Canine Enteric Coronaviruses: Emerging Viral Pathogens with Distinct Recombinant Spike Proteins
Abstract
:1. Introduction and Background
2. CCoV Structure and Genotyping
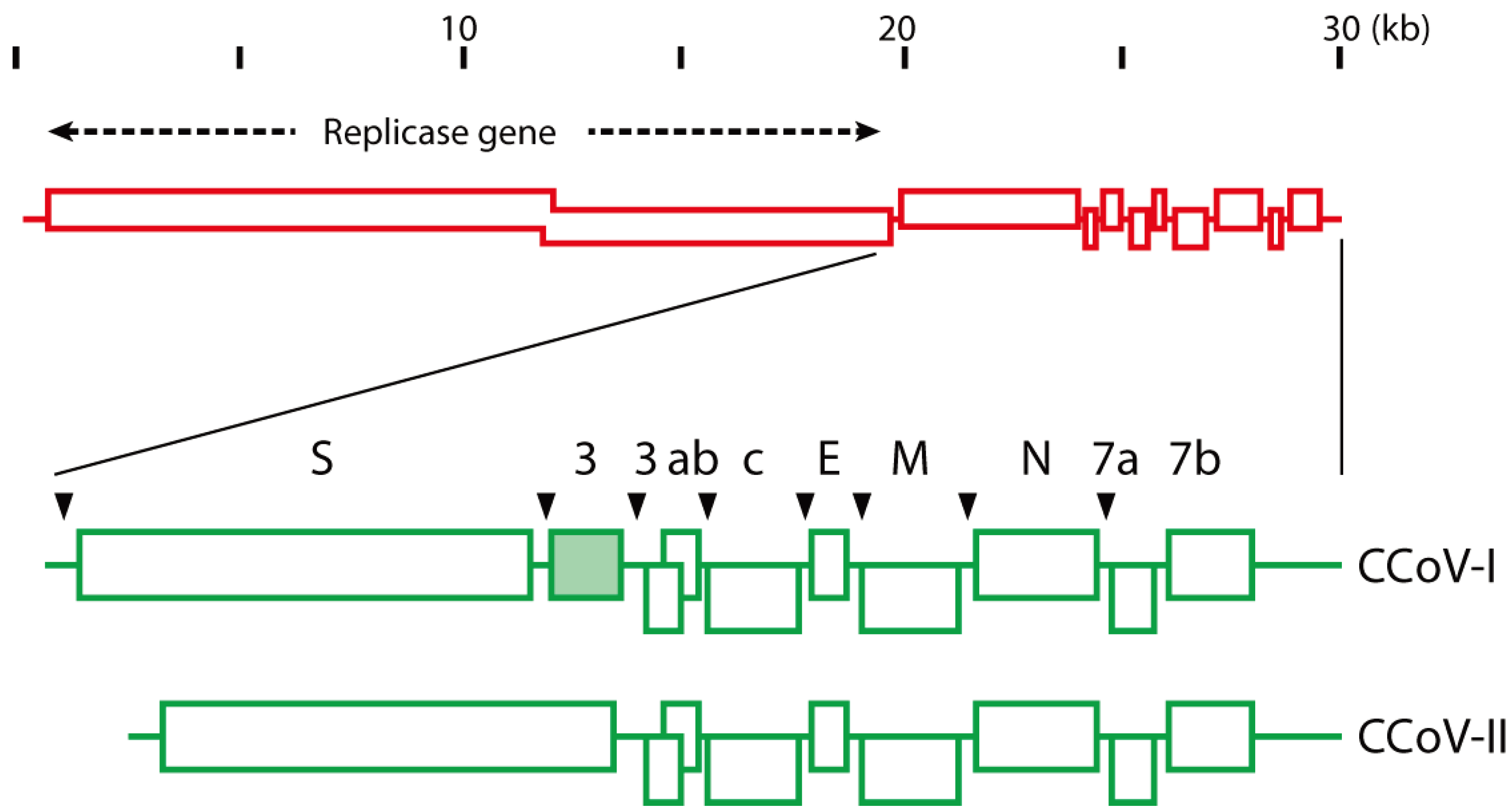
3. Functional Aspects of the CCoV Spike Protein
3.1. Receptor Binding and Host Tropism
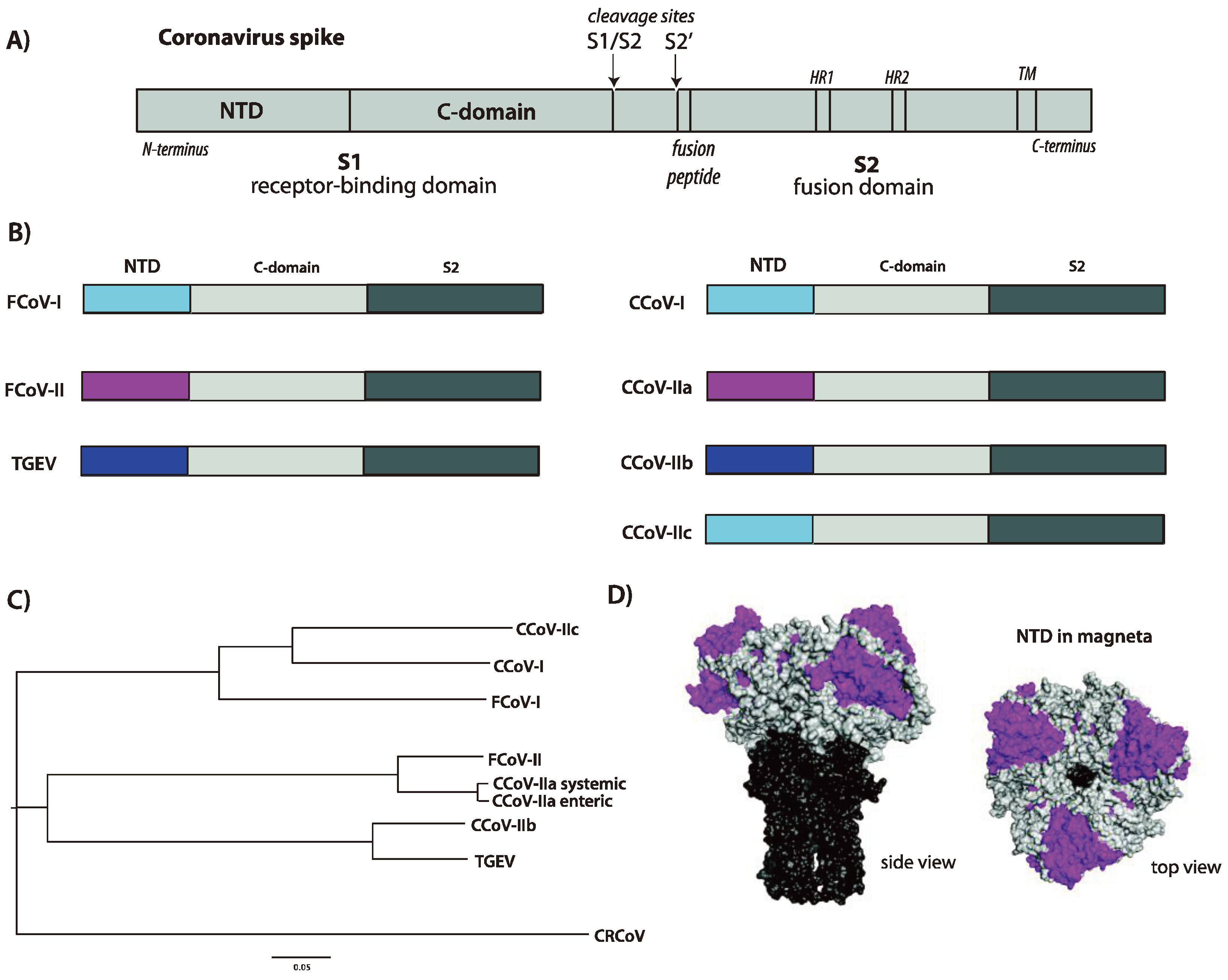
3.2. Activation by Proteolytic Cleavage
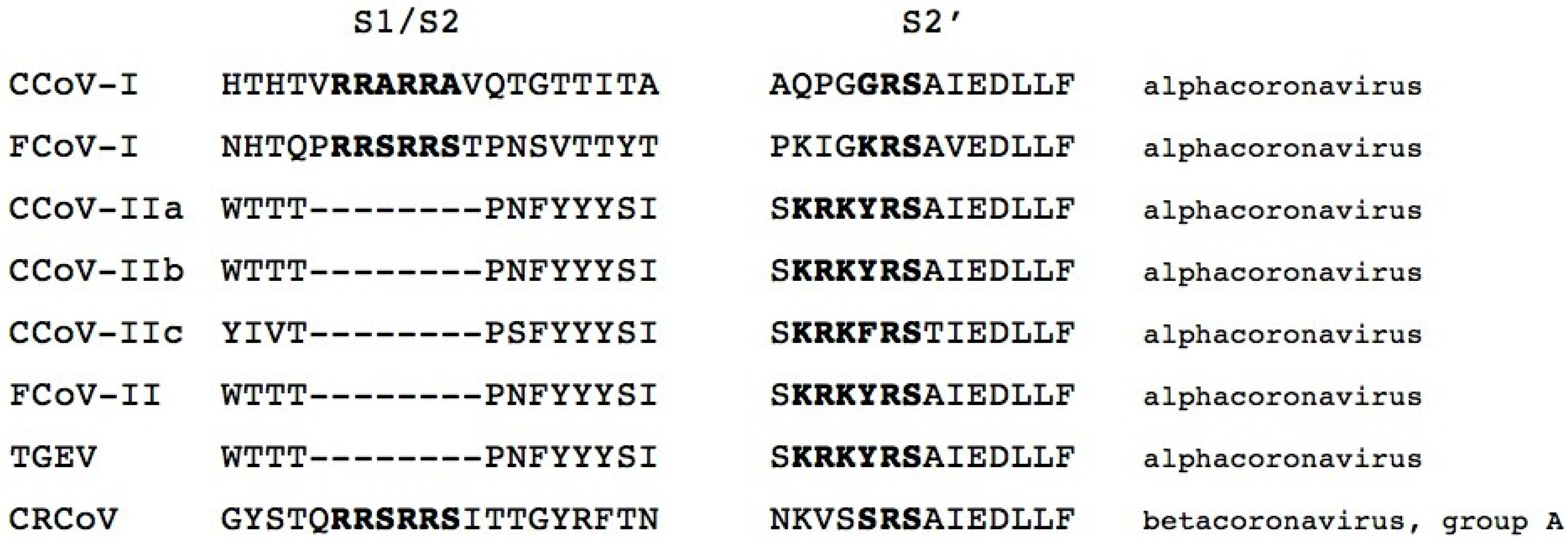

4. CCoV Pathogenesis and Clinical Presentation
5. Recombination and Emergence of CCoV Variants with Altered Pathogenicity
6. Laboratory Diagnosis
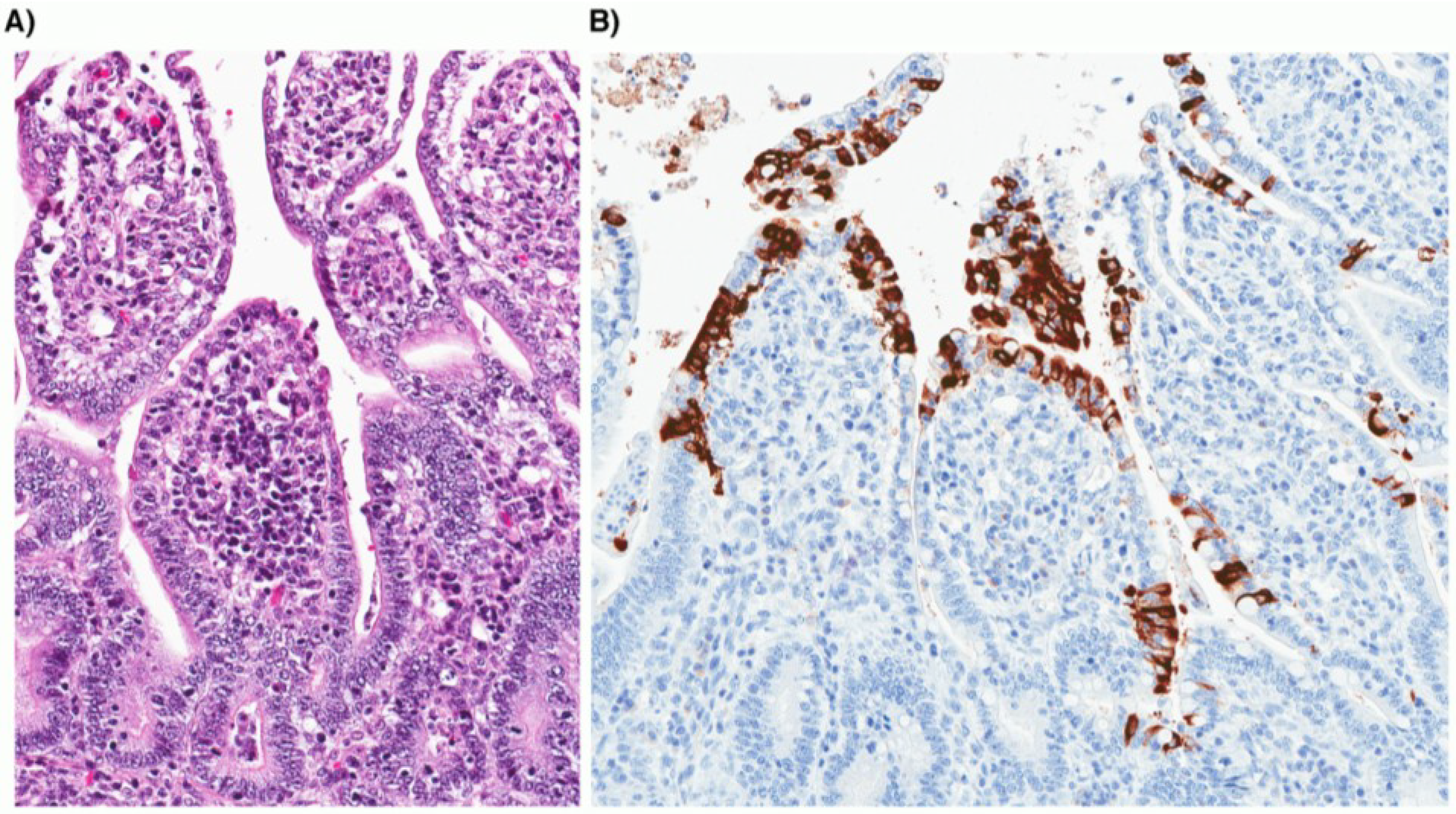
7. Vaccination and Treatment
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Binn, L.; Lazar, E.C.; Keenan, K.P.; Huxsoll, D.L.; Marchwicki, R.H.; Strano, A.J. Recovery and characterization of a coronavirus from military dogs with diarrhea. Proc. Annu. Meet. U. S. Anim. Health Assoc. 1974, 78, 359–366. [Google Scholar] [PubMed]
- Keenan, K.P.; Jervis, H.R.; Marchwicki, R.H.; Binn, L.N. Intestinal infection of neonatal dogs with canine coronavirus 1-71: Studies by virologic, histologic, histochemical, and immunofluorescent techniques. Am. J. Vet. Res. 1976, 37, 247–256. [Google Scholar] [PubMed]
- Perlman, S.; Gallagher, T.; Snijder, E.J. Nidoviruses; ASM Press: Washington, DC, USA, 2008. [Google Scholar]
- Lai, M.M.C.; Holmes, K.V. Coronaviridae: The Viruses and Their Replication. In Fields Virology; Knipe, D.M., Howely, P.M., Eds.; Lippincott Wilkins and Williams: Philadelphia, PA, USA, 2001. [Google Scholar]
- Hogue, B.G.; Machamer, C.E. Coronavirus Structural Proteins and Virus Assembly. In Nidoviruses; Perlman, S., Gallagher, T., Snijder, E.J., Eds.; ASM Press: Washington, DC, USA, 2008; pp. 179–200. [Google Scholar]
- Bosch, B.J.; Rottier, P.J. Nidovirus Entry into Cells. In Nidoviruses; Perlman, S., Gallagher, T., Snijder, E.J., Eds.; ASM Press: Washington, DC, USA, 2008; pp. 157–178. [Google Scholar]
