Lipophilic Peptide Dendrimers for Delivery of Splice-Switching Oligonucleotides
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Cell and Culture Conditions
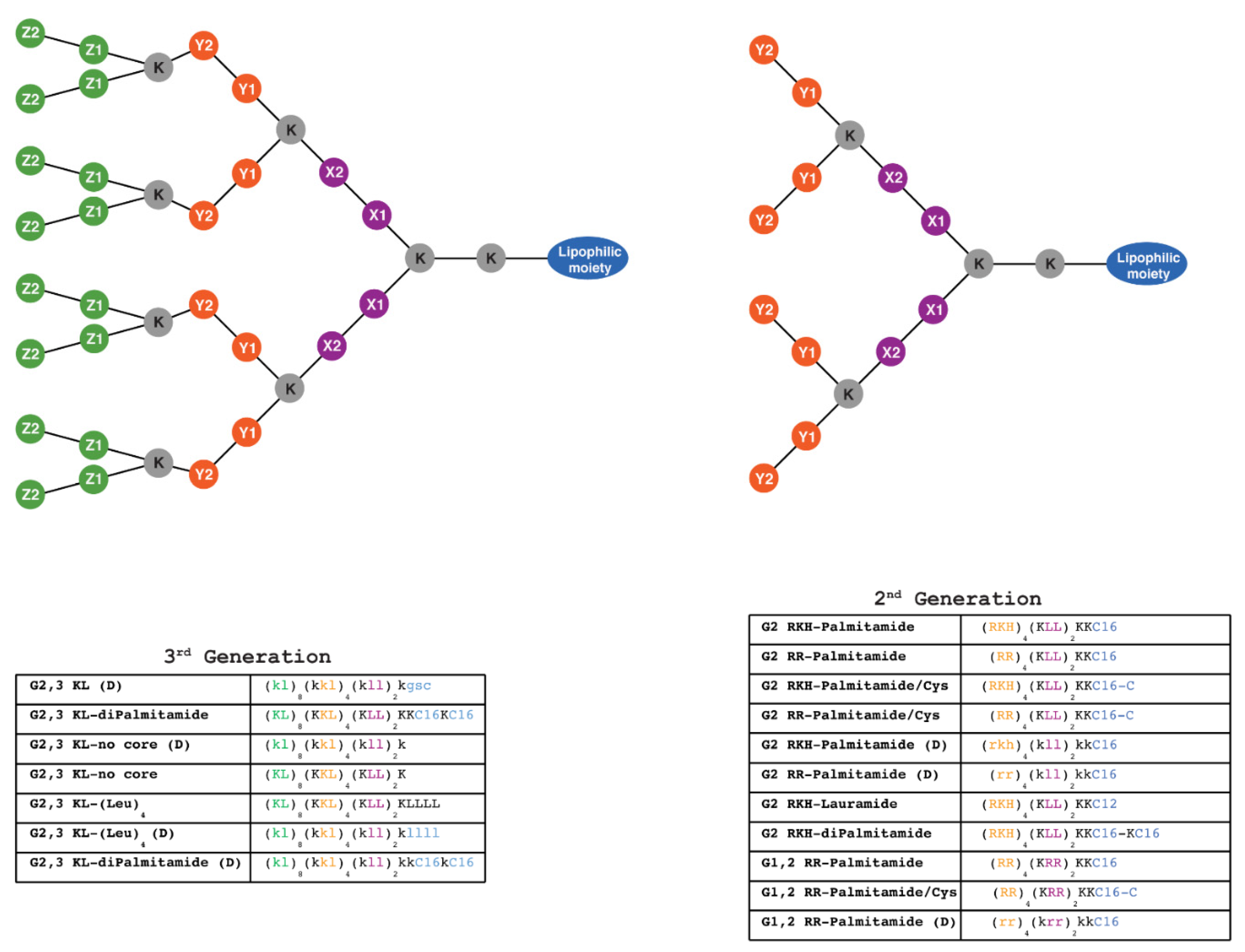
2.3. Synthesis and Characterization of Peptide Dendrimers
2.4. ON Transfection of Reporter Cells in Serum-Containing Media
2.5. Luciferase Assay
2.6. Live Cell Imaging by Fluorescence Microscopy
2.7. Live Cell Imaging by Confocal Microscopy
2.8. Particle Size Measurement
2.9. Cell Viability
2.10. Data Analysis
3. Results and Discussion
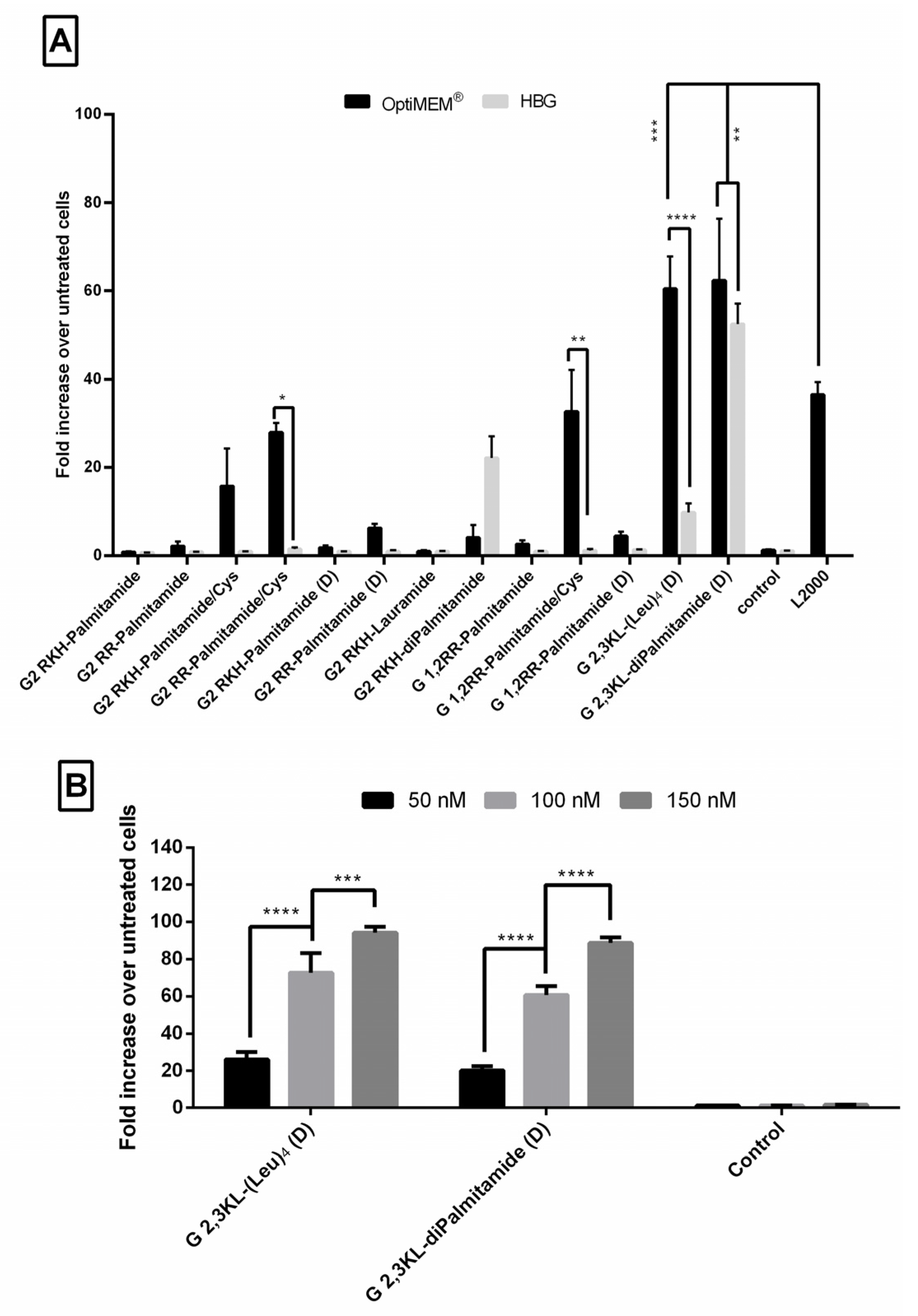
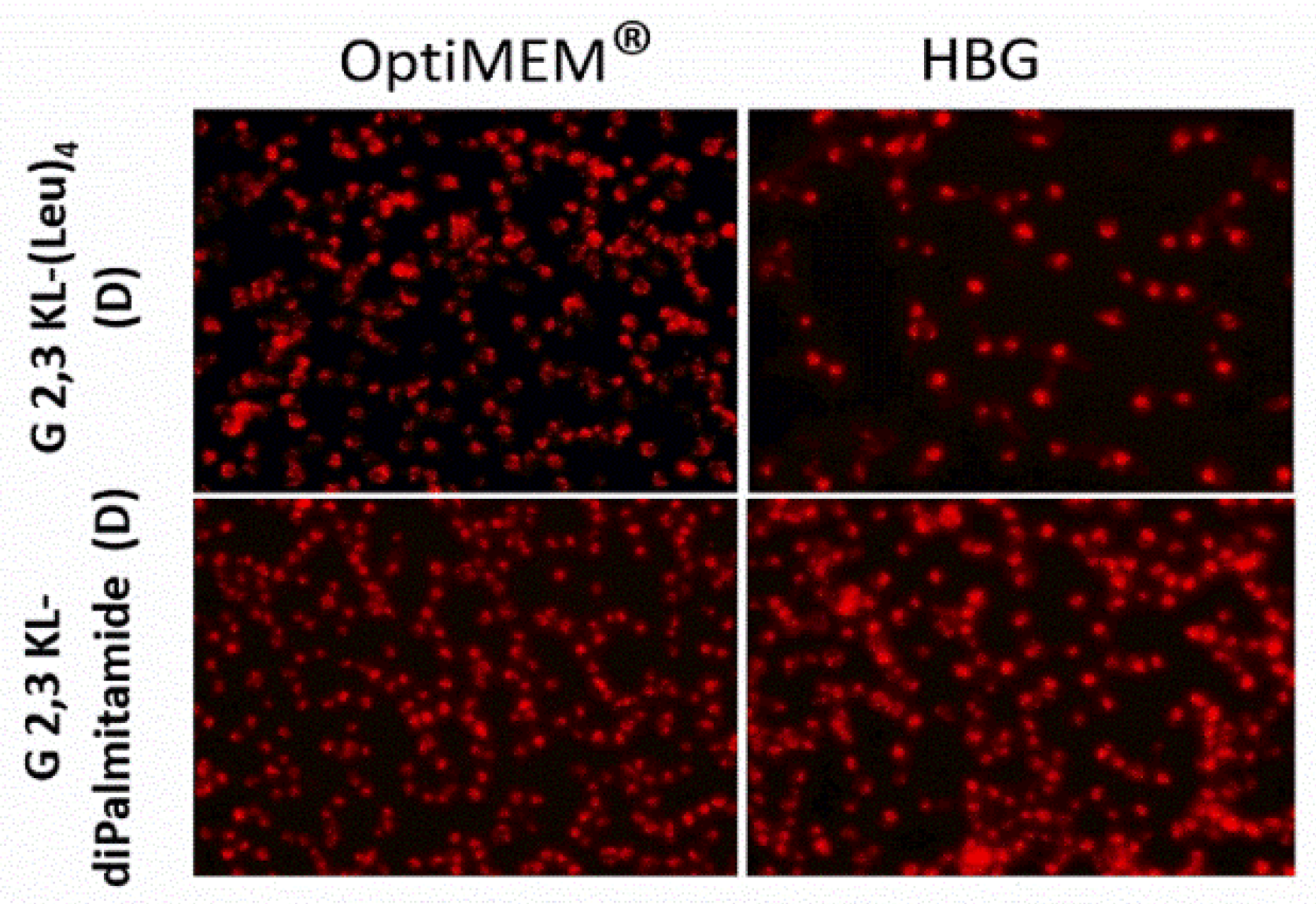
3.1. Lipophilic Peptide Dendrimers Surpass Lipofectamine 2000 (L2000) in Transfecting Reporter Cells in Serum-Containing Media
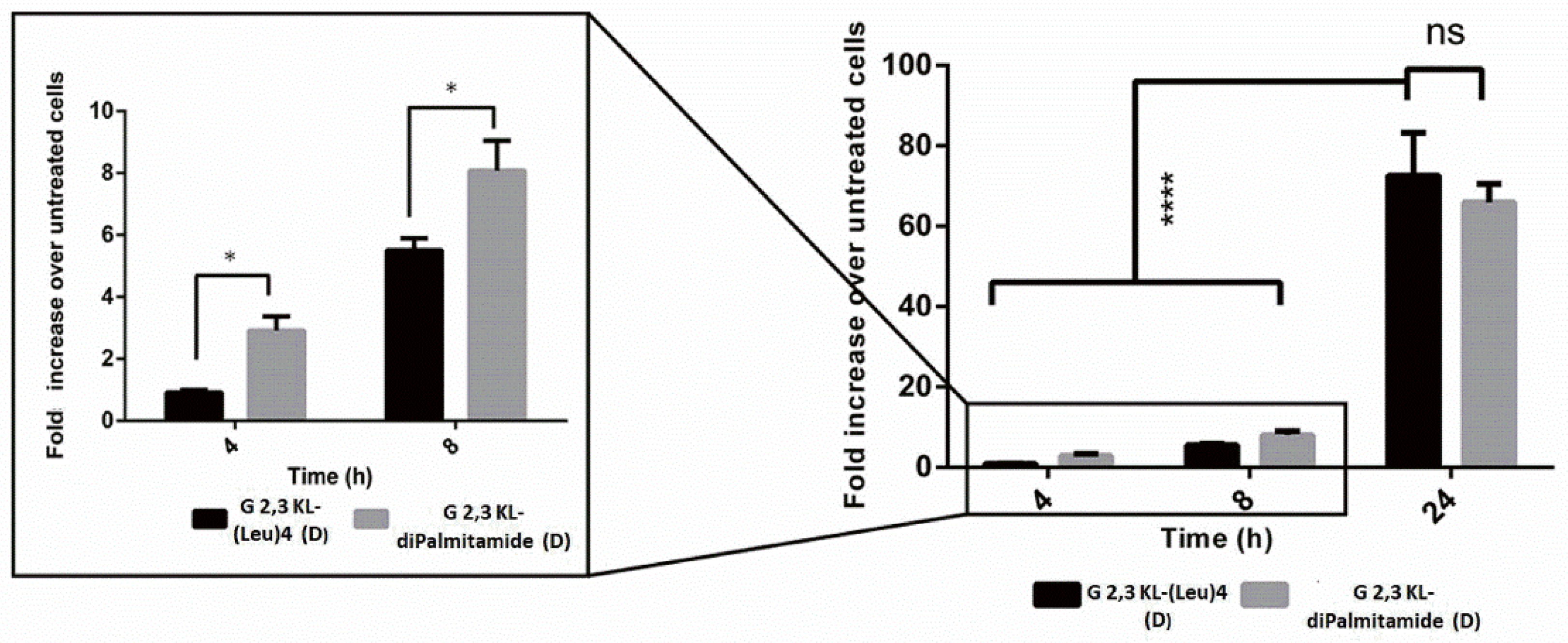
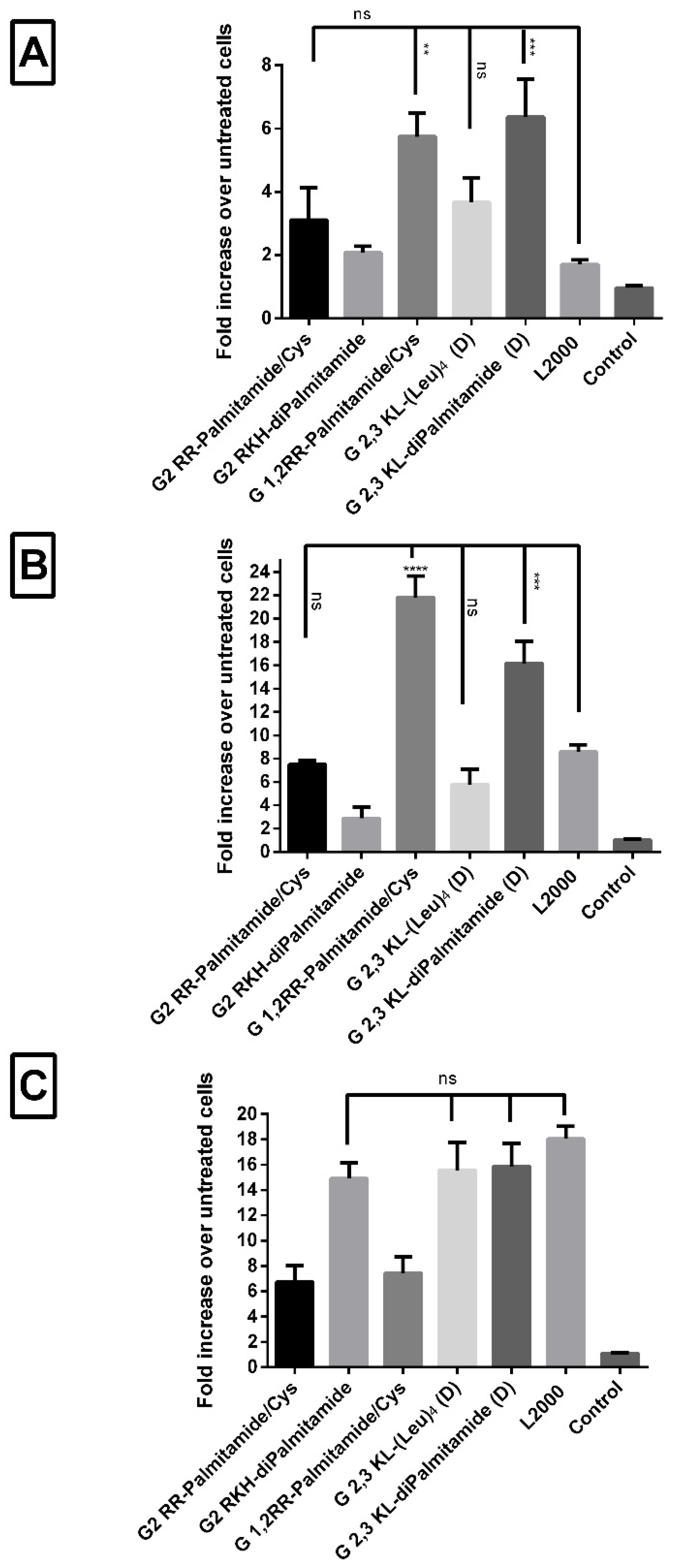
3.2. Cellular Uptake of Dendrimers/ON Complexes and Their Activity at Different Time-Points
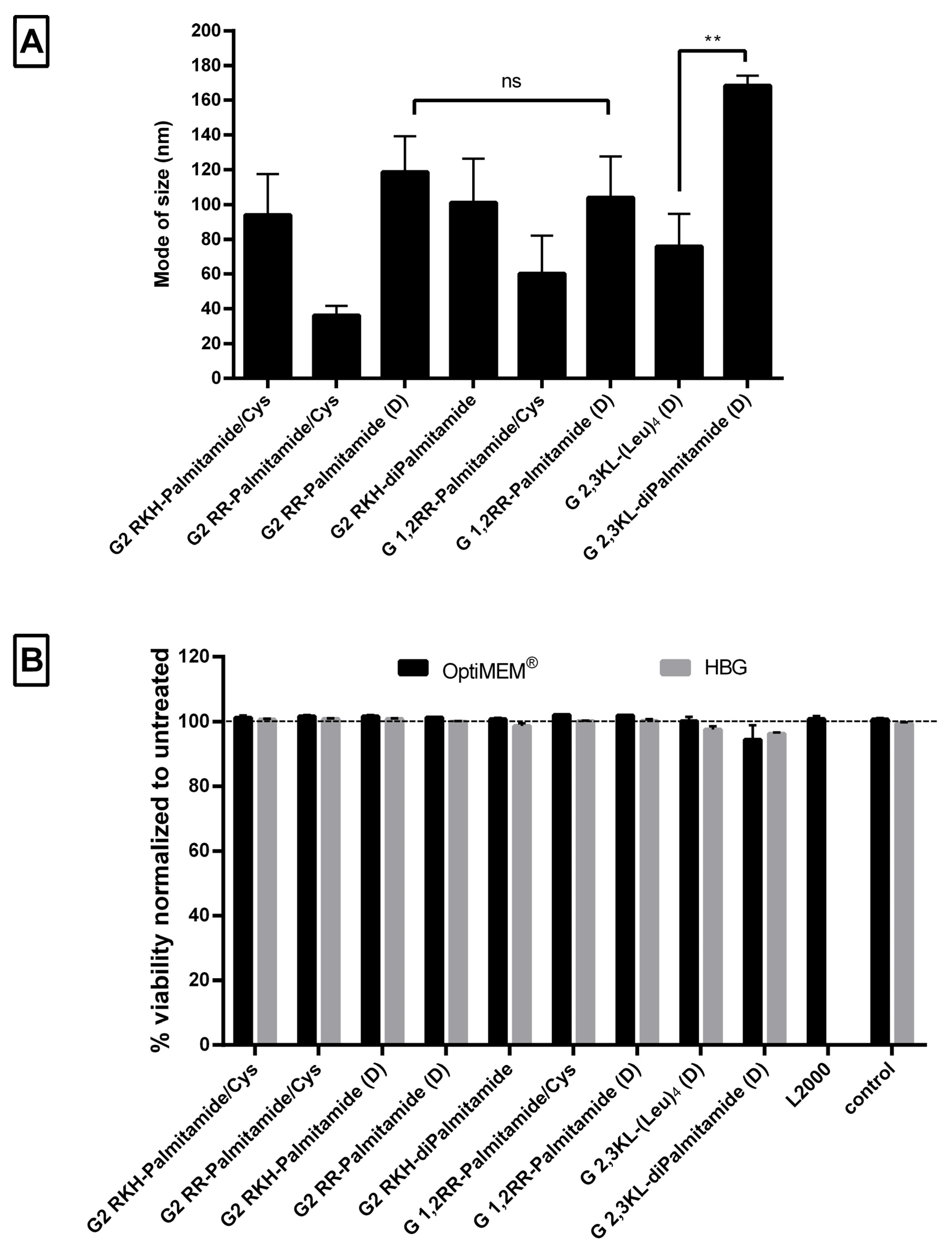
3.3. Physical Characterization and Viability of Lipophilic Peptide Dendrimers
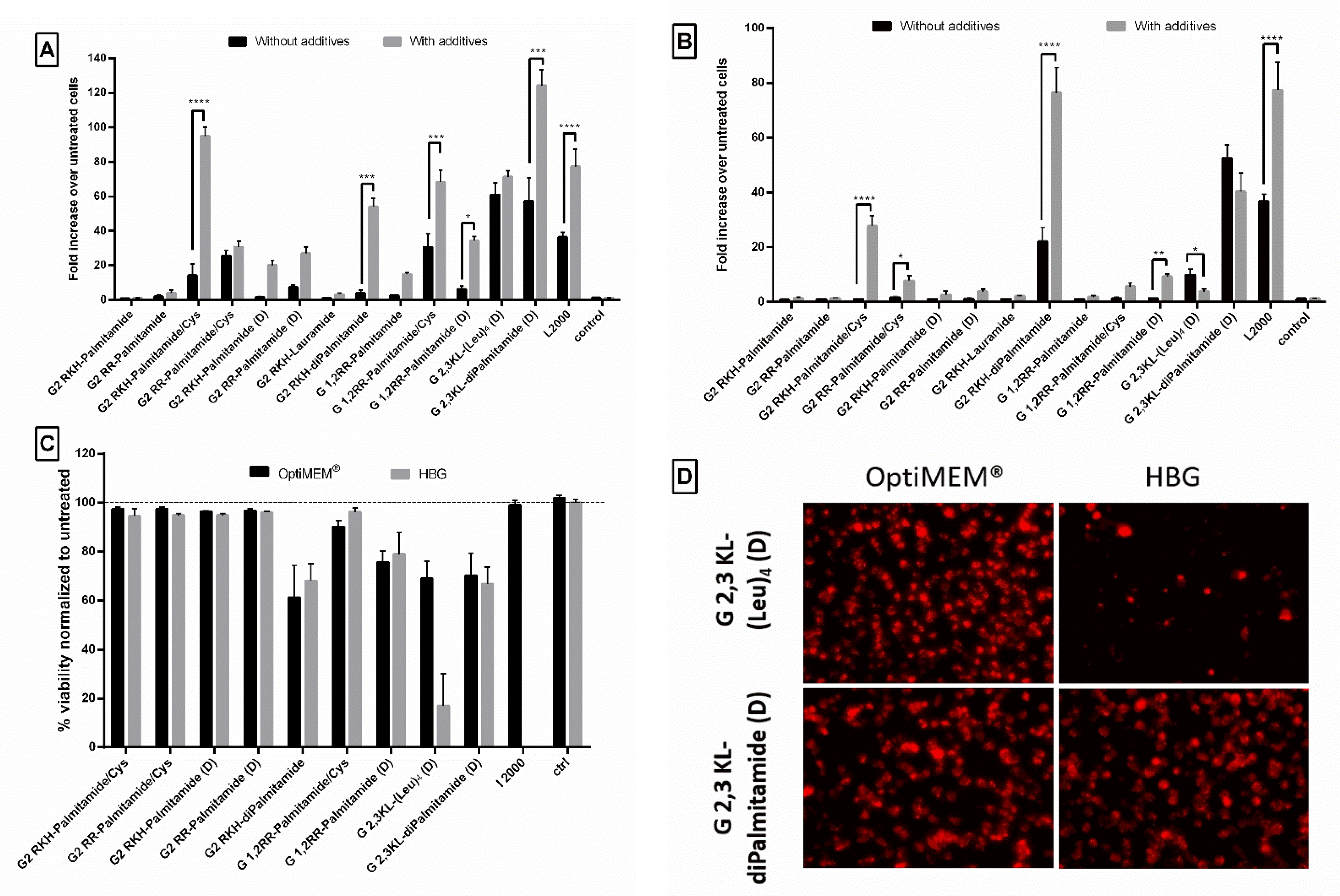
3.4. Lipophilic Peptide Dendrimers Are Similar or Surpass the Transfection Efficiency of Lipofectamine (L2000) in Various Reporter Cells
3.5. The Polymeric Excipient Polyvinylalcohol 18 Enhances Transfection Efficiency of Complexes in HeLa Luc/705 Reporter Cells in Serum-Containing Media
3.6. Intracellular Distribution by Confocal Microscopy
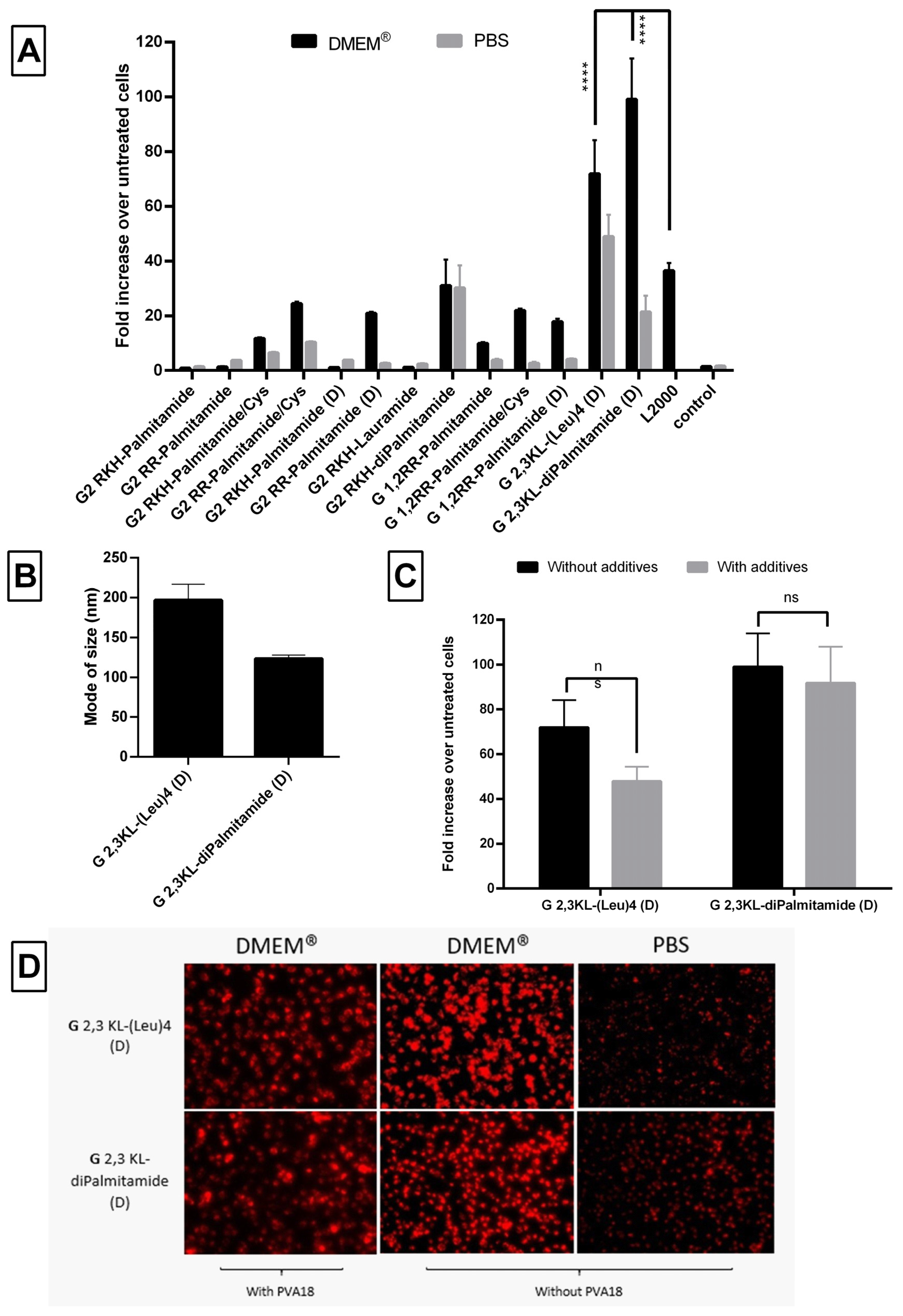
3.7. Effect of Different Buffers on Transfection Efficiency of the Lipophilic Peptide Dendrimers
4. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ANOVA | Analysis of Variance |
| CO2 | Carbon Dioxide |
| DAPI | 4′,6-diamidino-2-phenylindole |
| DMEM | Dulbecco’s modified Eagle’s medium |
| DNA | Deoxyribonucleic Acid |
| DOPE | Dioleoyl-phosphatidylethanolamine |
| DOTMA | 1,2-di-O-octadecenyl-3-trimethylammonium propane |
| FBS | Fetal bovine serum |
| HBG | HEPES Buffered Glucose |
| HEPES | 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid |
| HPLC | High Performance Liquid Chromatography |
| L2000 | Lipofectamine 2000 |
| LSD | Least Significant Difference |
| N/P ratio | Nitrogen/Phosphate ratio |
| NTA | Nanoparticles Tracking Analysis |
| ON | Oligonucleotide |
| OptiMEM® | Reduced-Serum Medium is an improved Minimal Essential Medium (MEM) |
| PAMAM | Polyamidoamine |
| PBS | Phosphate Buffered Saline |
| PEI | Polyethyleneimine |
| PEO | polyethylene oxide |
| PPI | Polypropyleneimine |
| PVA | Polyvinylalcohol |
| RLU | Relative Light Unit |
| SEM | Standard error of the mean |
| SiRNA | Small interfering RNA |
| SPPS | Solid-phase peptide synthesis |
| WST-1 | (2-(4-Iodophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium sodium salt |
| λem | Emission wavelength |
| λex | Excitation wavelength |
References
- Mintzer, M.A.; Simanek, E.E. Nonviral Vectors for Gene Delivery. Chem. Rev. 2009, 109, 259–302. [Google Scholar] [CrossRef]
- Pack, D.W.; Hoffman, A.S.; Pun, S.; Stayton, P.S. Design and development of polymers for gene delivery. Nat. Rev. Drug Discov. 2005, 4, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Gigante, A.; Li, M.; Junghänel, S.; Hirschhäuser, C.; Knauer, S.; Schmuck, C. Non-viral transfection vectors: Are hybrid materials the way forward? Medchemcomm 2019, 10, 1692–1718. [Google Scholar] [CrossRef] [PubMed]
- Buhleier, E.; Wehner, W.; Vögtle, F. “Cascade”- and “Nonskid-Chain-like” Syntheses of Molecular Cavity Topologies. Synthesis (Stuttg) 1978, 155–158. [Google Scholar] [CrossRef]
- Tomalia, D.A.; Baker, H.; Hall, M.; Kallos, G.; Martin, S.; Ryder, J.; Smith, P. Dendritic Macromolecules:1 Synthesis of Starburst Dendrimers. Macromolecules 1986, 19, 2466–2468. [Google Scholar] [CrossRef]
- Tomalia, D.A.; Baker, H.; Dewald, J.; Hall, M.; Kallos, G.; Martin, S.; Roeck, J.; Ryder, J.; Smith, P. A New Class of Polymers: Starburst-Dendritic Macromolecules. Polym. J. 1985, 17, 117–132. [Google Scholar] [CrossRef]
