Abstract
The discovery of infectious particles that challenge conventional thoughts concerning “what is a virus” has led to the evolution a new field of study in the past decade. Here, we review knowledge and information concerning “giant viruses”, with a focus not only on some of the best studied systems, but also provide an effort to illuminate systems yet to be better resolved. We conclude by demonstrating that there is an abundance of new host–virus systems that fall into this “giant” category, demonstrating that this field of inquiry presents great opportunities for future research.
1. Introduction: Defining Giant Viruses
In their editorial introduction to the “Giant Viruses” special issue of Virology, Fischer and Condit [1] stated “It is commonly agreed upon that these are double-stranded DNA (dsDNA) viruses with genome sizes beyond 200 kb pairs, and particles that do not pass through a 0.2-µm pore-size filter”. This definition illustrates the two striking features of giant viruses: their genome and particle size are both larger than has been historically considered for viruses. Beyond their breaking of previous paradigms, how giant viruses are defined remains contentious. Our goal in assembling this synthesis is to provide a “primer” for students of microbiology whom are interested in knowing more about these atypical viruses, and to establish a set of boundaries for their discussion. While not exhaustive, this overview addresses many of the main ideas that, for now, are current within a rapidly expanding field.
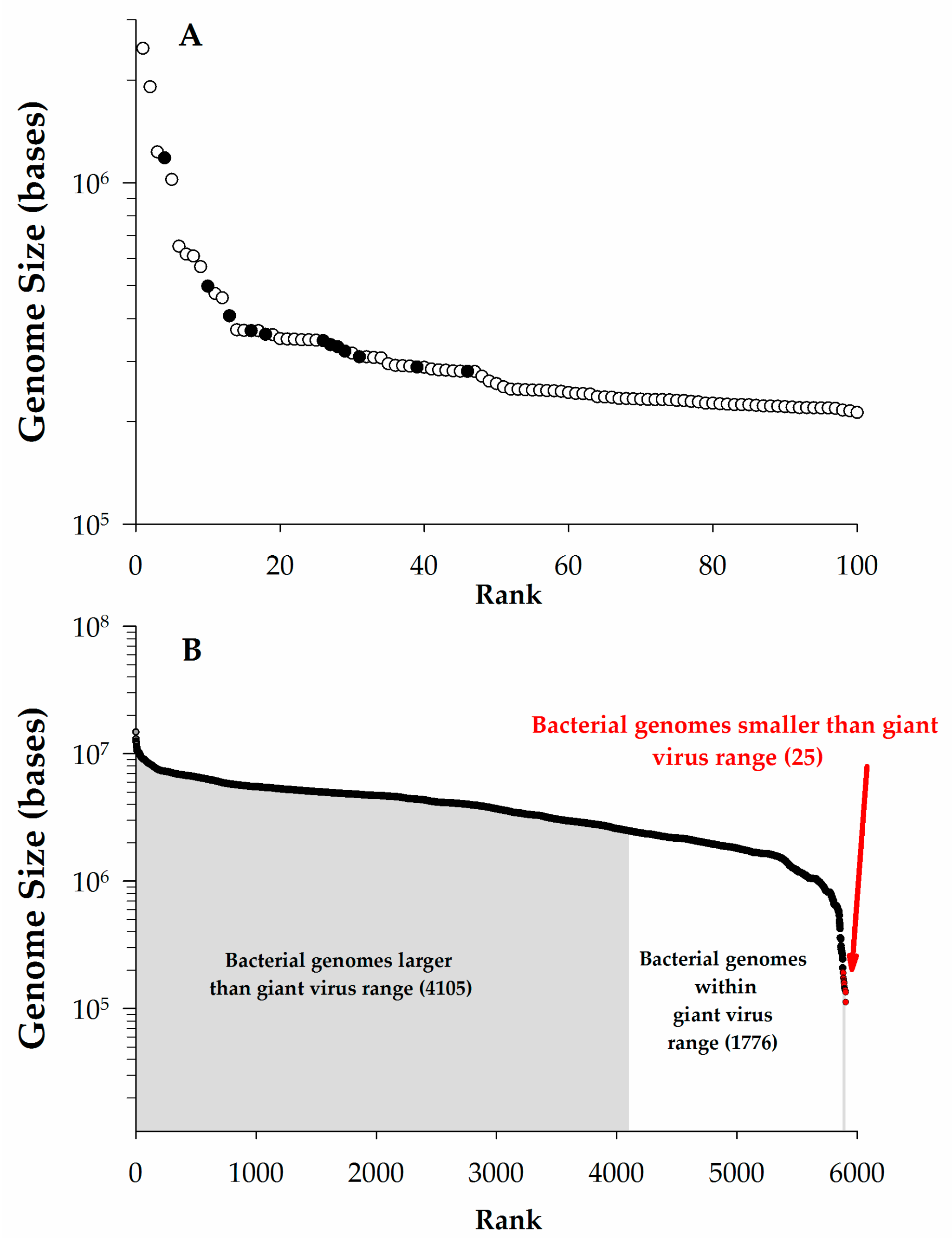
Some definitions of giant viruses focus only on genome size with lower limits ranging from undefined [2] to stringent (280 kb or 300 kb) cutoffs [3,4]. Other efforts have focused on the virus particle, suggesting they should be larger than 100 nm [2] or need be easily visible by light microscopy (>300 nm) [5]. One problem with establishing a particular definition for either genome or particle size is that, as additional large viruses are isolated, the rationale may no longer be justified (e.g., Aureococcus anophagefferens virus (AaV), a close phylogenetic relative of Mimivirus, is only ~140 nm in diameter) [6]. Indeed, a previous definition proposed a genome minimum of 280 kb due to a notable inflection point in a rank order plot of virus genome size [3]. However, in re-examining the largest 100 complete virus genomes in the National Center for Biotechnology Information’s (NCBI) genome database, this gap is no longer present and a change in slope now occurs at ~400 kb (Figure 1A). This undersampling of giant viruses has resulted in a lack of sufficient information to describe their general characteristics [7,8]. While the vagaries of this definition will fade over time, herein we consider viruses ‘giant’ if their genome is larger than 200 kb. Moreover, this review will focus primarily on giants that infect single-celled eukaryotes.
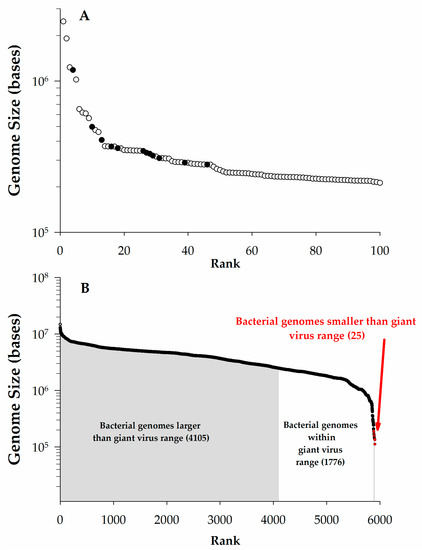
Figure 1.
The scale of giant virus genomes. (A). Genome size vs. rank plot for the largest 100 complete viral genomes as of January 2016 from National Center for Biotechnology Information (NCBI). Data points noted (●) were previously used in discussion by Claverie et al. [3] to define giants viruses as having genomes > 280 kb, open circles (○) represent additional data; (B). Genome size vs. rank order of completed bacterial genomes in NCBI as of January 2016. Sizes are color-coded to match the ranges of giant virus genomes.
Using a cutoff of genomic content >200 kb pairs (kbp), ~2.2% (115/5356) of all of the completed virus genomes in NCBI fall within the realm of giants (Figure 1A). To date, all of these giants have genomes consisting of double-stranded DNA: the largest complete genome for other nucleic acid-type viruses is that of the double-stranded RNA (dsRNA) Dendrolimus punctatus cypovirus 22 (32.75 kbp) [9]. Perhaps more surprising is that this genome size range for giant viruses overlaps with more than ~one third of the complete prokaryotic genomes in NCBI (Figure 1B), as well as the genome sizes of several small eukaryotes [10]. This includes the smallest free-living archaeon (Methanothermus fervidus, 1.2 Mb) and the smallest free-living bacterium (Candidatus Actinomarina minuta, estimated ~700 kbp) [11]. While we will not consider them beyond the occasional passing mention in this article, it should be noted that several bacteriophages have genomes exceeding the 200 kbp genome size (see Table 1), and therefore qualify as giants. These phages infect both Gram-positive and -negative bacteria, including cyanobacteria [12,13].

Table 1.
Comparison of host and viral genome size and GC content. All data was collected from the NCBI repository.
As with observed ranges in genomic size, there is also a wide range of GC content of these viruses relative to the small eukaryotes they infect (Table 1). On average, mobile elements such as phage and plasmids are more AT-rich than their host, but usually by only ~5% [14]. In contrast, Emiliania huxleyi virus (EhV) and AaV, which infect eukaryotic algae, have GC contents that are 24.3% and 38.7% lower than their hosts nuclear genomes, respectively [15,16], while the chloroviruses (freshwater viruses infecting Chlorella) have GC contents that are ~21% lower than their host’s nuclear genome. While not a defining feature of all large viruses, this GC difference raises interesting questions concerning the scavenging of nucleotides during the infection cycle. Construction of new viruses is in some cases thought to depend on materials “scavenged” from the host cell, yet in the case of these viruses there would seem to be a discrepancy in terms of what would be available for scavenging. An interest side note to this is that mitochondrial and chloroplast genomes are often observed to have such relative low GC content genomes, similar to these viruses [14,15], implying a potential for scavenged materials from organelles to be important in the construction of new virus particles.
The current size range for giant virus particles varies from our operationally defined ~200 nm to >1500 nm in diameter [5], although as noted, phylogenetic relatives to these giants exist that are only ~140 nm. Indeed, the upper limit of this range is larger than for several bacteria and archaea (Figure 1B), redefining how we think about the relative size of prokaryotes and viruses. These large particle diameters may be needed to house their large genomes (see below), but it has been argued that there are other evolutionary pressures for these virus particles to retain large physical sizes [5]. For example, viruses infecting Acanthamoeba are internalized via phagocytosis, and it has been shown that this process works less efficiently on smaller (<600 nm) particles [16]. Additionally, based on standard contact kinetics, a larger particle size may increase the probability of contact between the virus and its host in the environment [17].
