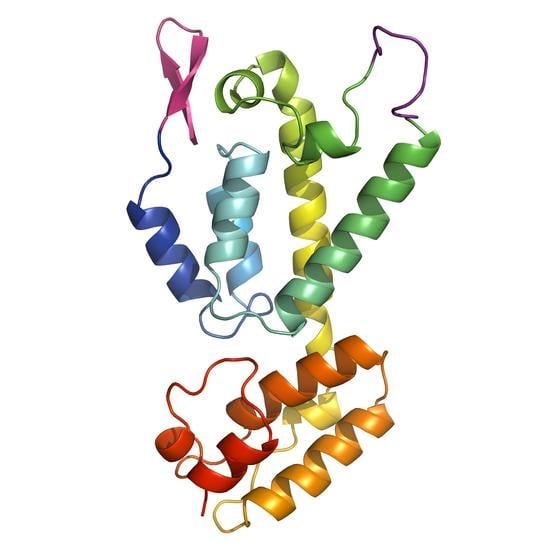
Crystal Structure of the Full-Length Feline Immunodeficiency Virus Capsid Protein Shows an N-Terminal β-Hairpin in the Absence of N-Terminal Proline
Abstract
:1. Introduction
2. Materials and Methods
2.1. Construction of Recombinant Plasmid Encoding the FIV Capsid Protein
2.2. Expression and Purification of FIV CA Protein
2.3. Removal of the 6 × His Tag
2.4. Crystallization of the FIV CA Protein
2.5. X-ray Data Collection and Structure Determination
3. Results
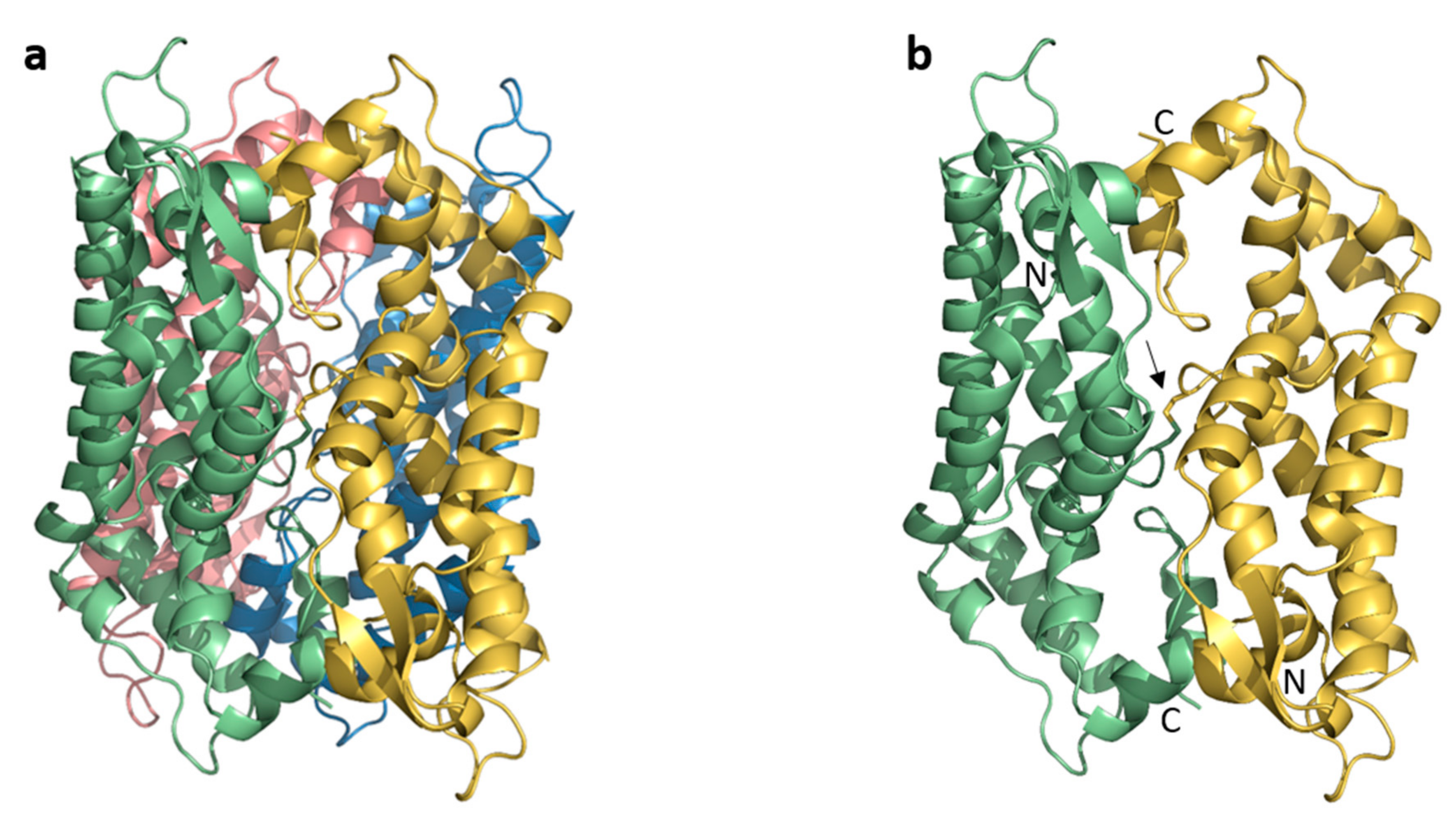
3.1. Asymmetric Unit and Crystal Packing
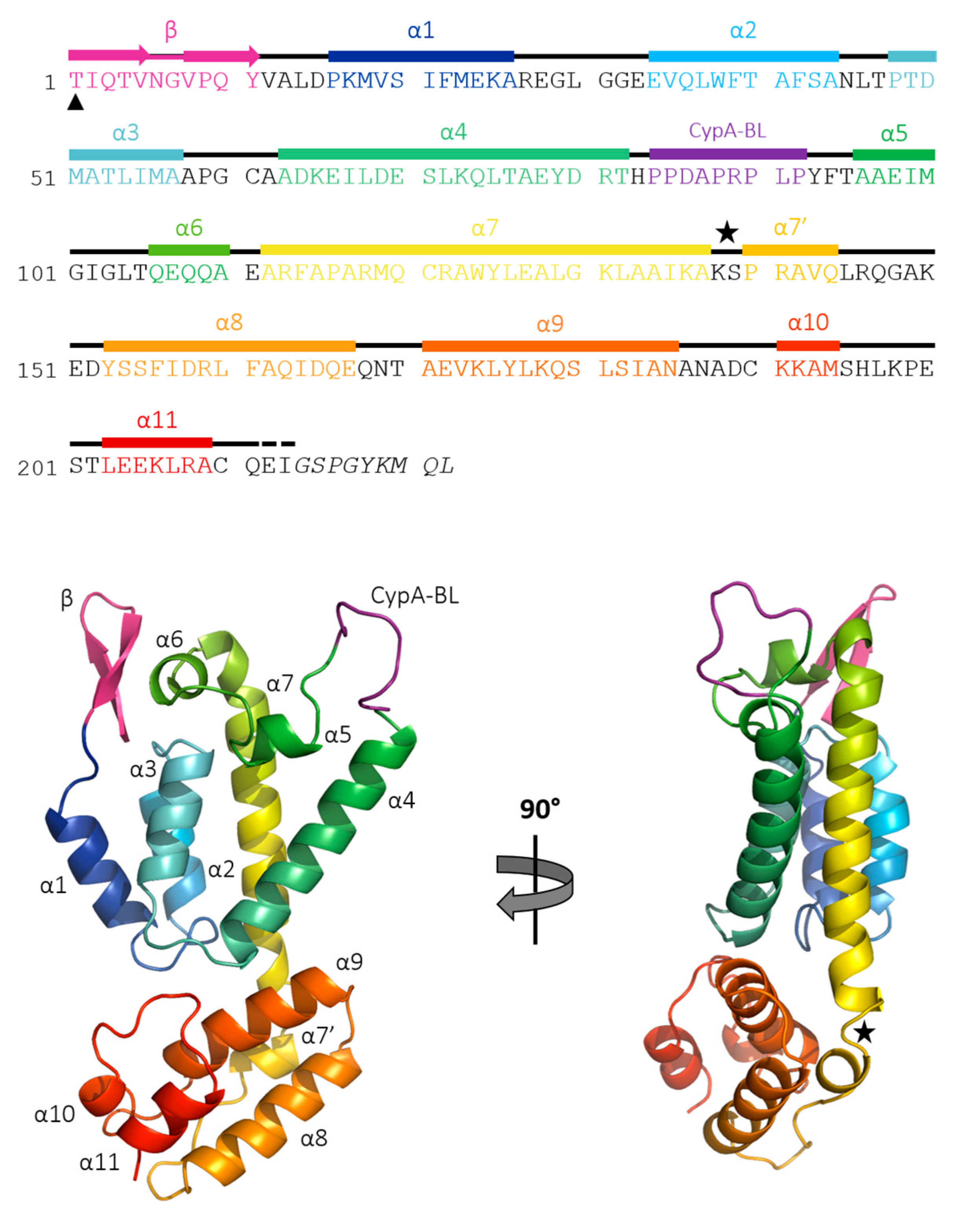
3.2. The Crystal Structure of Full Length FIV CA
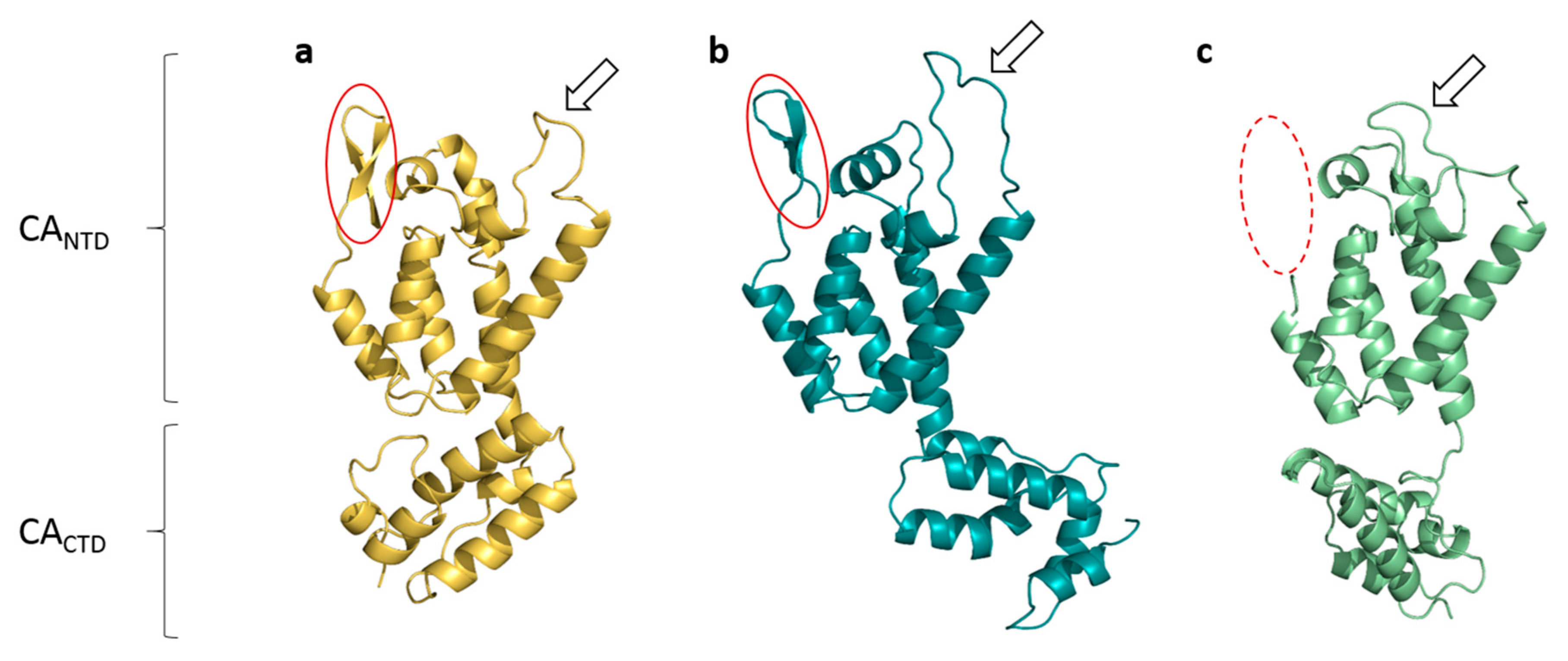
3.3. Comparisons with Known Structures of Lentiviral CAs
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lee, J.; Malmberg, J.L.; Wood, B.A.; Hladky, S.; Troyer, R.; Roelke, M.; Cunningham, M.; McBride, R.; Vickers, W.; Boyce, W.; et al. Feline immunodeficiency virus cross-species transmission: Implications for emergence of new lentiviral infections. J. Virol. 2016, 91. [Google Scholar] [CrossRef] [PubMed]
- Bendinelli, M.; Pistello, M.; Lombardi, S.; Poli, A.; Garzelli, C.; Matteucci, D.; Ceccherini-Nelli, L.; Malvaldi, G.; Tozzini, F. Feline immunodeficiency virus: An interesting model for AIDS studies and an important cat pathogen. Clin. Microbiol. Rev. 1995, 8, 87–112. [Google Scholar] [PubMed]
- Troyer, J.L.; Pecon-Slattery, J.; Roelke, M.E.; Johnson, W.; VandeWoude, S.; Vazquez-Salat, N.; Brown, M.; Frank, L.; Woodroffe, R.; Winterbach, C.; et al. Seroprevalence and genomic divergence of circulating strains of feline immunodeficiency virus among Felidae and Hyaenidae species. J. Virol. 2005, 79, 8282–8294. [Google Scholar] [CrossRef] [PubMed]
- Clements, J.E.; Zink, M.C. Molecular biology and pathogenesis of animal lentivirus infections. Clin. Microbiol. Rev. 1996, 9, 100–117. [Google Scholar] [PubMed]
- Luttge, B.G.; Freed, E.O. FIV Gag: Virus assembly and host-cell interactions. Vet. Immunol. Immunopathol. 2010, 134, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Burkhard, M.J.; Dean, G.A. Transmission and immunopathogenesis of FIV in cats as a model for HIV. Curr. HIV Res. 2003, 1, 15–29. [Google Scholar] [CrossRef] [PubMed]