- Medina, S.H.; El-Sayed, M.E.H. Dendrimers as carriers for delivery of chemotherapeutic agents. Chem. Rev. 2009, 109, 3141–3157. [Google Scholar] [CrossRef] [PubMed]
- Lakshminarayanan, A.; Ravi, V.K.; Tatineni, R.; Rajesh, Y.B.R.D.; Maingi, V.; Vasu, K.S.; Madhusudhan, N.; Maiti, P.K.; Sood, A.K.; Das, S.; et al. Efficient dendrimer-DNA complexation and gene delivery vector properties of nitrogen-core poly(propyl ether imine) dendrimer in mammalian cells. Bioconjug. Chem. 2013, 24, 1612–1623. [Google Scholar] [CrossRef] [PubMed]
- Kaminskas, L.M.; Boyd, B.J.; Porter, C.J.H. Dendrimer pharmacokinetics: The effect of size, structure and surface characteristics on ADME properties. Nanomedicine 2011, 6, 1063–1084. [Google Scholar] [CrossRef]
- NanoComposix. Nanocomposix’s guide to dynamic light scattering measurement and analysis. In Proceedings of the Protocols & White Papers; NanoComposix: San Diego, CA, USA, 2015; Available online: https://cdn.shopify.com/s/files/1/0257/8237/files/nanoComposix_Guidelines_for_DLS_Measurements_and_Analysis.pdf?13692 (accessed on 15 January 2021).
- Danaei, M.; Dehghankhold, M.; Ataei, S.; Hasanzadeh Davarani, F.; Javanmard, R.; Dokhani, A.; Khorasani, S.; Mozafari, M. Impact of Particle Size and Polydispersity Index on the Clinical Applications of Lipidic Nanocarrier Systems. Pharmaceutics 2018, 10, 57. [Google Scholar] [CrossRef]
- Tomalia, D.A.; Fréchet, J.M.J. Discovery of dendrimers and dendritic polymers: A brief historical perspective. J. Polym. Sci. Part A Polym. Chem. 2002, 40, 2719–2728. [Google Scholar] [CrossRef]
- Kesharwani, P.; Jain, K.; Jain, N.K. Dendrimer as nanocarrier for drug delivery. Prog. Polym. Sci. 2014, 39, 268–307. [Google Scholar] [CrossRef]
- Sapra, R.; Verma, R.P.; Maurya, G.P.; Dhawan, S.; Babu, J.; Haridas, V. Designer Peptide and Protein Dendrimers: A Cross-Sectional Analysis. Chem. Rev. 2019, 119, 11391–11441. [Google Scholar] [CrossRef] [PubMed]