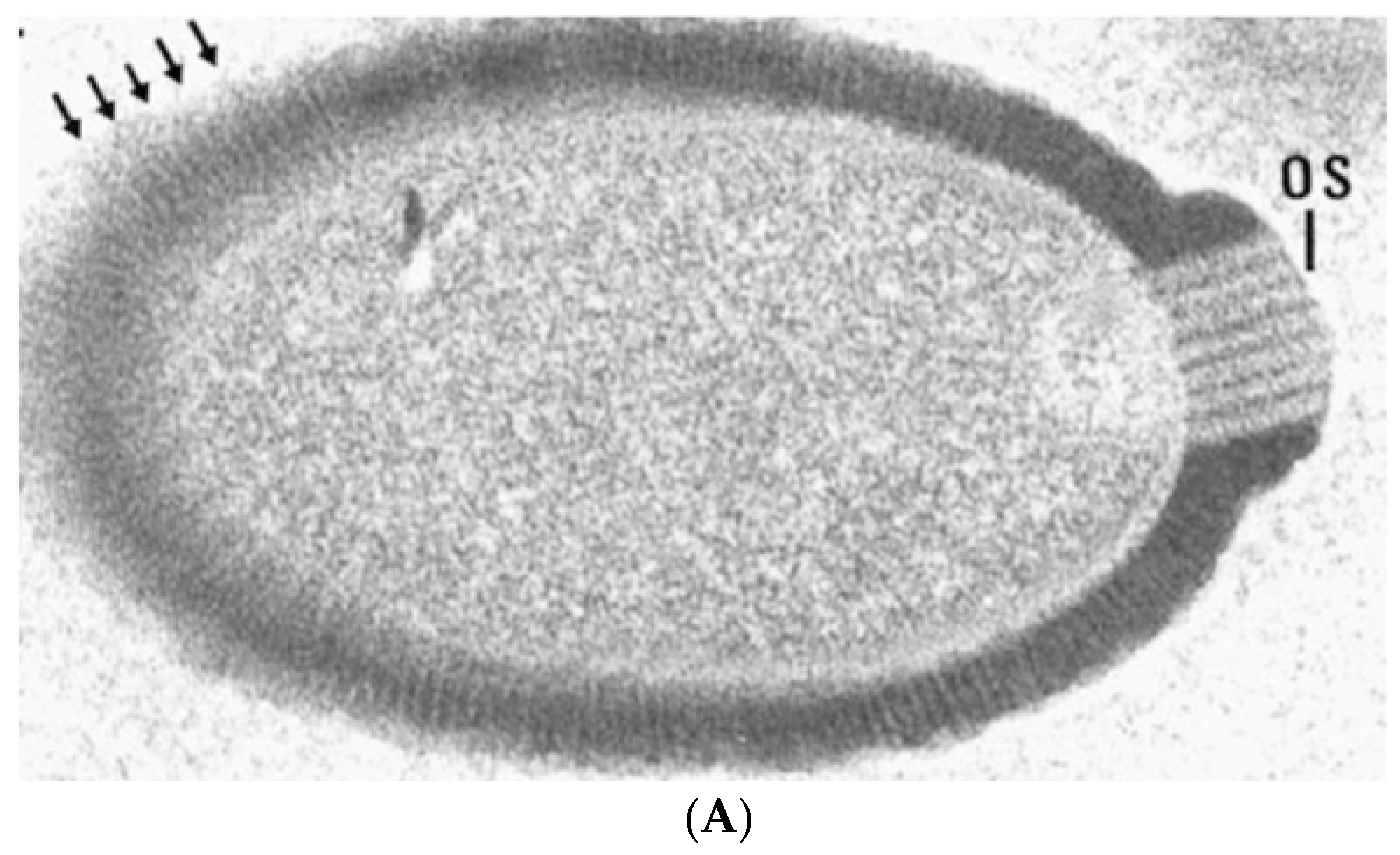
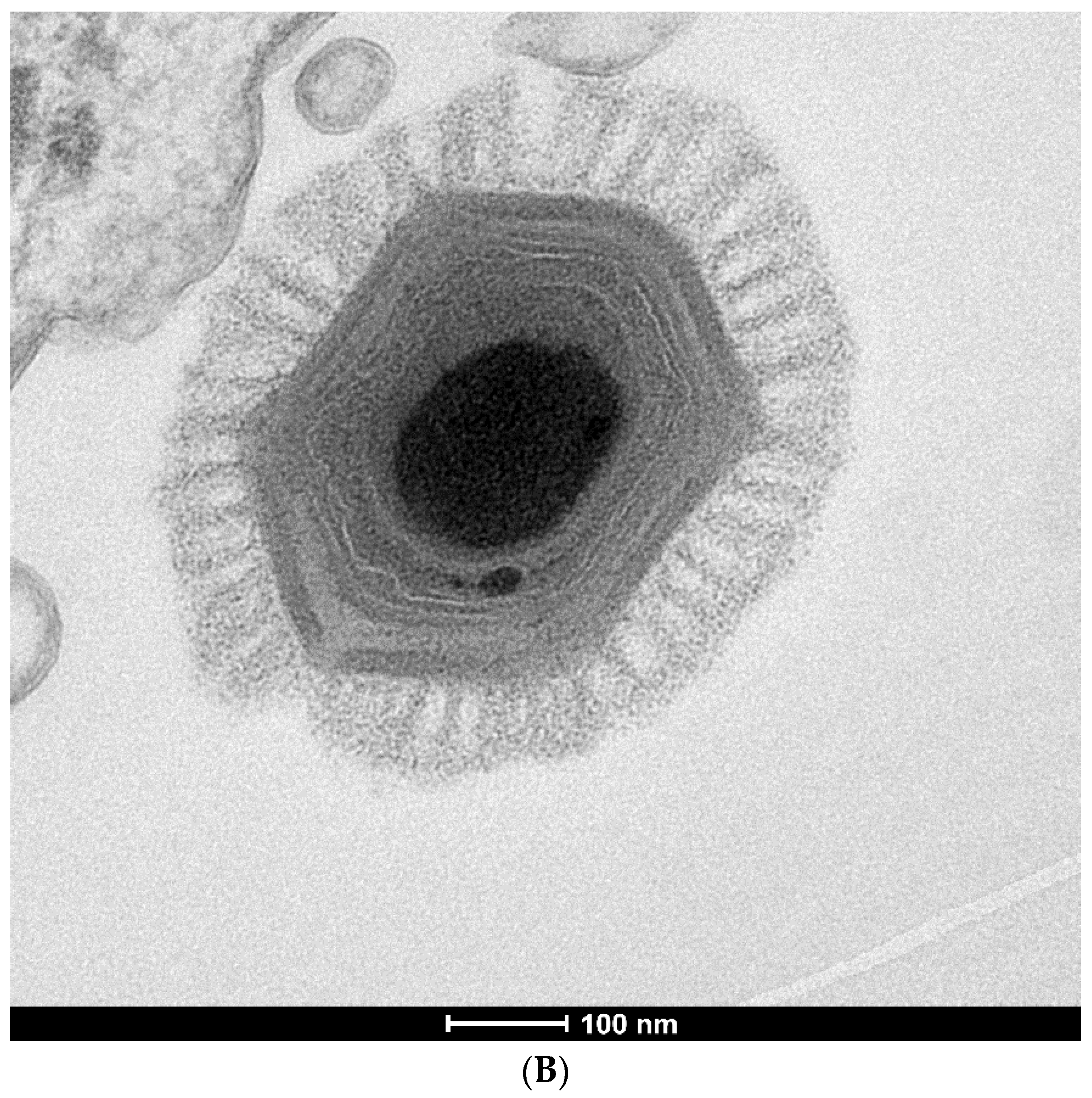
In addition to a tremendous variation in genome and particle size, giant viruses also have highly diverse morphologies that can be broadly categorized into two groups: ovoid and icosahedral (Figure 2). These morphological differences correspond to the structural proteins that make the virion capsids; icosahedrons are built by homologous β-barrel jelly-roll Major Capsid Proteins (MCPs) with minor capsid proteins acting as scaffolds connecting trisymmetrons and the outer capsid to the inner membrane surrounding the viral genome [18]. In contrast, ovoid viruses encode phylogenetically distant (Mollivirus) to unconvincing (Pandoravirus and Pithovirus) homologs to MCP [19,20,21]. It is unclear how the virion shape provides a selective advantage, since both types have been isolated in similar habitats.
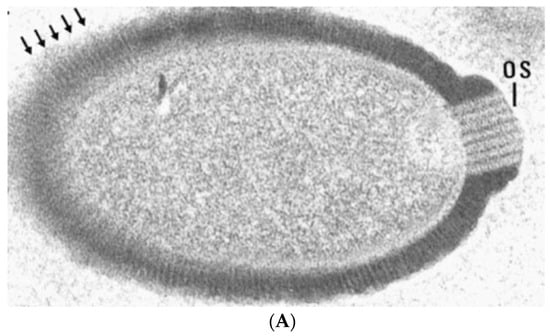
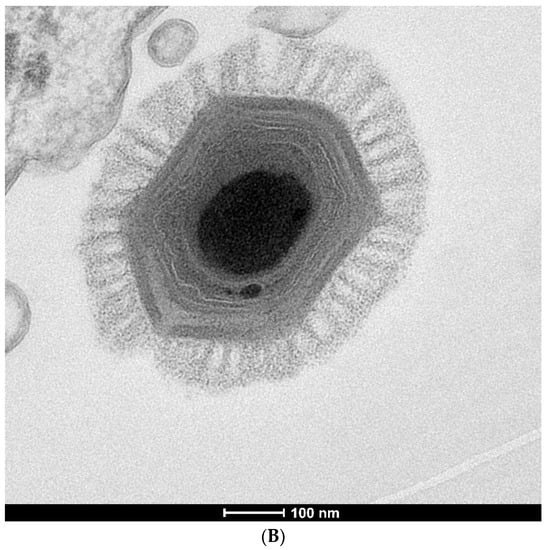
Figure 2.
Transmission electron micrographs of giant virus particles. (A) Pithovirus, as seen in Michel et al. [22]. Originally identified as a KC5/2 parasite, the image shows the electron dense viral wall consisting of perpendicularly oriented fibers or microtubules (arrows), and a marked ostiole (os) located at the apical end of the cell. Reprinted with permission—original magnification at 85,000×; (B) Megavirus chilensis. Image courtesy of Professors Chantal Abergel and Jean-Michel Claverie.
