Requirements within the Ebola Viral Glycoprotein for Tetherin Antagonism
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Lines, Plasmid Vectors and Antibodies
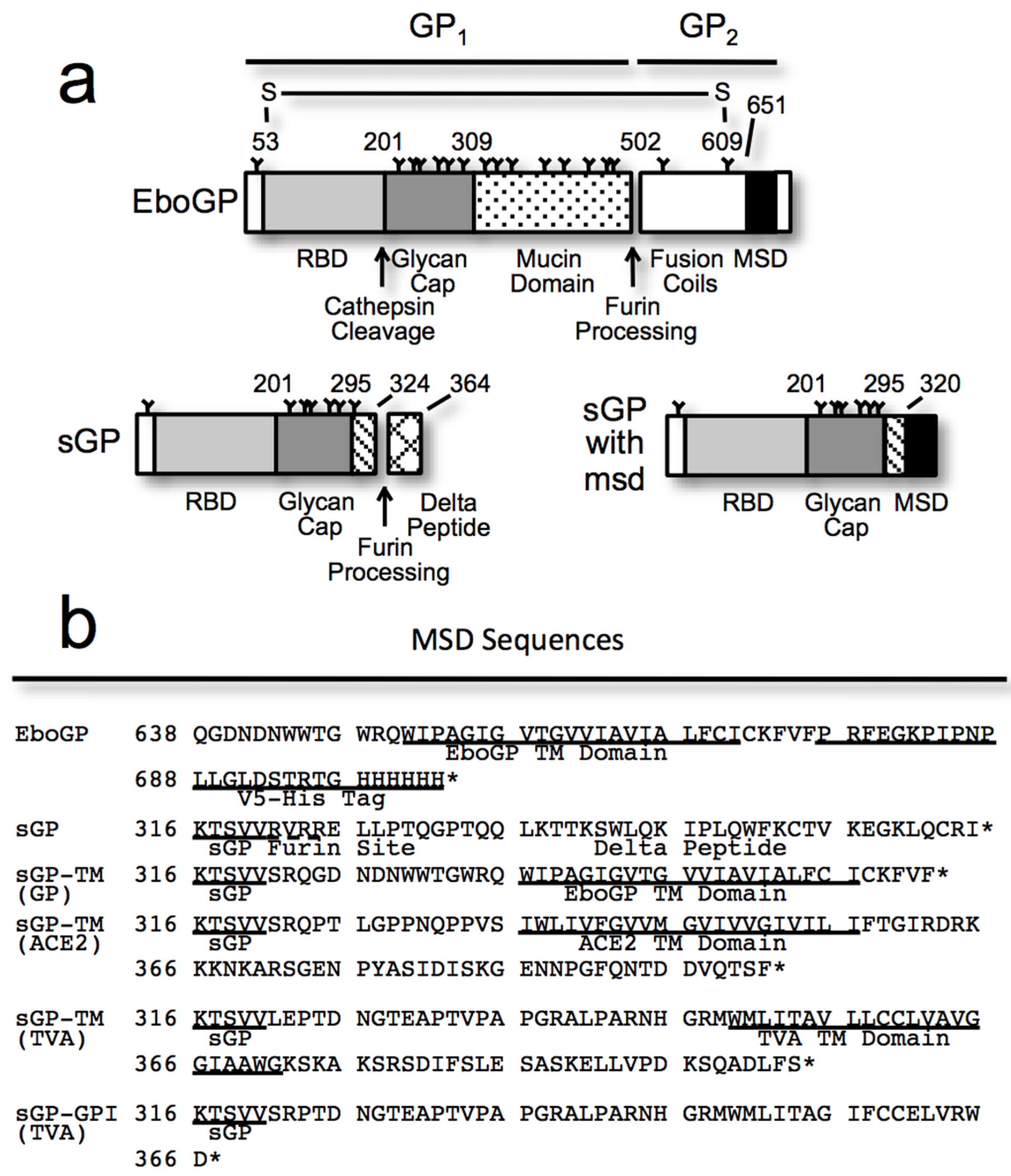
2.2. Virus-Like Particle (VLP) Budding Assay
2.3. Immunoprecipitaion Assay
2.4. Immunoblot Analysis
2.5. Flow Cytometry Analysis
3. Results
3.1. Requirements within EboGP for Tetherin Antagonism
3.2. Chimeras Reveal a Specific Requirement for the EboGP Membrane Spanning Domain (msd)
3.3. The Ebola msd Requirement is not Specific for Filoviral Budding
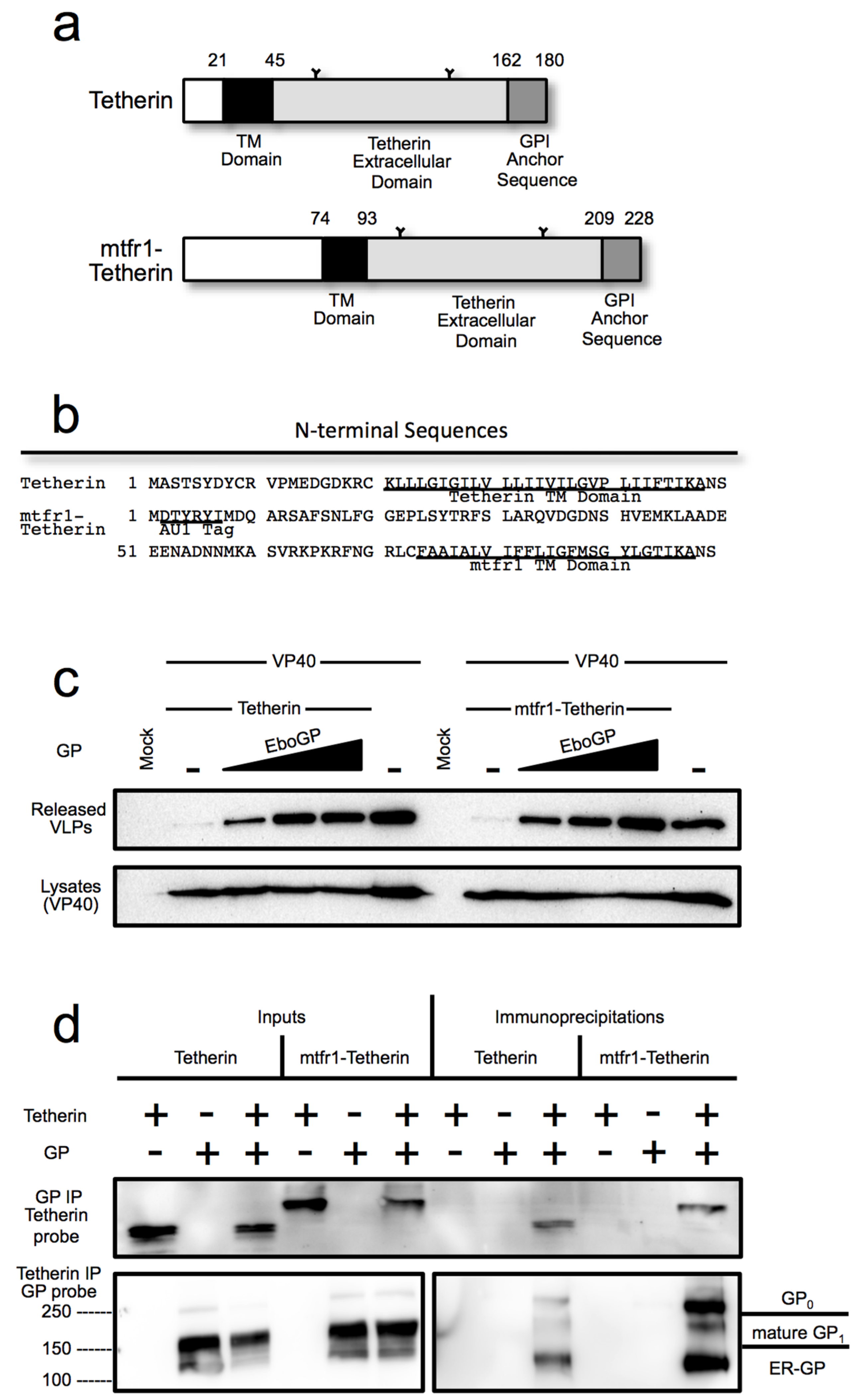
3.4. The Tetherin Amino-Terminal Region is not Required for EboGP Recognition
3.5. The Glycan Cap of EboGP is Required to Antagonize Tetherin
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Neil, S.J.; Zang, T.; Bieniasz, P.D. Tetherin inhibits retrovirus release and is antagonized by HIV-1 Vpu. Nature 2008, 451, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Kaletsky, R.L.; Francica, J.R.; Agrawal-Gamse, C.; Bates, P. Tetherin-mediated restriction of filovirus budding is antagonized by the Ebola glycoprotein. Proc. Natl. Acad. Sci. USA 2009, 106, 2886–2891. [Google Scholar] [CrossRef] [PubMed]
- Sakuma, T.; Noda, T.; Urata, S.; Kawaoka, Y.; Yasuda, J. Inhibition of Lassa and Marburg virus production by tetherin. J. Virol. 2009, 83, 2382–2385. [Google Scholar] [CrossRef] [PubMed]
- Pardieu, C.; Vigan, R.; Wilson, S.J.; Calvi, A.; Zang, T.; Bieniasz, P.; Kellam, P.; Towers, G.J.; Neil, S.J. The RING-CH ligase K5 antagonizes restriction of KSHV and HIV-1 particle release by mediating ubiquitin-dependent endosomal degradation of tetherin. PLoS Pathog. 2010, 6, e1000843. [Google Scholar] [CrossRef] [PubMed]
- Yondola, M.A.; Fernandes, F.; Belicha-Villanueva, A.; Uccelini, M.; Gao, Q.; Carter, C.; Palese, P. Budding capability of the influenza virus neuraminidase can be modulated by tetherin. J. Virol. 2011, 85, 2480–2491. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.H.; Maric, M.; Madison, M.N.; Maury, W.; Roller, R.J.; Okeoma, C.M. BST-2/tetherin-mediated restriction of chikungunya (CHIKV) VLP budding is counteracted by CHIKV non-structural protein 1 (nsp1). Virology 2013, 438, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Kupzig, S.; Korolchuk, V.; Rollason, R.; Sugden, A.; Wilde, A.; Banting, G. BST-2/HM1.24 is a raft-associated apical membrane protein with an unusual topology. Traffic 2003, 4, 694–709. [Google Scholar] [CrossRef] [PubMed]
- Hammonds, J.; Wang, J.J.; Yi, H.; Spearman, P. Immunoelectron microscopic evidence for tetherin/BST2 as the physical bridge between HIV-1 virions and the plasma membrane. PLoS Pathog. 2010, 6, e1000749. [Google Scholar] [CrossRef] [PubMed]
- Venkatesh, S.; Bieniasz, P.D. Mechanism of HIV-1 virion entrapment by tetherin. PLoS Pathog. 2013, 9, e1003483. [Google Scholar] [CrossRef] [PubMed]
- Van Damme, N.; Goff, D.; Katsura, C.; Jorgenson, R.L.; Mitchell, R.; Johnson, M.C.; Stephens, E.B.; Guatelli, J. The interferon-induced protein BST-2 restricts HIV-1 release and is downregulated from the cell surface by the viral Vpu protein. Cell Host Microbe 2008, 3, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Mansouri, M.; Viswanathan, K.; Douglas, J.L.; Hines, J.; Gustin, J.; Moses, A.V.; Fruh, K. Molecular mechanism of BST2/tetherin downregulation by K5/MIR2 of kaposi’s sarcoma-associated herpesvirus. J. Virol. 2009, 83, 9672–9681. [Google Scholar] [CrossRef] [PubMed]
- Hauser, H.; Lopez, L.A.; Yang, S.J.; Oldenburg, J.E.; Exline, C.M.; Guatelli, J.C.; Cannon, P.M. HIV-1 vpu and HIV-2 env counteract BST-2/tetherin by sequestration in a perinuclear compartment. Retrovirology 2010, 7. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.K.; Mlcochova, P.; Pelchen-Matthews, A.; Petit, S.J.; Mattiuzzo, G.; Pillay, D.; Takeuchi, Y.; Marsh, M.; Towers, G.J. Simian immunodeficiency virus envelope glycoprotein counteracts tetherin/BST-2/CD317 by intracellular sequestration. Proc. Natl. Acad. Sci. USA 2009, 106, 20889–20894. [Google Scholar] [CrossRef] [PubMed]
- Leroy, E.M.; Kumulungui, B.; Pourrut, X.; Rouquet, P.; Hassanin, A.; Yaba, P.; Delicat, A.; Paweska, J.T.; Gonzalez, J.P.; Swanepoel, R. Fruit bats as reservoirs of Ebola virus. Nature 2005, 438, 575–576. [Google Scholar] [CrossRef] [PubMed]
