The Operophtera brumata Nucleopolyhedrovirus (OpbuNPV) Represents an Early, Divergent Lineage within Genus Alphabaculovirus
Abstract
1. Introduction
2. Materials and Methods
2.1. Virus Production and Isolation
2.2. Electron Microscopy
2.3. Viral DNA Isolation and Sequencing
2.4. Genome Sequence Analysis and Feature Annotation
2.5. Sequence Comparison and Phylogeny
3. Results
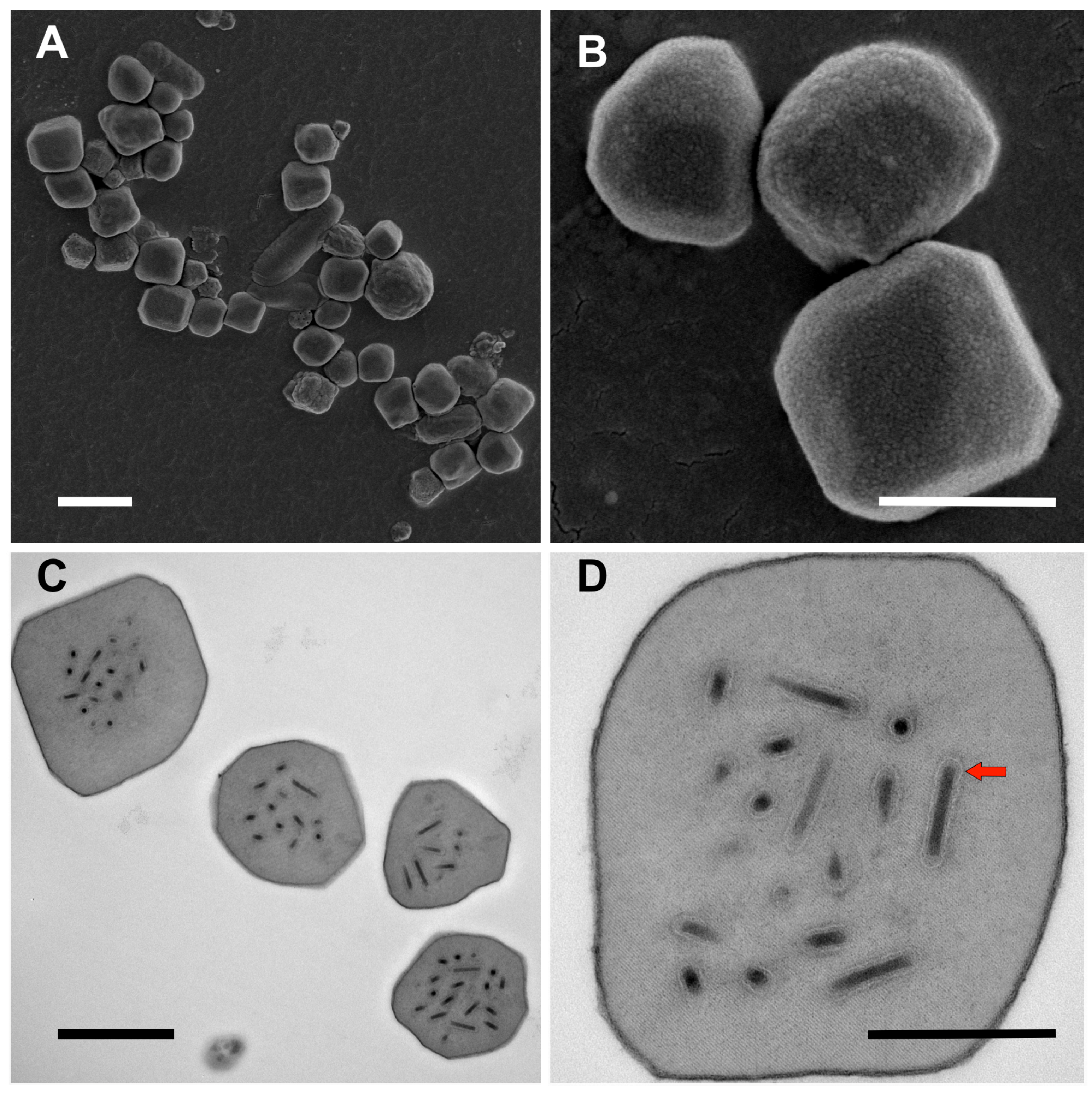
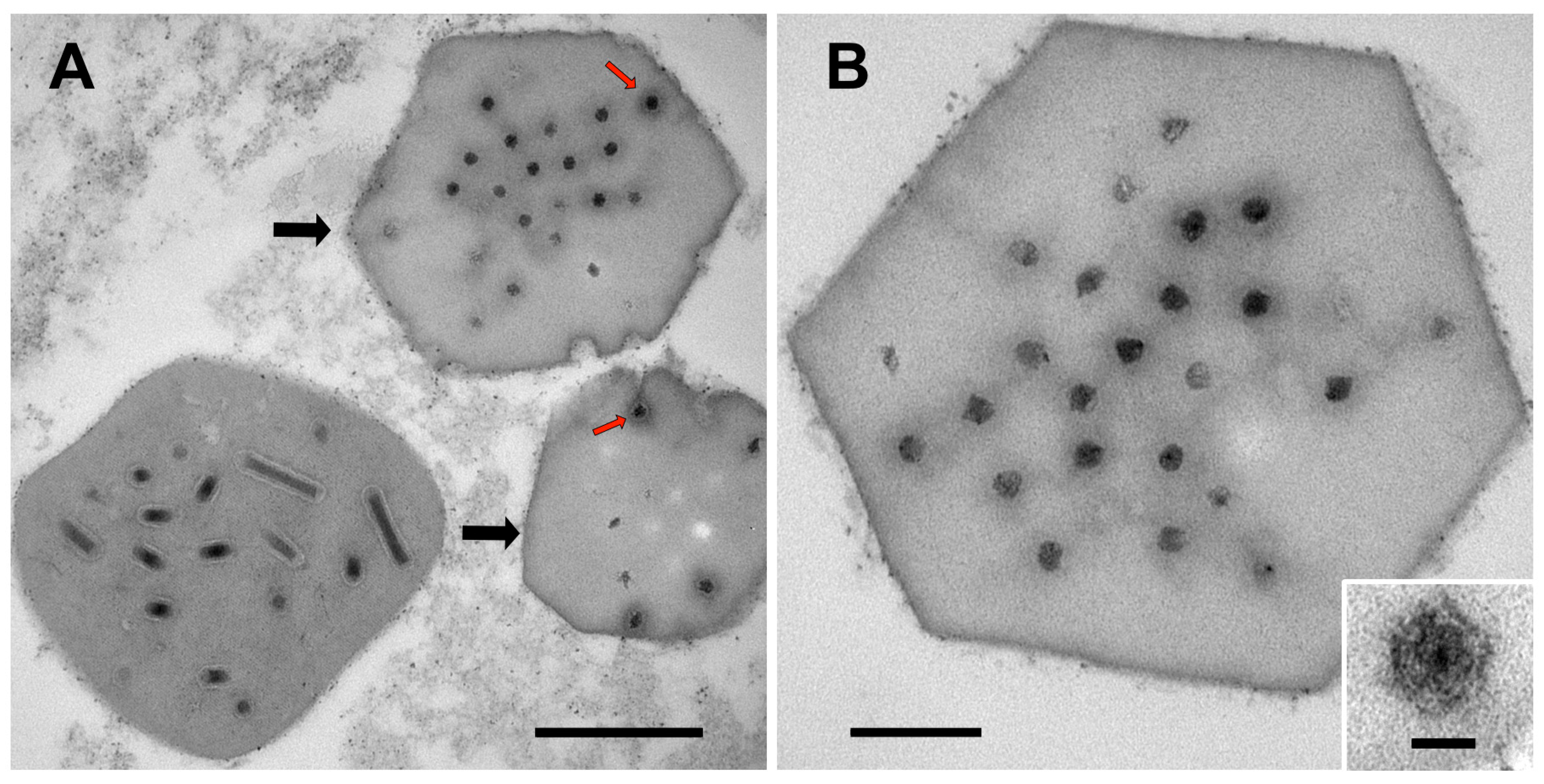
3.1. Ultrastructure of OBs from Winter Moth Larvae
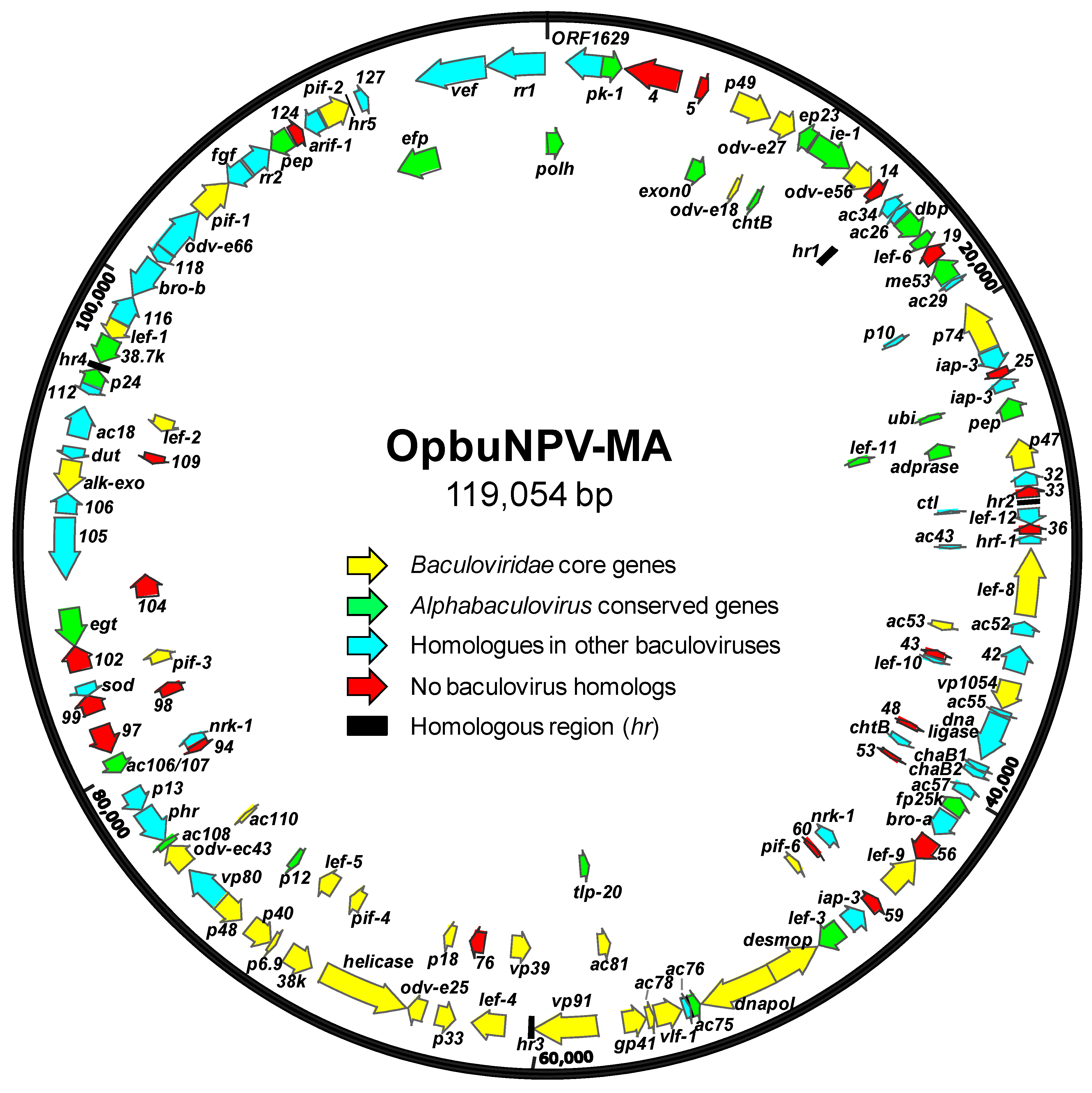
3.2. Properties of the OpbuNPV-MA Genome Sequence
3.3. OpbuNPV-MA ORF Content
3.1.1. Core Genes, bro Genes, and Unique ORFs
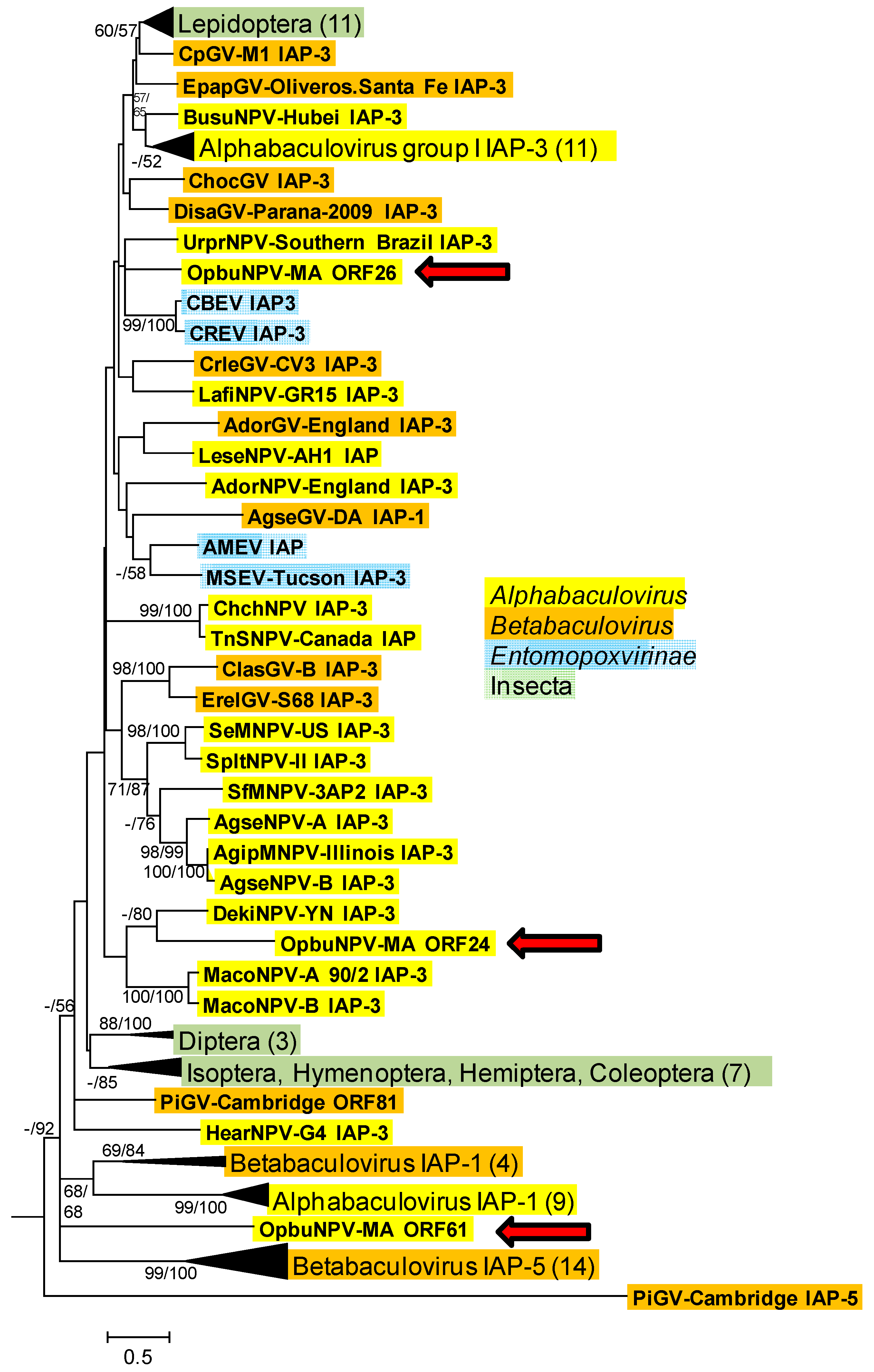
3.1.2. Inhibitor-Of-Apoptosis Protein (iap) Genes