Another mysterious aspect of these giant virus particles are the unique biochemical and morphological features. Virus–host interactions are thought to be facilitated in one of two ways: adsorption to the host cell wall, as is typical of algal host–virus systems [23], or phagocytosis by a protist host. These interactions often involve unique structures. For example, Mimivirus and its close relatives (Megavirus, Marseillevirus, Lausannevirus, and Moumouvirus) are characterized by proteinaceous fibers anchored to the icosahedron capsid [24,25] that are covered in glycolinkages [26,27,28]. It has been hypothesized that these fibers work in tandem with the large size of the viruses to facilitate phagocytosis, as they appear to have a similar composition to peptidoglycan and thus help mimic a bacterium (indeed, the name Mimivirus comes from “Mimicking Microbe” [29]). Additionally, the fibrous glycoproteins enable viral adsorption to diverse organisms ranging from bacteria and fungi to arthropods [30], implying a potential for both environmental dispersion and an incidental infection strategy in amoeba. Phycodnaviridae may also use unique structures to gain access to their host, though their mode of entry is typically by adsorption/injection, as opposed to phagocytosis. For example, the Chlorovirus capsid contains one spike located at a unique vertex of the icosahedral capsid that must be oriented towards the host cell surface to initiate infection [31]. Similarly, Mimivirus and its relatives utilize a five-pointed vertex called the ‘stargate’ structure that permits the first step in activating infection. Infection is initiated by fusion of the internal viral lipid membrane to the phagosomal membrane [24,32], which differs from algal viruses that fuse with the host cell membrane. This fusion event is observed in all giant viruses despite differences in structural features or infection strategies [20]. Whether these features are the result of homologous or convergent evolution remains to be determined, though given the breadth of physiological variation in the taxa, conservation of this mechanism is a compelling argument for monophyly.
2. Non-Structural Components of the Virion
An anomaly among the giant viruses are several viruses that include EhV, which have a lipid envelope outside of the capsid. These viruses include a Phaeocystis globosa virus (PgV-07T) [33] and several viruses infecting Micromonas pussila [34] which allows for a unique mode of infection and provides protection from environmental stressors [35]. This may be vital to the survival and transmission of these viruses, as they are ingested and transported across blooms by copepods [36]. An additional role of this lipid envelope and its associated proteins is an assumed association with recognition of the host and initiation of infection.
The nucleocytoplasmic large DNA viruses (NCLDVs) (described below) package a variety of proteins inside their capsids encoded by either viral or host genomes that are deployed immediately upon infection. For example, the seven proteomes of giant virus particles currently available (below) contain proteins predicted to combat oxidative stress, presumably because viral infections have been shown to generate Reactive Oxygen Species (ROS) that can inhibit viral replication [37]. Interestingly, Pandoravirus salinus carries one viral-encoded oxidoreductase, as well as three host-derived proteins predicted to combat ROS [19]. P. bursaria chlorella virus-1 (PBCV-1) and the more recently described Megavirus chilensis package homologous Cu-Zn superoxide dismutases [38,39]. In M. chilensis, this protein is remarkable for having the unique ability to fold and incorporate key metallic cofactors without the aid of chaperone proteins [39]. Additionally, C. roenbergensis virus (CroV) and Mimivirus both package novel sulfhydryl oxidases that may function in the formation of disulfide bonds [40,41]. These sulfhydryl oxidases as well as other protein disulfide isomerases present in CroV could aid in protein folding or viral entry similar to those found in retroviruses [42,43,44].
3. Gauging the Host Range of Giant Viruses in Nature
One concern regarding giant virus isolation using Acanthamoeba spp. is that while these are permissive, they may not be the natural hosts. Genomic analyses have been used in an attempt to determine natural hosts. In Mimivirus, most of the genes horizontally transferred from eukaryotes originated from amoeba, indicating amoebae are most likely the natural host of Mimivirus, but alternative hosts are still possible [45]. Indeed, their unique size and independence from host machinery may allow giant viruses to infect a wide range of hosts, which makes the search for the natural host more challenging. In addition to amoeba, NCLDVs have been reported to infect mice [46] and the symbiotic zooxanthelle of corals [47]. Giant viruses have also been isolated from human blood [48] and have been found in the human virome [49], indicating a potential role in human health (or at least a route of exposure). Indeed, the recent finding of Acanthocystis turfacea chlorella virus 1 (ATCV-1) from human oropharyngeal samples is intriguing: subsequent analyses have shown consistency between the presence of these viruses and reduced cognitive function in humans and mice [49].
4. Creating (an) Order from the Chaos: The Nucleocytoplasmic Large DNA Viruses
The NCLDV classification was created to define a monophyletic group of families that, when initially conceived, included Asfarviridae, Phycodnaviridae, Poxviridae, and Iridoviridae [50]. The rationale for this grouping was based on a conserved core of (1) nine genes hypothesized as representative of a common NCLDV ancestor and (2) a total of 22 more genes found in at least three of the four constituent viral families. The name is a reference to the replicative strategies of the included families as they replicate in both the nucleus and cytoplasm (phycodnaviruses, asfarvavirus and iridovirus) [51,52,53], or totally within the cytoplasm (poxviruses) [54]. A NCLDV ancestor has been hypothesized to have originated early in evolutionary history, possibly contemporaneously with early eukaryotic evolution, as suggested by the broad host range of NCLDV members [55]. However, the nature of this ancestral NCLDV remains unclear. Due to non-orthologous displacement of core genes [55] and potential reductive evolution [5] it is especially difficult to estimate the approximate genome size of any common ancestor and whether it would qualify as a giant virus when compared with modern giants. Indeed given theories on genome size variability, such as the genomic accordion [56,57], it is likely that predecessors of a variety of genome sizes existed. Moreover, it has been argued that mobile genetic elements encoded by virophages and transpovirions may have contributed significantly to the size of the NCLDV genome [58,59]. Therefore, the ancestral NCLDV may have been much smaller in genome size than modern representatives, and the mechanism by which it expanded its genome may have resulted in the wide range of genome sizes seen in current NCLDV members [60].
The NCLDV classification is not without its shortcomings. As new members are added to the group, the “nucleocytoplasmic” distinction of replicative strategies becomes less useful due to the increasing diversity of virion production. Many NCLDV families utilize a nucleocytoplasmic route for replication, including Asfarviridae [52], Iridoviridae and Ascoviridae [53], Phycodnaviridae [51], and Pandoraviridae. Other families, like Poxviridae [54], Mimiviridae [61], Marseilleviridae [62] and Pithoviridae [5], begin and complete their replication cycles exclusively in the cytoplasm, encoding the replication and transcription machinery necessary to produce virions without nuclear involvement. From a taxonomic perspective, the NCLDV group does not follow the naming conventions of (and is not recognized by) the International Committee on Taxonomy of Viruses (ICTV), as the classification lacks context within a larger hierarchy. To rectify this, Colson et al. proposed to reclassify NCLDVs within the new viral order Megavirales [63,64] based on the presence of conserved ancestral genes and a large icosahedral capsid composed of a homologous β-barrel jelly roll protein. This classification scheme, however, excludes the Poxviridae and Ascoviridae, [65] as well as Pandoravirus, Pithovirus and Mollivirus. In addition, the Megavirales classification required the capacity to assemble viral factories within the cytoplasm of host cells [62,66,67,68,69], a feature found in RNA viruses [70] but not seen in DNA viruses outside of the NCLDV group [64]. Currently (as of December 2016), the Megavirales is not considered a classification by the ICTV.
Most recently, the NCLDV genome size range has expanded to include genomes from 100 kb to 2.77 Mb encoding from 110 to 2556 genes [19,60]. The ten groups of NCLDV (Phycodnaviridae, Poxviridae, Asfarviridae, Ascoviridae, Iridoviridae, Mimiviridae, Marseilleviridae, Pandoraviridae, Pithovirus, and Mollivirus) infect a broad spectrum of hosts. In keeping with the NCLDV group’s high degree of variability regarding particle size and host range, these viruses also display varying degrees of reliance on host metabolism and machinery, resulting in a limited number of highly conserved or “core” genes (e.g., see [71]). Yet despite these variances in NCLDV traits, common ground does exist. There are genes conserved amongst all available NCLDV genomes that are crucial for viral production or virion structure, such as the D5R packaging ATPase, D13L major capsid protein, and the B family DNA polymerase.
Comparative analyses of the genes conserved amongst different giant virus families has historically supported the monophyletic nature of the NCLDV group, and recent efforts to determine the clusters of orthologous groups (COGs) for giant viruses support their monophyly [72]. The conserved genes further provide potential markers that might be used in the discovery of novel NCLDVs and the determination of phylogenetic relationships between more closely related taxa [60]. For example, Moniruzzaman and colleagues [73] demonstrated an expanded level of diversity of the algal-specific members of the Mimiviridae by targeting the conserved MCP gene in this clade. However, this approach has its limits; the three recently discovered representatives of Pandoravirus lack the major capsid protein and the D5R helicase, as well as a number of other core NCLDV genes [19]. Indeed only 17 of the 49 inferred ancestral NCLDV genes were found in at least one of the Pandoravirus genomes, calling into question their inclusion in the giant virus clade despite their particle and genome size [74].
5. Viruses as a Possible Fourth Domain of Life
Initially viruses were defined by their intrinsic filterability away from cellular life forms [75,76,77], a definition subsequently refined to include their lack of ribosomes, a host-dependent metabolic strategy, and replication by means other than binary fission [78]. That the unique capabilities of NCLDVs still fit well within the latter definition, after fifty years of discovery and scientific scrutiny, highlights a fundamental difference between cellular organisms and viral particles. However, giant viruses do challenge these distinctions. Independent of their size, which invalidates the informal 0.2-μm filter cutoff, NCLDVs are remarkably cell-like in virion structure and gene content. In addition to their protein coat, membrane, and genome, Mimivirus and Marseillevirus particles contain messenger RNA molecules, making them the only viruses, to date, that contain both types of nucleic acid [79]. Moreover, several viruses encode genes involved in translational processes, such as varying numbers of aminoacyl-transfer RNA (tRNA) synthetases [69,80]. Indeed, we hypothesize these proteins may be useful in overcoming differences in GC content seen between some viruses and hosts (Table 1), but this has yet to be empirically demonstrated.
