Investigating the Contribution of Drug-Metabolizing Enzymes in Drug-Drug Interactions of Dapivirine and Miconazole
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals and Reagents
2.2. Ultra-Performance Liquid Chromatography Tandem Mass Spectrometry (LC-MS/MS)
2.3. Reaction Phenotyping Assay
2.4. Enzyme Inhibition Assay
2.5. Enzyme Induction Assay
2.6. Real Time RT-PCR
3. Results
3.1. Evaluation of DPV Metabolism in CYP and UGT Enzymes and the Impact of MIC
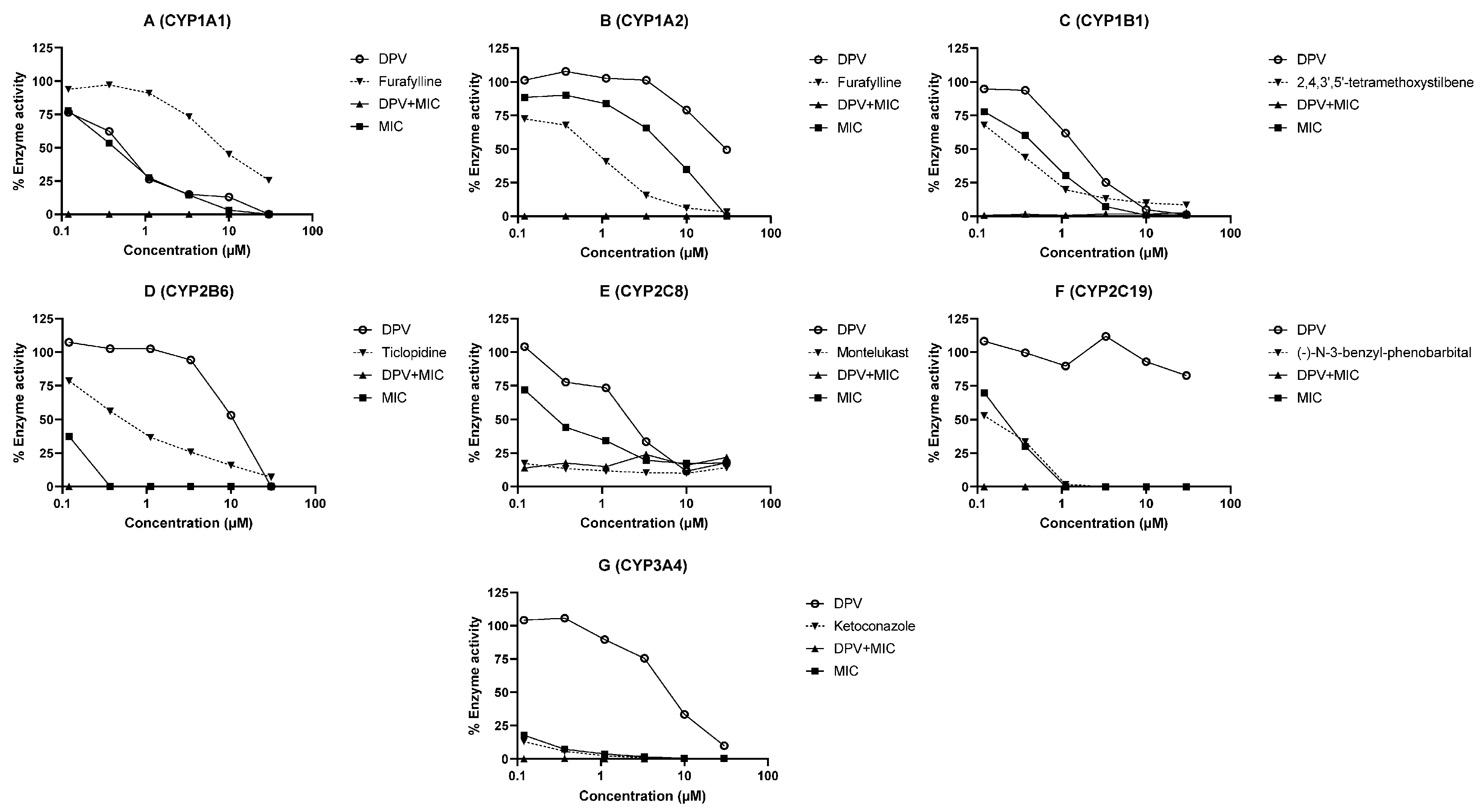
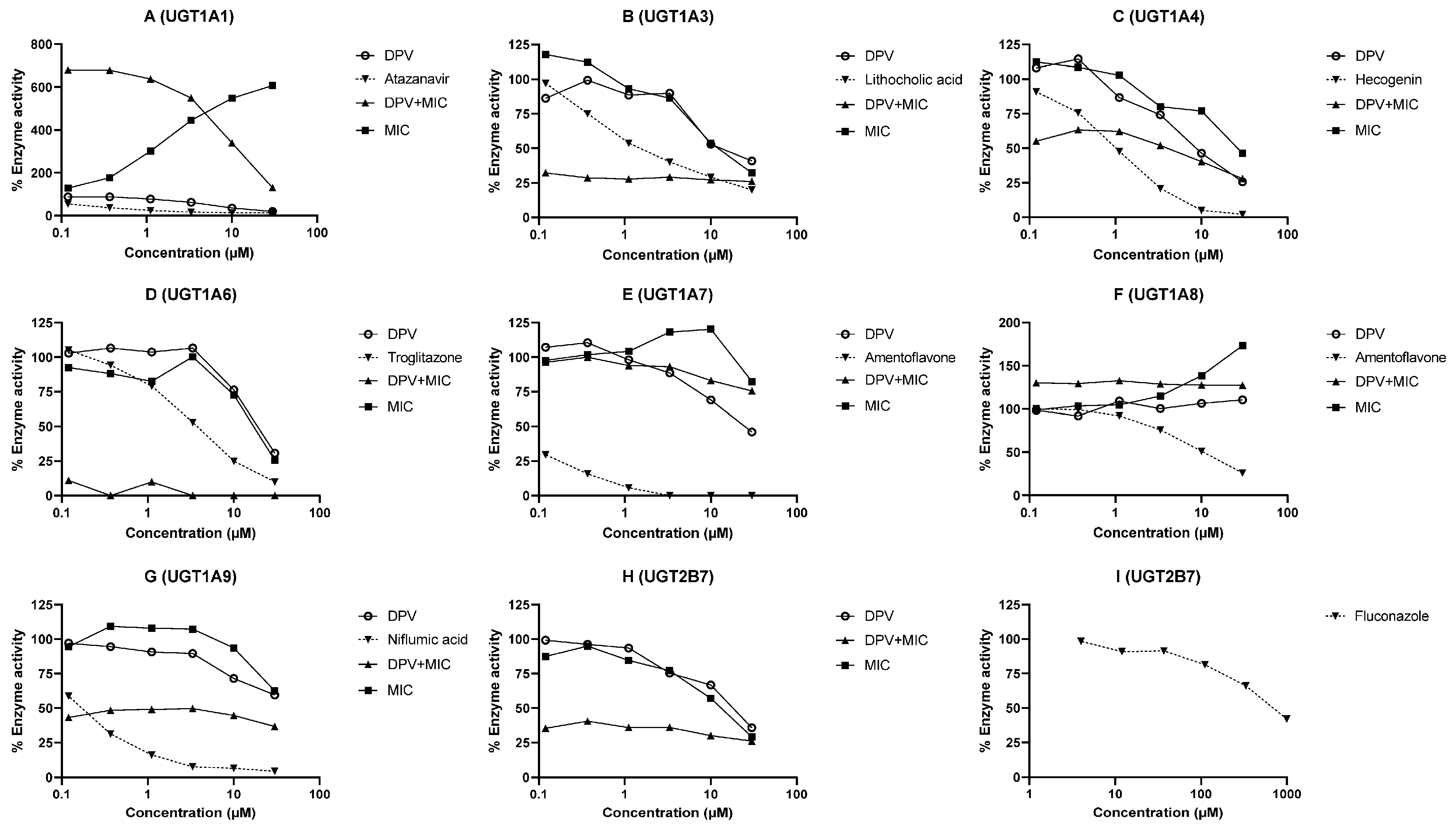
3.2. Evaluation of DPV as an Inhibitor of CYP and UGT Enzymes and the Impact of MIC
3.3. Evaluation of DPV as an Inducer of CYP Enzymes in the Hepatocytes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- UNAIDS. Global HIV & AIDS Statistics—2020 Fact Sheet. Available online: http://www.unaids.org/en/resources/fact-sheet (accessed on 1 April 2021).