- Elder, J.H.; Lin, Y.C.; Fink, E.; Grant, C.K. Feline immunodeficiency virus (FIV) as a model for study of lentivirus infections: Parallels with HIV. Curr. HIV Res. 2010, 8, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Kenyon, J.C.; Lever, A.M. The molecular biology of feline immunodeficiency virus (FIV). Viruses 2011, 3, 2192–2213. [Google Scholar] [CrossRef] [PubMed]
- Abdusetir Cerfoglio, J.C.; Gonzalez, S.A.; Affranchino, J.L. Structural elements in the Gag polyprotein of feline immunodeficiency virus involved in Gag self-association and assembly. J. Gen. Virol. 2014, 95, 2050–2059. [Google Scholar] [CrossRef] [PubMed]
- Freed, E.O. HIV-1 gag proteins: Diverse functions in the virus life cycle. Virology 1998, 251, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Pal, R.; Reitz, M.S., Jr.; Tschachler, E.; Gallo, R.C.; Sarngadharan, M.G.; Veronese, F.D. Myristoylation of gag proteins of HIV-1 plays an important role in virus assembly. AIDS Res. Hum. Retrovir. 1990, 6, 721–730. [Google Scholar] [CrossRef] [PubMed]
- Provitera, P.; Bouamr, F.; Murray, D.; Carter, C.; Scarlata, S. Binding of equine infectious anemia virus matrix protein to membrane bilayers involves multiple interactions. J. Mol. Biol. 2000, 296, 887–898. [Google Scholar] [CrossRef] [PubMed]
- Elder, J.H.; Schnolzer, M.; Hasselkus-Light, C.S.; Henson, M.; Lerner, D.A.; Phillips, T.R.; Wagaman, P.C.; Kent, S.B. Identification of proteolytic processing sites within the Gag and Pol polyproteins of feline immunodeficiency virus. J. Virol. 1993, 67, 1869–1876. [Google Scholar] [PubMed]
- Brown, L.A.; Cox, C.; Baptiste, J.; Summers, H.; Button, R.; Bahlow, K.; Spurrier, V.; Kyser, J.; Luttge, B.G.; Kuo, L.; et al. NMR structure of the myristylated feline immunodeficiency virus matrix protein. Viruses 2015, 7, 2210–2229. [Google Scholar] [CrossRef] [PubMed]