- Pourrut, X.; Kumulungui, B.; Wittmann, T.; Moussavou, G.; Delicat, A.; Yaba, P.; Nkoghe, D.; Gonzalez, J.P.; Leroy, E.M. The natural history of Ebola virus in africa. Microbes Infect. 2005, 7, 1005–1014. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, A.; Fiet, C.; Belpois-Duchamp, C.; Tiv, M.; Astruc, K.; Aho Glele, L.S. Case fatality rates of Ebola virus diseases: A meta-analysis of world health organization data. Med. Mal. Infect. 2014, 44, 412–416. [Google Scholar] [CrossRef] [PubMed]
- Johnson, K.M.; Lange, J.V.; Webb, P.A.; Murphy, F.A. Isolation and partial characterisation of a new virus causing acute haemorrhagic fever in Zaire. Lancet 1977, 1, 569–571. [Google Scholar] [CrossRef]
- Georges, A.J.; Leroy, E.M.; Renaut, A.A.; Benissan, C.T.; Nabias, R.J.; Ngoc, M.T.; Obiang, P.I.; Lepage, J.P.; Bertherat, E.J.; Benoni, D.D.; et al. Ebola hemorrhagic fever outbreaks in gabon, 1994–1997: Epidemiologic and health control issues. J. Infect. Dis. 1999, 179 (Suppl. S1), S65–S75. [Google Scholar] [CrossRef] [PubMed]
- Bwaka, M.A.; Bonnet, M.J.; Calain, P.; Colebunders, R.; De Roo, A.; Guimard, Y.; Katwiki, K.R.; Kibadi, K.; Kipasa, M.A.; Kuvula, K.J.; et al. Ebola hemorrhagic fever in Kikwit, Democratic Republic of the Congo: Clinical observations in 103 patients. J. Infect. Dis. 1999, 179 (Suppl. S1), S1–S7. [Google Scholar] [CrossRef] [PubMed]
- Volchkov, V.E.; Feldmann, H.; Volchkova, V.A.; Klenk, H.D. Processing of the Ebola virus glycoprotein by the proprotein convertase furin. Proc. Natl. Acad. Sci. USA 1998, 95, 5762–5767. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, A.; Yang, Z.Y.; Xu, L.; Nabel, G.J.; Crews, T.; Peters, C.J. Biochemical analysis of the secreted and virion glycoproteins of ebola virus. J. Virol. 1998, 72, 6442–6447. [Google Scholar] [PubMed]
- Lee, J.E.; Fusco, M.L.; Hessell, A.J.; Oswald, W.B.; Burton, D.R.; Saphire, E.O. Structure of the Ebola virus glycoprotein bound to an antibody from a human survivor. Nature 2008, 454, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Dube, D.; Brecher, M.B.; Delos, S.E.; Rose, S.C.; Park, E.W.; Schornberg, K.L.; Kuhn, J.H.; White, J.M. The primed ebolavirus glycoprotein (19-kilodalton GP1,2): Sequence and residues critical for host cell binding. J. Virol. 2009, 83, 2883–2891. [Google Scholar] [CrossRef] [PubMed]