- Goldstein, R.H.; Streed, C.G., Jr.; Cahill, S.R. Being PrEPared—Preexposure Prophylaxis and HIV Disparities. N. Engl. J. Med. 2018, 379, 1293–1295. [Google Scholar] [CrossRef] [PubMed]
- Ogbuagu, O.; Ruane, P.J.; Podzamczer, D.; Salazar, L.C.; Henry, K.; Asmuth, D.M.; Wohl, D.; Gilson, R.; Shao, Y.; Ebrahimi, R.; et al. Long-term safety and efficacy of emtricitabine and tenofovir alafenamide vs emtricitabine and tenofovir disoproxil fumarate for HIV-1 pre-exposure prophylaxis: Week 96 results from a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet HIV 2021, 8, e397–e407. [Google Scholar] [CrossRef]
- Nel, A.; van Niekerk, N.; Kapiga, S.; Bekker, L.G.; Gama, C.; Gill, K.; Kamali, A.; Kotze, P.; Louw, C.; Mabude, Z.; et al. Safety and Efficacy of a Dapivirine Vaginal Ring for HIV Prevention in Women. N. Engl. J. Med. 2016, 375, 2133–2143. [Google Scholar] [CrossRef] [PubMed]
- Baeten, J.M.; Palanee-Phillips, T.; Brown, E.R.; Schwartz, K.; Soto-Torres, L.E.; Govender, V.; Mgodi, N.M.; Matovu Kiweewa, F.; Nair, G.; Mhlanga, F.; et al. Use of a Vaginal Ring Containing Dapivirine for HIV-1 Prevention in Women. N. Engl. J. Med. 2016, 375, 2121–2132. [Google Scholar] [CrossRef] [PubMed]
- Nel, A.; van Niekerk, N.; Van Baelen, B.; Malherbe, M.; Mans, W.; Carter, A.; Steytler, J.; van der Ryst, E.; Craig, C.; Louw, C.; et al. Safety, adherence, and HIV-1 seroconversion among women using the dapivirine vaginal ring (DREAM): An open-label, extension study. Lancet HIV 2021, 8, e77–e86. [Google Scholar] [CrossRef]
- Baeten, J.M.; Palanee-Phillips, T.; Mgodi, N.M.; Mayo, A.J.; Szydlo, D.W.; Ramjee, G.; Gati Mirembe, B.; Mhlanga, F.; Hunidzarira, P.; Mansoor, L.E.; et al. Safety, uptake, and use of a dapivirine vaginal ring for HIV-1 prevention in African women (HOPE): An open-label, extension study. Lancet HIV 2021, 8, e87–e95. [Google Scholar] [CrossRef]
- EMA. Vaginal Ring to Reduce the Risk of HIV Infection for Women in Non-EU Countries with High Disease Burden. Available online: https://www.ema.europa.eu/en/news/vaginal-ring-reduce-risk-hiv-infection-women-non-eu-countries-high-disease-burden (accessed on 6 April 2021).
- WHO. WHO Recommends the Dapivirine Vaginal Ring as a New Choice for HIV Prevention for Women at Substantial Risk of HIV Infection. Available online: https://www.who.int/news/item/26-01-2021-who-recommends-the-dapivirine-vaginal-ring-as-a-new-choice-for-hiv-prevention-for-women-at-substantial-risk-of-hiv-infection (accessed on 6 April 2021).
