Cetacean Morbillivirus: Current Knowledge and Future Directions
Abstract
:1. Introduction
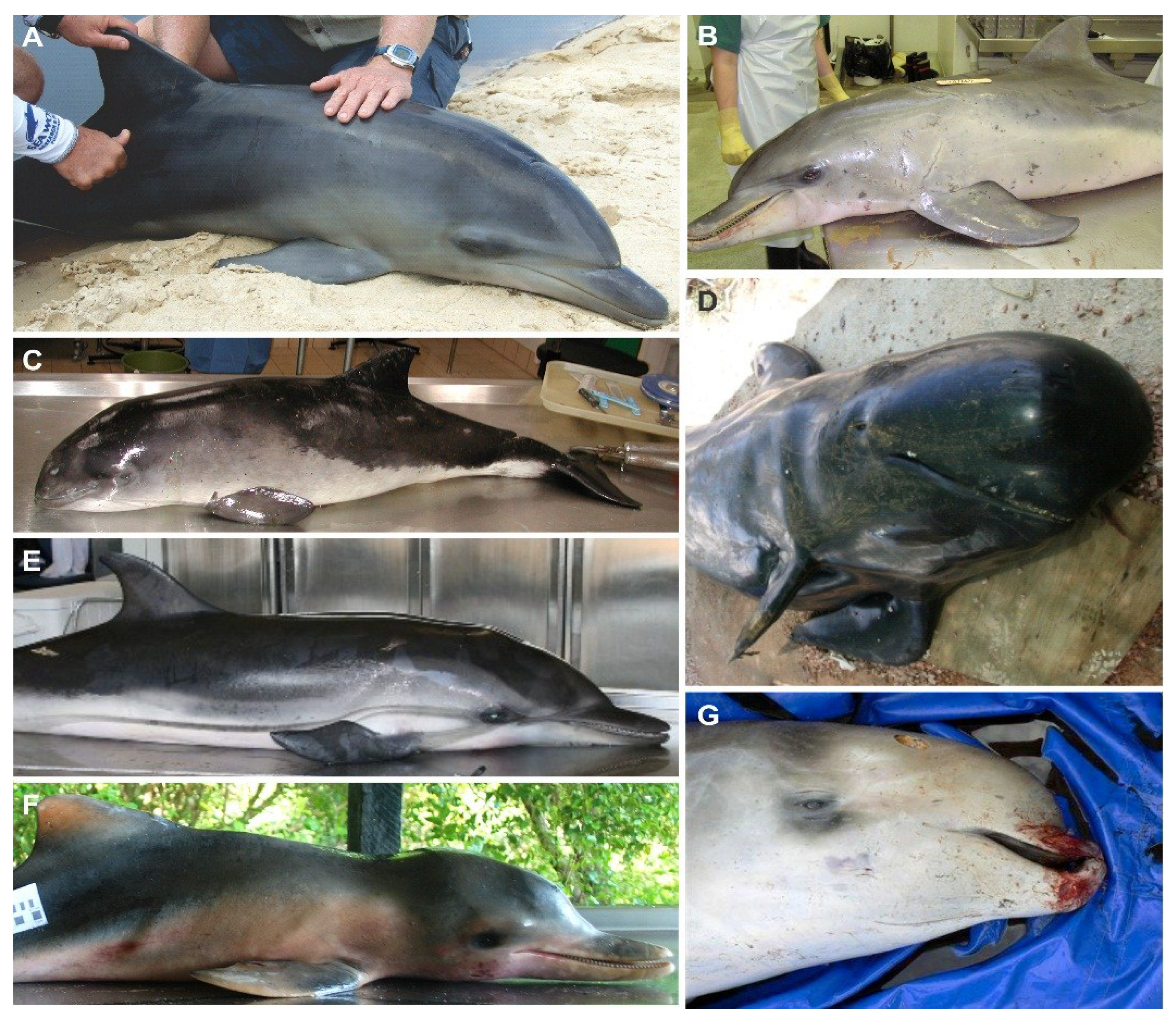

2. Antigenic and Molecular Characteristics of CeMV
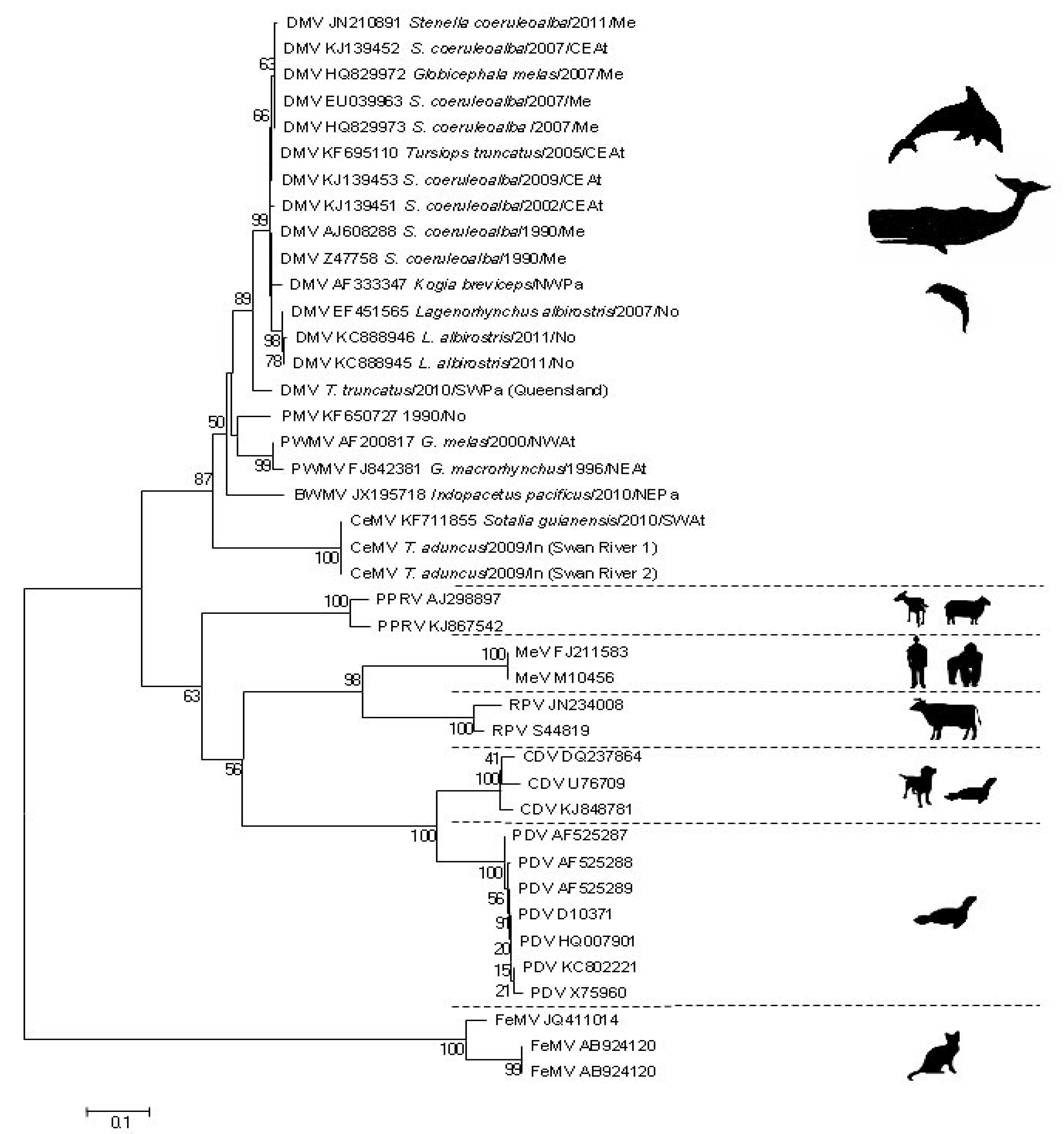
3. Mechanisms of Cellular Entry and Receptors
4. Diagnosis
4.1. Histology and Immunohistochemistry
4.2. Virus Isolation
4.3. Serology
4.4. Reverse Transcription Polymerase Chain Reaction
5. Pathology and Pathogenesis of CeMV Infection
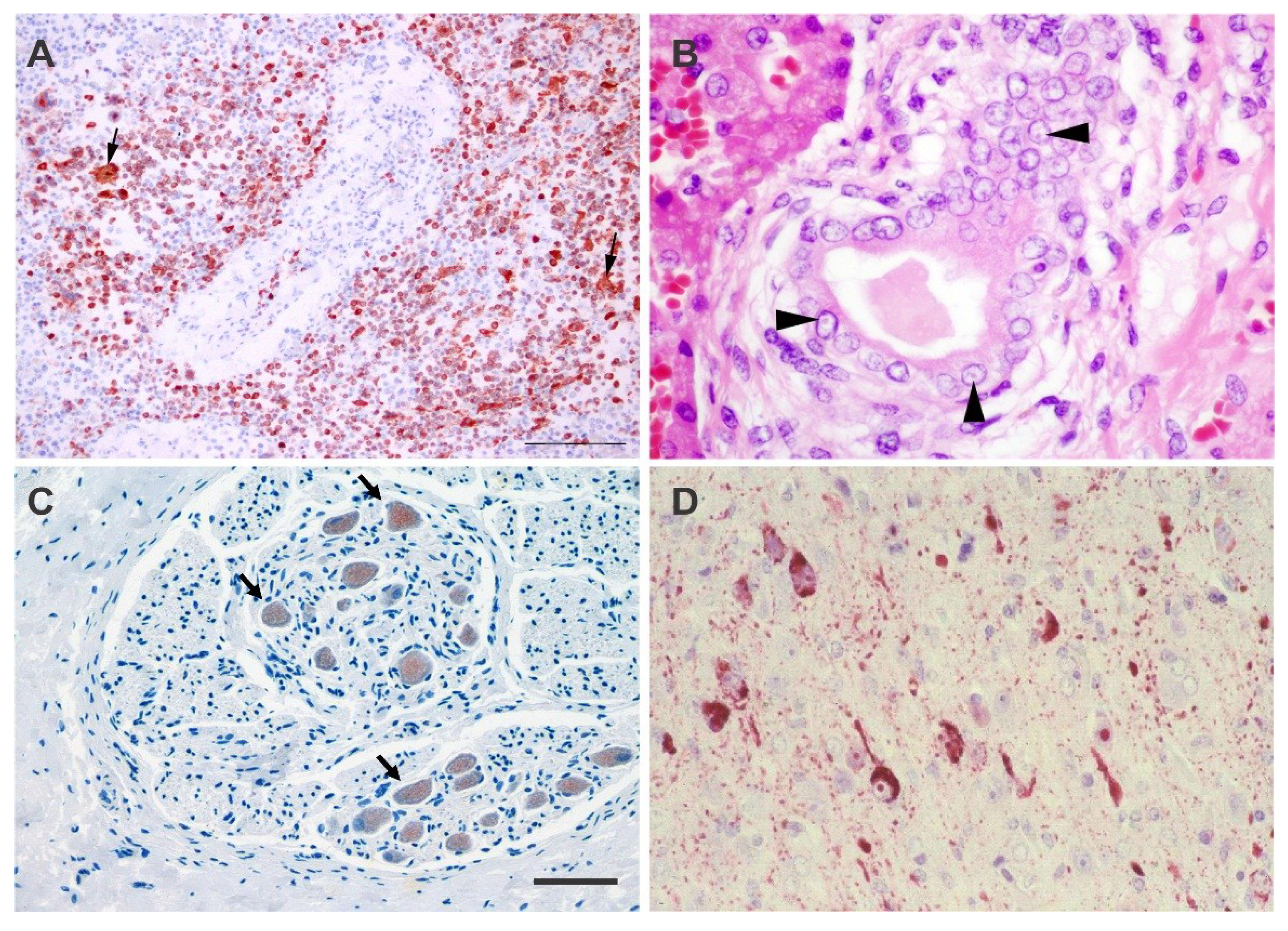
5.1. Acute, Systemic Disease
5.2. Sub-Acute Systemic Disease
5.3. Chronic Systemic Infection
5.4. Chronic, Localized CeMV Encephalitis
5.5. Subclinical Infection
5.6. Immune Function and CeMV Infections
5.7. CeMV Transmission
5.7.1. Horizontal Transmission
5.7.2. Evidence for Vertical Transmission
6. Outbreaks of Disease and Epidemiology
| Ocean Provinces/Species | Years | Countries | Epidemiolo-gical Status | Diagnosis | Virus | Literature Cited | |
|---|---|---|---|---|---|---|---|
| Eastern Atlantic & North Sea | |||||||
| Phocoena phocoena | 1988–1990 | N. Ireland, UK, Netherlands | periodic mortalities | VI, IHC, S, RT-PCR | PMV | [1,22,53], [54,61,63] | |
| Delphinus delphis | 1988–1990 | UK, Netherlands | unknown | S | CeMV | [22,61,63] | |
| Lagenorhynchus albirostris | 1988–1990, 2007, 2011 | Germany, Netherlands | periodic mortalities | S, IHC, RT-PCR | DMV | [22,34,61,125] | |
| Balaenoptera physalus | 1983 | Iceland | unknown | S | CeMV | [17] | |
| B. physalus | 1997–1998 | Belgium, France | periodic mortalities | IHC | unknown | [126] | |
| Tursiops truncatus | 1999 | Kent, UK | unknown | S | CeMV | [63] | |
| Globicephala macrorhynchus | 1996 | Canary Islands | periodic mortalities | RT-PCR | PWM | [30] | |
| T. truncatus | 2005 | Canary Islands | periodic mortalities | IHC, RT-PCR | DMV | [69] | |
| S. coeruleoalba | 2002–2011 | Canary Islands | periodic mortalities | IHQ, RT-PCR | DMV | [33] | |
| D. delphis | 2007 | Canary Islands | periodic mortalities | IHQ, RT-PCR | DMV | [33] | |
| Mediterranean Sea | |||||||
| S. coeruleoalba | 1990–1992 | Spain, France, Italy, Greece | epidemic | VI, IHC, S, RT-PCR | DMV | [2,3,21,58,127] | |
| S. coeruleoalba | 2006–2008 | Spain, France, Italy | epidemic | IHC, RT-PCR | DMV | [32,66,67] | |
| T. truncatus | 1994; 2007–2008, 2011 | Israel, Spain, France, Italy | periodic mortalities | IHC, RT-PCR, S | DMV | [32,63,66,128] | |
| D. delphis | 1990 | Italy | unknown | S | CeMV | [21] | |
| Globicephala melas | 2006–2007 | Spain, France | epidemic | IHQ, RT-PCR | DMV | [88] | |
| Grampus griseus | 1997, 1999 | Valencia, Spain | unknown | S | CeMV | [63] | |
| Balaenoptera acutorostrata | 1993 | Tuscany, Italy | unknown | S | unknown | [58] | |
| B. physalus | 2011 | Tuscany, Italy | periodic mortalities | RT-PCR | DMV | [89] | |