- Netherton, C.L.; Wileman, T. Virus factories, double membrane vesicles and viroplasm generated in animal cells. Curr. Opin. Virol. 2011, 1, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Masters, P.S.; Perlman, S. Coronaviridae. In Fields Virology, 6th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott, Williams and Wilkins: Philadelphia, PA, USA, 2013; pp. 825–858. [Google Scholar]
- Denison, M.R.; Graham, R.L.; Donaldson, E.F.; Eckerle, L.D.; Baric, R.S. Coronaviruses: An RNA proofreading machine regulates replication fidelity and diversity. RNA Biol. 2011, 8, 270–279. [Google Scholar] [CrossRef] [PubMed]
- Eaton, B.T.; Wang, L.F. Bats, civets and the emergence of SARS. Curr. Top Microbiol. Immunol. 2007, 315, 325–344. [Google Scholar] [PubMed]
- King, A.M.Q.; Lefkowitz, E.; Adams, M.J.; Carstens, E.B. Virus Taxonomy: IXth Report of the International Committee on Taxonomy of Viruses; Elsevier: Amsterdam, The Netherlands, 2011. [Google Scholar]
- Woo, P.C.; Huang, Y.; Lau, S.K.; Yuen, K.Y. Coronavirus genomics and bioinformatics analysis. Viruses 2010, 2, 1804–1820. [Google Scholar] [CrossRef] [PubMed]
- Erles, K.; Toomey, C.; Brooks, H.W.; Brownlie, J. Detection of a group 2 coronavirus in dogs with canine infectious respiratory disease. Virology 2003, 310, 216–223. [Google Scholar] [CrossRef] [PubMed]
- Erles, K.; Shiu, K.B.; Brownlie, J. Isolation and sequence analysis of canine respiratory coronavirus. Virus Res. 2007, 124, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Decaro, N.; Buonavoglia, C. An update on canine coronaviruses: Viral evolution and pathobiology. Vet. Microbiol. 2008, 132, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Le Poder, S. Feline and canine coronaviruses: Common genetic and pathobiological features. Adv. Virol. 2011, 2011, 11. [Google Scholar]