- Freeman, E.C.; Weiland, L.M.; Meng, W.S. Modeling the proton sponge hypothesis: Examining proton sponge effectiveness for enhancing intracellular gene delivery through multiscale modeling. J. Biomater. Sci. Polym. Ed. 2013, 24, 398–416. [Google Scholar] [CrossRef] [PubMed]
- Aurelia Chis, A.; Dobrea, C.; Morgovan, C.; Arseniu, A.M.; Rus, L.L.; Butuca, A.; Juncan, A.M.; Totan, M.; Vonica-Tincu, A.L.; Cormos, G.; et al. Applications and Limitations of Dendrimers in Biomedicine. Molecules 2020, 25, 3982. [Google Scholar] [CrossRef] [PubMed]
- Esfand, R.; Tomalia, D.A. Poly(amidoamine) (PAMAM) dendrimers: From biomimicry to drug delivery and biomedical applications. Drug Discov. Today 2001, 6, 427–436. [Google Scholar] [CrossRef]
- Roberts, J.C.; Bhalgat, M.K.; Zera, R.T. Preliminary biological evaluation of polyamidoamine (PAMAM) StarburstTM dendrimers. J. Biomed. Mater. Res. 1996, 30, 53–65. [Google Scholar] [CrossRef]
- Malik, N.; Wiwattanapatapee, R.; Klopsch, R.; Lorenz, K.; Frey, H.; Weener, J.W.; Meijer, E.W.; Paulus, W.; Duncan, R. Dendrimers: Relationship between structure and biocompatibility in vitro, and preliminary studies on the biodistribution of 125I-labelled polyamidoamine dendrimers in vivo. J. Control. Release 2000, 65, 133–148. [Google Scholar] [CrossRef]
- Jevprasesphant, R.; Penny, J.; Jalal, R.; Attwood, D.; Mckeown, N.B.; Emanuele, A.D. The influence of surface modification on the cytotoxicity of PAMAM dendrimers. Int. J. Pharm. 2003, 252, 263–266. [Google Scholar] [CrossRef]
- Trabulo, S.; Cardoso, A.L.; Mano, M.; de Lima, M.C.P. Cell-penetrating peptides-mechanisms of cellular uptake and generation of delivery systems. Pharmaceuticals 2010, 3, 961. [Google Scholar] [CrossRef]
- Eggimann, G.A.; Blattes, E.; Buschor, S.; Biswas, R.; Kammer, S.M.; Darbre, T.; Reymond, J.-L. Designed cell penetrating peptide dendrimers efficiently internalize cargo into cells. Chem. Commun. (Camb) 2014, 50, 7254–7257. [Google Scholar] [CrossRef] [PubMed]