- Nel, A.; Haazen, W.; Russell, M.; Nuttall, J.; Van Niekerk, N.; Treijtel, N. Drug-drug interactions between the Dapivirine Vaginal Ring (Ring-004) and miconazole nitrate vaginal capsule (Gyno-Daktarin®). AIDS Res. Hum. Retrovir. 2014, 30, A144. [Google Scholar] [CrossRef]
- Niwa, T.; Inoue-Yamamoto, S.; Shiraga, T.; Takagi, A. Effect of antifungal drugs on cytochrome P450 (CYP) 1A2, CYP2D6, and CYP2E1 activities in human liver microsomes. Biol. Pharm. Bull. 2005, 28, 1813–1816. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Niwa, T.; Shiraga, T.; Takagi, A. Effect of antifungal drugs on cytochrome P450 (CYP) 2C9, CYP2C19, and CYP3A4 activities in human liver microsomes. Biol. Pharm. Bull. 2005, 28, 1805–1808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Niwa, T.; Imagawa, Y.; Yamazaki, H. Drug interactions between nine antifungal agents and drugs metabolized by human cytochromes P450. Curr. Drug Metab. 2014, 15, 651–679. [Google Scholar] [CrossRef]
- Barnhart, K. Safety and efficacy of bedtime versus daytime administration of the miconazole nitrate 1200 mg vaginal ovule insert to treat vulvovaginal candidiasis. Curr. Med. Res. Opin. 2005, 21, 127–134. [Google Scholar] [CrossRef]
- Bourcier, K.; Hyland, R.; Kempshall, S.; Jones, R.; Maximilien, J.; Irvine, N.; Jones, B. Investigation into UDP-glucuronosyltransferase (UGT) enzyme kinetics of imidazole- and triazole-containing antifungal drugs in human liver microsomes and recombinant UGT enzymes. Drug Metab. Dispos. 2010, 38, 923–929. [Google Scholar] [CrossRef] [Green Version]
- Almazroo, O.A.; Miah, M.K.; Venkataramanan, R. Drug metabolism in the liver. Clin. Liver Dis. 2017, 21, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Hu, M.; Cost, M.; Poloyac, S.; Rohan, L. Short communication: Expression of transporters and metabolizing enzymes in the female lower genital tract: Implications for microbicide research. AIDS Res. Hum. Retrovir. 2013, 29, 1496–1503. [Google Scholar] [CrossRef] [PubMed]
- Hu, M.; Zhou, T.; Pearlman, A.P.; Paton, D.L.; Rohan, L.C. Comparative Expression Analysis of Cytochrome P450 1A1, Cytochrome P450 1B1 and Nuclear Receptors in the Female Genital and Colorectal Tissues of Human and Pigtailed Macaque. BAOJ Pharm. Sci. 2016, 2, 16. [Google Scholar]
- To, E.E.; Hendrix, C.W.; Bumpus, N.N. Dissimilarities in the metabolism of antiretroviral drugs used in HIV pre-exposure prophylaxis in colon and vagina tissues. Biochem. Pharm. 2013, 86, 979–990. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, M.; Valicherla, G.R.; Zhou, T.; Hillier, S.L.; Rohan, L.C. Expression, Activity, and Regulation of Phosphorylating Enzymes in Tissues and Cells Relevant to HIV-1 Sexual Transmission. AIDS Res. Hum. Retrovir. in print. 2021. [Google Scholar] [CrossRef] [PubMed]
- USFDA. In Vitro Drug Interaction Studies—Cytochrome P450 Enzyme- and Transporter-Mediated Drug Interactions Guidance for Industry; USFDA: Silver Spring, MD, USA, 2020.
- EMA. Guideline on the Investigation of Drug Interactions; EMA: Amsterdam, The Netherlands, 2012.
- USFDA. Bioanalytical Method Validation Guidance for Industry; USFDA: Silver Spring, MD, USA, 2018.
- Haupt, L.J.; Kazmi, F.; Ogilvie, B.W.; Buckley, D.B.; Smith, B.D.; Leatherman, S.; Paris, B.; Parkinson, O.; Parkinson, A. The Reliability of Estimating Ki Values for Direct, Reversible Inhibition of Cytochrome P450 Enzymes from Corresponding IC50 Values: A Retrospective Analysis of 343 Experiments. Drug Metab. Dispos. 2015, 43, 1744–1750. [Google Scholar] [CrossRef] [Green Version]