| Northwestern Atlantic | |||||||
| T. truncatus | 1982 | Florida, USA | epidemic | S, IHC | CeMV | [25,129] | |
| T. truncatus | 1987–1988 | East coast USA | epidemic | IHC, RT-PCR | CeMV | [57,92] | |
| T. truncatus | 1993–1994 | Gulf of Mexico, USA | epidemic | IHC, RT-PCR | CeMV | [25,57,92] | |
| T. truncatus | 2003–2007 | Florida, USA | unknown | S, IHC | CeMV | [64,65] | |
| T. truncatus | 2013–2014 | East coast USA | epidemic | IHC, RT-PCR | DMV | [71,130] | |
| T. truncatus | 1992–1994 | East coast USA | endemic | S | CeMV | [25] | |
| G. melas | 1982–1993 | Northeast coast USA | endemic | S | CeMV | [23] | |
| G. macrorhynchus | 1986–1994 | Florida, USA | endemic | S | CeMV | [23] | |
| G. melas | late nineties | New Jersey, USA | periodic mortalities | IHC, RT-PCR | PWM | [4] | |
| S. coeruleoalba | 1991–1993 | Northeast coast USA | unknown | S | CeMV | [24] | |
| Stenella frontalis | 1993 | Northeast coast USA | unknown | S | CeMV | [24] | |
| D. delphis | 1980–1994 | Northeast coast USA | possibly endemic | S | CeMV | [24] | |
| Lagenorhynchus acutus | 1985–1993 | Northeast coast USA | unknown | S | CeMV | [24] | |
| Kogia breviceps | 1983–1991 | Southeast coast USA | unknown | S | CeMV | [24] | |
| Feresa attenuata | 1983 | Southeast coast USA | unknown | S | CeMV | [24] | |
| Pseudorca crassidens | 1982–1988 | Southeast coast USA | possibly endemic | S | CeMV | [24] | |
| Lagenodelphis hosei | 1994 | Gulf of Mexico, USA | possibly endemic | S | CeMV | [24] | |
| P. phocoena | 1993–1994 | East coast, Canada | unknown | S | CeMV | [24] | |
| Southwestern Atlantic | |||||||
| L. hosei | 1999 | Puerto Madryn, Argentina | unknown | S | CeMV | [63] | |
| L. hosei | 1999 | Rio de Janeiro, Brazil | unknown | S | CeMV | [63] | |
| Sotalia guianensis | 2010 | Espirito Santo, Brazil | unknown | IHC, RT-PCR | CeMV NL | [6] | |
| Eastern Pacific | |||||||
| Lagenorhynchus obscurus | 1993–1995 | Central Peru | endemic | S | CeMV | [62]’ | |
| T. truncatus | 1993–1995 | Central Peru | endemic | S | CeMV | [62] | |
| Delphinus capensis | 1993–1995 | Central Peru | endemic | S | CeMV | [62] | |
| D. delphis | 1995–1997 | California, USA | unknown | S, IHC, RT-PCR | CeMV | [4,75] | |
| Indopacetus pacificus | 2010 | Hawaii, USA | unknown | HC, RT-PCR | BWMV | [5] | |
| Physeter macrocephalus | 2011 | Hawaii, USA | unknown | RT-PCR | BWMV | [121] | |
| Western Pacific | |||||||
| Lagenorhynchus obliquidens | 1998 | Miyazaki, Japan | unknown | IHC | unknown | [106] | |
| K. breviceps | 2009 | SW Taiwan | periodic mortalities | IHC, RT-PCR | DMV | [70] | |
| G. melas | 1997 | Northland, New Zealand | endemic | S | CeMV | [63] | |
| T. truncatus | 1997 | Tasmania, Australia | unknown | S | CeMV | [63] | |