- Pratelli, A.; Martella, V.; Pistello, M.; Elia, G.; Decaro, N.; Buonavoglia, D.; Camero, M.; Tempesta, M.; Buonavoglia, C. Identification of coronaviruses in dogs that segregate separately from the canine coronavirus genotype. J. Virol. Methods 2003, 107, 213–222. [Google Scholar] [CrossRef] [PubMed]
- Haijema, B.J.; Rottier, P.J.; de Groot, R.J. Feline Coronaviruses: A Tale of Two-Faced Types. In Coronaviruses. Molecular and Cellular Biology; Thiel, V., Ed.; Caister Academic Press: Norfolk, UK, 2007; pp. 183–203. [Google Scholar]
- Lorusso, A.; Decaro, N.; Schellen, P.; Rottier, P.J.; Buonavoglia, C.; Haijema, B.J.; de Groot, R.J. Gain, preservation, and loss of a group 1a coronavirus accessory glycoprotein. J. Virol. 2008, 82, 10312–10317. [Google Scholar] [CrossRef] [PubMed]
- Pratelli, A.; Martella, V.; Elia, G.; Decaro, N.; Aliberti, A.; Buonavoglia, D.; Tempesta, M.; Buonavoglia, C. Variation of the sequence in the gene encoding for transmembrane protein M of canine coronavirus (CCV). Mol. Cell. Probes 2001, 15, 229–233. [Google Scholar] [CrossRef] [PubMed]
- Pratelli, A.; Martella, V.; Decaro, N.; Tinelli, A.; Camero, M.; Cirone, F.; Elia, G.; Cavalli, A.; Corrente, M.; Greco, G.; et al. Genetic diversity of a canine coronavirus detected in pups with diarrhoea in Italy. J. Virol. Methods 2003, 110, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Decaro, N.; Martella, V.; Ricci, D.; Elia, G.; Desario, C.; Campolo, M.; Cavaliere, N.; di Trani, L.; Tempesta, M.; Buonavoglia, C. Genotype-specific fluorogenic RT-PCR assays for the detection and quantitation of canine coronavirus type I and type II RNA in faecal samples of dogs. J. Virol. Methods 2005, 130, 72–78. [Google Scholar] [CrossRef] [PubMed]