3.1.3. DNA Ligase III and Host Range Factor-1 (hrf-1)
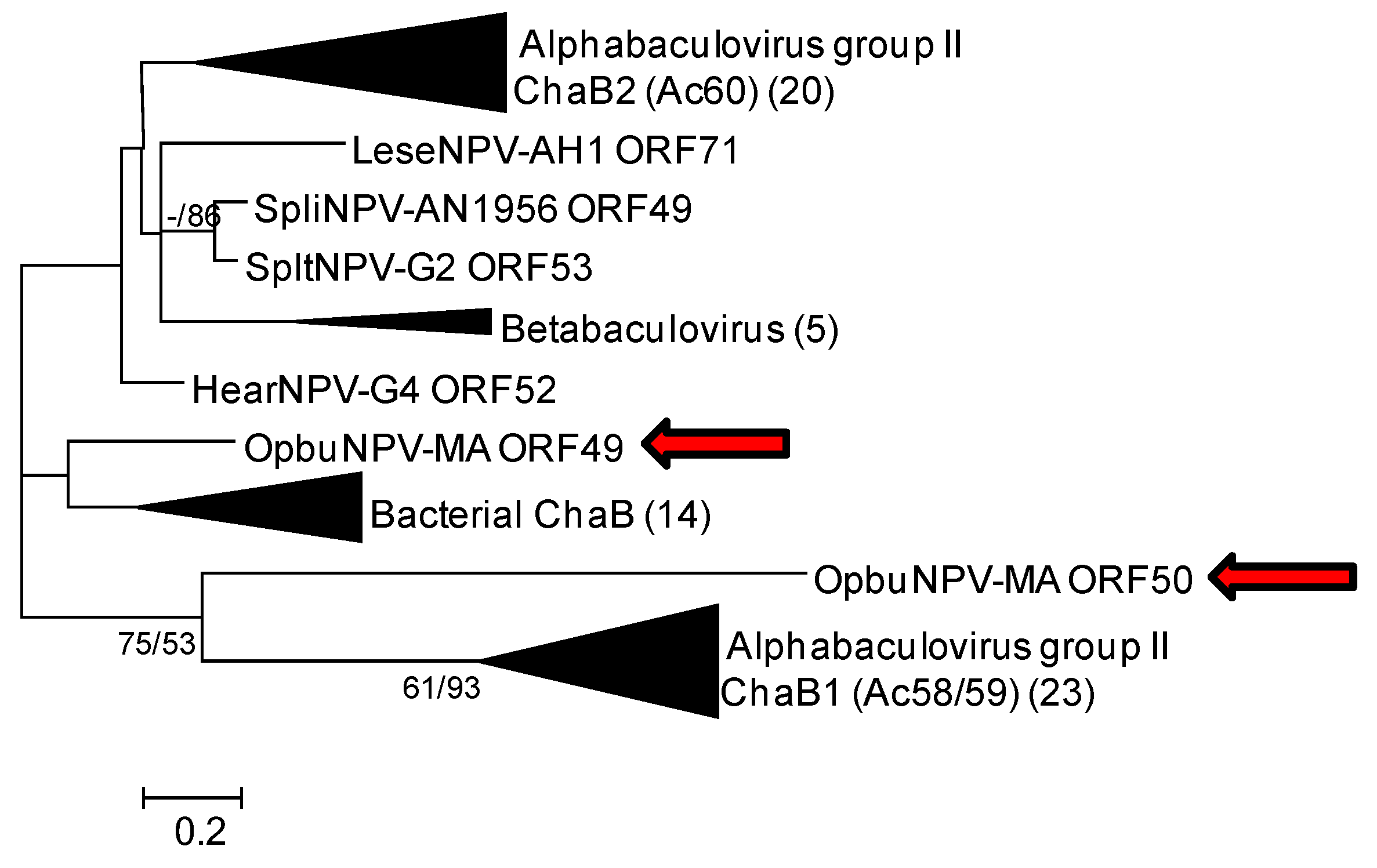
3.1.4. chaB (ac58/59 and ac60) Genes
3.1.5. Two ORFs with Sequence Similarity to Nicotinamide Riboside Kinase 1 (nrk1)
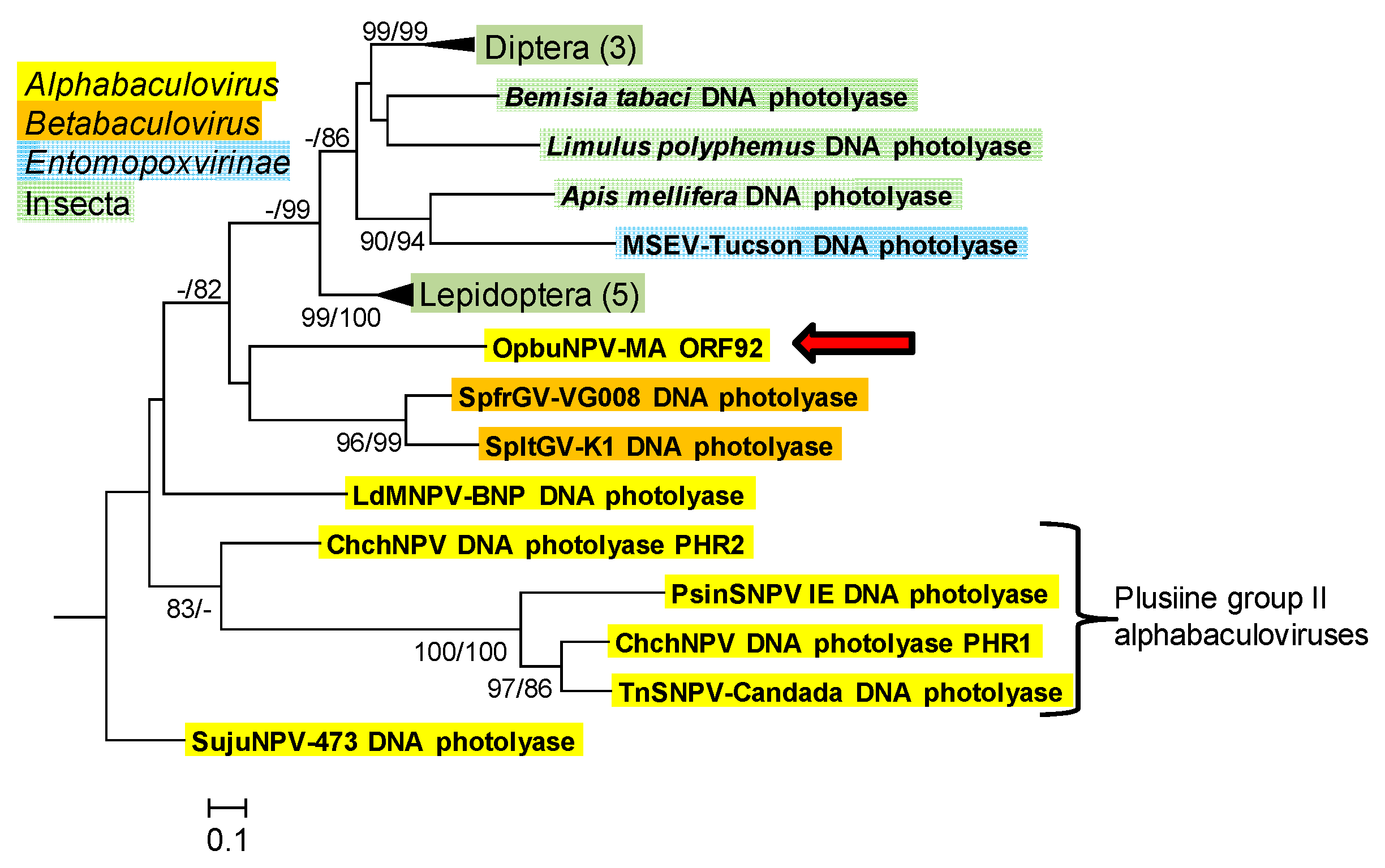
3.1.6. DNA Photolyase (phr)
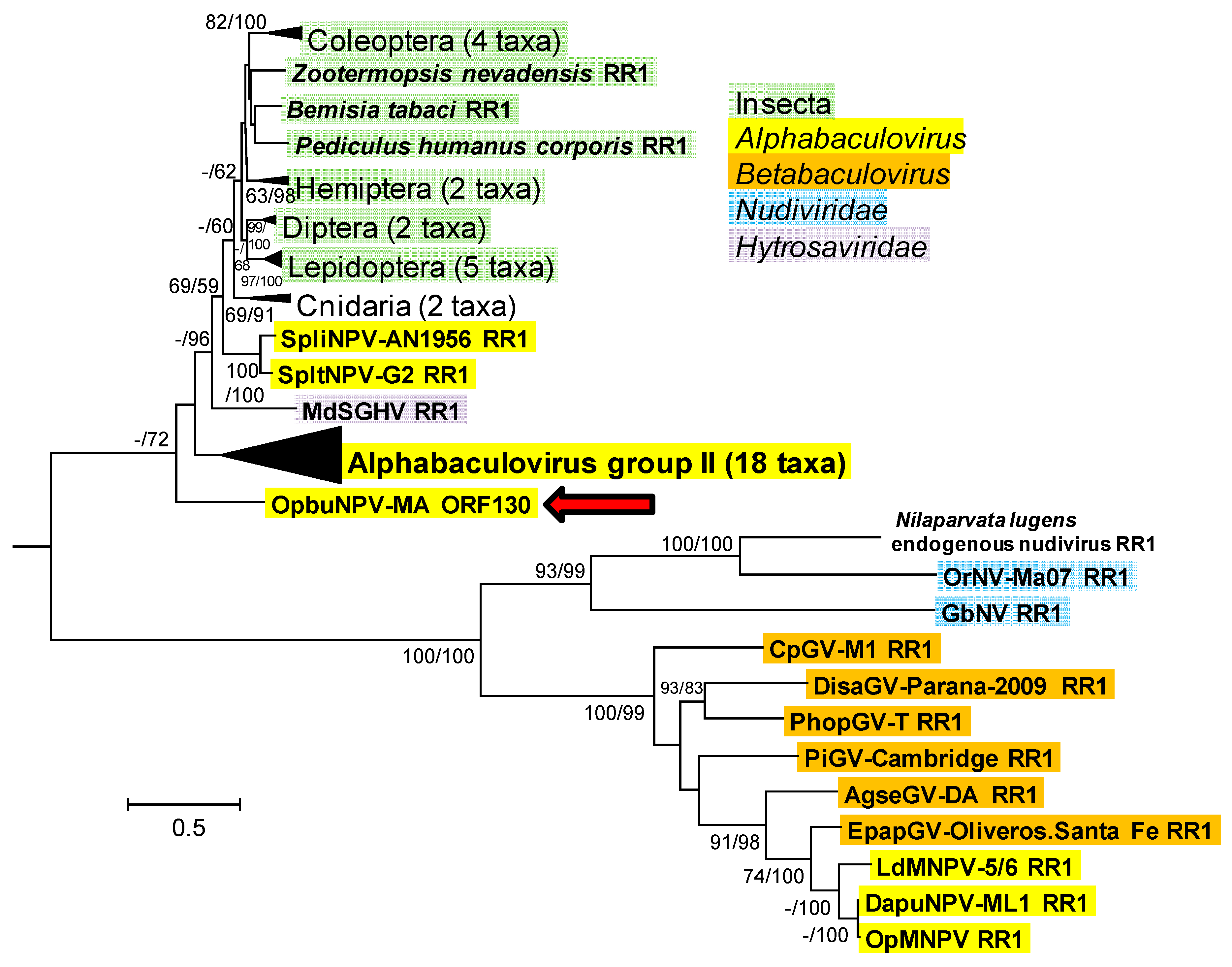
3.1.7. Ribonucleotide Reductase Large Subunit (rr1), Small Subunit (rr2), and dUTPase (dut)