- Provitera, P.; El-Maghrabi, R.; Scarlata, S. The effect of HIV-1 Gag myristoylation on membrane binding. Biophys. Chem. 2006, 119, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Erickson-Viitanen, S.; Manfredi, J.; Viitanen, P.; Tribe, D.E.; Tritch, R.; Hutchison, C.A., 3rd; Loeb, D.D.; Swanstrom, R. Cleavage of HIV-1 gag polyprotein synthesized in vitro: Sequential cleavage by the viral protease. AIDS Res. Hum. Retrovir. 1989, 5, 577–591. [Google Scholar] [CrossRef] [PubMed]
- Campbell, E.M.; Hope, T.J. HIV-1 capsid: The multifaceted key player in HIV-1 infection. Nat. Rev. Microbiol. 2015, 13, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Ganser-Pornillos, B.K.; von Schwedler, U.K.; Stray, K.M.; Aiken, C.; Sundquist, W.I. Assembly properties of the human immunodeficiency virus type 1 CA protein. J. Virol. 2004, 78, 2545–2552. [Google Scholar] [CrossRef] [PubMed]
- Nath, M.D.; Peterson, D.L. In vitro assembly of feline immunodeficiency virus capsid protein: Biological role of conserved cysteines. Arch Biochem. Biophys. 2001, 392, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Gelderblom, H.R.; Ozel, M.; Pauli, G. Morphogenesis and morphology of HIV. Structure-function relations. Arch. Virol. 1989, 106, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.; Loeliger, E.; Kinde, I.; Kyere, S.; Mayo, K.; Barklis, E.; Sun, Y.; Huang, M.; Summers, M.F. Antiviral inhibition of the HIV-1 capsid protein. J. Mol. Biol. 2003, 327, 1013–1020. [Google Scholar] [CrossRef]
- Jin, Z.; Jin, L.; Peterson, D.L.; Lawson, C.L. Model for lentivirus capsid core assembly based on crystal dimers of EIAV. p26. J. Mol. Biol. 1999, 286, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Momany, C.; Kovari, L.C.; Prongay, A.J.; Keller, W.; Gitti, R.K.; Lee, B.M.; Gorbalenya, A.E.; Tong, L.; McClure, J.; Ehrlich, L.S.; et al. Crystal structure of dimeric HIV-1 capsid protein. Nat. Struct. Biol. 1996, 3, 763–770. [Google Scholar] [CrossRef] [PubMed]
- Pornillos, O.; Ganser-Pornillos, B.K.; Yeager, M. Atomic-level modelling of the HIV capsid. Nature 2011, 469, 424–427. [Google Scholar] [CrossRef] [PubMed]
- Mattei, S.; Glass, B.; Hagen, W.J.; Krausslich, H.G.; Briggs, J.A. The structure and flexibility of conical HIV-1 capsids determined within intact virions. Science 2016, 354, 1434–1437. [Google Scholar] [CrossRef] [PubMed]