- Decaro, N.; Buonavoglia, C. Canine coronavirus: Not only an enteric pathogen. Vet. Clin. N. Am. Small Anim. Pract. 2011, 41, 1121–1132. [Google Scholar] [CrossRef]
- Buonavoglia, C.; Decaro, N.; Martella, V.; Elia, G.; Campolo, M.; Desario, C.; Castagnaro, M.; Tempesta, M. Canine coronavirus highly pathogenic for dogs. Emerg. Infect. Dis. 2006, 12, 492–494. [Google Scholar] [CrossRef] [PubMed]
- Ntafis, V.; Mari, V.; Decaro, N.; Papanastassopoulou, M.; Papaioannou, N.; Mpatziou, R.; Buonavoglia, C.; Xylouri, E. Isolation, tissue distribution and molecular characterization of two recombinant canine coronavirus strains. Vet. Microbiol. 2011, 151, 238–244. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Ma, G.; Lu, C.; Wen, H. Detection of canine coronaviruses genotype I and II in raised Canidae animals in China. Berl. Munch. Tierarztl. Wochenschr. 2006, 119, 35–39. [Google Scholar] [PubMed]
- Soma, T.; Ohinata, T.; Ishii, H.; Takahashi, T.; Taharaguchi, S.; Hara, M. Detection and genotyping of canine coronavirus RNA in diarrheic dogs in Japan. Res. Vet. Sci. 2011, 90, 205–207. [Google Scholar] [CrossRef] [PubMed]
- Costa, E.M.; de Castro, T.X.; Bottino Fde, O.; Garcia Rde, C. Molecular characterization of canine coronavirus strains circulating in Brazil. Vet. Microbiol. 2013, 168, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Regan, A.D.; Millet, J.K.; Tse, L.P.; Chillag, Z.; Rinaldi, V.D.; Licitra, B.N.; Dubovi, E.J.; Town, C.D.; Whittaker, G.R. Characterization of a recombinant canine coronavirus with a distinct receptor-binding (S1) domain. Virology 2012, 430, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Escutenaire, S.; Isaksson, M.; Renstrom, L.H.; Klingeborn, B.; Buonavoglia, C.; Berg, M.; Belak, S.; Thoren, P. Characterization of divergent and atypical canine coronaviruses from Sweden. Arch. Virol. 2007, 152, 1507–1514. [Google Scholar] [CrossRef] [PubMed]
