438–445. [Google Scholar] [CrossRef]
- Marazuela, E.G.; Prado, N.; Moro, E.; Fernández-García, H.; Villalba, M.; Rodríguez, R.; Batanero, E. Intranasal vaccination with poly(lactide-co-glycolide) microparticles containing a peptide T of Ole e 1 prevents mice against sensitization. Clin. Exp Allergy 2008, 38, 520–528. [Google Scholar] [CrossRef] [PubMed]
- Benedé, S.; Ramos-Soriano, J.; Palomares, F.; Losada, J.; Mascaraque, A.; López-Rodríguez, J.C.; Rojo, J.; Mayorga, C.; Villalba, M.; Batanero, E. Peptide Glycodendrimers as Potential Vaccines for Olive Pollen Allergy. Mol. Pharm. 2020, 17, 827–836. [Google Scholar] [CrossRef]
- Twaroch, T.E.; Focke, M.; Civaj, V.; Weber, M.; Balic, N.; Mari, A.; Ferrara, R.; Quirce, S.; Spitzauer, S.; Swoboda, I.; et al. Carrier-bound, nonallergenic Ole e 1 peptides for vaccination against olive pollen allergy. J. Allergy Clin. Immunol. 2011, 128, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Cárdaba, B.; Llanes, E.; Chacártegui, M.; Sastre, B.; López, E.; Mollá, R.; del Pozo, V.; Florido, F.; Quiralte, J.; Palomino, P.; et al. Modulation of Allergic Response by Gene–Environment Interaction: Olive Pollen Allergy. J. Investig. Allergol. Clin. Immunol. 2007, 17 (Suppl. S1), 83–87. [Google Scholar] [CrossRef]
- Calzada, D.; Aguerri, M.; Baos, S.; Montaner, D.; Mata, M.; Dopazo, J.; Quiralte, J.; Florido, F.; Lahoz, C.; Cárdaba, B. Therapeutic targets for olive pollen allergy defined by gene markers modulated by Ole e 1-derived peptides. Mol. Immunol. 2015, 64, 252–261. [Google Scholar] [CrossRef]
- Sokolowska, M.; Akdis, C.A. Highlights in immune response, microbiome and precision medicine in allergic disease and asthma. Curr. Opin. Immunol. 2017, 48, iv–ix. [Google Scholar] [CrossRef] [PubMed]
- Incorvaia, C.; Al-Ahmad, M.; Ansotegui, I.J.; Arasi, S.; Bachert, C.; Bos, C.; Bousquet, J.; Bozek, A.; Caimmi, D.; Calderón, M.A.; et al. Personalized medicine for allergy treatment: Allergen immunotherapy still a unique and unmatched model. Allergy 2021, 76, 1041–1052. [Google Scholar] [CrossRef]
- Calzada, D.; Cremades-Jimeno, L.; Pedro, M.Á.; Baos, S.; Rial, M.; Sastre, J.; Quiralte, J.; Florido, F.; Lahoz, C.; Cárdaba, B. Therapeutic potential of peptides from Ole e 1 in olive-pollen allergy. Sci. Rep. 2019, 4, 15942. [Google Scholar] [CrossRef]
- Tulaeva, I.; Kratzer, B.; Campana, R.; Curin, M.; van Hage, M.; Karsonova, A.; Riabova, K.; Karaulov, A.; Khaitov, M.; Pickl, W.F.; et al. Preventive Allergen-Specific Vaccination against Allergy: Mission Possible? Front. Immunol. 2020, 7, 1368. [Google Scholar] [CrossRef]
- Akinfenwa, O.; Rodríguez-Domínguez, A.; Vrtala, S.; Valenta, R.; Campana, R. Novel vaccines for allergen-specific immunotherapy. Curr. Opin. Allergy Clin. Immunol. 2021, 1, 86–99. [Google Scholar] [CrossRef]
- Layhadi, J.A.; Eguiluz-Gracia, I.; Shamji, M.H. Role of IL-35 in sublingual allergen immunotherapy. Curr. Opin. Allergy Clin. Immunol. 2019, 19, 12–17. [Google Scholar] [CrossRef]
- Hu, D. Role of Anti-inflammatory Cytokines IL-35 and IL-37 in Asthma. Inflammation 2017, 40, 697–707. [Google Scholar] [CrossRef] [PubMed]
- Niedbala, W.; Wei, X.Q.; Cai, B.; Hueber, A.J.; Leung, B.P.; McInnes, I.B.; Liew, F.Y. IL-35 is a novel cytokine with therapeutic effects against collagen-induced arthritis through the expansion of regulatory T cells and suppression of Th17 cells. Eur. J. Immunol. 2007, 37, 3021–3029. [Google Scholar] [CrossRef] [PubMed]
- Su, L.C.; Liu, X.Y.; Huang, A.F.; Xu, W.D. Emerging role of IL-35 in inflammatory autoimmune diseases. Autoimmun. Rev. 2018, 17, 665–673. [Google Scholar] [CrossRef]