- Lv, X.; Zhang, J.B.; Wang, X.X.; Hu, W.Z.; Shi, Y.S.; Liu, S.W.; Hao, D.C.; Zhang, W.D.; Ge, G.B.; Hou, J.; et al. Amentoflavone is a potent broad-spectrum inhibitor of human UDP-glucuronosyltransferases. Chem. Biol. Interact. 2018, 284, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Joo, J.; Lee, B.; Lee, T.; Liu, K.H. Screening of six UGT enzyme activities in human liver microsomes using liquid chromatography/triple quadrupole mass spectrometry. Rapid Commun. Mass. Spectrom. 2014, 28, 2405–2414. [Google Scholar] [CrossRef] [PubMed]
- Valicherla, G.R.; Mishra, A.; Lenkalapelly, S.; Jillela, B.; Francis, F.M.; Rajagopalan, L.; Srivastava, P. Investigation of the inhibition of eight major human cytochrome P450 isozymes by a probe substrate cocktail in vitro with emphasis on CYP2E1. Xenobiotica 2019, 49, 1396–1402. [Google Scholar] [CrossRef] [PubMed]
- Walsky, R.L.; Obach, R.S.; Gaman, E.A.; Gleeson, J.P.; Proctor, W.R. Selective inhibition of human cytochrome P4502C8 by montelukast. Drug Metab. Dispos. 2005, 33, 413–418. [Google Scholar] [CrossRef] [PubMed]
- Farin, F.M.; Bigler, L.G.; Oda, D.; McDougall, J.K.; Omiecinski, C.J. Expression of cytochrome P450 and microsomal epoxide hydrolase in cervical and oral epithelial cells immortalized by human papillomavirus type 16 E6/E7 genes. Carcinogenesis 1995, 16, 1391–1401. [Google Scholar] [CrossRef]
- Patel, K.R.; Astley, S.; Adams, D.J.; Lacey, C.J.; Ali, S.W.; Wells, M. Expression of cytochrome P450 enzymes in the cervix. An immunohistochemical study. Int. J. Gynecol. Cancer 1993, 3, 159–163. [Google Scholar] [CrossRef] [PubMed]
- Verhoeven, C.H.; van den Heuvel, M.W.; Mulders, T.M.; Dieben, T.O. The contraceptive vaginal ring, NuvaRing, and antimycotic co-medication. Contraception 2004, 69, 129–132. [Google Scholar] [CrossRef] [PubMed]
- Simmons, K.B.; Kumar, N.; Plagianos, M.; Roberts, K.; Hoskin, E.; Han, L.; Alami, M.; Creasy, G.; Variano, B.; Merkatz, R. Effects of concurrent vaginal miconazole treatment on the absorption and exposure of Nestorone(R) (segesterone acetate) and ethinyl estradiol delivered from a contraceptive vaginal ring: A randomized, crossover drug-drug interaction study. Contraception 2018, 97, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.A.; Panther, L.; Marzinke, M.A.; Hendrix, C.W.; Hoesley, C.J.; van der Straten, A.; Husnik, M.J.; Soto-Torres, L.; Nel, A.; Johnson, S.; et al. Phase 1 Safety, Pharmacokinetics, and Pharmacodynamics of Dapivirine and Maraviroc Vaginal Rings: A Double-Blind Randomized Trial. J. Acquir. Immune Defic. Syndr. 2015, 70, 242–249. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Achilles, S.; Kelly, C.; Blithe, D.; Long, J.; Richardson, B.; Devlin, B.; Hendrix, C.; Poloyac, S.; Marzinke, M.; Singh, D. Pharmacokinetics, safety, and vaginal bleeding associated with continuous versus cyclic 90-day use of dapivirine and levonorgestrel vaginal rings for multipurpose prevention of HIV and pregnancy. J. Int. AIDS Soc. 2021, 24, 15–17. [Google Scholar]


| Enzyme | % DPV Remaining at 60 min | Known Substrate (% Remaining at 60 min) | |
|---|---|---|---|
| without MIC | with MIC | ||
| rCYP1A1 | 40.38 | 83.38 | Phenacetin (40.98) |
| rCYP1A2 | 94.10 | - | Phenacetin (65.76) |
| rCYP1B1 | 97.14 | - | 7-ethoxyresorufin (35.11) |
| rCYP2B6 | 96.46 | - | Bupropion (69.59) |
| rCYP2C8 | 100.99 | - | Amodiaquine (0.92) |
| rCYP2C19 | 96.03 | - | Mephenytoin (61.67) |
| rCYP3A4 | 73.73 | 96.24 | Midazolam (4.02) |