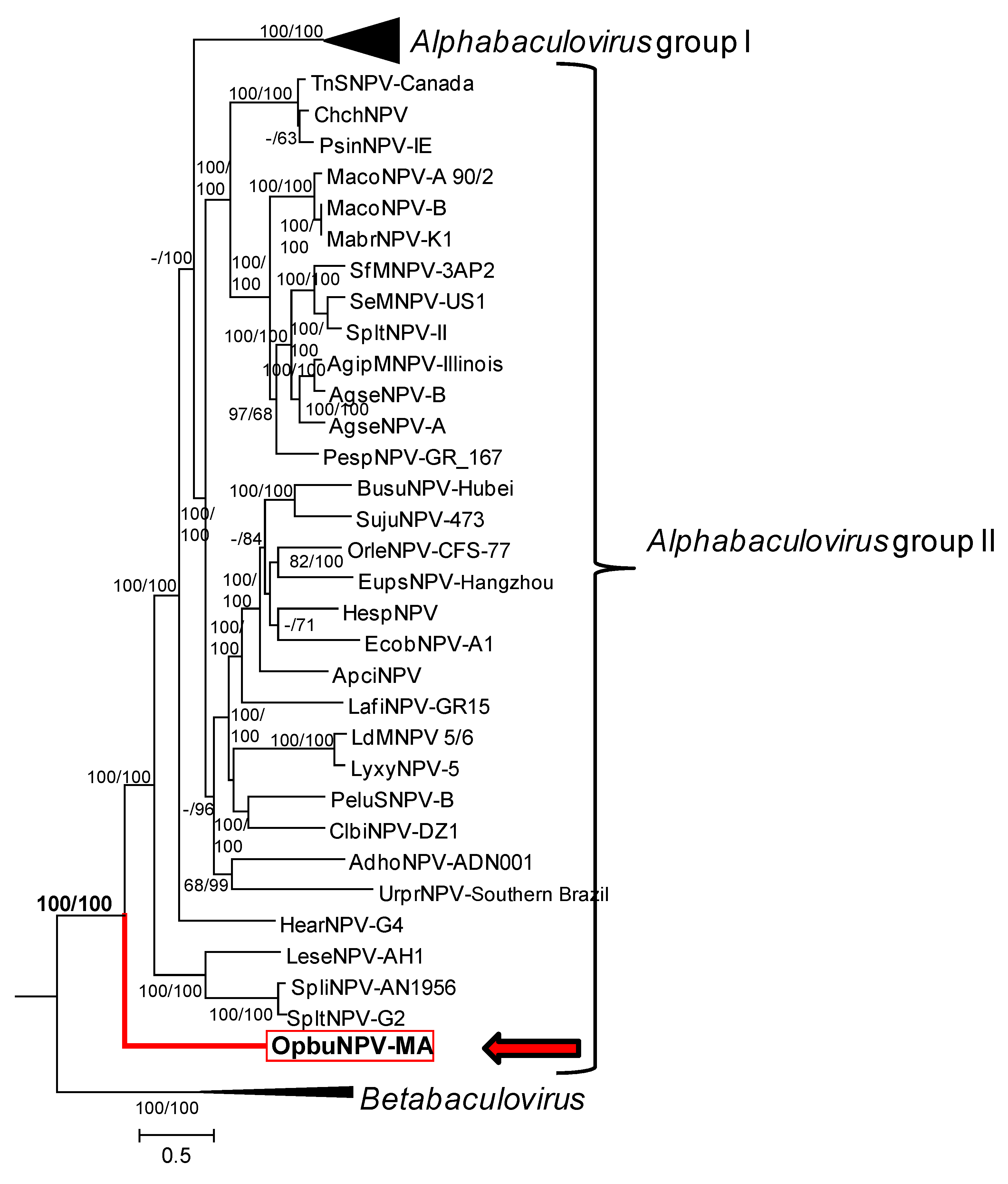
3.4. Relationship of OpbuNPV-MA to Other Baculoviruses
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Herniou, E.A.; Arif, B.M.; Becnel, J.J.; Blissard, G.W.; Bonning, B.; Harrison, R.L.; Jehle, J.A.; Theilmann, D.A.; Vlak, J.M. Baculoviridae. In Virus taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Adams, M.J., Eds.; Elsevier: Oxford, UK, 2011; pp. 163–174. [Google Scholar]
- Harrison, R.L.; Hoover, K. Baculoviruses and other occluded insect viruses. In Insect Pathology, 2nd ed.; Vega, F.E., Kaya, H.K., Eds.; Academic Press: Boston, MA, USA, 2012; pp. 73–131. [Google Scholar]
- Rohrmann, G.F. Baculovirus Molecular Biology, 3rd ed.; National Center for Biotechnology Information (US): Bethesda, MD, USA, 2013.
