Synthesis and Optimization of Mesoporous Silica Nanoparticles for Ruthenium Polypyridyl Drug Delivery
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials and Reagents
2.2. Synthesis of Mesoporous Silica Nanoparticles
2.3. Experimental Design
2.4. In Vitro Drug Loading and Release
2.5. Cytotoxicity Studies
2.6. Cell Cycle Analysis
3. Results
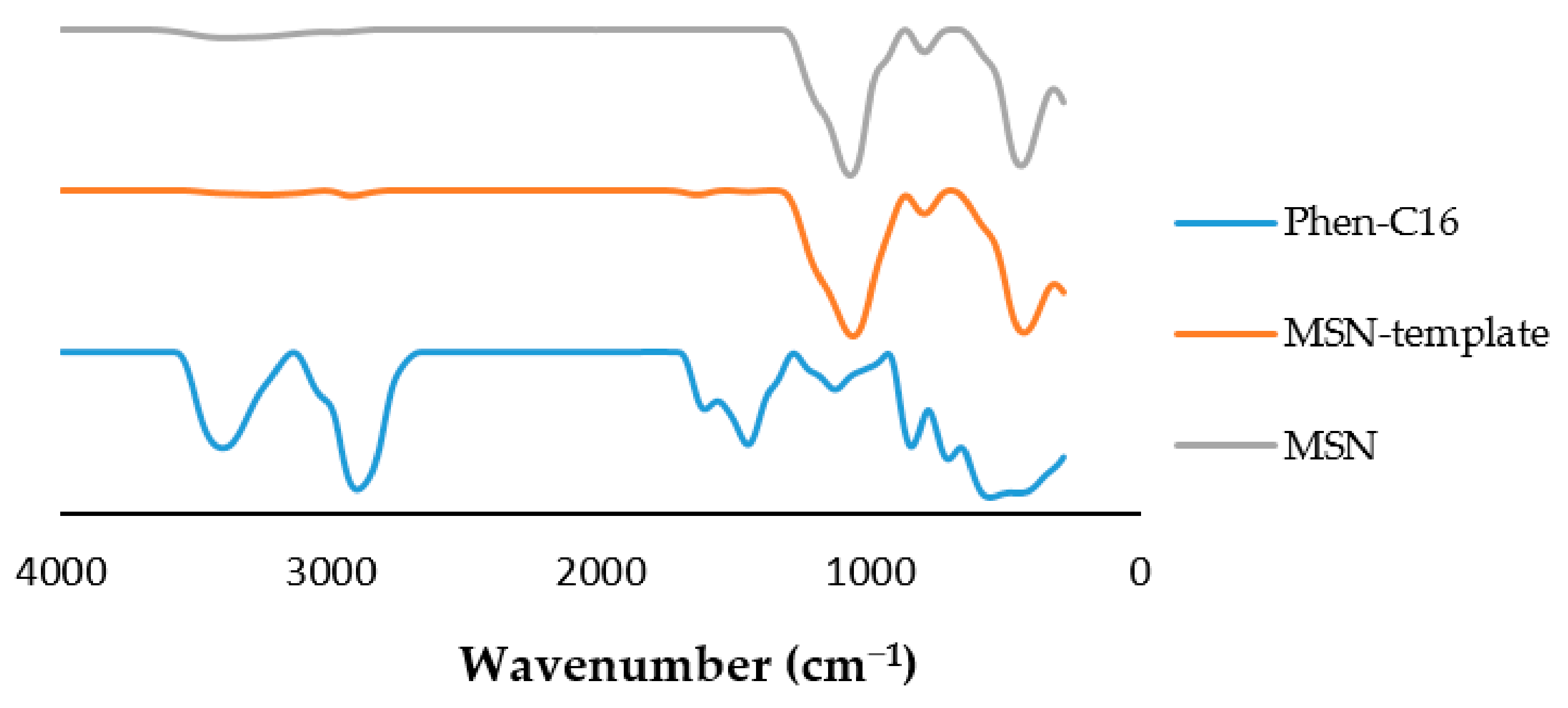
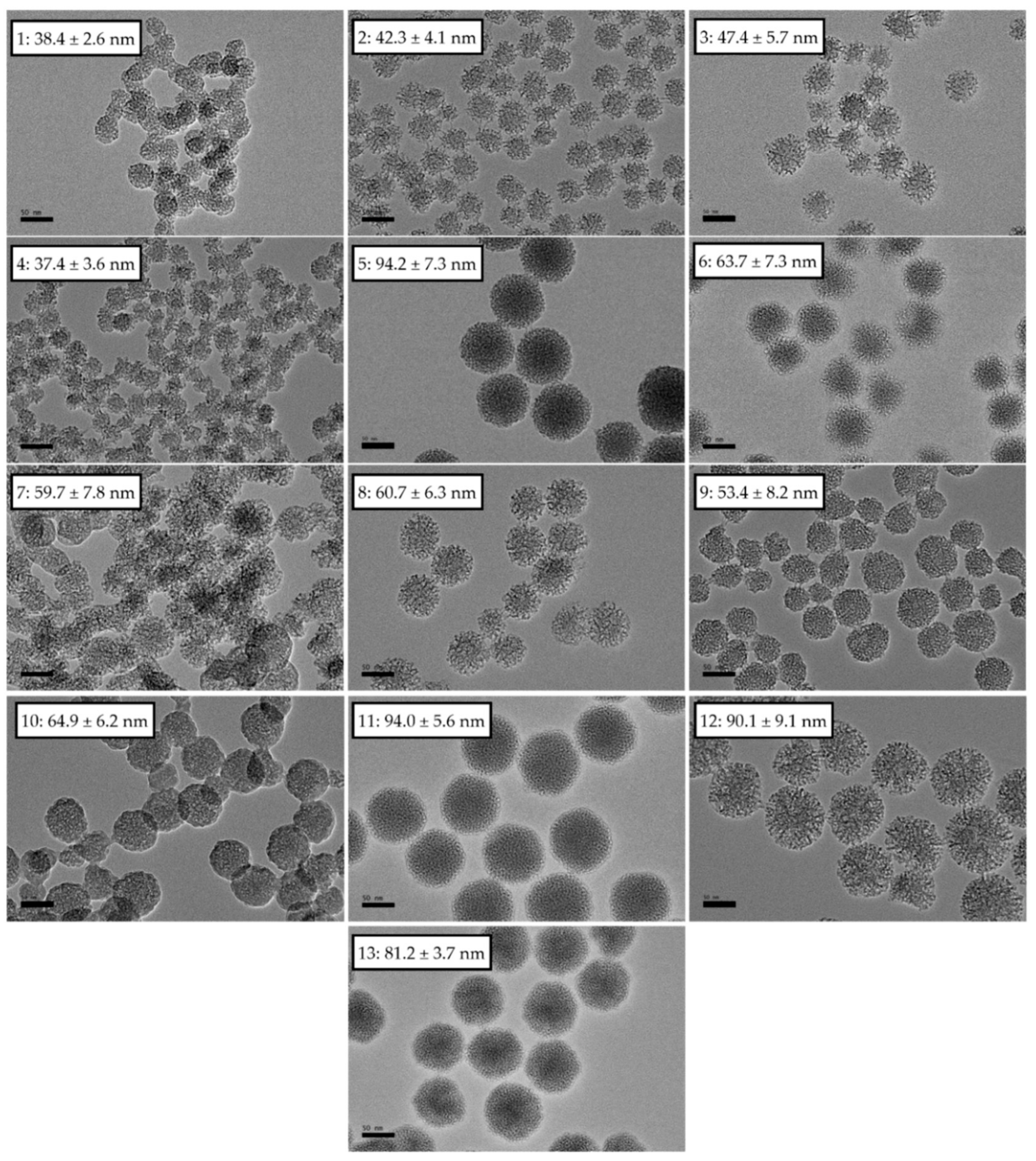
3.1. Physical Characterization
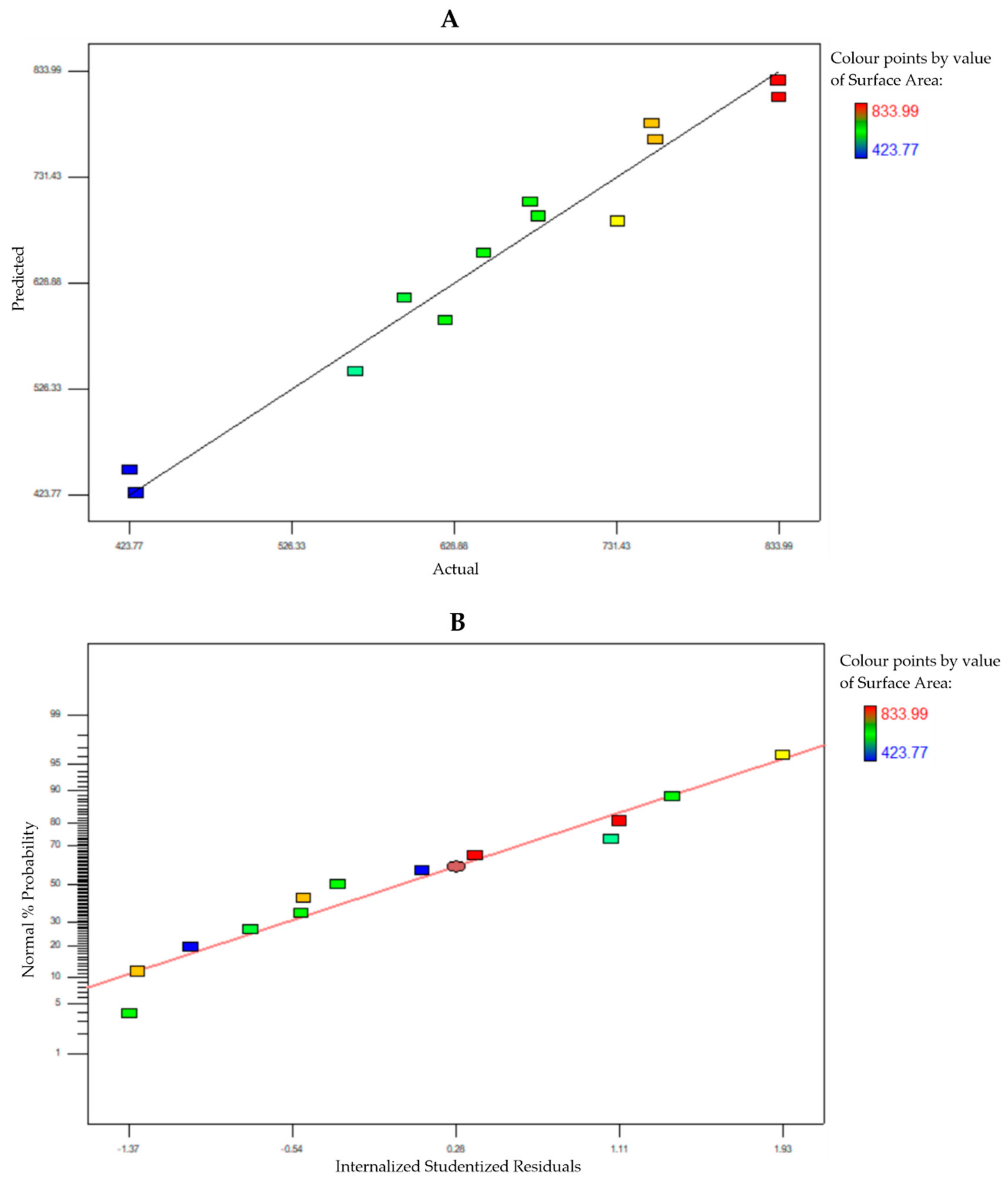
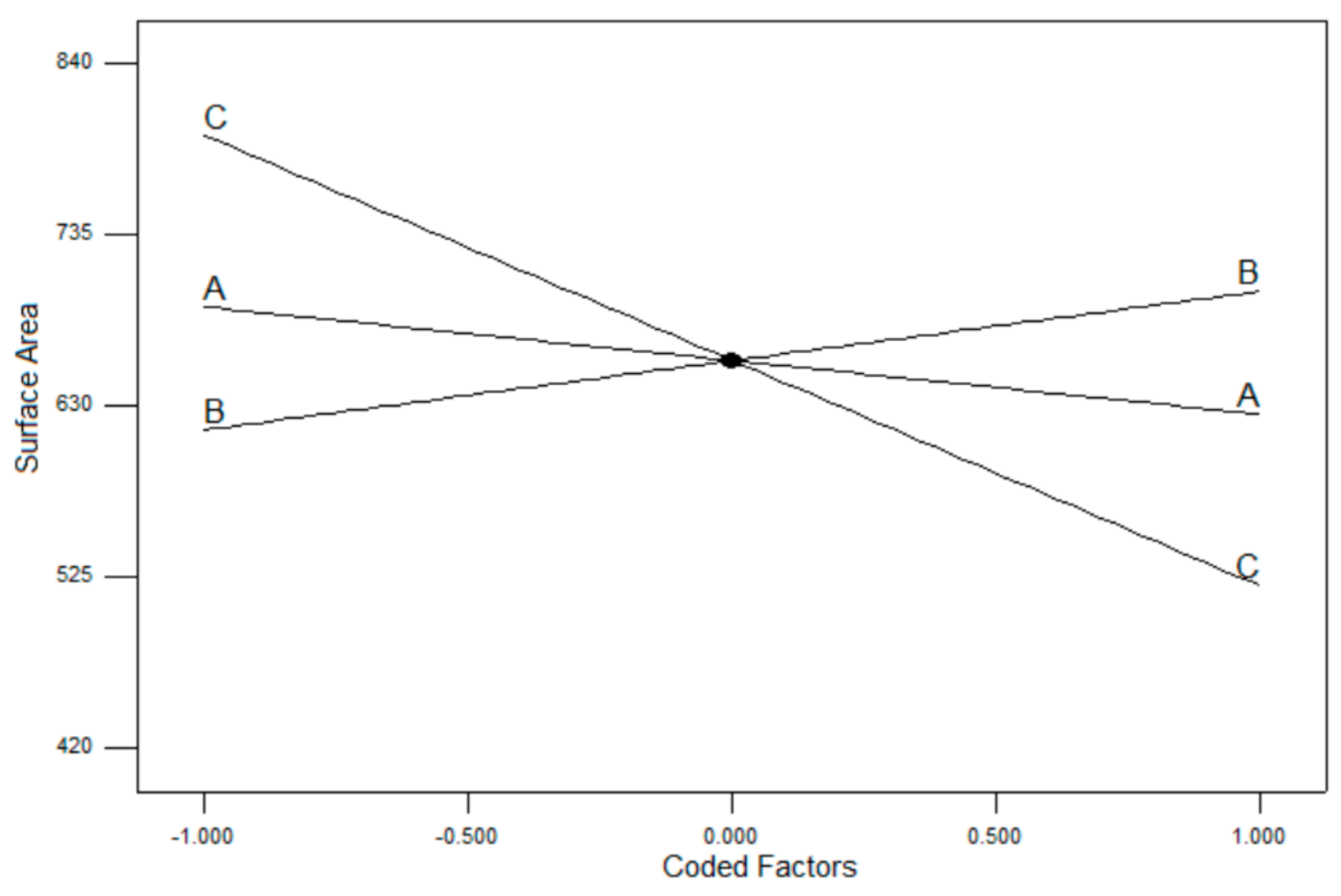
3.2. Box–Behnken Design Surface Response
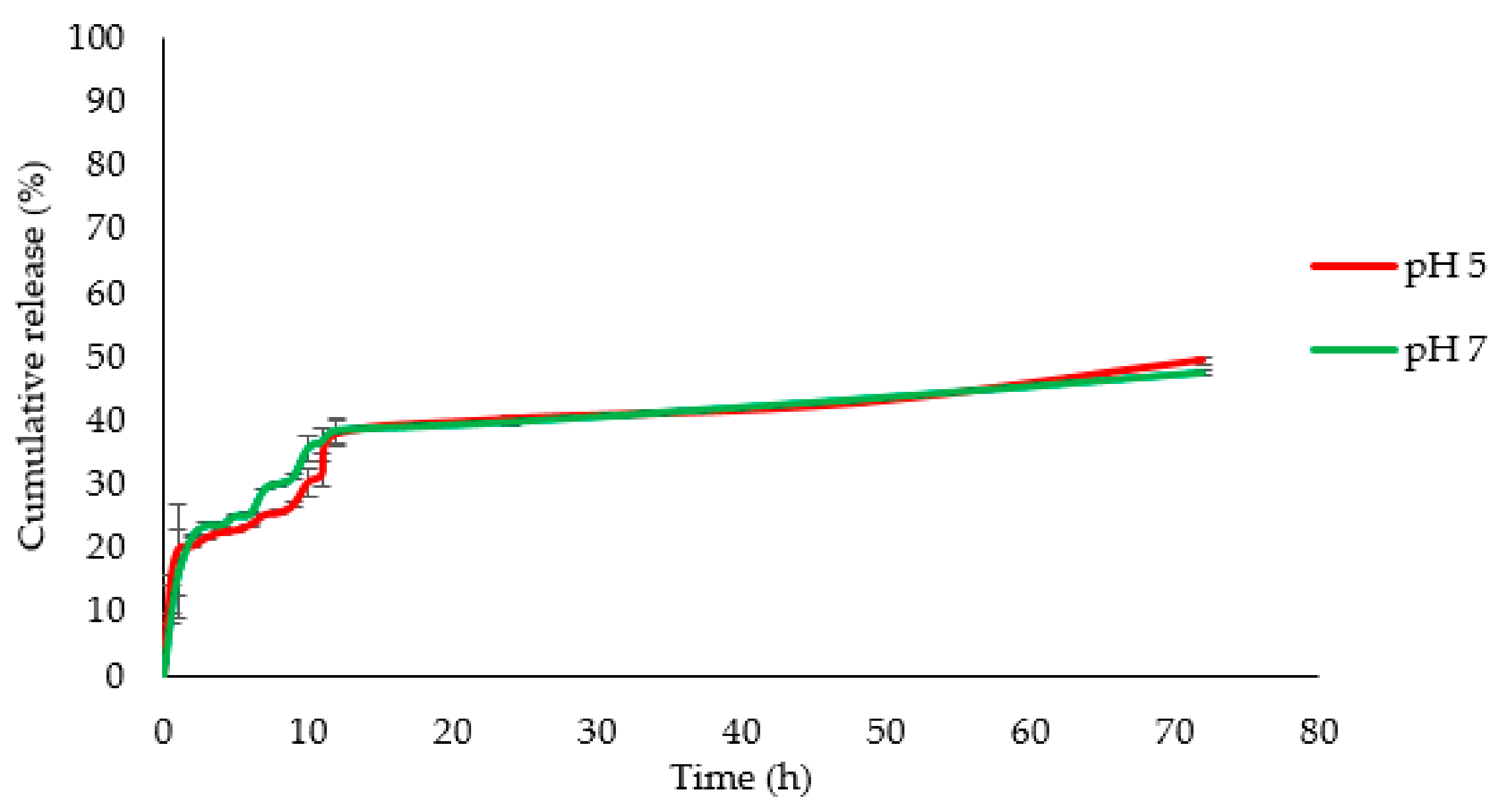
3.3. Drug Delivery
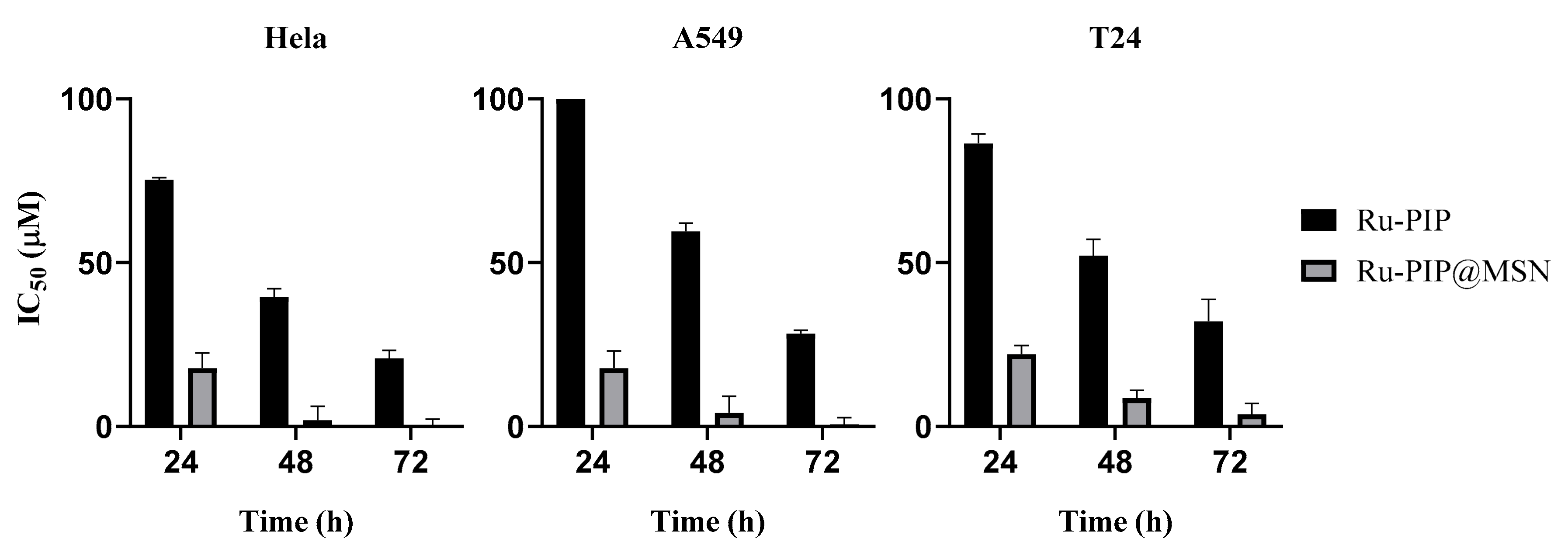
3.4. Cytotoxicity of Ru-PIP and Ru-PIP@MSNs
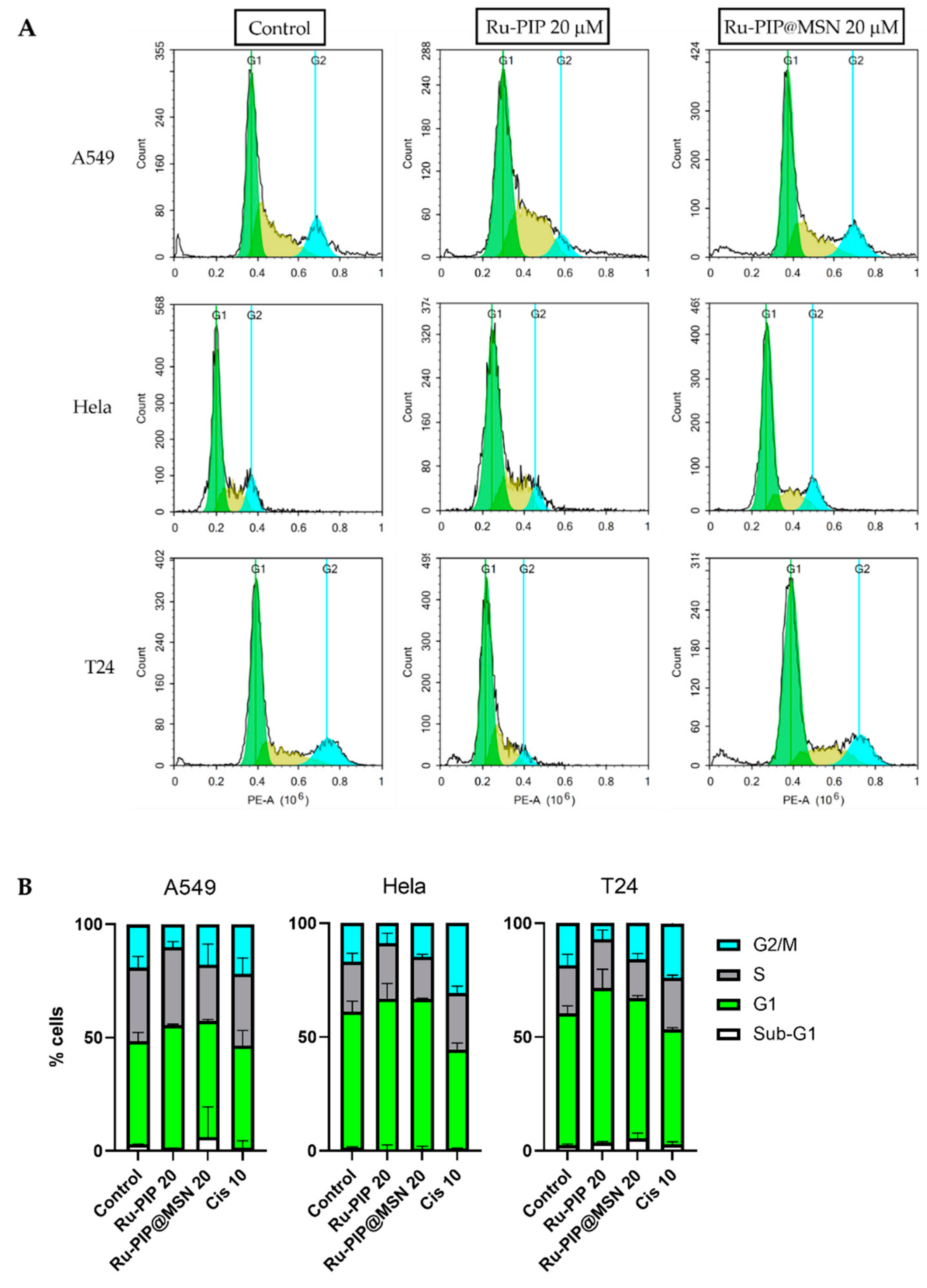
3.5. Cell Cycle Distribution
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Clarke, M.J. Ruthenium Chemistry Pertaining to the Design of Anticancer Agents. Prog. Clin. Biochem. Med. 1989, 10, 25–39. [Google Scholar]
- Notoro, A.; Gasser, G. Monomeric and dimeric coordinatively saturated and substitutionally inert Ru(ii) polypyridyl complexes as anticancer drug candidates. Chem. Soc. Rev. 2017, 46, 7317–7337. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Gupta, P.; Chen, Y.; Wang, E.; Ji, L.; Chao, H.; Chen, Z.-S. The development of anticancer ruthenium(ii) complexes: From single molecule compounds to nanomaterials. Chem. Soc. Rev. 2017, 46, 5771–5804. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Huang, Y.; Zhu, H.; Pang, G.; Zheng, W.; Wong, Y.-S.; Chen, T. Cancer-targeted monodisperse mesoporous silica nanoparticles as carrier of ruthenium polypyridyl complexes to enhance theranostic effects. Adv. Funct. Mater. 2014, 24, 2754–2763. [Google Scholar] [CrossRef]