| HLM (Phase 1) | 37.14 | 88.39 | Midazolam (0.00) |
| rUGT1A1 | 98.04 | - | SN38 (74.87) |
| rUGT1A3 | 109.17 | - | Chenodeoxycholic acid (59.44) |
| rUGT1A4 | 95.95 | - | Trifluoperazine (73.10) |
| rUGT1A6 | 100.48 | - | 4-methylumbelliferone (5.26) |
| rUGT1A7 | 107.08 | - | 4-methylumbelliferone (70.94) |
| rUGT1A8 | 99.20 | - | 4-methylumbelliferone (49.90) |
| rUGT1A9 | 104.45 | - | Mycophenolic acid (0.00) |
| rUGT2B7 | 102.37 | - | Zidovudine (74.41) |
| HLM (Phase 2) | 97.15 | 95.59 | 4-methylumbelliferone (0.82) |
| Enzyme | IC50 (μM) | Cmax/Ki | ||||
|---|---|---|---|---|---|---|
| DPV | MIC | DPV and MIC Combination | Known Inhibitor | DPV | MIC | |
| CYP1A1 | 0.50 | 0.43 | <0.12 | Furafylline (4.76) | <0.01 | 0.08 |
| CYP1A2 | 10.53 | 5.11 | <0.12 | Furafylline (0.58) | <0.01 | <0.01 |
| CYP1B1 | 1.55 | 0.47 | <0.12 | 2,4,3′,5′-tetramethoxystilbene (0.22) | <0.01 | 0.07 |
| CYP2B6 | 10.07 | 0.12 | <0.12 | Ticlopidine (0.47) | <0.01 | 0.30 |
| CYP2C8 | 1.41 | 0.21 | <0.12 | Montelukast (0.0017) | <0.01 | 0.17 |
| CYP2C19 | >30 | 0.21 | <0.12 | (-)-N-3-benzyl-phenobarbital (0.148) | <0.01 | 0.17 |
| CYP3A4 | 4.90 | 0.02 | <0.12 | Ketoconazole (0.012) | <0.01 | 1.82 |
| UGT1A1 | 5.20 | Apparent activation with MIC | Atazanavir (0.11) | <0.01 | N.A. | |
| UGT1A3 | 16.98 | 13.64 | <0.12 | Lithocholic acid (0.94) | <0.01 | <0.01 |
| UGT1A4 | 9.45 | 26.46 | 2.05 | Hecogenin (0.96) | <0.01 | <0.01 |
| UGT1A6 | 19.59 | 17.07 | <0.12 | Troglitazone (3.77) | <0.01 | <0.01 |
| UGT1A7 | 24.15 | >30 | >30 | Amentoflavone (0.0461) | <0.01 | <0.01 |
| UGT1A8 | >30 | Apparent activation with MIC | Amentoflavone (5.57) | <0.01 | N.A. | |
| UGT1A9 | >30 | >30 | <0.12 | Niflumic acid (0.15) | <0.01 | <0.01 |
| UGT2B7 | 17.22 | 12.32 | <0.12 | Fluconazole (712.3) | <0.01 | <0.01 |
| Enzyme | Test Compound | Enzyme Activity in Fold Induction | ||
|---|---|---|---|---|
| HU2036 | HU8373 | HU2068 | ||
| CYP1A2 | DPV | 0.89 ± 0.09 | 1.50 a | 0.87 ± 0.26 |
| Omeprazole | 14.02 ± 2.43 | 32.70 ± 6.67 | 35.54 ± 2.03 | |
| CYP2B6 | DPV | 0.81 ± 0.07 | 0.99 ± 0.10 | 0.87 ± 0.07 |
| Phenobarbital | 7.63 ±0.82 | 10.74 ± 0.61 | 6.43 ± 1.02 | |
| CYP3A4 | DPV | 0.81 ± 0.15 | 0.74 ± 0.07 | 0.93 ± 0.14 |
| Rifampicin | 16.93 ± 0.42 | 41.16 ± 2.43 | 8.70 ± 0.83 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Valicherla, G.R.; Graebing, P.; Zhang, J.; Zheng, R.; Nuttall, J.; Silvera, P.; Rohan, L.C. Investigating the Contribution of Drug-Metabolizing Enzymes in Drug-Drug Interactions of Dapivirine and Miconazole. Pharmaceutics 2021, 13, 2193. https://doi.org/10.3390/pharmaceutics13122193
Valicherla GR, Graebing P, Zhang J, Zheng R, Nuttall J, Silvera P, Rohan LC. Investigating the Contribution of Drug-Metabolizing Enzymes in Drug-Drug Interactions of Dapivirine and Miconazole. Pharmaceutics. 2021; 13(12):2193. https://doi.org/10.3390/pharmaceutics13122193
Chicago/Turabian StyleValicherla, Guru Raghavendra, Phillip Graebing, Junmei Zhang, Ruohui Zheng, Jeremy Nuttall, Peter Silvera, and Lisa Cencia Rohan. 2021. "Investigating the Contribution of Drug-Metabolizing Enzymes in Drug-Drug Interactions of Dapivirine and Miconazole" Pharmaceutics 13, no. 12: 2193. https://doi.org/10.3390/pharmaceutics13122193
APA StyleValicherla, G. R., Graebing, P., Zhang, J., Zheng, R., Nuttall, J., Silvera, P., & Rohan, L. C. (2021). Investigating the Contribution of Drug-Metabolizing Enzymes in Drug-Drug Interactions of Dapivirine and Miconazole. Pharmaceutics, 13(12), 2193. https://doi.org/10.3390/pharmaceutics13122193