- Barrientos, L.G.; Martin, A.M.; Rollin, P.E.; Sanchez, A. Disulfide bond assignment of the Ebola virus secreted glycoprotein SGP. Biochem. Biophys. Res. Commun. 2004, 323, 696–702. [Google Scholar] [CrossRef] [PubMed]
- Falzarano, D.; Krokhin, O.; Wahl-Jensen, V.; Seebach, J.; Wolf, K.; Schnittler, H.J.; Feldmann, H. Structure-function analysis of the soluble glycoprotein, sGP, of Ebola virus. Chembiochem 2006, 7, 1605–1611. [Google Scholar] [CrossRef] [PubMed]
- Lopez, L.A.; Yang, S.J.; Exline, C.M.; Rengarajan, S.; Haworth, K.G.; Cannon, P.M. Anti-tetherin activities of HIV-1 Vpu and Ebola virus glycoprotein do not involve removal of tetherin from lipid rafts. J. Virol. 2012, 86, 5467–5480. [Google Scholar] [CrossRef] [PubMed]
- Lopez, L.A.; Yang, S.J.; Hauser, H.; Exline, C.M.; Haworth, K.G.; Oldenburg, J.; Cannon, P.M. Ebola virus glycoprotein counteracts BST-2/tetherin restriction in a sequence-independent manner that does not require tetherin surface removal. J. Virol. 2010, 84, 7243–7255. [Google Scholar] [CrossRef] [PubMed]
- Gustin, J.K.; Bai, Y.; Moses, A.V.; Douglas, J.L. Ebola virus glycoprotein promotes enhanced viral egress by preventing Ebola VP40 from associating with the host restriction factor BST2/tetherin. J. Infect. Dis. 2015, 211 (Suppl. S2), S181–S190. [Google Scholar] [CrossRef] [PubMed]
- Kuhl, A.; Banning, C.; Marzi, A.; Votteler, J.; Steffen, I.; Bertram, S.; Glowacka, I.; Konrad, A.; Sturzl, M.; Guo, J.T.; et al. The ebola virus glycoprotein and HIV-1 vpu employ different strategies to counteract the antiviral factor tetherin. J. Infect. Dis. 2011, 204 (Suppl. S3), S850–S860. [Google Scholar] [CrossRef] [PubMed]
- Gnirss, K.; Fiedler, M.; Kramer-Kuhl, A.; Bolduan, S.; Mittler, E.; Becker, S.; Schindler, M.; Pohlmann, S. Analysis of determinants in filovirus glycoproteins required for tetherin antagonism. Viruses 2014, 6, 1654–1671. [Google Scholar] [CrossRef] [PubMed]