The discovery of translational machinery (including that mentioned above) encoded in select virus genomes allows for comparisons to traditionally “cellular” functions normally associated with the three domains of life. Sequence alignments comparing multiple genes involved in DNA replication and repair, transcription, and translation shared between cellular organisms and NCLDVs appear to show deeply branching relationships as ancient as the domain Eukarya. It was subsequently hypothesized that giant viruses evolved from a cellular common ancestor belonging to a currently extinct fourth domain of life, unique from Bacteria, Archaea, and Eukarya [63,81]. Seemingly in support of this hypothesis is the abundance of coding sequences (ORFans) in giant viral genomes with no known homologues in the other domains.
These ideas have proven somewhat controversial, as direct sequence comparison of genes conserved among cellular organisms with virus-encoded homologs is problematic. As selective pressures on similar genes within viruses and their hosts are likely different, accelerated sequence divergence in viruses may exaggerate their perceived distance from the derived gene [82]. Subsequent alignments accounting for compositional heterogeneity and homoplasy place giant virus genes with eukaryotes [83]. While it has been countered that giant virus genes do not evolve more quickly than their cellular counterparts, this has yet to be demonstrated outside of a single example within Marseilleviridae [5,84]. Indeed, an overabundance of viral open reading frames (ORFs) without known homologues is not a problem unique to giant viruses [85]. These observations and others have led to the alternative hypothesis: gene content within the different NCLDV families suggests that their genomes have been built up from smaller viruses over time, rather than by loss of unnecessary genes by an ancient cellular ancestor [72]. As some of the current NCLDVs replicate in phagotrophs like Acanthamoeba and Cafeteria roenbergensis, it was hypothesized that smaller viruses may incorporate genetic material from other organisms phagocytosed by the host.
6. Giant Viruses in the Environment
While surveys are not yet exhaustive, giant viruses appear to be found in all environments. Since the discovery of Mimivirus from a water cooling tower [86], giant viruses have been found in locations where amoebae normally thrive, including seawater, soil, aerosols, and man-made aquatic environments such as sewage, fountains and air conditioners [87], in addition to harsh, unexpected ecosystems such as permafrost [20]. Lastly, giant viruses or their DNA sequences have been observed in animals such as dinoflagellate-associated coral [47], arthropods, and humans [49,88].
A powerful tool in the identification of putative new viruses are environmental metagenomic studies (Table 2), though most have not focused specifically on giant viruses until recently [21]. Current research suggests giant viruses only comprise a small percentage of viruses (<1%) in most samples. However, virus densities can fluctuate based on contact with their host: for example, Chlorella viruses are much more abundant when their hosts, normally sequestered as endosymbionts of Paramecium bursaria, are made available as a consequence of predatory activity on the Paramecium [89]. Regardless, it is clear some families tend to be more common than others: in marine metagenomics samples Phycodnaviridae-related sequences were found to be highest in abundance, followed by Mimiviridae [90,91]. It is also clear that these viruses are persistent: the discovery of 30,000-year-old Mollivirus particles in permafrost suggests that giant viruses can survive, under the correct conditions, for long periods of time [21]. When combined with other tools such as flow cytometry sorting of either individual particles [92] or infected hosts [93], these new approaches will begin to shed significant light on the natural diversity of these populations.

Table 2.
Comparison of giant virus reads to total viral reads in shotgun metagenomic studies from different environments.
To date, much of the focus on giant viruses has been on their genomics rather than their influence on the environments in which they persist. Several large, dsDNA viruses including EhV [97], PgV [98], AaV [6,97] and Heterosigma akashiwo virus (HaV) [99] are associated with algal blooms, although only a few have been directly shown to infect and lyse the phytoplankton involved with the bloom in situ [97,100]. Algal blooms occur on large geographical scales and result in significant influxes of atmospheric carbon into the world’s oceans. Viruses, particularly bacteriophages, are known drivers of dissolved organic matter (DOM) release back into the environment via a process known as the “viral shunt” [101,102]. With the large biomass of algae associated with these blooms, virus-mediated collapse by giant viruses may also be an important driver of dissolved and particulate organic matter release. Giant viruses that infect algae may be likened to bacteriophages in terms of participating in the viral shunt, and the release of nutrients back into the environment may be an important part of the ecological cycle in aquatic systems [102].
A recent estimate suggested that giant viruses available in culture were infectious to at least 22 different algal species [103]. Globally, it has been proposed that there are more than 350,000 algal species [104]. Given the possibility that all algae may be infected with one or more viruses [105,106], the possibility of a collection of unknown giants remains very real, and indeed molecular data point to at least a broad diversity within the known groups [107,108]. Building on the above, it is clear from a survey of the literature that researchers identified candidate protist-giant virus systems well before Mimivirus was documented (Table 3). In the late 1960s and early 1970s, the expanded availability of transmission electron microscopes to researchers resulted in a series of observations concerning the presence of large virus-like particles inside algal cells [107,109]. In many cases, these virus–host systems have been largely ignored by the scientific community, creating a broad spectrum of opportunities for researchers to begin to cultivate these plankton in an effort to isolate and characterize new giant viruses. Given the expansive putative host-range that has been observed, it is likely that many of these viruses could fill in knowledge gaps concerning the diversity and potential function of these particles. Indeed, one example of how new hosts can be used to discover new viruses are the Faustovirus, recently discovered using Vermamoeba (a protist found in both humans and natural systems) as a screen [110]: unique to these viruses is a collection of genes three times larger than the other members of the Asfarviridae family.

Table 3.
A chronological list of organisms shown in the literature to contain viruses consistent with the giant virus size class.
And while it is obvious that there is a dearth of knowledge concerning giant viruses that infect algae in the environment, there is an even larger knowledge gap regarding giant viruses infecting heterotrophic eukaryotes. The most studied of these viruses is CroV, which infects the heterotrophic grazer Cafeteria roenbergensis [111]. Given this organism is a grazer of primary producers it is possible that infection of this organism by CroV could have effects on lower trophic level organisms. It has been shown that grazing can be an important driver of algal bloom decline [112], so it stands to reason that the effects giant viruses have on mixo/heterotrophic-plankton are critical to understanding bloom dynamics. Almost no information, at this time, is available to discuss the impacts of these infections, but they will most likely result in interesting discoveries and further our understanding of how giant viruses alter the microbial food web.
7. Intimate Interactions with the Host: Eco-Evolutionary Consequences
Only recently have we come to appreciate the possibility of gene transfer between giant viruses and their hosts. A large proportion of giant virus genes comes from diverse sources, including from their eukaryotic hosts [127]. In EhV, seven genes involved in sphingolipid biosynthesis pathway were putatively transferred from the host algae [128]. Upon infection, the host sphingolipid biosynthesis pathway is downregulated concomitant with the upregulation of the corresponding viral genes, leading to increased production of viral glycosphingolipids (vGSLs) [129]. EhV particles are covered by vGSLs, and this unique lipid molecule ultimately induces programmed cell death (PCD) in infected hosts [130].
A genome wide phylogenetic study of AaV identified a number of genes having their highest phylogenetic affinity to host (Aureococcus) homologs, [71]. This agrees with observations made by earlier studies on several other giant viruses [127,131]. While gene acquisition may be one of the evolutionary strategies of giant viruses, how these genes confer ecological advantages remains largely unknown. As the vast majority of viruses harbor streamlined genomes with few genes, the enormous genetic resource of giant viruses poses a paradox in terms of energetic cost of replication. Closer inspection of a number of sequenced eukaryotic genomes revealed a large number of genes originated from giant viruses [132,133]. In a recent study, large genomic islands, putatively derived from both giant viruses and a virophage, were found in Bigelowiella natans, a Cryptomonad algae [134]. In another study, “core” genes from giant viruses were detected in eight protists and a metazoan (Hydra magnipapillata) genome [132]. Remarkably, a 400-kb region in the H. magnipapillata was putatively identified to be of viral origin [132]. Major capsid gene phylogeny indicated the genes were likely from a Mimiviridae family member. Giant virus particles and marker genes have also recently also been observed associated with zooxanthellae from the genus Symbiodinium, a dinoflagellate typically found closely associated with corals [47]. Giant virus-like genes were also found in several other protists [133,135] and some plant genomes, namely Physcomitrella patens and Selaginella moellendorffii [136]. The role of giant virus-derived genes in host remain an open question.
Host–virus interactions result in an evolutionary arms race—leading to the emergence of new diversity in the host and virus population [137]. Hosts of giant viruses have evolved a variety of defense mechanisms against giant viruses. An elegant example is the ‘Cheshire cat’ strategy adopted by Emiliania huxleyi [138]. The diploid calcified cells of E. huxleyi are susceptible to EhV infection, while the haploid stage is ‘invisible’ to infection. It has been suggested that during the decline of the Brown tide blooms, a virus-resistant population of the Aureococcus persists, maintaining a relatively high abundance of Aureococcus even after the demise of the bloom [73,97].
8. Virophage
Another interesting characteristic of some giant viruses is their susceptibility to infection by other bioactive particles, termed “virophage”. The first virophage to be isolated was named Sputnik [59], which replicates within the viral factory used by Mamavirus within Acanthamoeba castellanii. Because of this, Sputnik only replicates within A. castellanii co-infected with Mamavirus. Infection by the virophage causes abnormal capsid structure of Mamavirus, increasing capsid size and causing abnormal fiber localization on its surface, suggesting a parasitic relationship between the two [59]. Co-incubation of Sputnik and Mamavirus decreased infective Mamavirus particle titers by approximately 70% and increased the survival rate of the A. castellanii [59]. Similar virophages have been found infecting other giant viruses as well [139,140,141,142]. The discovery of “viruses that infect viruses” has strengthened the argument that viruses are living entities [143]. Some classes of Mimivirus appear to have developed a CRISPR-CAS-like system suggested to combat these virophages, called the Mimivirus virophage resistant element (MIMIVIRE) [144]. Interestingly, a number of genes homologous to those in the MIMIVIRE system are present in other giant viruses, suggesting that the MIMIVIRE-like defense systems might not be exclusive to Mimivirus [134,145]. Other interpretations, however, are questioning these conclusions [146]. Much has yet to be learned in these systems, but virophages may act like both ‘provirophage’ and ‘provirus’, depending on the genomic context at multiple levels.
9. Conclusions
The discovery of Mimivirus has driven both the nascence and evolution of a new area of scientific inquiry. Giant viruses are now the topics of evolutionary, ecological and biotechnological inquiries. Moreover, broad-scale efforts to identify new virus–host systems, ranging from classic culture-based approaches to newer bioinformatics efforts to link viruses and their hosts [147] will soon provide a larger data base of information concerning the key features of these novel virus particles. Indeed, a survey of older literature (Table 3) clearly demonstrates that there are many virus–host systems that have been observed but are yet to be isolated and characterized. Moving forward, there is little doubt that the study of giant viruses will shed new light not only on virus–host relationships, but also on key evolutionary processes including the natural occurrence rates of transduction and horizontal gene transfer.