- Jacques, D.A.; McEwan, W.A.; Hilditch, L.; Price, A.J.; Towers, G.J.; James, L.C. HIV-1 uses dynamic capsid pores to import nucleotides and fuel encapsidated DNA synthesis. Nature 2016, 536, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Ganser-Pornillos, B.K.; Yeager, M.; Sundquist, W.I. The structural biology of HIV assembly. Curr. Opin. Struct. Biol. 2008, 18, 203–217. [Google Scholar] [CrossRef] [PubMed]
- Pornillos, O.; Ganser-Pornillos, B.K.; Banumathi, S.; Hua, Y.; Yeager, M. Disulfide bond stabilization of the hexameric capsomer of human immunodeficiency virus. J. Mol. Biol. 2010, 401, 985–995. [Google Scholar] [CrossRef] [PubMed]
- Berthet-Colominas, C.; Monaco, S.; Novelli, A.; Sibai, G.; Mallet, F.; Cusack, S. Head-to-tail dimers and interdomain flexibility revealed by the crystal structure of HIV-1 capsid protein (p24) complexed with a monoclonal antibody Fab. EMBO J. 1999, 18, 1124–1136. [Google Scholar] [CrossRef] [PubMed]
- Bharat, T.A.; Castillo Menendez, L.R.; Hagen, W.J.; Lux, V.; Igonet, S.; Schorb, M.; Schur, F.K.; Krausslich, H.G.; Briggs, J.A. Cryo-electron microscopy of tubular arrays of HIV-1 Gag resolves structures essential for immature virus assembly. Proc. Natl. Acad. Sci. USA 2014, 111, 8233–8238. [Google Scholar] [CrossRef] [PubMed]
- Mammano, F.; Ohagen, A.; Hoglund, S.; Gottlinger, H.G. Role of the major homology region of human immunodeficiency virus type 1 in virion morphogenesis. J. Virol. 1994, 68, 4927–4936. [Google Scholar] [PubMed]
- Bocanegra, R.; Fuertes, M.A.; Rodriguez-Huete, A.; Neira, J.L.; Mateu, M.G. Biophysical analysis of the MHR motif in folding and domain swapping of the HIV capsid protein C-terminal domain. Biophys. J. 2015, 108, 338–349. [Google Scholar] [CrossRef] [PubMed]
- Khwaja, A.; Galilee, M.; Marx, A.; Alian, A. Structure of FIV capsid C-terminal domain demonstrates lentiviral evasion of genetic fragility by coevolved substitutions. Sci. Rep. 2016, 6, 24957. [Google Scholar] [CrossRef] [PubMed]