| Peponocephala electra | 2005–2007 | NE Australia | endemic | S | CeMV | [90] | |
| Tursiops aduncus | 2005–2010 | NE Australia | unknown | S | CeMV | [90] | |
| L. hosei | 2006 | NE Australia | unknown | S | CeMV | [90] | |
| T. truncatus | 2009–2010 | Queensland, Australia | periodic mortalities | S, IHC, RT-PCR | DMV | [68,90] | |
| Indian Ocean | |||||||
| D. delphis | 1999 | East London, South Africa | unknown | S | CeMV | [63] | |
| T. aduncus | 2009 | Western Australia | periodic mortalitie s | IHC, RT-PCR | CeMV NL | [7] | |
| Southern Ocean | |||||||
| T. aduncus | 2012–2013 | South Australia | unknown | IHC, RT-PCR | CeMV NL | [51,131] | |
| T. truncatus | 2013 | South Australia | unknown | IHC, RT-PCR | CeMV NL | [51,131] | |
| D. delphis | 2012–2013 | South Australia | unknown | RT-PCR | CeMV NL | [51,131] | |
6.1. Europe
6.1.1. North Sea, NE Atlantic, and CE Atlantic
6.1.2. Mediterranean Sea
6.1.3. Black Sea
6.2. North America
6.2.1. Atlantic Coast
6.2.2. Gulf of Mexico
6.2.3. North Pacific
6.3. South America
6.4. Asia and Australasia
6.4.1. Asia
6.4.2. Australasia
7. Conclusions
Acknowledgements
Author Contributions
Conflicts of Interest
References and Notes
- McCullough, S.J.; McNeilly, F.; Allan, G.M.; Kennedy, S.; Smyth, J.A.; Cosby, S.L.; McQuaid, S.; Rima, B.K. Isolation and characterisation of a porpoise morbillivirus. Arch. Virol. 1991, 118, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Domingo, M.; Ferrer, L.; Pumarola, M.; Marco, A.; Plana, J.; Kennedy, S.; McAliskey, M.; Rima, B.K. Morbillivirus in dolphins. Nature 1990, 348, 21. [Google Scholar] [CrossRef] [PubMed]
- Van Bressem, M.-F.; Visser, I.K.G.; van de Bildt, M.W.; Teppema, J.S.; Raga, J.A.; Osterhaus, A.D.M.E. Morbillivirus infection in Mediterranean striped dolphins (Stenella coeruleoalba). Vet Rec. 1991, 129, 471–472. [Google Scholar]
- Taubenberger, J.K.; Tsai, M.M.; Atkin, T.J.; Fanning, T.G.; Krafft, A.E.; Moeller, R.B.; Kodsi, S.E.; Mense, M.G.; Lipscomb, T.P. Molecular genetic evidence of a novel morbillivirus in a long-finned pilot whale (Globicephala melas). Emerg. Infect. Dis. 2000, 6, 42–45. [Google Scholar] [CrossRef] [PubMed]
- West, K.L.; Sanchez, S.; Rotstein, D.; Robertson, K.M.; Dennison, S.; Levine, G.; Davis, N.; Schofield, D.; Potter, C.W.; Jensen, B. A Longman’s beaked whale (Indopacetus pacificus) strands in Maui, Hawaii, with first case of morbillivirus in the central Pacific. Mar. Mamm. Sci. 2013, 29, 767–776. [Google Scholar]
- Groch, K.R.; Colosio, A.C.; Marcondes, M.C.; Zucca, D.; Díaz-Delgado, J.; Niemeyer, C.; Marigo, J.; Brandão, P.E.; Fernández, A.; Luiz Catão-Dias, J. Novel cetacean morbillivirus in Guiana dolphin, Brazil. Emerg. Infect. Dis. 2014, 20, 511–513. [Google Scholar] [CrossRef] [PubMed]