- Krempl, C.; Schultze, B.; Laude, H.; Herrler, G. Point mutations in the S protein connect the sialic acid binding activity with the enteropathogenicity of transmissible gastroenteritis coronavirus. J. Virol. 1997, 71, 3285–3287. [Google Scholar] [PubMed]
- Schultze, B.; Krempl, C.; Ballesteros, M.L.; Shaw, L.; Schauer, R.; Enjuanes, L.; Herrler, G. Transmissible Gastroenteritis Coronavirus, but Not the Related Porcine Respiratory Coronavirus, Has a Sialic Acid (N-Glycolylneuraminic Acid) Binding Activity? J. Virol. 1996, 70, 5634–5637. [Google Scholar]
- Decaro, N.; Campolo, M.; Lorusso, A.; Desario, C.; Mari, V.; Colaianni, M.L.; Elia, G.; Martella, V.; Buonavoglia, C. Experimental infection of dogs with a novel strain of canine coronavirus causing systemic disease and lymphopenia. Vet. Microbiol. 2008, 128, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Decaro, N.; Cordonnier, N.; Demeter, Z.; Egberink, H.; Elia, G.; Grellet, A.; le Poder, S.; Mari, V.; Martella, V.; Ntafis, V.; et al. European surveillance for pantropic canine coronavirus. J. Clin. Microbiol. 2012, 51, 83–88. [Google Scholar] [CrossRef] [PubMed]
- Marinaro, M.; Mari, V.; Bellacicco, A.L.; Tarsitano, E.; Elia, G.; Losurdo, M.; Rezza, G.; Buonavoglia, C.; Decaro, N. Prolonged depletion of circulating CD4+ T lymphocytes and acute monocytosis after pantropic canine coronavirus infection in dogs. Virus Res. 2010, 152, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Decaro, N.; Martella, V.; Elia, G.; Campolo, M.; Desario, C.; Cirone, F.; Tempesta, M.; Buonavoglia, C. Molecular characterisation of the virulent canine coronavirus CB/05 strain. Virus Res. 2007, 125, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Zappulli, V.; Caliari, D.; Cavicchioli, L.; Tinelli, A.; Castagnaro, M. Systemic fatal type II coronavirus infection in a dog: Pathological findings and immunohistochemistry. Res. Vet. Sci. 2008, 84, 278–282. [Google Scholar] [CrossRef] [PubMed]