- Lainé, A.L.; Passirani, C. Novel metal-based anticancer drugs: A new challenge in drug delivery. Curr. Opin. Pharmacol. 2012, 12, 420–426. [Google Scholar] [CrossRef] [PubMed]
- Peppas, L.B. Recent advances on the use of biodegradable microparticles and nanoparticles in controlled drug-delivery. Int. J. Pharm. 1995, 116, 1–9. [Google Scholar] [CrossRef]
- Klichko, Y.; Liong, M.; Choi, E.; Angelos, S.; Nel, A.; Stoddart, F.; Tamano, F.; Zink, J. Mesostructured Silica for Optical Functionality, Nanomachines, and Drug Delivery. J. Am. Ceram. Soc. 2014, 8, 21–27. [Google Scholar] [CrossRef]
- Sun, X. Mesoporous Silica Nanoparticles for Applications in Drug Delivery and Catalysis. Ph.D. Thesis, Iowa State University, Ames, IA, USA, 2012. [Google Scholar]
- Tarn, D.; Ashley, C.E.; Xue, M.; Carner, E.C.; Zink, J.I.; Brinker, C.J. Mesoporous Silica Nanoparticle Nanocarriers—Biofunctionality and Biocompatibility. Acc. Chem. Res. 2013, 46, 792–801. [Google Scholar] [CrossRef]
- Liu, Q.; Xu, N.; Liu, L.; Zhang, Y.; Shen, C.; Shezad, K.; Zhang, L.; Zhu, J.; Tao, J. Dacarbazine-Loaded Hollow Mesoporous Silica Nanoparticles Grafted with Folic Acid for Enhancing Antimetastatic Melanoma Response. ACS Appl. Mater. Interfaces 2017, 9, 21673–21687. [Google Scholar] [CrossRef]
- Moreira, A.F.; Dias, D.R.; Correia, I.J. Stimuli-responsive mesoporous silica nanoparticles for cancer therapy: A review. Microporous Mesoporous Mater. 2016, 236, 141–157. [Google Scholar] [CrossRef]