Acknowledgments
The authors wish to thank the Department of Microbiology at the University of Tennessee: this review was assembled as part of a graduate course (Virology 604) instructed by S.W.W. and G.R.L. Funds to support publication were received from the Kenneth & Blaire Mossman Endowment to the University of Tennessee (S.W.W.).
Author Contributions
S.W.W. and G.R.L. conceived this exercise for the Department of Microbiology graduate journal club in virology. All authors participated in research and crafting of the document.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Fischer, M.G.; Condit, R.C. Editorial introduction to “giant viruses” special issue of virology. Virology 2014, 466–467, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Durzyriska, J. Giant viruses: Enfants terribles in the microbal world. Future Virol. 2015, 10, 795–806. [Google Scholar] [CrossRef]
- Claverie, J.M.; Ogata, H.; Audic, S.; Abergel, C.; Suhre, K.; Fournier, P.E. Mimivirus and the emerging concept of “giant” virus. Virus Res. 2006, 117, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T. Giant viruses in the environment: Their origins and evolution. Curr. Opin. Virol. 2011, 1, 58–62. [Google Scholar] [CrossRef] [PubMed]
- Abergel, C.; Legendre, M.; Claverie, J.M. The rapidly expanding universe of giant viruses: Mimivirus, Pandoravirus, Pithovirus and Mollivirus. FEMS Microbiol. Rev. 2015, 39, 779–796. [Google Scholar] [CrossRef] [PubMed]
- Gastrich, M.D.; Leigh-Bell, J.A.; Gobler, C.; Anderson, O.R.; Wilhelm, S.W. Viruses as potential regulators of regional brown tide blooms caused by the alga, Aureococcus anophagefferens: A comparison of bloom years 1999–2000 and 2002. Estuaries 2004, 27, 112–119. [Google Scholar] [CrossRef]
- Serwer, P.; Hayes, S.J.; Thomas, J.A.; Hardies, S.C. Propagating the missing bacteriophages: A large bacteriophage in a new class. Virol. J. 2007, 4, 21. [Google Scholar] [CrossRef] [PubMed]
- Martínez, J.M.; Swan, B.K.; Wilson, W.H. Marine viruses, a genetic reservoir revealed by targeted viromics. ISME J. 2014, 8, 1079–1088. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Qin, T.; Xiao, Y.; Qin, F.; Lei, C.; Sun, X. Genomic and biological characterization of a new cypovirus isolated from Dendrolimus punctatus. PLoS ONE 2014, 9, e113201. [Google Scholar] [CrossRef] [PubMed]
- Corradi, N.; Pombert, J.-F.; Farinelli, L.; Didier, E.S.; Keeling, P.J. The complete sequence of the smallest known nuclear genome from the microsporidian Encephalitozoon intestinalis. Nat. Commun. 2010, 1, 77. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Cano, D.J.; Reyes-Prieto, M.; Martínez-Romero, E.; Partida-Martínez, L.P.; Latorre, A.; Moya, A.; Delaye, L. Evolution of small prokaryotic genomes. Front. Microbiol. 2015, 5, 742. [Google Scholar] [CrossRef] [PubMed]
- Mesyanzhinov, V.V.; Robben, J.; Grymonprez, B.; Kostyuchenko, V.A.; Bourkaltseva, M.V.; Sykilinda, N.N.; Krylov, V.N.; Volckaert, G. The genome of bacteriophage φKZ of Pseudomonas aeruginosa. J. Mol. Biol. 2002, 317, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.B.; Coleman, M.L.; Weigele, P.; Rohwer, F.; Chisholm, S.W. Three Prochlorococcus cyanophage genomes: Signature features and ecological interpretations. PLoS Biol. 2005, 3, e144. [Google Scholar] [CrossRef] [PubMed]
- Ong, H.C.; Wilhelm, S.W.; Gobler, C.J.; Bullerjahn, G.; Jacobs, M.A.; McKay, J.; Sims, E.H.; Gillett, W.G.; Zhou, Y.; Haugen, E.; et al. Analyses of the complete chloroplast genome of two members of the pelagophyceae: Aureococcus anophagefferens CCMP1984 and Aureoumbra lagunesis CCMP1507. J. Phycol. 2010, 46, 602–615. [Google Scholar] [CrossRef]
- Orsini, M.; Costelli, C.; Malavasi, V.; Cusano, R.; Alessandro, C.; Angius, A.; Cao, G. Complete sequence and characterization of mitochondrial and chloroplast genome of Chlorella variabilis NC64A. Mitochondrial DNA A 2015, 27, 3128–3130. [Google Scholar]
- Korn, E.D.; Weisman, R.A. Phagocytosis of latex beads by Acanthomoeba. J. Cell Biol. 1967, 34, 219–227. [Google Scholar] [CrossRef]
- Murray, A.G.; Jackson, G.A. Viral dynamics: A model of the effects of size, shape, motion and abundance of single-celled planktonic organisms and other particles. Mar. Ecol. Prog. Ser. 1992, 89, 103–116. [Google Scholar] [CrossRef]
- Klose, T.; Rossmann, M.G. Structure of large dsDNA viruses. Biol. Chem. 2014, 395, 711–719. [Google Scholar] [CrossRef] [PubMed]
- Philippe, N.; Legendre, M.; Doutre, G.; Coute, Y.; Poirot, O.; Lescot, M.; Arslan, D.; Seltzer, V.; Bertaux, L.; Bruley, C.; et al. Pandoraviruses: Amoeba viruses with genomes up to 2.5 mb reaching that of parasitic eukaryotes. Science 2013, 341, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Legendre, M.; Bartoli, J.; Shmakova, L.; Jeudy, S.; Labadie, K.; Adrait, A.; Lescot, M.; Poirot, O.; Bertaux, L.; Bruley, C.; et al. Thirty-thousand-year-old distant relative of giant icosahedral DNA viruses with a Pandoravirus morphology. Proc. Natl. Acad. Sci. USA 2014, 111, 4274–4279. [Google Scholar] [CrossRef] [PubMed]
- Legendre, M.; Lartigue, A.; Bertaux, L.; Jeudy, S.; Bartoli, J.; Lescot, M.; Alempic, J.M.; Ramus, C.; Bruley, C.; Labadie, K.; et al. In-depth study of Mollivirus sibericum, a new 30,000-y-old giant virus infecting Acanthamoeba. Proc. Natl. Acad. Sci. USA 2015, 112, E5327–E5335. [Google Scholar] [CrossRef] [PubMed]
- Pearson, H. ‘Virophage’ suggests viruses are alive. Nature 2008, 454, 677. [Google Scholar] [CrossRef] [PubMed]
- Wilson, W.H.; Van Etten, J.L.; Allen, M.J. The phycodnaviridae: The story of how tiny giants rule the world. In Lesser Known Large dsDNA Viruses; VanEtten, J.L., Ed.; Springer Science & Business Media: Berlin, Germany, 2009; Volume 328, pp. 1–42. [Google Scholar]
- Xiao, C.; Rossmann, M.G. Structures of giant icosahedral eukaryotic dsDNA viruses. Curr. Opin. Virol. 2011, 1, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.A.; Chipman, P.R.; Battisti, A.J.; Bowman, V.D.; Renesto, P.; Raoult, D.; Rossmann, M.G. Cryo-electron microscopy of the giant mimivirus. J. Mol. Biol. 2005, 353, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Chothi, M.P.; Duncan, G.A.; Armirotti, A.; Abergel, C.; Gurnon, J.R.; Van Etten, J.L.; Bernardi, C.; Damonte, G.; Tonetti, M. Identification of an l-rhamnose synthetic pathway in two nucleocytoplasmic large DNA viruses. J. Virol. 2010, 84, 8829–8838. [Google Scholar] [CrossRef] [PubMed]
- Tonetti, M.; Chothi, M.P.; Abergel, C.; Seltzer, V.; Gurnon, J.; Van Etten, J.L. Glycosylation in nucleo-cytoplasmic large DNA viruses (NCLDV). FEBS J. 2011, 278, 420. [Google Scholar]
- Piacente, F.; Gaglianone, M.; Laugieri, M.E.; Tonetti, M.G. The autonomous glycosylation of large DNA viruses. Int. J. Mol. Sci. 2015, 16, 29315–29328. [Google Scholar] [CrossRef] [PubMed]
- Raoult, D. Viruses reconsidered. Scientist 2014, 28, 41. [Google Scholar]
- Rodrigues, R.A.L.; Silva, L.K.D.; Dornas, F.P.; de Oliveira, D.B.; Magalhaes, T.F.F.; Santos, D.A.; Costa, A.O.; Farias, L.D.; Magalhaes, P.P.; Bonjardim, C.A.; et al. Mimivirus fibrils are important for viral attachment to the microbial world by a diverse glycoside interaction repertoire. J. Virol. 2015, 89, 11812–11819. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.Z.; Xiang, Y.; Dunigan, D.D.; Klose, T.; Chipman, P.R.; Van Etten, J.L.; Rossmann, M.G. Three-dimensional structure and function of the Paramecium bursaria chlorella virus capsid. Proc. Natl. Acad. Sci. USA 2011, 108, 14837–14842. [Google Scholar] [CrossRef] [PubMed]
- Suzan-Monti, M.; La Scola, B.; Raoult, D. Genomic and evolutionary aspects of mimivirus. Virus Res. 2006, 117, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Maat, D.S.; Bale, N.J.; Hopmans, E.C.; Baudoux, A.C.; Damste, J.S.S.; Schouten, S.; Brussaard, C.P.D. Acquisition of intact polar lipids from the prymnesiophyte Phaeocystis globosa by its lytic virus PgV-07t. Biogeosciences 2014, 11, 185–194. [Google Scholar] [CrossRef]
- Martinez-Martinez, J.; Boere, A.; Gilg, I.C.; van Lent, J.W.M.; Witte, H.J.; van Bleijswijk, J.D.L.; Brussaard, C.P.D. New lipid envelop-containing dsDNA virus isolates infecting Micromonas pusilla reveal a separate phylogenetic group. Aquat. Microb. Ecol. 2015, 74, 17–28. [Google Scholar] [CrossRef]