- Decaro, N.; Mari, V.; Campolo, M.; Lorusso, A.; Camero, M.; Elia, G.; Martella, V.; Cordioli, P.; Enjuanes, L.; Buonavoglia, C. Recombinant canine coronaviruses related to transmissible gastroenteritis virus of Swine are circulating in dogs. J. Virol. 2009, 83, 1532–1537. [Google Scholar] [CrossRef] [PubMed]
- Wesley, R.D. The S gene of canine coronavirus, strain UCD-1, is more closely related to the S gene of transmissible gastroenteritis virus than to that of feline infectious peritonitis virus. Virus Res. 1999, 61, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Carmichael, L.E. Canine viral vaccines at a turning point—A personal perspective. Adv. Vet. Med. 1999, 41, 289–307. [Google Scholar] [PubMed]
- Gallagher, T.; Buchmeier, M.J. Coronavirus spike proteins in viral entry and pathogenesis. Virology 2001, 279, 371–374. [Google Scholar] [CrossRef] [PubMed]
- Wentworth, D.; Holmes, K.V. Molecular determinants of species specificity in the coronavirus receptor aminopeptidase N (CD13): Influence of N-linked glycosylation. J. Virol. 2001, 75, 9741–9752. [Google Scholar] [CrossRef] [PubMed]
- Tresnan, D.B.; Levis, R.; Holmes, K.V. Feline aminopeptidase N serves as a receptor for feline, canine, porcine, and human coronaviruses in serogroup I. J. Virol. 1996, 70, 8669–8674. [Google Scholar] [PubMed]
- Hofmann, H.; Pyrc, K.; van der Hoek, L.; Geier, M.; Berkhout, B.; Pohlmann, S. Human coronavirus NL63 employs the severe acute respiratory syndrome coronavirus receptor for cellular entry. Proc. Natl. Acad. Sci. USA 2005, 102, 7988–7993. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Tree Figure Drawing Tool, v.1.4.0.; Andrew Rambaut, I.O.E.B., University of Edinburgh: Edinburgh, UK, 2006–2012.