- Roland, J.; Embree, D.G. Biological control of the winter moth. Ann. Rev. Entomol. 1995, 40, 475–492. [Google Scholar] [CrossRef]
- Elkinton, J.; Boettner, G.; Liebhold, A.; Gwiazdowski, R. Biology, Spread, and Biological Control of Winter Moth in the Eastern United States; Service, U.F., Ed.; Forest Health Technology Enterprise Team: Morgantown, West Virginia, USA, 2015. [Google Scholar]
- Embree, D.G. The role of introduced parasites in the control of the winter moth in Nova Scotia. Can. Entomol. 1966, 98, 1159–1168. [Google Scholar] [CrossRef]
- Graham, R.I.; Tyne, W.I.; Possee, R.D.; Sait, S.M.; Hails, R.S. Genetically variable nucleopolyhedroviruses isolated from spatially separate populations of the winter moth Operophtera brumata (Lepidoptera: Geometridae) in Orkney. J. Invertebr. Pathol. 2004, 87, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Burand, J.P.; Kim, W.; Welch, A.; Elkinton, J.S. Identification of a nucleopolyhedrovirus in winter moth populations from Massachusetts. J. Invertebr. Pathol. 2011, 108, 217–219. [Google Scholar] [CrossRef] [PubMed]
- Broadley, H.J.; Boucher, M.; Burand, J.P.; Elkinton, J.S. The phylogenetic relationship and cross-infection of nucleopolyhedroviruses between the invasive winter moth (Operophtera brumata) and its native congener, bruce spanworm (O. bruceata). J. Invertebr. Pathol. 2017, 143, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, J.C.; Tonks, N.V.; Kaupp, W.J. Viruses to control winter moth, Operophtera brumata, (Lepidoptera: Geometridae). J. Entomol. Soc. B. C. 1981, 78, 17–24. [Google Scholar]
- Guo, F.-B.; Zhang, C.-T. ZCURVE_V: A new self-training system for recognizing protein-coding genes in viral and phage genomes. BMC Bioinform. 2006, 7, 9. [Google Scholar] [CrossRef] [PubMed]
- Alva, V.; Nam, S.Z.; Soding, J.; Lupas, A.N. The MPI bioinformatics toolkit as an integrative platform for advanced protein sequence and structure analysis. Nucleic Acids Res. 2016, 44, W410–415. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. Clustal w: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. Raxml version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Attoui, H.; Mertens, P.P.C.; Becnel, J.; Belaganahalli, S.; Bergoin, M.; Brussaard, C.P.; Chappell, J.D.; Ciarlet, M.; del Vas, M.; Dermody, T.S.; et al. Reoviridae. In Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Adams, M.J., Eds.; Elsevier: Oxford, UK, 2011; pp. 541–637. [Google Scholar]
- Graham, R.I.; Rao, S.; Possee, R.D.; Sait, S.M.; Mertens, P.P.; Hails, R.S. Detection and characterisation of three novel species of reovirus (Reoviridae), isolated from geographically separate populations of the winter moth Operophtera brumata (Lepidoptera: Geometridae) on Orkney. J. Invertebr. Pathol. 2006, 91, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Graham, R.I.; Rao, S.; Sait, S.M.; Mertens, P.P.; Hails, R.S.; Possee, R.D. Characterisation and partial sequence analysis of two novel cypoviruses isolated from the winter moth Operophtera brumata (Lepidoptera: Geometridae). Virus Genes 2007, 35, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Van Oers, M.M.; Vlak, J.M. Baculovirus genomics. Curr. Drug Targets 2007, 8, 1051–1068. [Google Scholar] [CrossRef] [PubMed]
- Chateigner, A.; Bezier, A.; Labrousse, C.; Jiolle, D.; Barbe, V.; Herniou, E.A. Ultra deep sequencing of a baculovirus population reveals widespread genomic variations. Viruses 2015, 7, 3625–3646. [Google Scholar] [CrossRef] [PubMed]
- Garavaglia, M.J.; Miele, S.A.; Iserte, J.A.; Belaich, M.N.; Ghiringhelli, P.D. The ac53, ac78, ac101, and ac103 genes are newly discovered core genes in the family Baculoviridae. J. Virol. 2012, 86, 12069–12079. [Google Scholar] [CrossRef] [PubMed]
- Javed, M.A.; Biswas, S.; Willis, L.G.; Harris, S.; Pritchard, C.; van Oers, M.M.; Donly, B.C.; Erlandson, M.A.; Hegedus, D.D.; Theilmann, D.A. Autographa californica multiple nucleopolyhedrovirus ac83 is a per os infectivity factor (PIF) protein required for occlusion-derived virus (ODV) and budded virus nucleocapsid assembly as well as assembly of the PIF complex in ODV envelopes. J. Virol. 2017, 91, e02115–02116. [Google Scholar] [CrossRef] [PubMed]
- Bideshi, D.K.; Renault, S.; Stasiak, K.; Federici, B.A.; Bigot, Y. Phylogenetic analysis and possible function of bro-like genes, a multigene family widespread among large double-stranded DNA viruses of invertebrates and bacteria. J. Gen. Virol. 2003, 84, 2531–2544. [Google Scholar] [CrossRef] [PubMed]
- Derks, M.F.; Smit, S.; Salis, L.; Schijlen, E.; Bossers, A.; Mateman, C.; Pijl, A.S.; de Ridder, D.; Groenen, M.A.; Visser, M.E.; et al. The genome of winter moth (Operophtera brumata) provides a genomic perspective on sexual dimorphism and phenology. Genome Biol. Evol. 2015, 7, 2321–2332. [Google Scholar] [CrossRef] [PubMed]