- Serriere, J.; Fenel, D.; Schoehn, G.; Gouet, P.; Guillon, C. Biophysical characterization of the feline immunodeficiency virus p24 capsid protein conformation and in vitro capsid assembly. PLoS ONE 2013, 8, e56424. [Google Scholar] [CrossRef] [PubMed]
- Kabsch, W. XDS. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Keegan, R.M.; Winn, M.D. MrBUMP: An automated pipeline for molecular replacement. Acta Crystallogr. Sect. D Biol. Crystallogr. 2008, 64, 119–124. [Google Scholar] [CrossRef] [PubMed]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.; McCoy, A.; et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. Sect. D Biol. Crystallogr. 2011, 67, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Goldstone, D.C.; Yap, M.W.; Robertson, L.E.; Haire, L.F.; Taylor, W.R.; Katzourakis, A.; Stoye, J.P.; Taylor, I.A. Structural and functional analysis of prehistoric lentiviruses uncovers an ancient molecular interface. Cell Host Microbe 2010, 8, 248–259. [Google Scholar] [CrossRef] [PubMed]
- Adams, P.D.; Afonine, P.V.; Bunkóczi, G.; Chen, V.B.; Echols, N.; Headd, J.J.; Hung, L.-W.; Jain, S.; Kapral, G.J.; Grosse Kunstleve, R.W.; et al. The Phenix software for automated determination of macromolecular structures. Methods 2011, 55, 94–106. [Google Scholar] [CrossRef] [PubMed]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and development of Coot. Acta Crystallogr. Sec. D Biol. Crystallogr. 2010, 66, 486–501. [Google Scholar] [CrossRef] [PubMed]
- DeLano, Warren. The PyMol Molecular Graphics Systems. Available online: http://pymol.sourceforge.net/overview/index.htm (accessed on 8 December 2017).
- Krissinel, E.; Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 2007, 372, 774–797. [Google Scholar] [CrossRef] [PubMed]
- Du, S.; Betts, L.; Yang, R.; Shi, H.; Concel, J.; Ahn, J.; Aiken, C.; Zhang, P.; Yeh, J.I. Structure of the HIV-1 Full-Length Capsid in a Conformationally-Trapped Unassembled State Induced by Small-Molecule Binding. J. Mol. Biol. 2011, 406, 371–386. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Perilla, J.R.; Yufenyuy, E.L.; Meng, X.; Chen, B.; Ning, J.; Ahn, J.; Gronenborn, A.M.; Schulten, K.; Aiken, C.; et al. Mature HIV-1 capsid structure by cryo-electron microscopy and all-atom molecular dynamics. Nature 2013, 497, 643–646. [Google Scholar] [CrossRef] [PubMed]