- Stephens., N.; Duignan, P.J.; Wang, J.; Bingham, J.; Finn, H.; Bejder, L.; Patterson, A.P.; Holyoake, C. Cetacean morbillivirus in coastal Indo-Pacific bottlenose dolphins, Western Australia. Emerg. Infect. Dis. 2014, 20, 666–670. [Google Scholar] [CrossRef] [PubMed]
- Barrett, T. Morbillivirus infections, with special emphasis on morbilliviruses of carnivores. Vet. Microbiol. 1999, 69, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Ohishi, K.; Suzuki, R.; Maruyama, T. Host-Virus Specificity of the Morbillivirus Receptor, SLAM, in Marine Mammals: Risk Assessment of infection based on three-dimensional models. In New Approaches to the Study of Marine Mammals; Romero, A., Keith, E.O., Eds.; InTech: Rijeka, Croatia, 2012; pp. 183–204. [Google Scholar]
- Hall, A.J. Morbilliviruses in marine mammals. Trends Microbiol. 1995, 3, 4–9. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lau, S.K.; Wong, B.H.; Fan, R.Y.; Wong, A.Y.; Zhang, A.J.; Wu, Y.; Choi, G.K.; Li, K.S.; Hui, J.; et al. Feline morbillivirus, a previously undescribed paramyxovirus associated with tubulointerstitial nephritis in domestic cats. Proc. Natl. Acad. Sci. USA 2012, 109, 5435–5440. [Google Scholar] [CrossRef] [PubMed]
- Delpeut, S.; Noyce, R.S.; Richardson, C.D. The tumor-associated marker, PVRL4 (nectin-4), is the epithelial receptor for morbilliviruses. Viruses 2014, 6, 2268–2286. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, Y.; Ohishi, K.; Suzuki, R.; Tajima, Y.; Yamada, T.; Kakizoe, Y.; Bando, T.; Fujise, Y.; Taru, H.; Murayama, T.; et al. Amino acid sequence variations of signaling lymphocyte activation molecule and mortality caused by morbillivirus infection in cetaceans. Microbiol. Immunol. 2013, 57, 624–632. [Google Scholar] [PubMed]
- Barrett, T.; Blixenkrone-Møller, M.; di Guardo, G.; Domingo, M.; Duignan, P.; Hall, A.; Mamaev, L.; Osterhaus, A.D.M.E. Morbilliviruses in aquatic mammals: Report on round table discussion. Vet. Microbiol. 1995, 44, 261–265. [Google Scholar] [CrossRef] [PubMed]
- Osterhaus, A.D.M.E.; de Swart, R.L.; Vos, H.W.; Ross, P.S.; Kenter, M.J.H.; Barrett, T. Morbillivirus infections of aquatic mammals: Newly identified members of the genus. Vet. Microb. 1995, 44, 219–227. [Google Scholar] [CrossRef]
- Barrett, T.; Visser, I.K.G.; Mamaev, L.; Goatley, L.; van Bressem, M.-F.; Osterhaus, A.D.M.E. Dolphin and porpoise morbilliviruses are genetically distinct from phocine distemper virus. Virology 1993, 193, 1010–1012. [Google Scholar] [CrossRef] [PubMed]
- Blixenkrone-Möller, M.; Bolt, G.; Gottschalk, E.; Kenter, M. Comparative analysis of the gene encoding the nucleocapsid protein of dolphin morbillivirus reveals its distant evolutionary relationship to measles virus and ruminant morbilliviruses. J. Gen. Virol. 1994, 75, 2829–2834. [Google Scholar] [CrossRef] [PubMed]