- Shchipunov, Y.A.; Burtseva, Y.V.; Karpenko, T.Y.; Shevchenko, N.M.; Zvyagintseva, T.N. Highly efficient immobilization of endo-1,3-β-d-glucanases (laminarinases) from marine mollusks in novel hybrid polysaccharide-silica nanocomposites with regulated composition. J. Mol. Catal. B Enzym. 2006, 40, 16–23. [Google Scholar] [CrossRef]
- Trewyn, B.G.; Slowing, I.I.; Chen, H.; Lin, V.S. Synthesis and Functionalization of a Mesoporous Silica Nanoparticle Based on the Sol–Gel Process and Applications in Controlled Release. Acc. Chem. Res. 2007, 40, 846–853. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, Q.; Han, N.; Ling, B.; Li, J.; Liu, J.; Che, E.; Hu, L.; Zhang, Q.; Hiang, T.; et al. Mesoporous silica nanoparticles in drug delivery and biomedical applications. Nanomed. Nanotechnol. Biol. Med. 2015, 11, 313–327. [Google Scholar] [CrossRef] [PubMed]
- He, Q.; Shi, J.; Zhu, M.; Chen, Y.; Chen, F. The three-stage in vitro degradation behavior of mesoporous silica in simulated body fluid. Microporous Mesoporous Mater. 2010, 131, 314–320. [Google Scholar] [CrossRef]
- Hudson, S.P.; Padera, R.F.; Langer, R.; Kohane, D.S. The biocompatibility of mesoporous silicates. Biomaterials 2008, 29, 4045–4055. [Google Scholar] [CrossRef]
- Castillo, R.R.; Colilla, M.; Vallet-Regí, M. Advances in mesoporous silica-based nanocarriers for co-delivery and combination therapy against cancer. Expert Opin. Drug Deliv. 2017, 14, 229–243. [Google Scholar] [CrossRef]
- Alothman, Z.A. A Review: Fundamental Aspects of Silicate Mesoporous Materials. Materials 2012, 5, 2874–2902. [Google Scholar] [CrossRef]
- Yamada, H.; Urata, C.; Ujiie, H.; Yamauchi, Y.; Kuroda, K. Preparation of aqueous colloidal mesostructured and mesoporous silica nanoparticles with controlled particle size in a very wide range from 20 nm to 700 nm. Nanoscale 2013, 5, 6145. [Google Scholar] [CrossRef]