- Binley, J.M.; Sanders, R.W.; Clas, B.; Schuelke, N.; Master, A.; Guo, Y.; Kajumo, F.; Anselma, D.J.; Maddon, P.J.; Olson, W.C.; et al. A recombinant human immunodeficiency virus type 1 envelope glycoprotein complex stabilized by an intermolecular disulfide bond between the gp120 and gp41 subunits is an antigenic mimic of the trimeric virion-associated structure. J. Virol. 2000, 74, 627–643. [Google Scholar] [CrossRef] [PubMed]
- Haines, K.M.; Vande Burgt, N.H.; Francica, J.R.; Kaletsky, R.L.; Bates, P. Chinese hamster ovary cell lines selected for resistance to ebolavirus glycoprotein mediated infection are defective for NPC1 expression. Virology 2012, 432, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Kaletsky, R.L.; Simmons, G.; Bates, P. Proteolysis of the Ebola virus glycoproteins enhances virus binding and infectivity. J. Virol. 2007, 81, 13378–13384. [Google Scholar] [CrossRef] [PubMed]
- Masuyama, N.; Kuronita, T.; Tanaka, R.; Muto, T.; Hirota, Y.; Takigawa, A.; Fujita, H.; Aso, Y.; Amano, J.; Tanaka, Y. HM1.24 is internalized from lipid rafts by clathrin-mediated endocytosis through interaction with alpha-adaptin. J. Biol. Chem. 2009, 284, 15927–15941. [Google Scholar] [CrossRef] [PubMed]
- Bavari, S.; Bosio, C.M.; Wiegand, E.; Ruthel, G.; Will, A.B.; Geisbert, T.W.; Hevey, M.; Schmaljohn, C.; Schmaljohn, A.; Aman, M.J. Lipid raft microdomains: A gateway for compartmentalized trafficking of Ebola and Marburg viruses. J. Exp. Med. 2002, 195, 593–602. [Google Scholar] [CrossRef] [PubMed]
- Bates, P.; Young, J.A.; Varmus, H.E. A receptor for subgroup a rous sarcoma virus is related to the low density lipoprotein receptor. Cell 1993, 74, 1043–1051. [Google Scholar] [CrossRef]
- Narayan, S.; Barnard, R.J.; Young, J.A. Two retroviral entry pathways distinguished by lipid raft association of the viral receptor and differences in viral infectivity. J. Virol. 2003, 77, 1977–1983. [Google Scholar] [CrossRef] [PubMed]
- Ito, H.; Watanabe, S.; Takada, A.; Kawaoka, Y. Ebola virus glycoprotein: Proteolytic processing, acylation, cell tropism, and detection of neutralizing antibodies. J. Virol. 2001, 75, 1576–1580. [Google Scholar] [CrossRef] [PubMed]