- Bernini, A.; Spiga, O.; Ciutti, A.; Chiellini, S.; Bracci, L.; Yan, X.; Zheng, B.; Huang, J.; He, M.L.; Song, H.D.; et al. Prediction of quaternary assembly of SARS coronavirus peplomer. Biochem. Biophys. Res. Commun. 2004, 325, 1210–1214. [Google Scholar] [CrossRef] [PubMed]
- Dye, C.; Temperton, N.; Siddell, S.G. Type I feline coronavirus spike glycoprotein fails to recognize aminopeptidase N as a functional receptor on feline cell lines. J. Gen. Virol. 2007, 88, 1753–1760. [Google Scholar] [CrossRef] [PubMed]
- Hohdatsu, T.; Izumiya, Y.; Yokoyama, Y.; Kida, K.; Koyama, H. Differences in virus receptor for type I and type II feline infectious peritonitis virus. Arch. Virol. 1998, 143, 839–850. [Google Scholar] [CrossRef] [PubMed]
- Van Hamme, E.; Desmarets, L.; Dewerchin, H.L.; Nauwynck, H.J. Intriguing interplay between feline infectious peritonitis virus and its receptors during entry in primary feline monocytes. Virus Res. 2011, 160, 32–39. [Google Scholar]
- Regan, A.D.; Ousterout, D.G.; Whittaker, G.R. Feline lectin activity is critical for the cellular entry of feline infectious peritonitis virus. J. Virol. 2010, 84, 7917–7921. [Google Scholar] [CrossRef] [PubMed]
- Regan, A.D.; Whittaker, G.R. Utilization of DC-SIGN for entry of feline coronaviruses into host cells. J. Virol. 2008, 82, 11992–11996. [Google Scholar] [CrossRef] [PubMed]
- Klenk, H.-D.; Garten, W. Activation cleavage of viral spike proteins by host proteases. In Cellular Receptors for Animal Viruses; Wimmer, E., Ed.; Cold Spring Harbor Press: Cold Spring Harbor, NY, USA, 1994; pp. 241–280. [Google Scholar]
- Belouzard, S.; Chu, V.C.; Whittaker, G.R. Activation of the SARS coronavirus spike protein via sequential proteolytic cleavage at two distinct sites. Proc. Natl. Acad. Sci. USA 2009, 106, 5871–5876. [Google Scholar] [CrossRef] [PubMed]
- Madu, I.G.; Roth, S.L.; Belouzard, S.; Whittaker, G.R. Characterization of a highly conserved domain within the severe acute respiratory syndrome coronavirus spike protein S2 domain with characteristics of a viral fusion peptide. J. Virol. 2009, 83, 7411–7421. [Google Scholar] [CrossRef] [PubMed]
- Wicht, O.; Burkard, C.; de Haan, C.A.; van Kuppeveld, F.J.; Rottier, P.J.; Bosch, B.J. Identification and Characterization of a Proteolytically Primed Form of the Murine Coronavirus Spike Proteins after Fusion with the Target Cell. J. Virol. 2014, 88, 4943–4952. [Google Scholar] [CrossRef] [PubMed]
- Belouzard, S.; Millet, J.K.; Licitra, B.N.; Whittaker, G.R. Mechanisms of coronavirus cell entry mediated by the viral spike protein. Viruses 2012, 4, 1011–1033. [Google Scholar] [CrossRef] [PubMed]
- Licitra, B.N.; Millet, J.K.; Regan, A.D.; Hamilton, B.S.; Rinaldi, V.; Duhamel, G.E.; Whittaker, G.R. Mutation in spike protein cleavage site and pathogenesis of feline coronavirus. Emerg. Infect. Dis. 2013, 19, 1066–1073. [Google Scholar] [CrossRef] [PubMed]
- Greene, C.E.; Decaro, N. Canine Viral Enteritis. In Infectious Diseases of the Dog and Cat; Greene, C.E., Ed.; Elsevier Saunders: St. Louis, MO, USA, 2012. [Google Scholar]
- Pollock, R.V.; Carmichael, L.E. Canine viral enteritis. Vet. Clin. N. Am. Small Anim. Pract. 1983, 13, 551–566. [Google Scholar]
- Saif, L.J.; Theil, K.W. Viral Diarrheas of Man and Animals; CRC Press: Boca Raton, FL, USA, 1989. [Google Scholar]
- Pratelli, A.; Tempesta, M.; Roperto, F.P.; Sagazio, P.; Carmichael, L.; Buonavoglia, C. Fatal coronavirus infection in puppies following canine parvovirus 2b infection. J. Vet. Diagn. Investig. 1999, 11, 550–553. [Google Scholar] [CrossRef]
- Apple, M.J.G. Does canine coronavirus augment the effects of parvovirus infection? Vet. Med. 1988, 36, 360–366. [Google Scholar]
- Evermann, J.; Abbott, J.R.; Han, S. Canine coronavirus-associated puppy mortality without evidence of concurrent canine parvovirus infection. J. Vet. Diagn. Investig. 2005, 17, 610–614. [Google Scholar] [CrossRef]