- Chen, Q.; Han, L.; Gao, C.; Che, S. Synthesis of monodispersed mesoporous silica spheres (MMSSs) with controlled particle size using gemini surfactant. Microporous Mesoporous Mater. 2010, 128, 203–212. [Google Scholar] [CrossRef]
- Kobler, J.; Möller, K.; Bein, T. Colloidal Suspensions of Functionalized Mesoporous Silica Nanoparticles. ACS Nano 2008, 2, 791–799. [Google Scholar] [CrossRef]
- Archariyapanyakul, P.; Pangkumhang, B.; Khamdahsag, P.; Tanboonchuy, V. Synthesis of Silica-supported Nanoiron for Cr (VI) Removal: Application of Box-Behnken Statistical Design (BBD). Sains Malays. 2017, 46, 655–665. [Google Scholar] [CrossRef]
- Mei, X.; Liu, R.; Shen, F.; Wu, H. Optimization of Fermentation Conditions for the Production of Ethanol from Stalk Juice of Sweet Sorghum by Immobilized Yeast Using Response Surface Methodology. Energy Fuels 2009, 23, 487–491. [Google Scholar] [CrossRef]
- Bala, N.; Napiah, M.; Kamaruddin, I.; Danlami, N. Optimization of Nanocomposite Modified Asphalt Mixtures Fatigue Life using Response Surface Methodology Optimization of Nanocomposite Modified Asphalt Mixtures Fatigue Life using Response Surface Methodology. IOP Conf. Ser. Earth Environ. Sci. 2018, 140, 012064. [Google Scholar] [CrossRef]
- Gill, M.R.; Harun, S.N.; Halder, S.; Boghozian, R.A.; Ramdan, K.; Ahmad, H.; Vallis, K.A. A ruthenium polypyridyl intercalator stalls DNA replication forks, radiosensitizes human cancer cells and is enhanced by Chk1 inhibition. Sci. Rep. 2016, 6, 31973. [Google Scholar] [CrossRef] [PubMed]