- Mackinder, L.C.M.; Worthy, C.A.; Biggi, G.; Hall, M.; Ryan, K.P.; Varsani, A.; Harper, G.M.; Wilson, W.H.; Brownlee, C.; Schroeder, D.C. A unicellular algal virus, Emiliania huxleyi virus 86, exploits an animal-like infection strategy. J. Gen. Virol. 2009, 90, 2306–2316. [Google Scholar] [CrossRef] [PubMed]
- Frada, M.J.; Schatz, D.; Farstey, V.; Ossolinski, J.E.; Sabanay, H.; Ben-Dor, S.; Koren, I.; Vardi, A. Zooplankton may serve as transmission vectors for viruses infecting algal blooms in the ocean. Curr. Biol. 2014, 24, 2592–2597. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, K.B. Oxidative stress during viral infection: A review. Free Radic. Biol. Med. 1996, 21, 641–649. [Google Scholar] [CrossRef]
- Kang, M.; Duncan, G.A.; Kuszynski, C.; Oyler, G.; Zheng, J.Y.; Becker, D.F.; Van Etten, J.L. Chlorovirus PBCV-1 encodes an active copper-zinc superoxide dismutase. J. Virol. 2014, 88, 12541–12550. [Google Scholar] [CrossRef] [PubMed]
- Lartigue, A.; Burlat, B.; Coutard, B.; Chaspoul, F.; Claverie, J.M.; Abergel, C. The Megavirus chilensis Cu,Zn-superoxide dismutase: The first viral structure of a typical cellular copper chaperone-independent hyperstable dimeric enzyme. J. Virol. 2015, 89, 824–832. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.G.; Kelly, I.; Foster, L.J.; Suttle, C.A. The virion of Cafeteria roenbergensis virus (CroV) contains a complex suite of proteins for transcription and DNA repair. Virology 2014, 466, 82–94. [Google Scholar] [CrossRef] [PubMed]
- Hakim, M.; Ezerina, D.; Alon, A.; Vonshak, O.; Fass, D. Exploring ORFan domains in giant viruses: Structure of mimivirus sulfhydryl oxidase R596. PLoS ONE 2012, 7, e50649. [Google Scholar] [CrossRef] [PubMed]
- Apperizeller-Herzog, C.; Ellgaard, L. The human PDI family: Versatility packed into a single fold. Biochim. Biophys. Acta 2008, 1783, 535–548. [Google Scholar] [CrossRef] [PubMed]
- Ryser, H.J.P.; Levy, E.M.; Mandel, R.; Disciullo, G.J. Inhibition of human-immunodeficiency-virus infection by agents that interfere with thiol-disulfide interchange upon virus-receptor interaction. Proc. Natl. Acad. Sci. USA 1994, 91, 4559–4563. [Google Scholar] [CrossRef] [PubMed]
- Schelhaas, M.; Malmstrom, J.; Pelkmans, L.; Haugstetter, J.; Ellgaard, L.; Grunewald, K.; Helenius, A. Simian virus 40 depends on ER protein folding and quality control factors for entry into host cells. Cell 2007, 131, 516–529. [Google Scholar] [CrossRef] [PubMed]
- Moreira, D.; Brochier-Armanet, C. Giant viruses, giant chimeras: The multiple evolutionary histories of mimivirus genes. BMC Evol. Biol. 2008, 8, 12. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; La Scola, B.; Lepidi, H.; Raoult, D. Pneumonia in mice inoculated experimentally with Acanthamoeba polyphaga mimivirus. Microb. Pathog. 2007, 42, 56–61. [Google Scholar] [CrossRef] [PubMed]
- Correa, A.M.S.; Ainsworth, T.D.; Rosales, S.M.; Thurber, A.R.; Butler, C.R.; Vega Thurber, R.L. Viral outbreak in corals associated with an in situ bleaching event: Atypical herpes-like viruses and a new megavirus infecting symbiodinium. Front. Microbiol. 2016, 7, 127. [Google Scholar] [CrossRef] [PubMed]
- Popgeorgiev, N.; Boyer, M.; Fancello, L.; Monteil, S.; Robert, C.; Rivet, R.; Nappez, C.; Azza, S.; Chiaroni, J.; Raoult, D.; et al. Marseillevirus-like virus recovered from blood donated by asymptomatic humans. J. Infect. Dis. 2013, 208, 1042–1050. [Google Scholar] [CrossRef] [PubMed]
- Yolken, R.H.; Jones-Brando, L.; Dunigan, D.D.; Kannan, G.; Dickerson, F.; Severance, E.; Sabunciyan, S.; Talbot, C.C.; Prandovszky, E.; Gurnon, J.R.; et al. Chlorovirus ATCV-1 is part of the human oropharyngeal virome and is associated with changes in cognitive functions in humans and mice. Proc. Natl. Acad. Sci. USA 2014, 111, 16106–16111. [Google Scholar] [CrossRef] [PubMed]
- Iyer, L.M.; Aravind, L.; Koonin, E.V. Common origin of four diverse families of large eukaryotic DNA viruses. J. Virol. 2001, 75, 11720–11734. [Google Scholar] [CrossRef] [PubMed]
- Van Etten, J.L.; Meints, R.H. Giant viruses infecting algae. Annu. Rev. Microbiol. 1999, 53, 447–494. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Beato, R.; Salas, M.L.; Vinuela, E.; Salas, J. Role of the host cell nucleus in the replication of african swine fever virus DNA. Virology 1992, 188, 637–649. [Google Scholar] [CrossRef]
- Goorha, R. Frog virus-3 DNA-replication occurs in 2 stages. J. Virol. 1982, 43, 519–528. [Google Scholar] [PubMed]
- Moss, B. Poxviridae: The Viruses and Their Replication; Lippincott-Raven Publishers: Philadelphia, PA, USA, 1996; pp. 1163–1197. [Google Scholar]
- Koonin, E.V.; Yutin, N. Origin and evolution of eukaryotic large nucleo-cytoplasmic DNA viruses. Intervirology 2010, 53, 284–292. [Google Scholar] [CrossRef] [PubMed]
- Elde, N.C.; Child, S.J.; Eickbush, M.T.; Kitzman, J.O.; Rogers, K.S.; Shendure, J.; Geballe, A.P.; Malik, H.S. Poxviruses deploy genomic accordions to adapt rapidly against host antiviral defenses. Cell 2012, 150, 831–841. [Google Scholar] [CrossRef] [PubMed]
- Filee, J. Route of NCLDV evolution: The genoic accordion. Curr. Opin. Virol. 2013, 3, 595–599. [Google Scholar] [CrossRef] [PubMed]
- Desnues, C.; La Scola, B.; Yutin, N.; Fournous, G.; Robert, C.; Azza, S.; Jardot, P.; Monteil, S.; Campocasso, A.; Koonin, E.V.; et al. Provirophages and transpovirons as the diverse mobilome of giant viruses. Proc. Natl. Acad. Sci. USA 2012, 109, 18078–18083. [Google Scholar] [CrossRef] [PubMed]
- La Scola, B.; Desnues, C.; Pagnier, I.; Robert, C.; Barrassi, L.; Fournous, G.; Merchat, M.; Suzan-Monti, M.; Forterre, P.; Koonin, E.; et al. The virophage as a unique parasite of the giant mimivirus. Nature 2008, 455, 100–104. [Google Scholar] [CrossRef] [PubMed]
- Yutin, N.; Wolf, Y.I.; Raoult, D.; Koonin, E.V. Eukaryotic large nucleo-cytoplasmic DNA viruses: Clusters of orthologous genes and reconstruction of viral genome evolution. Virol. J. 2009, 6, 13. [Google Scholar] [CrossRef] [PubMed]
- Claverie, J.M.; Abergel, C. Mimivirus and its virophage. In Annual Review of Genetics; Annual Reviews: Palo Alto, CA, USA, 2009; Volume 43, pp. 49–66. [Google Scholar]
- Aherfi, S.; La Scola, B.; Pagnier, I.; Raoult, D.; Colson, P. The expanding family Marseilleviridae. Virology 2014, 466–467, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Colson, P.; de Lamballerie, X.; Fournous, G.; Raoult, D. Reclassification of giant viruses composing a fourth domain of life in the new order megavirales. Intervirology 2012, 55, 321–332. [Google Scholar] [CrossRef] [PubMed]
- Colson, P.; De Lamballerie, X.; Yutin, N.; Asgari, S.; Bigot, Y.; Bideshi, D.K.; Cheng, X.W.; Federici, B.A.; Van Etten, J.L.; Koonin, E.V.; et al. “Megavirales”, a proposed new order for eukaryotic nucleocytoplasmic large DNA viruses. Arch. Virol. 2013, 158, 2517–2521. [Google Scholar] [CrossRef] [PubMed]
- Krupovic, M.; Bamford, D.H. Virus evolution: How far does the double beta-barrel viral lineage extend? Nat. Rev. Microbiol. 2008, 6, 941–948. [Google Scholar] [CrossRef] [PubMed]
- Condit, R.C. Vaccinia, Inc.—Probing the functional substructure of poxviral replication factories. Cell Host Microbe 2007, 2, 205–207. [Google Scholar] [CrossRef] [PubMed]
- Netherton, C.; Moffat, K.; Brooks, E.; Wileman, T. A guide to viral inclusions, membrane rearrangements, factories, and viroplasm produced during virus replication. Adv. Virus Res. 2007, 70, 101–182. [Google Scholar] [PubMed]
- Mutsafi, Y.; Zauberman, N.; Sabanay, I.; Minsky, A. Vaccinia-like cytoplasmic replication of the giant mimivirus. Proc. Natl. Acad. Sci. USA 2010, 107, 5978–5982. [Google Scholar] [CrossRef] [PubMed]
- Boyer, M.; Yutin, N.; Pagnier, I.; Barrassi, L.; Fournous, G.; Espinosa, L.; Robert, C.; Azza, S.; Sun, S.Y.; Rossmann, M.G.; et al. Giant marseillevirus highlights the role of amoebae as a melting pot in emergence of chimeric microorganisms. Proc. Natl. Acad. Sci. USA 2009, 106, 21848–21853. [Google Scholar] [CrossRef] [PubMed]
- Netherton, C.L.; Wileman, T. Virus factories, double membrane vesicles and viroplasm generated in animal cells. Curr. Opin. Virol. 2011, 1, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Moniruzzaman, M.; LeCleir, G.R.; Brown, C.M.; Gobler, C.J.; Bidle, K.D.; Wilson, W.H.; Wilhelm, S.W. Genome of the brown tide virus (AaV), the little giant of the megaviridae, elucidates NCLDV genome expansion and host-virus coevolution. Virology 2014, 466–467, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Yutin, N.; Wolf, Y.I.; Koonin, E.V. Origin of giant viruses from smaller DNA viruses not from a fourth domain of cellular life. Virology 2014, 466–467, 38–52. [Google Scholar] [CrossRef] [PubMed]