- Clem, R.J. Viral IAPs, then and now. Semim. Cell Dev. Biol. 2015, 39, 72–79. [Google Scholar] [CrossRef] [PubMed]
- Kuzio, J.; Pearson, M.N.; Harwood, S.H.; Funk, C.J.; Evans, J.T.; Slavicek, J.M.; Rohrmann, G.F. Sequence and analysis of the genome of a baculovirus pathogenic for Lymantria dispar. Virology 1999, 253, 17–34. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.J.; Quentin, M.E.; Brennan, L.A.; Kukel, C.; Thiem, S.M. Lymantria dispar nucleopolyhedrovirus HRF-1 expands the larval host range of Autographa californica nucleopolyhedrovirus. J. Virol. 1998, 72, 2526–2531. [Google Scholar] [PubMed]
- Ishikawa, H.; Ikeda, M.; Alves, C.A.; Thiem, S.M.; Kobayashi, M. Host range factor 1 from Lymantria dispar nucleopolyhedrovirus (NPV) is an essential viral factor required for productive infection of NPVs in IPLB-Ld652Y cells derived from L. dispar. J. Virol. 2004, 78, 12703–12708. [Google Scholar] [CrossRef] [PubMed]
- Pearson, M.N.; Rohrmann, G.F. Characterization of a baculovirus-encoded ATP-dependent DNA ligase. J. Virol. 1998, 72, 9142–9149. [Google Scholar] [PubMed]
- Krejmer, M.; Skrzecz, I.; Wasag, B.; Szewczyk, B.; Rabalski, L. The genome of Dasychira pudibunda nucleopolyhedrovirus (DapuNPV) reveals novel genetic connection between baculoviruses infecting moths of the Lymantriidae family. BMC Genom. 2015, 16, 759. [Google Scholar] [CrossRef] [PubMed]
- Ahrens, C.H.; Russell, R.L.; Funk, C.J.; Evans, J.T.; Harwood, S.H.; Rohrmann, G.F. The sequence of the Orgyia pseudotsugata multinucleocapsid nuclear polyhedrosis virus genome. Virology 1997, 229, 381–399. [Google Scholar] [CrossRef] [PubMed]
- Nai, Y.S.; Wu, C.Y.; Wang, T.C.; Chen, Y.R.; Lau, W.H.; Lo, C.F.; Tsai, M.F.; Wang, C.H. Genomic sequencing and analyses of Lymantria xylina multiple nucleopolyhedrovirus. BMC Genom. 2010, 11, 116. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yin, F.; Zhu, Z.; Hou, D.; Wang, J.; Zhang, L.; Wang, M.; Wang, H.; Hu, Z.; Deng, F. Genomic sequencing and analysis of Sucra jujuba nucleopolyhedrovirus. PLoS ONE 2014, 9, e110023. [Google Scholar] [CrossRef] [PubMed]
- Thumbi, D.K.; Eveleigh, R.J.; Lucarotti, C.J.; Lapointe, R.; Graham, R.I.; Pavlik, L.; Lauzon, H.A.; Arif, B.M. Complete sequence, analysis and organization of the Orgyia leucostigma nucleopolyhedrovirus genome. Viruses 2011, 3, 2301–2327. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Li, Z.; Chen, W.; Liu, C.; Huang, H.; Yang, K.; Pang, Y. Characterization of Spodoptera exigua multicapsid nucleopolyhedrovirus orf100 and orf101, two homologues of E. coli ChaB. Virus Res. 2006, 121, 42–50. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, L.; Yu, H.; Li, S.; Pang, Y. Characterization of two homologues of ChaB in Spodoptera litura multicapsid nucleopolyhedrovirus. Gene 2006, 372, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Magni, G.; Amici, A.; Emanuelli, M.; Orsomando, G.; Raffaelli, N.; Ruggieri, S. Enzymology of NAD+ homeostasis in man. Cell. Mol. Life Sci. 2004, 61, 19–34. [Google Scholar] [CrossRef] [PubMed]
- Ratajczak, J.; Joffraud, M.; Trammell, S.A.; Ras, R.; Canela, N.; Boutant, M.; Kulkarni, S.S.; Rodrigues, M.; Redpath, P.; Migaud, M.E.; et al. Nrk1 controls nicotinamide mononucleotide and nicotinamide riboside metabolism in mammalian cells. Nat. Commun. 2016, 7, 13103. [Google Scholar] [CrossRef] [PubMed]
- Cuartas, P.E.; Barrera, G.P.; Belaich, M.N.; Barreto, E.; Ghiringhelli, P.D.; Villamizar, L.F. The complete sequence of the first Spodoptera frugiperda betabaculovirus genome: A natural multiple recombinant virus. Viruses 2015, 7, 394–421. [Google Scholar] [CrossRef] [PubMed]
- Willis, L.G.; Seipp, R.; Stewart, T.M.; Erlandson, M.A.; Theilmann, D.A. Sequence analysis of the complete genome of Trichoplusia ni single nucleopolyhedrovirus and the identification of a baculoviral photolyase gene. Virology 2005, 338, 209–226. [Google Scholar] [CrossRef] [PubMed]
- Biernat, M.A.; Ros, V.I.; Vlak, J.M.; van Oers, M.M. Baculovirus cyclobutane pyrimidine dimer photolyases show a close relationship with lepidopteran host homologues. Insect Mol. Biol. 2011, 20, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Choi, J.Y.; Roh, J.Y.; Liu, Q.; Tao, X.Y.; Park, J.B.; Kim, J.S.; Je, Y.H. Genomic sequence analysis of granulovirus isolated from the tobacco cutworm, Spodoptera litura. PLoS ONE 2011, 6, e28163. [Google Scholar] [CrossRef] [PubMed]
- Van Oers, M.M.; Lampen, M.H.; Bajek, M.I.; Vlak, J.M.; Eker, A.P. Active DNA photolyase encoded by a baculovirus from the insect Chrysodeixis chalcites. DNA Repair 2008, 7, 1309–1318. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Vlak, J.M.; van Oers, M.M. Conservation of DNA photolyase genes in group Ⅱ nucleopolyhedroviruses infecting plusiine insects. Virus Res. 2008, 136, 58–64. [Google Scholar] [CrossRef] [PubMed]