- Yusoh, N.A.; Leong, S.W.; Chia, S.L.; Harun, S.N.; Rahman, M.B.A.; Vallis, K.A.; Gill, M.R.; Ahmad, H. Metallointercalator [Ru(dppz)2(PIP)]2+ Renders BRCA Wild-Type Triple-Negative Breast Cancer Cells Hypersensitive to PARP Inhibition. ACS Chem. Biol. 2020, 15, 378–387. [Google Scholar] [CrossRef] [PubMed]
- Dayana, E.; Isa, M.; Basyaruddin, M.; Rahman, A.; Ahmad, H. Monodispersed mesoporous silica nanospheres based on pyridinium ionic liquids. J. Porous Mater. 2018, 25, 1439–1446. [Google Scholar]
- Dayana, E.; Isa, M.; Ahmad, H.; Basyaruddin, M.; Rahman, A. Optimization of Synthesis Parameters of Mesoporous Silica Nanoparticles Based on Ionic Liquid by Experimental Design and its Application as a Drug Delivery Agent. J. Nanomater. 2019, 2019, 4982054. [Google Scholar]
- Marek, J.; Stodulka, P.; Cabal, J.; Soukup, O.; Pohanka, M.; Korabecny, J.; Musilek, K.; Kuca, K. Preparation of the Pyridinium Salts Differing in the Length of the N-Alkyl Substituent. Molecules 2010, 15, 1967–1972. [Google Scholar] [CrossRef]
- Lv, X.; Zhang, L.; Xing, F.; Lin, H. Controlled synthesis of monodispersed mesoporous silica nanoparticles: Particle size tuning and formation mechanism investigation. Microporous Mesoporous Mater. 2016, 225, 238–244. [Google Scholar] [CrossRef]
- Möller, K.; Kobler, J.; Bein, T. Colloidal suspensions of nanometer-sized mesoporous silica. Adv. Funct. Mater. 2007, 17, 605–612. [Google Scholar] [CrossRef]
- Innocenzi, P. From the Precursor to a Sol. In The Sol to Gel Transition; Springe International Publishing: Cham, Switzerland, 2016; pp. 7–25. [Google Scholar]
- Kruk, M.; Jaroniec, M. Determination of Pore Size and Pore Wall Structure of MCM-41 by Using Nitrogen Adsorption, Transmission Electron Microscopy, and X-ray Diffraction. J. Phys. Chem. B 2000, 104, 292–301. [Google Scholar] [CrossRef]