- Moniruzzaman, M.; Gann, E.R.; LeCleir, G.R.; Kang, Y.; Gobler, C.J.; Wilhelm, S.W. Diversity and dynamics of algal megaviridae members during a harmful brown tide caused by the pelagophyte, Aureococcus anophagefferens. FEMS Microbiol. Ecol. 2016, 92, fiw058. [Google Scholar] [CrossRef] [PubMed]
- Yutin, N.; Koonin, E.V. Pandoraviruses are highly derived phycodnaviruses. Biol. Direct 2013, 8, 8. [Google Scholar] [CrossRef] [PubMed]
- Lwoff, A. The concept of a virus. J. Gen. Microbiol. 1957, 17, 239–253. [Google Scholar] [CrossRef] [PubMed]
- Twort, F.W. An investigation on the nature of ultra-microscopic viruses. Lancet 1915, 2, 1241–1243. [Google Scholar] [CrossRef]
- D’Herelle, F. Sur un microbe invisible antagonistic des bacilles dysenterique. C. R. Acad. Sci. Paris 1917, 165, 373–375. [Google Scholar]
- Lwoff, A.; Tournier, P. Classification of viruses. Annu. Rev. Microbiol. 1966, 20, 45–74. [Google Scholar] [CrossRef] [PubMed]
- Raoult, D.; La Scola, B.; Birtles, R. The discovery and characterization of mimivirus, the largest known virus and putative pneumonia agent. Clin. Infect. Dis. 2007, 45, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Raoult, D.; Audic, S.; Robert, C.; Abergel, C.; Renesto, P.; Ogata, H.; La Scola, B.; Suzan, M.; Claverie, J.M. The 1.2-megabase genome sequence of mimivirus. Science 2004, 306, 1344–1350. [Google Scholar] [CrossRef] [PubMed]
- Boyer, M.; Madoui, M.A.; Gimenez, G.; La Scola, B.; Raoult, D. Phylogenetic and phyletic studies of informational genes in genomes highlight existence of a 4th domain of life including giant viruses. PLoS ONE 2010, 5, e15530. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Inferring Phylogenies; Sinauer Associates: Sunderland, MA, USA, 2004. [Google Scholar]
- Williams, T.A.; Embley, T.M.; Heinz, E. Informational gene phylogenies do not support a fourth domain of life for nucleocytoplasmic large DNA viruses. PLoS ONE 2011, 6, e21080. [Google Scholar] [CrossRef] [PubMed]
- Doutre, G.; Philippe, N.; Abergel, C.; Claverie, J.M. Genome analysis of the first Marseilleviridae representative from Australia indicates that most of its genes contribute to virus fitness. J. Virol. 2014, 88, 14340–14349. [Google Scholar] [CrossRef] [PubMed]
- Chow, C.-E.T.; Winget, D.M.; White, R.A.; Hallam, S.J.; Suttle, C.A. Combining genomic sequencing methods to explore viral diversity and reveal potential virus-host interactions. Front. Microbiol. 2015, 6, 265. [Google Scholar] [CrossRef] [PubMed]
- La Scola, B.; Audic, S.; Robert, C.; Jungang, L.; de Lamballerie, X.; Drancourt, M.; Birtles, R.; Claverie, J.M.; Raoult, D. A giant virus in amoebae. Science 2003, 299, 2033. [Google Scholar] [CrossRef] [PubMed]
- Abrahão, J.S.; Dornas, F.P.; Silva, L.C.; Almeida, G.M.; Boratto, P.V.; Colson, P.; La Scola, B.; Kroon, E.G. Acanthamoeba polyphaga mimivirus and other giant viruses: An open field to outstanding discoveries. Virol. J. 2014, 11, 120. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Park, E.J.; Roh, S.W.; Bae, J.W. Diversity and abundance of single-stranded DNA viruses in human feces. Appl. Environ. Microbiol. 2011, 77, 8062–8070. [Google Scholar] [CrossRef] [PubMed]
- DeLong, J.P.; Al-Ameeli, Z.; Duncan, G.; Van Etten, J.L.; Dunigan, D.D. Predators catalyze an increase in chloroviruses by foraging on the symbiotic hosts of zoochlorellae. Proc. Natl. Acad. Sci. USA 2016, 113, 13780–13784. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, D.M.; Mushegian, A.R.; Dolja, V.V.; Koonin, E.V. New dimensions of the virus world discovered through metagenomics. Trends Microbiol. 2010, 18, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Williamson, S.J.; Allen, L.Z.; Lorenzi, H.A.; Fadrosh, D.W.; Brami, D.; Thiagarajan, M.; McCrow, J.P.; Tovchigrechko, A.; Yooseph, S.; Venter, J.C. Metagenomic exploration of viruses throughout the Indian Ocean. PLoS ONE 2012, 7, e42047. [Google Scholar] [CrossRef] [PubMed]
- Khalil, J.Y.B.; Langlois, T.; Andreani, J.; Sorraing, J.-M.; Raoult, D.; Carmoin, L.; La Scola, B. Flow cytometry sorting to separate viable giant viruses from amoeba co-culture supernatants. Front. Cell. Infect. Microbiol. 2016, 6, 202. [Google Scholar] [CrossRef] [PubMed]
- Martinez Martinez, J.; Poulton, N.J.; Stepanauskas, R.; Sieracki, M.E.; Wilson, W.H. Targeted sorting of single virus-infected cells of the coccolithophore Emiliania huxleyi. PLoS ONE 2011, 6, e22520. [Google Scholar] [CrossRef] [PubMed]
- Zablocki, O.; van Zyl, L.; Adriaenssens, E.M.; Rubagotti, E.; Tuffin, M.; Cary, S.C.; Cowana, D. High-level diversity of tailed phages, eukaryote-associated viruses, and virophage-like elements in the metaviromes of Antarctic soils. Appl. Environ. Microbiol. 2014, 80, 10. [Google Scholar] [CrossRef] [PubMed]
- Correa, A.M.; Welsh, R.M.; Vega Thurber, R.L. Unique nucleocytoplasmic dsDNA and +ssRNA viruses are associated with the dinoflagellate endosymbionts of corals. ISME J. 2013, 7, 13–27. [Google Scholar] [CrossRef] [PubMed]
- Lysholm, F.; Wetterbom, A.; Lindau, C.; Darban, H.; Bjerkner, A.; Fahlander, K.; Lindberg, A.M.; Persson, B.; Allander, T.; Andersson, B. Characterization of the viral microbiome in patients with severe lower respiratory tract infections, using metagenomic sequencing. PLoS ONE 2012, 7, e30875. [Google Scholar] [CrossRef] [PubMed]
- Brussaard, C.P.D.; Kuipers, B.; Veldhuis, M.J.W. A mesocosm study of Phaeocystis globosa population dynamics. Harmful Algae 2005, 4, 859–874. [Google Scholar] [CrossRef]
- Castberg, T.; Thyrhaug, R.; Larsen, A.; Sandaa, R.-A.; Heldal, M.; Van Etten, J.L.; Bratbak, G. Isolation and characterization of a virus that infects Emiliania huxleyi (Haptophyta). J. Phycol. 2002, 38, 767–774. [Google Scholar] [CrossRef]
- Gobler, C.J.; Anderson, O.R.; Gastrich, M.D.; Wilhelm, S.W. Ecological aspects of viral infection and lysis in the harmful brown tide alga Aureococcus anophagefferens. Aquat. Microb. Ecol. 2007, 47, 25–36. [Google Scholar] [CrossRef]
- Nagasaki, K.; Tarutani, K.; Yamaguchi, M. Growth characteristics of Heterosigma akashiwo virus and its possible use as a microbiological agent for red tide control. Appl. Environ. Microbiol. 1999, 65, 898–902. [Google Scholar] [PubMed]
- Brussaard, C.P.D.; Gast, G.J.; van Duyl, F.C.; Riegmen, R. Impact of phytoplankton bloom magnitude on a pelagic microbial food web. Mar. Ecol. Prog. Ser. 1996, 144, 211–221. [Google Scholar] [CrossRef]
- Weitz, J.S.; Wilhelm, S.W. Ocean viruses and their effects on microbial communities and biogeochemical cycles. F1000 Biol. Rep. 2012, 4, 17. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, S.W.; Suttle, C.A. Viruses and nutrient cycles in the sea. Bioscience 1999, 49, 781–788. [Google Scholar] [CrossRef]
- Nagasaki, K.; Bratbak, G. Isolation of viruses infecting photosynthetic and nonphotosynthetic protists. In Manual of Aquatic Viral Ecology; Wilhelm, S.W., Weinbauer, M.G., Suttle, C.A., Eds.; ASLO: Waco, TX, USA, 2010; pp. 92–101. [Google Scholar]
- Brodie, J.; Zuccarello, G.C. Systematics of the species rich algae: Red algal classificiation, phylogeny and speciation. In Reconstructing the Tree of Life. Taxonomy and Systematics of Species Rich Taxa; Hodkinson, T.R., Parnell, J.A.N., Eds.; CRC Press: New York, NY, USA, 2006; pp. 323–336. [Google Scholar]
- Short, S.M. The ecology of viruses that infect eukaryotic algae. Environ. Microbiol. 2012, 14, 2253–2271. [Google Scholar] [CrossRef] [PubMed]
- Johannessen, T.V.; Bratbak, G.; Larsen, A.; Ogata, H.; Egge, E.S.; Edvardsen, B.; Eikrem, W.; Sandaa, R.A. Characterisation of three novel giant viruses reveals huge diversity among viruses infecting Prymnesiales (Haptophyta). Virology 2015, 476, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, S.W.; Coy, S.R.; Gann, E.R.; Moniruzzaman, M.; Stough, J.M.A. Standing on the shoulders of giant viruses: 5 lessons learned about large viruses infecting small eukaryotes and the opportunities they create. PLoS Pathog. 2016, 12, e1005752. [Google Scholar] [CrossRef] [PubMed]