- Perez-Caballero, D.; Zang, T.; Ebrahimi, A.; McNatt, M.W.; Gregory, D.A.; Johnson, M.C.; Bieniasz, P.D. Tetherin inhibits HIV-1 release by directly tethering virions to cells. Cell 2009, 139, 499–511. [Google Scholar] [CrossRef] [PubMed]
- Carette, J.E.; Raaben, M.; Wong, A.C.; Herbert, A.S.; Obernosterer, G.; Mulherkar, N.; Kuehne, A.I.; Kranzusch, P.J.; Griffin, A.M.; Ruthel, G.; et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature 2011, 477, 340–343. [Google Scholar] [CrossRef] [PubMed]
- Cote, M.; Misasi, J.; Ren, T.; Bruchez, A.; Lee, K.; Filone, C.M.; Hensley, L.; Li, Q.; Ory, D.; Chandran, K.; et al. Small molecule inhibitors reveal Niemann-Pick C1 is essential for ebola virus infection. Nature 2011, 477, 344–348. [Google Scholar] [CrossRef] [PubMed]
- Chandran, K.; Sullivan, N.J.; Felbor, U.; Whelan, S.P.; Cunningham, J.M. Endosomal proteolysis of the Ebola virus glycoprotein is necessary for infection. Science 2005, 308, 1643–1645. [Google Scholar] [CrossRef] [PubMed]
- Schornberg, K.; Matsuyama, S.; Kabsch, K.; Delos, S.; Bouton, A.; White, J. Role of endosomal cathepsins in entry mediated by the Ebola virus glycoprotein. J. Virol. 2006, 80, 4174–4178. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.K.; Hue, S.; Schaller, T.; Verschoor, E.; Pillay, D.; Towers, G.J. Mutation of a single residue renders human tetherin resistant to HIV-1 Vpu-mediated depletion. PLoS Pathog. 2009, 5, e1000443. [Google Scholar] [CrossRef] [PubMed]
- Rong, L.; Zhang, J.; Lu, J.; Pan, Q.; Lorgeoux, R.P.; Aloysius, C.; Guo, F.; Liu, S.L.; Wainberg, M.A.; Liang, C. The transmembrane domain of BST-2 determines its sensitivity to down-modulation by human immunodeficiency virus type 1 vpu. J. Virol. 2009, 83, 7536–7546. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Ode, H.; Yoshida, T.; Sato, K.; Gee, P.; Yamamoto, S.P.; Ebina, H.; Strebel, K.; Sato, H.; Koyanagi, Y. Identification of amino acids in the human tetherin transmembrane domain responsible for HIV-1 Vpu interaction and susceptibility. J. Virol. 2011, 85, 932–945. [Google Scholar] [CrossRef] [PubMed]