- Zhang, J.; Ma, Y.; Shi, F.; Liu, L.; Deng, Y. Room temperature ionic liquids as templates in the synthesis of mesoporous silica via a sol–gel method. Microporous Mesoporous Mater. 2009, 119, 97–103. [Google Scholar] [CrossRef]
- Joglekar, A.M.; May, A.T. Product Excellence through Design of Experiments. Cereal Foods World. 1987, 37, 857–868. [Google Scholar]
- Qiu, P.; Cui, M.; Kang, K.; Park, B.; Son, Y.; Khim, E.; Jang, M.; Khim, J. Application of Box-Behnken design with response surface methodology for modeling and optimizing ultrasonic oxidation of arsenite with H2O2. Cent. Eur. J. Chem. 2014, 12, 164–172. [Google Scholar] [CrossRef]
- Permanadewi, I.; Kumoro, A.C.; Wardhani, D.H.; Aryanti, N. Modelling of controlled drug release in gastrointestinal tract simulation. J. Phys. Conf. Ser. 2019, 1295, 012063. [Google Scholar] [CrossRef]
- Yamada, H.; Urata, C.; Higashitamori, S.; Aoyama, Y.; Yamauchi, Y.; Kuroda, K. Critical roles of cationic surfactants in the preparation of colloidal mesostructured silica nanoparticles: Control of mesostructure, particle size, and dispersion. ACS Appl. Mater. Interfaces 2014, 6, 3491–3500. [Google Scholar] [CrossRef]
- Yadav, G.; Bansal, M.; Thakur, N.; Khare, P. Multilayer Tablets and Their Drug Release Kinetic Models for Oral Controlled Drug Delivery System. Middle-East J. Sci. Res. 2013, 16, 782–795. [Google Scholar]
- Ma, B.; He, L.; You, Y.; Mo, J.; Chen, T. Controlled synthesis and size effects of multifunctional mesoporous silica nanosystem for precise cancer therapy. Drug Deliv. 2018, 25, 293–306. [Google Scholar] [CrossRef]