- Stenström, P.; Manzanares, D.; Zhang, Y.; Ceña, V.; Malkoch, M. Evaluation of amino-functional polyester dendrimers based on Bis-MPA as nonviral vectors for siRNA delivery. Molecules 2018, 23, 2028. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Jin, R.; Wang, J.; Yue, D.; Jiang, Q.; Wu, Y.; Gu, Z. Bioreducible Fluorinated Peptide Dendrimers Capable of Circumventing Various Physiological Barriers for Highly Efficient and Safe Gene Delivery. ACS Appl. Mater. Interfaces 2016, 8, 5821–5832. [Google Scholar] [CrossRef] [PubMed]
- Welser, K.; Campbell, F.; Kudsiova, L.; Mohammadi, A.; Dawson, N.; Hart, S.L.; Barlow, D.J.; Hailes, H.C.; Lawrence, M.J.; Tabor, A.B. Gene delivery using ternary lipopolyplexes incorporating branched cationic peptides: The role of peptide sequence and branching. Mol. Pharm. 2013, 10, 127–141. [Google Scholar] [CrossRef] [PubMed]
- Lv, H.; Zhang, S.; Wang, B.; Cui, S.; Yan, J. Toxicity of cationic lipids and cationic polymers in gene delivery. J. Control. Release 2006, 114, 100–109. [Google Scholar] [CrossRef]
- Hafez, I.; Maurer, N.; Cullis, P. On the mechanism whereby cationic lipids promote intracellular delivery of polynucleic acids. Gene Ther. 2001, 8, 1188–1196. [Google Scholar] [CrossRef]
- Lampela, P.; Elomaa, M.; Ruponen, M.; Urtti, A.; Männistö, P.T.; Raasmaja, A. Different synergistic roles of small polyethylenimine and Dosper in gene delivery. J. Control. Release 2003, 88, 173–183. [Google Scholar] [CrossRef]
- Lampela, P.; Soininen, P.; Urtti, A.; Männistö, P.T.; Raasmaja, A. Synergism in gene delivery by small PEIs and three different nonviral vectors. Int. J. Pharm. 2004, 270, 175–184. [Google Scholar] [CrossRef]
- Heitz, M.; Kwok, A.; Eggimann, G.A.; Hollfelder, F.; Darbre, T.; Reymond, J. Peptide Dendrimer–Lipid Conjugates as DNA and siRNA Transfection Reagents: Role of Charge Distribution Across Generations. Chimia (Aarau) 2017, 71, 220–225. [Google Scholar] [CrossRef]
- Kwok, A.; Eggimann, G.A.; Reymond, J.-L.; Darbre, T.; Hollfelder, F. Peptide Dendrimer/Lipid Hybrid Systems Are Efficient DNA Transfection Reagents: Structure–Activity Relationships Highlight the Role of Charge Distribution Across Dendrimer Generations. ACS Nano 2013, 7, 4668–4682. [Google Scholar] [CrossRef]