- Gres, A.T.; Kirby, K.A.; KewalRamani, V.N.; Tanner, J.J.; Pornillos, O.; Sarafianos, S.G. X-ray crystal structures of native HIV-1 capsid protein reveal conformational variability. Science 2015, 349, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Byeon, I.J.; Meng, X.; Jung, J.; Zhao, G.; Yang, R.; Ahn, J.; Shi, J.; Concel, J.; Aiken, C.; Zhang, P.; et al. Structural convergence between Cryo-EM and NMR reveals intersubunit interactions critical for HIV-1 capsid function. Cell 2009, 139, 780–790. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhou, L.; Chen, L.; Dastidar, S.G.; Verma, C.; Li, J.; Tan, D.; Beuerman, R. Effect of structural parameters of peptides on dimer formation and highly oxidized side products in the oxidation of thiols of linear analogues of human beta-defensin 3 by DMSO. J. Pept. Sci. 2009, 15, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Gamble, T.R.; Yoo, S.; Vajdos, F.F.; von Schwedler, U.K.; Worthylake, D.K.; Wang, H.; McCutcheon, J.P.; Sundquist, W.I.; Hill, C.P. Structure of the carboxyl-terminal dimerization domain of the HIV-1 capsid protein. Science 1997, 278, 849–853. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.; Ndassa, Y.; Summers, M.F. Structure of the N-terminal 283-residue fragment of the immature HIV-1 Gag polyprotein. Nat. Struct. Biol. 2002, 9, 537–543. [Google Scholar] [CrossRef] [PubMed]
- Cortines, J.R.; Monroe, E.B.; Kang, S.; Prevelige, P.E. A retroviral chimeric capsid protein reveals the role of the N-terminal β-hairpin in mature core assembly. J. Mol. Biol. 2011, 410, 641–652. [Google Scholar] [CrossRef] [PubMed]
- Konno, M.; Ito, M.; Hayano, T.; Takahashi, N. The substrate-binding site in Escherichia coli cyclophilin A preferably recognizes a cis-proline isomer or a highly distorted form of the trans isomer. J. Mol. Biol. 1996, 256, 897–908. [Google Scholar] [CrossRef] [PubMed]
- Lin, T.Y.; Emerman, M. Cyclophilin A interacts with diverse lentiviral capsids. Retrovirology 2006, 3, 70. [Google Scholar] [CrossRef] [PubMed]