- Naylor, M.J.; Walia, C.S.; Mcorist, S.; Lehrbach, P.R.; Deane, E.M.; Harrison, G.A. Molecular characterization confirms the presence of a divergent strain of canine coronavirus (UWSMN-1) in Australia. J. Clin. Microbiol. 2002, 40, 3518–3522. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Morgado, J.M.; Poynter, S.; Morris, T.H. Molecular characterization of a virulent canine coronavirus BGF strain. Virus Res. 2004, 104, 27–31. [Google Scholar]
- Naylor, M.J.; Monckton, R.P.; Lehrbach, P.R.; Deane, E.M. Canine coronavirus in Australian dogs. Aust. Vet. J. 2001, 79, 116–119. [Google Scholar] [CrossRef] [PubMed]
- Naylor, M.J.; Harrison, G.A.; Monckton, R.P.; Mcorist, S.; Lehrbach, P.R.; Deane, E.M. Identification of canine coronavirus strains from feces by S gene nested PCR and molecular characterization of a new Australian isolate. J. Clin. Microbiol. 2001, 39, 1036–1041. [Google Scholar] [CrossRef] [PubMed]
- Zicola, A.; Jolly, S.; Mathijs, E.; Ziant, D.; Decaro, N.; Mari, V.; Thiry, E. Fatal outbreaks in dogs associated with pantropic canine coronavirus in France and Belgium. J. Small Anim. Pract. 2012, 53, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Graham, R.L.; Baric, R.S. Recombination, reservoirs, and the modular spike: Mechanisms of coronavirus cross-species transmission. J. Virol. 2010, 84, 3134–3146. [Google Scholar] [CrossRef] [PubMed]
- Herrewegh, A.A.; Smeenk, I.; Horzinek, M.C.; Rottier, P.J.; de Groot, R.J. Feline coronavirus type II strains 79–1683 and 79–1146 originate from a double recombination between feline coronavirus type I and canine coronavirus. J. Virol. 1998, 72, 4508–4514. [Google Scholar] [PubMed]
- Le Poder, S.; Pham-Hung D’alexandry D’orangiani, A.L.; Duarte, L.; Fournier, A.; Horhogea, C.; Pinhas, C.; Vabret, A.; Eloit, M. Infection of cats with atypical feline coronaviruses harbouring a truncated form of the canine type I non-structural ORF3 gene. Infect. Genet. Evol. 2013, 20, 488–494. [Google Scholar]
- Decaro, N.; Pratelli, A.; Campolo, M.; Elia, G.; Martella, V.; Tempesta, M.; Buonavoglia, C. Quantitation of canine coronavirus RNA in the faeces of dogs by TaqMan RT-PCR. J. Virol. Methods 2004, 119, 145–150. [Google Scholar] [CrossRef] [PubMed]
- Gizzi, A.B.; Oliveira, S.T.; Leutenegger, C.M.; Estrada, M.; Kozemjakin, D.A.; Stedile, R.; Marcondes, M.; Biondo, A.W. Presence of infectious agents and co-infections in diarrheic dogs determined with a real-time polymerase chain reaction-based panel. BMC Vet. Res. 2014, 10, 23. [Google Scholar] [CrossRef] [PubMed]
- Herrewegh, A.A.; de Groot, R.J.; Cepica, A.; Egberink, H.F.; Horzinek, M.C.; Rottier, P.J. Detection of feline coronavirus RNA in feces, tissues, and body fluids of naturally infected cats by reverse transcriptase PCR. J. Clin. Microbiol. 1995, 33, 684–689. [Google Scholar] [PubMed]
- Greene, C.E.; Levy, J.K. Immunoprophylaxis. In Infectious Diseases of the Dog and Cat; Greene, C.E., Ed.; Elsevier Saunders: St. Louis, MO, USA, 2012. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Licitra, B.N.; Duhamel, G.E.; Whittaker, G.R. Canine Enteric Coronaviruses: Emerging Viral Pathogens with Distinct Recombinant Spike Proteins. Viruses 2014, 6, 3363-3376. https://doi.org/10.3390/v6083363
Licitra BN, Duhamel GE, Whittaker GR. Canine Enteric Coronaviruses: Emerging Viral Pathogens with Distinct Recombinant Spike Proteins. Viruses. 2014; 6(8):3363-3376. https://doi.org/10.3390/v6083363
Chicago/Turabian StyleLicitra, Beth N., Gerald E. Duhamel, and Gary R. Whittaker. 2014. "Canine Enteric Coronaviruses: Emerging Viral Pathogens with Distinct Recombinant Spike Proteins" Viruses 6, no. 8: 3363-3376. https://doi.org/10.3390/v6083363
APA StyleLicitra, B. N., Duhamel, G. E., & Whittaker, G. R. (2014). Canine Enteric Coronaviruses: Emerging Viral Pathogens with Distinct Recombinant Spike Proteins. Viruses, 6(8), 3363-3376. https://doi.org/10.3390/v6083363