- Saher, O.; Rocha, C.S.J.; Zaghloul, E.M.; Wiklander, O.P.B.; Zamolo, S.; Heitz, M.; Ezzat, K.; Gupta, D.; Reymond, J.-L.; Zain, R.; et al. Novel peptide-dendrimer/lipid/oligonucleotide ternary complexes for efficient cellular uptake and improved splice-switching activity. Eur. J. Pharm. Biopharm. 2018, 132, 29–40. [Google Scholar] [CrossRef] [PubMed]
- Kwok, A.; Eggimann, G.A.; Heitz, M.; Reymond, J.L.; Hollfelder, F.; Darbre, T. Efficient Transfection of siRNA by Peptide Dendrimer–Lipid Conjugates. ChemBioChem 2016, 17, 2223–2229. [Google Scholar] [CrossRef] [PubMed]
- Heitz, M.; Javor, S.; Darbre, T.; Reymond, J.-L. Stereoselective pH Responsive Peptide Dendrimers for siRNA Transfection. Bioconjug. Chem. 2019, 30, 2165–2182. [Google Scholar] [CrossRef] [PubMed]
- Zamolo, S.J.; Darbre, T.; Reymond, J.-L. Transfecting tissue models with CRISPR/Cas9 plasmid DNA using peptide dendrimers. Chem. Commun. 2020. [Google Scholar] [CrossRef] [PubMed]
- Heitz, M.; Zamolo, S.; Javor, S.; Reymond, J.-L. Fluorescent Peptide Dendrimers for siRNA Transfection: Tracking pH Responsive Aggregation, siRNA Binding, and Cell Penetration. Bioconjug. Chem. 2020, 31, 1671–1684. [Google Scholar] [CrossRef]
- Rocha, C.S.J.; Lundin, K.E.; Behlke, M.A.; Zain, R.; EL Andaloussi, S.; Smith, C.E. Four Novel Splice-Switch Reporter Cell Lines: Distinct Impact of Oligonucleotide Chemistry and Delivery Vector on Biological Activity. Nucleic Acid Ther. 2016, 26, 381–391. [Google Scholar] [CrossRef]
- Kang, S.-H.; Cho, M.-J.; Kole, R. Up-Regulation of Luciferase Gene Expression with Antisense Oligonucleotides: Implications and Applications in Functional Assay Development †. Biochemistry 1998, 37, 6235–6239. [Google Scholar] [CrossRef]
- Dominski, Z.; Kole, R. Restoration of correct splicing in thalassemic pre-mRNA by antisense oligonucleotides. Proc. Natl. Acad. Sci. USA 1993, 90, 8673–8677. [Google Scholar] [CrossRef]