- Dambuza, I.M.; He, C.; Choi, J.K.; Yu, C.R.; Wang, R.; Mattapallil, M.J.; Wingfield, P.T.; Caspi, R.R.; Egwuagu, C.E. IL-12p35 induces expansion of IL-10 and IL-35-expressing regulatory B cells and ameliorates autoimmune disease. Nat. Commun. 2017, 28, 719. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.R.; Choi, J.K.; Uche, A.N.; Egwuagu, C.E. Production of IL-35 by Bregs is mediated through binding of BATF-IRF-4-IRF-8 complex to il12a and ebi3 promoter elements. J. Leukoc. Biol. 2018, 104, 1147–1157. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Zhang, Y.; Ye, J.; Wang, X.; Fu, X.; Yin, Y.; Wen, J.; Wu, X.; Xia, Z. IL-35 promoted STAT3 phosphorylation and IL-10 production in B cells, but its production was reduced in patients with coronary artery diseases. Hum. Immunol. 2018, 79, 869–875. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Deng, Y.; Chen, H.; Wu, X.; Cheng, S.; Xu, Y.; Xiong, W.; Xie, J. Decreased concentration of IL-35 in plasma of patients with asthma and COPD. Asian Pac. J. Allergy Immunol. 2014, 32, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.F.; Chen, Q.; Li, L.; Liu, J.M.; Zhang, G.P.; Zhu, X.H.; Wu, A.M.; Ke, J.W.; Dai, Y.L.; Wu, C.X. Effects of sublingual immunotherapy on serum IL-17 and IL-35 levels in children with allergic rhinitis or asthma. Zhongguo Dang Dai Er Ke Za Zhi 2014, 16, 1206–1210. [Google Scholar] [PubMed]
- Collison, L.W.; Delgoffe, G.M.; Guy, C.S.; Vignali, K.M.; Chaturvedi, V.; Fairweather, D.L.; Satoskar, A.R.K.; Garcia, C.; Hunter, C.A.; Drake, C.G.; et al. The composition and signaling of the IL-35 receptor are unconventional. Nat. Immunol. 2012, 13, 290–299. [Google Scholar] [CrossRef] [PubMed]
- Burchill, M.A.; Yang, J.; Vogtenhuber, C.; Blazar, B.R.; Farrar, M.A. IL-2 receptor beta-dependent STAT5 activation is required for the development of Foxp3+ regulatory T cells. J. Immunol. 2007, 1, 280–290. [Google Scholar] [CrossRef]
- Burchill, M.A.; Yang, J.; Vang, K.B.; Moon, J.J.; Chu, H.H.; Lio, C.W.; Vegoe, A.L.; Hsieh, C.S.; Jenkins, M.K.; Farrar, M.A. Linked T cell receptor and cytokine signaling govern the development of the regulatory T cell repertoire. Immunity 2008, 28, 112–121. [Google Scholar] [CrossRef]
- Yao, Z.; Kanno, Y.; Kerenyi, M.; Stephens, G.; Durant, L.; Watford, W.T.; Laurence, A.; Robinson, G.W.; Shevach, E.M.; Moriggl, R.; et al. Nonredundant roles for Stat5a/b in directly regulating Foxp3. Blood 2007, 109, 4368–4375. [Google Scholar] [CrossRef]
- Liu, H.; Zhang, W.; Tian, F.F.; Kun, A.; Zhou, W.B.; Xiao, B.; Li, J. IL-35 Is Involved in the Pathogenesis of Guillain-Barré Syndrome Through Its Influence on the Function of CD4+ T Cells. Immunol. Investig. 2015, 44, 566–577. [Google Scholar] [CrossRef]
| Peptide | Start | End | Combined Score | Immunogenicity Score | Peptide Core | Median Percentile Rank (7-Allele) | HLA-DRB1: 03:01 | HLA-DRB1: 07:01 | HLA-DRB1: 15:01 | HLA-DRB3: 01:01 | HLA-DRB3: 02:02 | HLA-DRB4: 01:01 | HLA-DRB5: 01:01 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ole e 1 sequence | TCRAGFITELSEFIP | 21 | 35 | 56.25 | 97.12 | FITELSEFI | 29.0 | 55.0 | 20.0 | 24.0 | 3.1 | 31.0 | 51.0 | 29.0 |
| SEFIPGASVRLQCRE | 31 | 45 | 56.20 | 98.51 | FIPGASVRL | 28.0 | 46.0 | 9.1 | 28.0 | 34.0 | 3.9 | 42.0 | 19.0 | |
| VGYTRAEGLYSMLVE | 56 | 70 | 58.36 | 99.42 | YTRAEGLYS | 31.0 | 71.0 | 17.0 | 33.0 | 31.0 | 15.0 | 64.0 | 11.0 | |
| EFCEITLISSGRKDC | 76 | 90 | 57.37 | 90.93 | ITLISSGRK | 35.0 | 35.0 | 35.0 | 11.0 | 83.0 | 39.0 | 58.0 | 0.9 | |