- Connor, M.J.O. Review Targeting the DNA Damage Response in Cancer. Mol. Cell. 2015, 60, 547–560. [Google Scholar]
- Giglia-Mari, G.; Zotter, A.; Vermeulen, W. DNA Damage Response. Cold Spring Harb. Perspect. Biol. 2011, 3, a000745. [Google Scholar] [CrossRef]
| Variables | Symbol | Levels | ||
|---|---|---|---|---|
| −1 | 0 | +1 | ||
| Mass of template (g) | A | 0.25 | 0.38 | 0.50 |
| Amount of TEA (g) | B | 0.06 | 0.08 | 1.00 |
| Temperature (°C) | C | 65 | 77.5 | 90 |
| Model | R2 | Adj-R2 | F-Value | p-Value |
|---|---|---|---|---|
| Linear | 0.8442 | 0.7923 | 16.250 | 0.0006 |
| 2FI | 0.9653 | 0.9307 | 6.990 | 0.0220 |
| Quadratic | 0.9670 | 0.8680 | 0.050 | 0.9825 |
| Run | Coded Variables | Real Variables | Response (m2g−1) | |||||
|---|---|---|---|---|---|---|---|---|
| A | B | C | A | B | C | Experimental | Predicted | |
| 1 | 0 | −1 | −1 | 0.38 | 0.06 | 65.00 | 753.61 | 783.556 |
| 2 | 0 | +1 | −1 | 0.38 | 0.10 | 65.00 | 833.99 | 809.008 |
| 3 | −1 | 0 | −1 | 0.25 | 0.08 | 65.00 | 756.34 | 767.387 |
| 4 | +1 | 0 | −1 | 0.50 | 0.08 | 65.00 | 833.71 | 825.177 |
| 5 | −1 | 0 | 0 | 0.38 | 0.08 | 77.50 | 647.62 | 658.101 |
| 6 | +1 | +1 | 0 | 0.50 | 0.10 | 77.50 | 676.82 | 707.715 |
| 7 | −1 | +1 | 0 | 0.25 | 0.10 | 77.50 | 682.14 | 693.455 |
| 8 | −1 | −1 | 0 | 0.25 | 0.06 | 77.50 | 732.19 | 688.577 |
| 9 | +1 | −1 | 0 | 0.50 | 0.06 | 77.50 | 566.69 | 542.657 |
| 10 | 0 | +1 | +1 | 0.38 | 0.10 | 90.00 | 623.18 | 592.161 |
| 11 | +1 | 0 | +1 | 0.50 | 0.08 | 90.00 | 427.69 | 425.195 |
| 12 | −1 | 0 | +1 | 0.25 | 0.08 | 90.00 | 597.56 | 614.645 |
| 13 | 0 | −1 | +1 | 0.38 | 0.06 | 90.00 | 423.77 | 447.678 |
| Terms | F-Value | p-Value | Characteristic |