- Bolt, G.; Blixenkrone-Möller, M.; Gottschalk, E.; Wishaupt, R.G.A.; Welsh, M.J.; Earle, P.J.A.; Rima, B.K. Nucleotide and deduced amino acid sequences of the matrix (M) and fusion (F) protein genes of cetacean morbilliviruses isolated isolated from a porpoise and a dolphin. Virus Res. 1994, 34, 291–304. [Google Scholar] [CrossRef] [PubMed]
- Rima, B.K.; Collin, A.M.; Earle, J.A. Completion of the sequence of a cetacean morbillivirus and comparative analysis of the complete genome sequences of four morbilliviruses. Virus Genes. 2005, 30, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Banyard, A.C.; Grant, R.J.; Romero, C.H.; Barrett, T. Sequence of the nucleocapsid gene and genome and antigenome promoters for an isolate of porpoise morbillivirus. Virus Res. 2008, 132, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Van Bressem, M.-F.; Visser, I.K.G.; de Swart, R.L.; Örvell, C.; Stanzani, L.; Androukaki, E.; Siakavara, K.; Osterhaus, A.D.M.E. Dolphin morbillivirus in different parts of the Mediterranean Sea. Arch. Virol. 1993, 129, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Visser, I.K.G.; van Bressem, M.-F.; de Swart, R.L.; van de Bildt, M.W.G.; Vos, H.W.; van der Heijden, R.W.J.; Saliki, J.T.; Örvell, C.; Kitching, P.; Kuiken, T.; et al. Characterization of morbilliviruses isolated from dolphins and porpoises in Europe. J. Gen. Virol. 1993, 74, 631–641. [Google Scholar] [CrossRef] [PubMed]
- Duignan, P.J.; House, C.; Geraci, J.R.; Early, G.; Copland, H.G.; Walsh, M.T.; Bossart, G.D.; Cray, C.; Sadove, S.; St. Aubin, D.J.; et al. Morbillivirus infection in two species of pilot whales (Globicephala sp.) from the western Atlantic. Mar. Mamm. Sci. 1995, 11, 150–162. [Google Scholar] [CrossRef]
- Duignan, P.J.; House, C.; Geraci, J.R.; Duffy, N.; Rima, B.K.; Walsh, M.T.; Early, G.; St Aubin, D.J.; Sadove, S.; Koopman, H.; et al. Morbillivirus infection in cetaceans of the western Atlantic. Vet. Microbiol. 1995, 44, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Duignan, P.J.; House, C.; Ode11, D.K.; Wells, R.S.; Hansen, W.; Walsh, M.T.; St. Aubin, D.J.; Rima, B.K.; Geraci, J.R. Morbillivirus in bottlenose dolphins: Evidence for recurrent epizootics in the western Atlantic and Gulf of Mexico. Mar. Mamm. Sci. 1996, 12, 499–515. [Google Scholar] [CrossRef]
- Rowles, T.R.; Schwacke, L.S.; Wells, R.S.; Saliki, J.T.; Hansen, L.; Hohn, A.; Townsend, F.; Sayre, R.A.; Hall, A.J. Evidence of susceptibility to morbillivirus infection in cetaceans from the United States. Mar. Mamm. Sci. 2011, 27, 1–19. [Google Scholar] [CrossRef]
- Banyard, A.C.; Tiwari, A.; Barrett, T. Morbillivirus infection in pilot whales: Strict protein requirement drives genetic conservation. Arch. Virol. 2011, 156, 1853–1839. [Google Scholar] [CrossRef] [PubMed]