| PSLKFILNTVNGTTR | 101 | 115 | 35.37 | 71.92 | LKFILNTVN | 11.0 | 27.0 | 18.0 | 14.0 | 8.4 | 0.01 | 11.0 | 7.0 |
| Elm Name | Name | Matched Sequence | Positions | ELM Description | Cell Compartment | Pattern | Probability |
|---|---|---|---|---|---|---|---|
| DEG_Nend_Nbox_1 | N-degron | FHIQGQVYCDTC | 1-2 | N-terminal motif that initiates protein degradation. | Cytosol | ^M{0,1}[FYLIW][^P] | 2.302 × 10−4 |
| LIG_PDZ_Class_3 | Class 3 PDZ domain ligands | FHIQGQVYCDTC | 7–12 | C-terminal class 3 PDZ-binding motif. | Cytosol, internal side of plasma membrane | ...[DE].[ACVILF]$ | 6.168 × 10−5 |
| LIG_SH2_STAT5 | STAT5 Src Homology 2 (SH2) domain ligand | FHIQGQVYCDTC | 8–11 | STAT5 Src Homology 2 (SH2) domain binding motif. | Cytosol | (Y)[VLTFIC].. | 3.296 × 10−3 |
| DEG_Nend_UBRbox_4 | N-degron | CRAGFITELSEF | 1–2 | N-terminal motif that initiates protein degradation. | Cytosol | ^M{0,1}(C). | 1.768 × 10−5 |
| LIG_PDZ_Class_1 | Class 1 PDZ domain ligands | CRAGFITELSEF | 7–12 | C-terminal class 1 PDZ-binding motif. | Cytosol, internal side of plasma membrane | ...[ST].[ACVILF]$ | 7.255 × 10−5 |
| DEG_Nend_UBRbox_3 | N-degron | NEIPTEGWAKPS | 1–2 | N-terminal motif that initiates protein degradation. | Cytosol | ^M{0,1}([NQ]). | 1.645 × 10−4 |
| LIG_LIR_Apic_2 | LC3-interacting region ligand | NEIPTEGWAKPS NEIPTEGWAKPS | 5–11 6–11 | Apicomplexa specific variant of the canonical LIR motif involved in autophagy. | Cytosol, cytoplasmic side of late endosome membrane | [EDST].{0,2}[WFY]..P | 3.371 × 10−3 |
| LIG_FHA_1 | FHA phosphopeptide ligands | TVNGTTRTVNPL | 4–10 | Phosphothreonine motif binding a subset of FHA domains. | Nucleus | ..(T)..[ILV]. | 8.662 × 10−3 |
| MOD_N-GLC_1 | Generic motif for N-glycosylation | TVNGTTRTVNPL | 2–7 | Generic motif for N-glycosylation. | Extracellular, Golgi apparatus, endoplasmic reticulum | .(N)[^P][ST].. | 5.018 × 10−3 |
| CLV_PCSK_SKI1_1 | Subtilisin/kexin isozyme-1 (SKI1) cleavage site | PLGFFKKEALPK | 7–11 | Subtilisin/kexin isozyme-1 (SKI1) cleavage site. | Endoplasmic reticulum lumen, endoplasmic reticulum, Golgi apparatus, extracellular | [RK].[AILMFV][LTKF]. | 6.821 × 10−3 |
| LIG_REV1ctd_RIR_1 | Rev1-interacting regions ligand | PLGFFKKEALPK | 2–10 | DNA repair proteins interact with the C-terminal domain of the Rev1 translesion synthesis scaffold (Rev1-Interacting Region). | Nucleoplasm, nucleus | ..FF[^P]{0,2}[KR]{1,2}[^P]{0,4} | 5.350 × 10−4 |
| MOD_SUMO_for_1 | Motif recognised for modification by SUMO-1 | PLGFFKKEALPK | 5–8 | Motif recognised for modification by SUMO-1 | Nucleus, PML body | [VILMAFP](K).E | 1.914 × 10−3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Calzada, D.; Cremades-Jimeno, L.; López-Ramos, M.; Cárdaba, B. Peptide Allergen Immunotherapy: A New Perspective in Olive-Pollen Allergy. Pharmaceutics 2021, 13, 1007. https://doi.org/10.3390/pharmaceutics13071007
Calzada D, Cremades-Jimeno L, López-Ramos M, Cárdaba B. Peptide Allergen Immunotherapy: A New Perspective in Olive-Pollen Allergy. Pharmaceutics. 2021; 13(7):1007. https://doi.org/10.3390/pharmaceutics13071007
Chicago/Turabian StyleCalzada, David, Lucía Cremades-Jimeno, María López-Ramos, and Blanca Cárdaba. 2021. "Peptide Allergen Immunotherapy: A New Perspective in Olive-Pollen Allergy" Pharmaceutics 13, no. 7: 1007. https://doi.org/10.3390/pharmaceutics13071007
APA StyleCalzada, D., Cremades-Jimeno, L., López-Ramos, M., & Cárdaba, B. (2021). Peptide Allergen Immunotherapy: A New Perspective in Olive-Pollen Allergy. Pharmaceutics, 13(7), 1007. https://doi.org/10.3390/pharmaceutics13071007