|---|---|---|---|
| Model | 27.85 | 0.0004 | Significant |
| A (mass of template) | 7.20 | 0.0363 | Significant |
| B (amount of TEA) | 12.00 | 0.0134 | Significant |
| C (reaction temperature) | 126.95 | <0.0001 | Significant |
| AB | 5.33 | 0.0604 | Not significant |
| AC | 12.70 | 0.0119 | Significant |
| BC | 2.94 | 0.1370 | Not significant |
| Sample | Correlation Coefficient of Model (R) | ||||
|---|---|---|---|---|---|
| Zero-Order | First-Order | Higuchi | Hixson-Crowell | Korsmeyer-Peppas | |
| pH 5 | 0.878 | 0.876 | 0.878 | 0.880 | 0.946 |
| pH 7 | 0.927 | 0.952 | 0.927 | 0.946 | 0.842 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Harun, S.N.; Ahmad, H.; Lim, H.N.; Chia, S.L.; Gill, M.R. Synthesis and Optimization of Mesoporous Silica Nanoparticles for Ruthenium Polypyridyl Drug Delivery. Pharmaceutics 2021, 13, 150. https://doi.org/10.3390/pharmaceutics13020150
Harun SN, Ahmad H, Lim HN, Chia SL, Gill MR. Synthesis and Optimization of Mesoporous Silica Nanoparticles for Ruthenium Polypyridyl Drug Delivery. Pharmaceutics. 2021; 13(2):150. https://doi.org/10.3390/pharmaceutics13020150
Chicago/Turabian StyleHarun, Siti Norain, Haslina Ahmad, Hong Ngee Lim, Suet Lin Chia, and Martin R. Gill. 2021. "Synthesis and Optimization of Mesoporous Silica Nanoparticles for Ruthenium Polypyridyl Drug Delivery" Pharmaceutics 13, no. 2: 150. https://doi.org/10.3390/pharmaceutics13020150
APA StyleHarun, S. N., Ahmad, H., Lim, H. N., Chia, S. L., & Gill, M. R. (2021). Synthesis and Optimization of Mesoporous Silica Nanoparticles for Ruthenium Polypyridyl Drug Delivery. Pharmaceutics, 13(2), 150. https://doi.org/10.3390/pharmaceutics13020150