- Moniruzzaman, M.; Wurch, L.L.; Alexander, H.; Dyhrman, S.T.; Gobler, C.J.; Wilhelm, S.W. Virus-host infection dynamics of marine single-celled eukaryotes resolved from metatranscriptomics. bioRxiv 2016. [Google Scholar] [CrossRef]
- Van Etten, J.L.; Lane, L.C.; Meints, R.H. Viruses and viruslike particles of eukaryotic algae. Microbiol. Rev. 1991, 55, 586–620. [Google Scholar] [PubMed]
- Reteno, D.G.; Benamar, S.; Kahlil, J.B.; Andreani, J.; Armstrong, N.; Klose, T.; Rossmann, M.G.; Colson, P.; Raoult, D.; La Scola, B. Faustovirus, an asfarvirus-related new lineage of giant viruses infecting amoebae. J. Virol. 2015, 89, 6585–6594. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.G.; Allen, M.J.; Wilson, W.H.; Suttle, C.A. Giant virus with a remarkable complement of genes infects marine zooplankton. Proc. Natl. Acad. Sci. USA 2010, 107, 19508–19513. [Google Scholar] [CrossRef] [PubMed]
- Schnepf, E.; Soeder, C.J.; Hegewald, E. Polyhedral virus-like particles lysing the aquatic phycomycete Aphedlidium sp., a parasite of the grean algae Scenedesmus armatus. Virology 1970, 42, 482–487. [Google Scholar] [CrossRef]
- Pickett-Heaps, J.D. A possible virus infection in the green alga Oedogonium. J. Phycol. 1972, 8, 44–47. [Google Scholar] [CrossRef]
- Toth, R.; Wilce, R.T. Viruslike particles in the marine alga Chorda tomentosa lyngye (Phaeophyceae). J. Phycol. 1972, 8, 126–130. [Google Scholar]
- Baker, J.R.J.; Evans, L.V. The ship fouling alga Ectocarpus. Protoplasma 1973, 77, 1–13. [Google Scholar] [CrossRef]
- Clitheroe, S.B.; Evans, L.V. Virus like particles in the brown algae Ectocarpus. J. Ultrastruct. Res. 1974, 49, 211–217. [Google Scholar] [CrossRef]
- Swale, E.M.F.; Belcher, J.H. A light and electron microscope study of the colourless flagellate Aulacomonas skuja. Arch. Microbiol. 1973, 92, 91–103. [Google Scholar] [CrossRef]
- Markey, D.R. A possivle virus infection in the brown alga Phlaiella littoralis. Protoplasma 1974, 80, 223–232. [Google Scholar] [CrossRef] [PubMed]
- Moestrup, O.; Thomsen, H.A. An ultrastructural study of the flagellate Phyramimonas orientalis with particular emphasis on golgi apparatus activity and the flagellar apparatus. Protoplasma 1974, 81, 247–269. [Google Scholar] [CrossRef]
- Gibbs, A.; Skotnicki, A.H.; Gardiner, J.E.; Walker, E.S.; Hollings, M. A tabamovirus of a green alga. Virology 1975, 64, 571–574. [Google Scholar] [CrossRef]
- Oliveira, L.; Bisalputra, T. A virus infection in the brown alga Sorocarpus uvaeformis (Lyngbye) pringsheim (Phaeophyta, Ecocarpales). Ann. Bot. 1978, 42, 439–445. [Google Scholar] [CrossRef]
- Sicko-Goad, L.; Walker, G. Viroplasm and large virus-like particles in the dinoflagellate Gymnodiunium uberrimum. Protoplasma 1979, 99, 203–210. [Google Scholar] [CrossRef]
- Dodds, J.A.; Cole, A. Microscopy and biology of Uronema gigas, a filamentous eucaryotic green alga, and its associated tailed virus-like particle. Virology 1980, 100, 156–165. [Google Scholar] [CrossRef]
- Preisig, H.R.; Hibberd, D.J. Virus-like particles and endophytic bacteria in Paraphysomonas and Chromophysomonas (Chrysophyceae). Nord. J. Bot. 1984, 4, 279–285. [Google Scholar] [CrossRef]
- Gowing, M.M. Large virus-like particles from vacuoles of phaeodarian radiolarians and from other marine samples. Mar. Ecol. Prog. Ser. 1993, 101, 33–43. [Google Scholar] [CrossRef]
- Smayda, T.J. Complexity in the eutrophication-harmful algal bloom relationship, with comment on the important of grazing. Harmful Algae 2008, 8, 140–151. [Google Scholar] [CrossRef]
- Filee, J.; Pouget, N.; Chandler, M. Phylogenetic evidence for extensive lateral acquisition of cellular genes by nucleocytoplasmic large DNA viruses. BMC Evol. Biol. 2008, 8, 320. [Google Scholar] [CrossRef] [PubMed]
- Monier, A.; Pagarete, A.; de Vargas, C.; Allen, M.J.; Read, B.; Claverie, J.-M.; Ogata, H. Horizontal gene transfer of an entire metabolic pathway between a eukaryotic alga and its DNA virus. Genome Res. 2009, 19, 1441–1449. [Google Scholar] [CrossRef] [PubMed]
- Rosenwasser, S.; Mausz, M.A.; Schatz, D.; Sheyn, U.; Malitsky, S.; Aharoni, A.; Weinstock, E.; Tzfadia, O.; Ben-Dor, S.; Feldmesser, E.; et al. Rewiring host lipid metabolism by large viruses determines the fate of Emiliania huxleyi, a bloom-forming alga in the ocean. Plant Cell Online 2014, 26, 2689–2707. [Google Scholar] [CrossRef] [PubMed]
- Vardi, A.; Van Mooy, B.A.S.; Fredricks, H.F.; Popendorf, K.J.; Ossolinski, J.E.; Haramaty, L.; Bidle, K.D. Viral glycosphingolipids induce lytic infection and cell death in marine phytoplankton. Science 2009, 326, 861–865. [Google Scholar] [CrossRef] [PubMed]
- Filee, J.; Siguier, P.; Chandler, M. I am what I eat and I eat what I am: Acquisition of bacterial genes by giant viruses. Trends Genet. 2007, 23, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Filee, J. Multiple occurrences of giant virus core genes acquired by eukaryotic genomes: The visible part of the iceberg? Virology 2014, 466–467, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Blanc, G.; Duncan, G.; Agarkova, I.; Borodovsky, M.; Gurnon, J.; Kuo, A.; Lindquist, E.; Lucas, S.; Pangilinan, J.; Polle, J.; et al. The Chlorella variabilis NC64A genome reveals adaptation to photosymbiosis, coevolution with viruses, and cryptic sex. Plant Cell 2010, 22, 2943–2955. [Google Scholar] [CrossRef] [PubMed]
- Blanc, G.; Gallot-Lavallee, L.; Maumus, F. Provirophages in the bigelowiella genome bear testimony to past encounters with giant viruses. Proc. Natl. Acad. Sci. USA 2015, 112, E5318–5326. [Google Scholar] [CrossRef] [PubMed]
- Read, B.A.; Kegel, J.; Klute, M.J.; Kuo, A.; Lefebvre, S.C.; Maumus, F.; Mayer, C.; Miller, J.; Monier, A.; Salamov, A.; et al. Pan genome of the phytoplankton Emiliania underpins its global distribution. Nature 2013, 499, 209–213. [Google Scholar] [CrossRef] [PubMed]
- Maumus, F.; Epert, A.; Nogué, F.; Blanc, G. Plant genomes enclose footprints of past infections by giant virus relatives. Nat. Commun. 2014, 5, 4268. [Google Scholar] [CrossRef] [PubMed]
- Breitbart, M. Marine viruses: Truth or dare. Ann Rev Mar Sci 2012, 4, 425–448. [Google Scholar] [CrossRef] [PubMed]
- Frada, M.; Probert, I.; Allen, M.J.; Wilson, W.H.; de Vargas, C. The “cheshire cat” escape strategy of the coccolithophore Emiliania huxleyi in response to viral infection. Proc. Natl. Acad. Sci. USA 2008, 105, 15944–15949. [Google Scholar] [CrossRef] [PubMed]
- Yau, S.; Lauro, F.M.; DeMaere, M.Z.; Brown, M.V.; Thomas, T.; Raftery, M.J.; Andrews-Pfannkoch, C.; Lewis, M.; Hoffman, J.M.; Gibson, J.A.; et al. Virophage control of antarctic algal host-virus dynamics. Proc. Natl. Acad. Sci. USA 2011, 108, 6163–6168. [Google Scholar] [CrossRef] [PubMed]
- Campos, R.K.; Boratto, P.V.; Assis, F.L.; Aguiar, E.R.; Silva, L.C.; Albarnaz, J.D.; Dornas, F.P.; Trindade, G.S.; Ferreira, P.P.; Marques, J.T.; et al. Samba virus: A novel mimivirus from a giant rain forest, the Brazilian amazon. Virol. J. 2014, 11, 95. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.G.; Suttle, C.A. A virophage at the origin of large DNA transposons. Science 2011, 332, 231–234. [Google Scholar] [CrossRef] [PubMed]
- Gaia, M.; Benamar, S.; Boughalmi, M.; Pagnier, I.; Croce, O.; Colson, P.; Raoult, D.; La Scola, B. Zamilon, a novel virophage with Mimiviridae host specificity. PLoS ONE 2014, 9, e94923. [Google Scholar] [CrossRef] [PubMed]
- Levasseur, A.; Bekliz, M.; Chabrière, E.; Pontarotti, P.; La Scola, B.; Raoult, D. Mimivire is a defence system in mimivirus that confers resistance to virophage. Nature 2016, 531, 249–252. [Google Scholar] [CrossRef] [PubMed]
- Santini, S.; Jeudy, S.; Bartoli, J.; Poirot, O.; Lescot, M.; Abergel, C.; Barbe, V.; Wommack, K.E.; Noordeloos, A.A.M.; Brussaard, C.P.D.; et al. Genome of Phaeocystis globosa virus PgV-16t highlights the common ancestry of the largest known DNA viruses infecting eukaryotes. Proc. Natl. Acad. Sci. USA 2013, 110, 10800–10805. [Google Scholar] [CrossRef] [PubMed]
- Claverie, J.M.; Abergel, C. CRISPR-CAS-like system in giant viruses: Why mimivire is not likely to be an adaptive immune system. Virol. Sin. 2016, 31, 193–196. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.B.; Weitz, J.S.; Wilhelm, S.W. Viral ecology comes of age. Environ. Microbiol. Repo. 2017, 9, 33–35. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).