- Van de Bildt, M.W.; Kuiken, T.; Osterhaus, A.D.M.E. Cetacean morbilliviruses are phylogenetically divergent. Arch. Virol. 2005, 150, 577–583. [Google Scholar] [CrossRef] [PubMed]
- King, A.M.Q.; Adams, M.J.; Carstens, E.B.; Lefkowitz, E.J. Virus Taxonomy: Classification and Nomenclature of Viruses: Ninth Report of the International Committee on Taxonomy of Viruses; Elsevier/Academic Press: London, UK, 2011. [Google Scholar]
- Bellière, E.N.; Esperón, F.; Fernández, A.; Arbelo, M.; Muñoz, M.J.; Sánchez-Vizcaíno, J.M. Phylogenetic analysis of a new Cetacean morbillivirus from a short-finned pilot whale stranded in the Canary Islands. Res. Vet. Sci. 2011, 90, 324–328. [Google Scholar]
- Bellière, E.N.; Esperón, F.; Sánchez-Vizcaíno, J.M. Genetic comparison among dolphin morbillivirus in the 1990–1992 and 2006–2008 Mediterranean outbreaks. Infect. Genet. Evol. 2011, 11, 1913–1920. [Google Scholar] [CrossRef] [PubMed]
- Keck, N.; Kwiatek, O.; Dhermain, F.; Dupraz, F.; Boulet, H.; Danes, C.; Laprie, C.; Perrin, A.; Godenir, J.; Micout, L.; et al. Resurgence of Morbillivirus infection in Mediterranean dolphins off the French coast. Vet. Rec. 2010, 166, 654–655. [Google Scholar] [CrossRef] [PubMed]
- Sierra, E.; Sánchez, S.; Saliki, J.T.; Blas-Machado, U.; Arbelo, M.; Zucca, D.; Fernández, A. Retrospective study of etiologic agents associated with nonsuppurative meningoencephalitis in stranded cetaceans in the Canary Islands. J. Clin. Microbiol. 2014, 52, 2390–2397. [Google Scholar] [CrossRef] [PubMed]
- Van Elk, C.E.; van de Bildt, M.W.; Jauniaux, T.; Hiemstra, S.; van Run, P.R.; Foster, G.; Meerbeek, J.; Osterhaus, A.D.M.E.; Kuiken, T. Is Dolphin Morbillivirus Virulent for White-Beaked Dolphins (Lagenorhynchus albirostris)? Vet. Pathol. 2014. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Milinkovitch, M.C.; Thewissen, J.G.M. Even-toed fingerprints on whale ancestry. Nature 1997, 388, 622–624. [Google Scholar]
- Nikaido, M.; Rooney, A.P.; Okada, N. Phylogenetic relationships among cetartiodactyls based on insertions of short and long interpersed elements: Hippopotamuses are the closest extant relatives of whales. Proc. Natl. Acad. Sci. USA 1999, 96, 10261–10266. [Google Scholar] [CrossRef] [PubMed]
- Barrett, T.; Rossiter, P.B. Rinderpest: The disease and its impact on humans and animals. Adv. Virus Res. 1999, 53, 89–110. [Google Scholar] [PubMed]
- Kumar, N.; Maherchandani, S.; Kashyap, S.K.; Singh, S.V.; Sharma, S.; Chaubey, K.K.; Ly, H. Peste des petits ruminants virus infection of small ruminants: A comprehensive review. Vet. Med. Int. 2014, 6, 2287–2327. [Google Scholar]
- Nollens, H.H.; Ruiz, C.; Walsh, M.T.; Gulland, F.M.D.; Bossart, G.; Jensen, E.E.; McBain, J.F.; Wellehan, J.F.X. Cross- reactivity of immunoglobulin G of whales and dolphins correlates with evolutionary distance of cytochrome B genes. Clin. Vacc. Immunol. 2008, 15, 1547–1554. [Google Scholar] [CrossRef]