- Saher, O.; Lehto, T.; Gissberg, O.; Gupta, D.; Gustafsson, O.; Andaloussi, S.E.L.; Darbre, T.; Lundin, K.E.; Smith, C.I.E.; Zain, R. Sugar and Polymer Excipients Enhance Uptake and Splice-Switching Activity of Peptide-Dendrimer/Lipid/Oligonucleotide Formulations. Pharmaceutics 2019, 11, 666. [Google Scholar] [CrossRef]
- Dufès, C.; Uchegbu, I.F.; Schätzlein, A.G. Dendrimers in gene delivery. Adv. Drug Deliv. Rev. 2005, 57, 2177–2202. [Google Scholar] [CrossRef]
- Boas, U.; Christensen, J.B.; Heegaard, P.M. Dendrimers in Medicine and Biotechnology. In Dendrimers in Medicine and Biotechnology. New Moleculat Tools; Royal Society of Chemistry: Cambridge, UK, 2007; p. 28. [Google Scholar]
- Wang, H.; Chang, H.; Zhang, Q.; Cheng, Y. Fabrication of Low-Generation Dendrimers into Nanostructures for Efficient and Nontoxic Gene Delivery. Top. Curr. Chem. 2017, 375, 62. [Google Scholar] [CrossRef] [PubMed]
- Haensler, J.; Szoka, F.C. Polyamidoamine cascade polymers mediate efficient transfection of cells in culture. Bioconjug. Chem. 1993, 4, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Muppalaneni, srinath Polyvinyl Alcohol in Medicine and Pharmacy: A Perspective. J. Dev. Drugs 2013, 2, 3. [CrossRef]
- Wang, W. Lyophilization and development of solid protein pharmaceuticals. Int. J. Pharm. 2000, 203, 1–60. [Google Scholar] [CrossRef]
- Wiśniewska, M.; Ostolska, I.; Szewczuk-Karpisz, K.; Chibowski, S.; Terpiłowski, K.; Gun’ko, V.M.; Zarko, V.I. Investigation of the polyvinyl alcohol stabilization mechanism and adsorption properties on the surface of ternary mixed nanooxide AST 50 (Al2O3–SiO2–TiO2). J. Nanoparticle Res. 2015, 17, 12. [Google Scholar] [CrossRef]
- Sumner, S.G.; Pringle, I.A.; Gill, D.R.; Hyde, S.C. Effect of Residual Polyvinyl Alcohol on Nanoparticle-Mediated Gene Transfection in Breast Cancer Cells. Mol. Ther. 2003, 7, S67. [Google Scholar] [CrossRef]