- Serriere, J.; Robert, X.; Perez, M.; Gouet, P.; Guillon, C. Biophysical characterization and crystal structure of the Feline Immunodeficiency Virus p15 matrix protein. Retrovirology 2013, 10, 64. [Google Scholar] [CrossRef] [PubMed]

| Data Collection | FIV CA |
|---|---|
| Space group | C121 |
| Unit cell parameters | |
| a, b, c (Å) | 122, 74, 77 |
| α, β, γ (°) | 90, 128, 90 |
| Resolution range (Å) | 38.5–1.6 (1.75–1.67) |
| Rsym (%) | 4.4 (77.6) |
| I/σI | 11.2 (1.0) |
| Completeness (%) | 92.2 (94.1) |
| Redundancy | 2.38 |
| CC (1/2) | 99.9 (51.6) |
| Refinement | |
| Resolution range (Å) | 38.5–1.67 |
| Number of unique reflections | 60,447 |
| Rwork (%) | 19.7 |
| Rfree (%) | 24.1 |
| Number of proteins atoms | 3331 |
| Number of water/glycerol | 699/6 |
| Mean B-factor (Å2) | 28.9 |
| Coordinate deviations | |
| RMSD bond lengths (Å) | 0.007 |
| RMSD angles (°) | 0.888 |
| PDB ID | 5NA2 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Folio, C.; Sierra, N.; Dujardin, M.; Alvarez, G.; Guillon, C. Crystal Structure of the Full-Length Feline Immunodeficiency Virus Capsid Protein Shows an N-Terminal β-Hairpin in the Absence of N-Terminal Proline. Viruses 2017, 9, 335. https://doi.org/10.3390/v9110335
Folio C, Sierra N, Dujardin M, Alvarez G, Guillon C. Crystal Structure of the Full-Length Feline Immunodeficiency Virus Capsid Protein Shows an N-Terminal β-Hairpin in the Absence of N-Terminal Proline. Viruses. 2017; 9(11):335. https://doi.org/10.3390/v9110335
Chicago/Turabian StyleFolio, Christelle, Natalia Sierra, Marie Dujardin, Guzman Alvarez, and Christophe Guillon. 2017. "Crystal Structure of the Full-Length Feline Immunodeficiency Virus Capsid Protein Shows an N-Terminal β-Hairpin in the Absence of N-Terminal Proline" Viruses 9, no. 11: 335. https://doi.org/10.3390/v9110335
APA StyleFolio, C., Sierra, N., Dujardin, M., Alvarez, G., & Guillon, C. (2017). Crystal Structure of the Full-Length Feline Immunodeficiency Virus Capsid Protein Shows an N-Terminal β-Hairpin in the Absence of N-Terminal Proline. Viruses, 9(11), 335. https://doi.org/10.3390/v9110335