- Wild, T.F.; Malvoisin, E.; Buckland, R. Measles virus: Both the haemagglutinin and fusion glycoproteins are required for fusion. J. Gen. Virol. 1991, 72, 439–442. [Google Scholar] [CrossRef] [PubMed]
- Melia, M.M.; Earle, J.P.; Abdullah, H.; Reaney, K.; Tangy, F.; Cosby, S.L. Use of SLAM and PVRL4 and identification of pro-HB-EGF as cell entry receptors for wild type phocine distemper virus. PLoS One 2014, 9, e106281. [Google Scholar] [CrossRef] [PubMed]
- Tatsuo, H.; Ono, N.; Yanagi, Y. Morbilliviruses use signaling lymphocyte activation molecules (CD150) as cellular receptors. J. Virol. 2001, 75, 5842–5850. [Google Scholar] [CrossRef] [PubMed]
- Muhlebach, M.D.; Mateo, M.; Sinn, P.L.; Prufer, S.; Uhlig, K.M.; Leonard, V.H.; Navaratnarajah, C.K.; Frenzke, M.; Wong, X.X.; Sawatsky, B.; et al. Adherens junction protein nectin-4 is the epithelial receptor for measles virus. Nature 2011, 480, 530–533. [Google Scholar] [PubMed]
- Noyce, R.S.; Bondre, D.G.; Ha, M.N.; Lin, L.T.; Sisson, G.; Tsao, M.S.; Richardson, C.D. Tumor cell marker PVRL4 (nectin 4) is an epithelial cell receptor for measles virus. PLoS Pathog. 2011, 7, e1002240. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, A.; Yoneda, M.; Ikeda, F.; Terao-Muto, Y.; Sato, H.; Kai, C. CD147/EMMPRIN acts as a functional entry receptor for measles virus on epithelial cells. J. Virol. 2010, 84, 4183–4193. [Google Scholar] [CrossRef] [PubMed]
- Baron, M.D. Wildtype rinderpest virus uses SLAM (CD150) as its receptor. J. Gen. Virol. 2005, 86, 1753–1787. [Google Scholar] [CrossRef] [PubMed]
- Adombi, C.M.; Lelenta, M.; Lamien, C.E.; Shamaki, D.; Koffi, Y.M.; Traoré, A.; Silber, R.; Couacy-Hymann, E.; Bodjo, S.C.; Djaman, J.A.; et al. Monkey CV1 cell line expressing the sheep-goat SLAM protein: A highly sensitive cell line for the isolation of peste des petits ruminants virus from pathological specimens. J. Virol. Methods 2011, 173, 306–313. [Google Scholar] [CrossRef]
- Ono, N.; Tatsuo, H.; Tanaka, K.; Minagawa, H.; Yanagi, Y. V domain of human SLAM (CDw150) is essential for its function as a measles virus receptor. J. Virol. 2001, 75, 1594–1600. [Google Scholar] [CrossRef] [PubMed]
- Bieringer, M.; Han, J.W.; Kendl, S.; Khosravi, M.; Plattet, P.; Schneider-Schaulies, J. Experimental adaptation of wild-type canine distemper virus (CDV) to the human entry receptor CD150. PLoS One 2013, 8, e57488. [Google Scholar] [CrossRef] [PubMed]
- Kemper, M.C.; Woolford, L.; Tomo, I.; Dickason, C.; Bastianello, S.; Gibbs, S.; Kelly, D.; Wang, J.; Bingham, J. Abnormally high dolphin mortalities linked to Morbillivirus in South Australia. In Proceedings of the 20 Biennial Conference on the Biology of Marine Mammals, Dunedin, New Zealand, 9–13 December 2013.