- Jordan, A.; Reichard, P. Ribonucleotide reductases. Annu. Rev. Biochem. 1998, 67, 71–98. [Google Scholar] [CrossRef] [PubMed]
- Nyman, P.O. Introduction. DUTPases. Curr. Protein. Pept. Sci. 2001, 2, 277–285. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Wang, H.; Mansky, L.M. Roles of uracil-DNA glycosylase and dUTPase in virus replication. J. Gen. Virol. 2002, 83, 2339–2345. [Google Scholar] [CrossRef] [PubMed]
- Shackelton, L.A.; Holmes, E.C. The evolution of large DNA viruses: Combining genomic information of viruses and their hosts. Trends Microbiol. 2004, 12, 458–465. [Google Scholar] [CrossRef] [PubMed]
- Harrison, R.L.; Rowley, D.L.; Mowery, J.; Bauchan, G.R.; Theilmann, D.A.; Rohrmann, G.F.; Erlandson, M.A. The complete genome sequence of a second distinct betabaculovirus from the true armyworm, Mythimna unipuncta. PLoS ONE 2017, 12, e0170510. [Google Scholar] [CrossRef] [PubMed]
- Theze, J.; Takatsuka, J.; Nakai, M.; Arif, B.; Herniou, E.A. Gene acquisition convergence between entomopoxviruses and baculoviruses. Viruses 2015, 7, 1960–1974. [Google Scholar] [CrossRef] [PubMed]
- Ardisson-Araujo, D.M.; Pereira, B.T.; Melo, F.L.; Ribeiro, B.M.; Bao, S.N.; de, A.Z.P.M.; Moscardi, F.; Kitajima, E.W.; Sosa-Gomez, D.R.; Wolff, J.L. A betabaculovirus encoding a gp64 homolog. BMC Genom. 2016, 17, 94. [Google Scholar] [CrossRef] [PubMed]
- Harper, J.D. Interactions between baculoviruses and other entompathogens, chemical pesticides, and parasitioids. In The Biology of Baculoviruses. Volume II, Practical Application for Insect Control; Granados, R.R., Federici, B.A., Eds.; CRC Press: Boca Raton, FL, USA, 1986; pp. 133–155. [Google Scholar]
- Laitinen, A.M.; Otvos, I.S.; Levin, D.B. Geographic distribution of cytoplasmic polyhedrosis virus infection in Douglas-fir tussock moth larvae, Orgyia pseudotsugata, in British Columbia. J. Invertebr. Pathol. 1996, 67, 229–235. [Google Scholar] [CrossRef] [PubMed]
- Belloncik, S.; Mori, H. Cypoviruses. In The Insect Viruses; Miller, L.K., Ball, L.A., Eds.; Plenum Press: New York, NY, USA, 1998; pp. 337–370. [Google Scholar]
- Bird, F.T. Infection and mortality of spruce budworm, Choristoneura fumiferana, and forest tent caterpillar, Malacosoma disstria, caused by nuclear and cytoplasmic polyhedrosis viruses. Can. Entomol. 1969, 101, 1269–1285. [Google Scholar] [CrossRef]
- Zwart, M.P.; Elena, S.F. Matters of size: Genetic bottlenecks in virus infection and their potential impact on evolution. Annu. Rev .Virol. 2015, 2, 161–179. [Google Scholar] [CrossRef] [PubMed]
- Smith, I.R.; Crook, N.E. In vivo isolation of baculovirus genotypes. Virology 1988, 166, 240–244. [Google Scholar] [CrossRef]
- Zwart, M.P.; Hemerik, L.; Cory, J.S.; de Visser, J.A.; Bianchi, F.J.; Van Oers, M.M.; Vlak, J.M.; Hoekstra, R.F.; Van der Werf, W. An experimental test of the independent action hypothesis in virus-insect pathosystems. Proc. Biol. Sci. 2009, 276, 2233–2242. [Google Scholar] [CrossRef] [PubMed]
- Hodgson, D.J.; Hitchman, R.B.; Vanbergen, A.J.; Hails, R.S.; Possee, R.D.; Cory, J.S. Host ecology determines the relative fitness of virus genotypes in mixed-genotype nucleopolyhedrovirus infections. J. Evol. Biol. 2004, 17, 1018–1025. [Google Scholar] [CrossRef] [PubMed]
- Clavijo, G.; Williams, T.; Simon, O.; Munoz, D.; Cerutti, M.; Lopez-Ferber, M.; Caballero, P. Mixtures of complete and pif1- and pif2-deficient genotypes are required for increased potency of an insect nucleopolyhedrovirus. J. Virol. 2009, 83, 5127–5136. [Google Scholar] [CrossRef] [PubMed]


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Harrison, R.L.; Rowley, D.L.; Mowery, J.D.; Bauchan, G.R.; Burand, J.P. The Operophtera brumata Nucleopolyhedrovirus (OpbuNPV) Represents an Early, Divergent Lineage within Genus Alphabaculovirus. Viruses 2017, 9, 307. https://doi.org/10.3390/v9100307
Harrison RL, Rowley DL, Mowery JD, Bauchan GR, Burand JP. The Operophtera brumata Nucleopolyhedrovirus (OpbuNPV) Represents an Early, Divergent Lineage within Genus Alphabaculovirus. Viruses. 2017; 9(10):307. https://doi.org/10.3390/v9100307
Chicago/Turabian StyleHarrison, Robert L., Daniel L. Rowley, Joseph D. Mowery, Gary R. Bauchan, and John P. Burand. 2017. "The Operophtera brumata Nucleopolyhedrovirus (OpbuNPV) Represents an Early, Divergent Lineage within Genus Alphabaculovirus" Viruses 9, no. 10: 307. https://doi.org/10.3390/v9100307
APA StyleHarrison, R. L., Rowley, D. L., Mowery, J. D., Bauchan, G. R., & Burand, J. P. (2017). The Operophtera brumata Nucleopolyhedrovirus (OpbuNPV) Represents an Early, Divergent Lineage within Genus Alphabaculovirus. Viruses, 9(10), 307. https://doi.org/10.3390/v9100307