- Orienti, I.; Zuccari, G.; Fini, A.; Rabasco, A.M.; Carosio, R.; Raffaghello, L.; Montaldo, P.G. Modified doxorubicin for improved encapsulation in PVA polymeric micelles. Drug Deliv. J. Deliv. Target. Ther. Agents 2005, 12, 15–20. [Google Scholar] [CrossRef]
- St Croix, C.M.; Shand, S.H.; Watkins, S.C. Confocal microscopy: Comparisons, applications, and problems. Biotechniques 2005, 39. [Google Scholar] [CrossRef]
- Hobro, A.J.; Smith, N.I. An evaluation of fixation methods: Spatial and compositional cellular changes observed by Raman imaging. Vib. Spectrosc. 2017, 91, 31–45. [Google Scholar] [CrossRef]
- Pichon, C.; Monsigny, M.; Roche, A.C. Intracellular localization of oligonucleotides: Influence of fixative protocols. Antisense Nucleic Acid Drug Dev. 1999, 9, 89–93. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Daralnakhla, H.; Saher, O.; Zamolo, S.; Bazaz, S.; P. Bost, J.; Heitz, M.; Lundin, K.E.; EL Andaloussi, S.; Darbre, T.; Reymond, J.-L.; et al. Lipophilic Peptide Dendrimers for Delivery of Splice-Switching Oligonucleotides. Pharmaceutics 2021, 13, 116. https://doi.org/10.3390/pharmaceutics13010116
Daralnakhla H, Saher O, Zamolo S, Bazaz S, P. Bost J, Heitz M, Lundin KE, EL Andaloussi S, Darbre T, Reymond J-L, et al. Lipophilic Peptide Dendrimers for Delivery of Splice-Switching Oligonucleotides. Pharmaceutics. 2021; 13(1):116. https://doi.org/10.3390/pharmaceutics13010116
Chicago/Turabian StyleDaralnakhla, Haneen, Osama Saher, Susanna Zamolo, Safa Bazaz, Jeremy P. Bost, Marc Heitz, Karin E. Lundin, Samir EL Andaloussi, Tamis Darbre, Jean-Louis Reymond, and et al. 2021. "Lipophilic Peptide Dendrimers for Delivery of Splice-Switching Oligonucleotides" Pharmaceutics 13, no. 1: 116. https://doi.org/10.3390/pharmaceutics13010116
APA StyleDaralnakhla, H., Saher, O., Zamolo, S., Bazaz, S., P. Bost, J., Heitz, M., Lundin, K. E., EL Andaloussi, S., Darbre, T., Reymond, J.-L., Zain, R., & Smith, C. I. E. (2021). Lipophilic Peptide Dendrimers for Delivery of Splice-Switching Oligonucleotides. Pharmaceutics, 13(1), 116. https://doi.org/10.3390/pharmaceutics13010116