- McNatt, M.W.; Zang, T.; Hatziioannou, T.; Bartlett, M.; Fofana, I.B.; Johnson, W.E.; Neil, S.J.; Bieniasz, P.D. Species-specific activity of HIV-1 Vpu and positive selection of tetherin transmembrane domain variants. PLoS Pathog. 2009, 5, e1000300. [Google Scholar] [CrossRef] [PubMed]
- Sauter, D.; Schindler, M.; Specht, A.; Landford, W.N.; Munch, J.; Kim, K.A.; Votteler, J.; Schubert, U.; Bibollet-Ruche, F.; Keele, B.F.; et al. Tetherin-driven adaptation of Vpu and Nef function and the evolution of pandemic and nonpandemic HIV-1 strains. Cell Host Microbe 2009, 6, 409–421. [Google Scholar] [CrossRef] [PubMed]
- Panchal, R.G.; Ruthel, G.; Kenny, T.A.; Kallstrom, G.H.; Lane, D.; Badie, S.S.; Li, L.; Bavari, S.; Aman, M.J. In vivo oligomerization and raft localization of Ebola virus protein VP40 during vesicular budding. Proc. Natl. Acad. Sci. USA 2003, 100, 15936–15941. [Google Scholar] [CrossRef] [PubMed]
- Fritz, J.V.; Tibroni, N.; Keppler, O.T.; Fackler, O.T. HIV-1 Vpu’s lipid raft association is dispensable for counteraction of the particle release restriction imposed by CD317/tetherin. Virology 2012, 424, 33–44. [Google Scholar] [CrossRef] [PubMed]
- Le Tortorec, A.; Neil, S.J. Antagonism to and intracellular sequestration of human tetherin by the human immunodeficiency virus type 2 envelope glycoprotein. J. Virol. 2009, 83, 11966–11978. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vande Burgt, N.H.; Kaletsky, R.L.; Bates, P. Requirements within the Ebola Viral Glycoprotein for Tetherin Antagonism. Viruses 2015, 7, 5587-5602. https://doi.org/10.3390/v7102888
Vande Burgt NH, Kaletsky RL, Bates P. Requirements within the Ebola Viral Glycoprotein for Tetherin Antagonism. Viruses. 2015; 7(10):5587-5602. https://doi.org/10.3390/v7102888
Chicago/Turabian StyleVande Burgt, Nathan H., Rachel L. Kaletsky, and Paul Bates. 2015. "Requirements within the Ebola Viral Glycoprotein for Tetherin Antagonism" Viruses 7, no. 10: 5587-5602. https://doi.org/10.3390/v7102888
APA StyleVande Burgt, N. H., Kaletsky, R. L., & Bates, P. (2015). Requirements within the Ebola Viral Glycoprotein for Tetherin Antagonism. Viruses, 7(10), 5587-5602. https://doi.org/10.3390/v7102888