- Van Bressem, M.-F.; Raga, J.-A.; di Guardo, G.; Jepson, P.D.; Duignan, P.J.; Siebert, U.; Barrett, T.; Santos, M.C.; Moreno, I.B.; Siciliano, S.; et al. Emerging infectious diseases in cetaceans worldwide and the possible role of environmental stressors. Dis. Aquat. Org. 2009, 86, 143–157. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, S.; Smyth, J.A.; Cush, P.F.; McCullough, S.J.; Allan, G.M.; McQuaid, S. Viral distemper now found in porpoises. Nature 1988, 336, 21. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, S.; Smyth, J.A.; Cush, P.F.; McAliskey, M.; McCullough, S.J.; Rima, B.K. Histopathologic and immunocytochemical studies of distemper in harbor porpoises. Vet. Pathol. 1991, 28, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Grant, R.J.; Banyard, A.C.; Barrett, T.; Saliki, J.T.; Romero, C.H. Real-time RT-PCR assays for the rapid and differential detection of dolphin and porpoise morbilliviruses. J. Virol. Methods. 2009, 156, 117–123. [Google Scholar] [CrossRef] [PubMed]
- Soto, S.; Alba, A.; Ganges, L.; Vidal, E.; Raga, J.A.; Alegre, F.; González, B.; Medina, P.; Zorrilla, I.; Martínez, J. Post-epizootic chronic dolphin morbillivirus infection in Mediterranean striped dolphins Stenella coeruleoalba. Dis. Aquat. Organ. 2011, 96, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Lipscomb, T.P.; Schulman, F.Y.; Moffett, D.; Kennedy, S. Morbilliviral disease in Atlantic bottlenose dolphins (Tursiops truncatus) from the 1987–1988 epizootic. J. Wildl. Dis. 1994, 30, 567–557. [Google Scholar] [CrossRef] [PubMed]
- Di Guardo, G.; Agrimi, U.; Morelli, L.; Cardeti, G.; Terracciano, G.; Kennedy, S. Post mortem investigations on cetaceans found stranded on the coasts of Italy between 1990 and 1993. Vet. Rec. 1995, 136, 439–442. [Google Scholar] [CrossRef] [PubMed]
- Domingo, M.; Visa, J.; Pumarola, M.; Marco, A.; Ferrer, L.; Rabanal, R.; Kennedy, S. Pathologic and immunocytochemical studies of morbillivirus infection in striped dolphins (Stenella coeruleoalba). Vet. Pathol. 1992, 29, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Duignan, P.J.; Geraci, J.R.; Raga, J.A.; Calzada, N. Pathology of morbillivirus infection in striped dolphins (Stenella coeruleoalba) from Valencia and Murcia, Spain. Can. J. Vet. Res. 1992, 56, 242–248. [Google Scholar] [PubMed]
- Van Bressem, M.-F.; Jepson, P.; Barrett, T. Further insight on the epidemiology of cetacean morbillivirus in the northeastern Atlantic. Mar. Mamm. Sci. 1998, 14, 605–613. [Google Scholar] [CrossRef]
- Van Bressem, M-F.; van Waerebeek, K.; Fleming, M.; Barrett, T. Serological evidence of morbillivirus infection in small cetaceans from the Southeast Pacific. Vet. Microb. 1998, 59, 89–98. [Google Scholar]
- Van Bressem, M-F.; van Waerebeek, K.; Jepson, P.D.; Raga, J.A.; Duignan, P.J.; Nielsen, O.; di Beneditto, A.P.; Siciliano, S.; Ramos, R.; Kant, W.; et al. An insight into the epidemiology of dolphin morbillivirus worldwide. Vet. Microb. 2001, 81, 287–304. [Google Scholar]
- Bossart, G.D.; Reif, J.S.; Schaefer, A.M.; Goldstein, J.; Fair, P.A.; Saliki, J.T. Morbillivirus infection in free-ranging Atlantic bottlenose dolphins (Tursiops truncatus) from the Southeastern United States: Seroepidemiologic and pathologic evidence of subclinical infection. Vet. Microbiol. 2010, 143, 160–166. [Google Scholar] [CrossRef] [PubMed]
- Bossart, G.D.; Romano, T.A.; Peden-Adams, M.M.; Schaefer, A.; McCulloch, S.; Goldstein, J.D.; Rice, C.D.; Saliki, J.T.; Fair, P.A.; Reif, J.S. Clinicoimmunopathologic findings in atlantic bottlenose dolphins Tursiops truncatus with positive cetacean morbillivirus antibody titers. Dis. Aquat. Organ. 2011, 97, 103–112. [Google Scholar] [CrossRef]
- Di Guardo, G.; di Francesco, C.E.; Eleni, C.; Cocumelli, C.; Scholl, F.; Casalone, C.; Peletto, S.; Mignone, W.; Tittarelli, C.; di Nocera, F.; et al. Morbillivirus infection in cetaceans stranded along the Italian coastline: Pathological, immunohistochemical and biomolecular findings. Res. Vet. Sci. 2013, 94, 132–137. [Google Scholar] [CrossRef] [PubMed]
- Raga, J.A.; Banyard, A.; Domingo, M.; Corteyn, M.; Van Bressem, M-F.; Fernández, M.; Aznar, F.J.; Barrett, T. Dolphin morbillivirus epizootic resurgence, Mediterranean Sea. Emerg. Infect. Dis. 2008, 14, 471–473. [Google Scholar] [CrossRef]
- Stone, B.M.; Blyde, D.J.; Saliki, J.T.; Blas-Machado, U.; Bingham, J.; Hyatt, A.; Wang, J.; Payne, J.; Crameri, S. Fatal cetacean morbillivirus infection in an Australian offshore bottlenose dolphin (Tursiops truncatus). Aust. Vet. J. 2011, 89, 452–457. [Google Scholar]
- Sierra, E.; Zucca, D.; Arbelo, M.; García-Álvarez, N.; Andrada, M.; Déniz, S.; Fernández, A. Fatal systemic morbillivirus infection in bottlenose dolphin, Canary Islands, Spain. Emerg. Infect. Dis. 2014, 20, 269–271. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.C.; Pang, V.F.; Jeng, C.R.; Chou, L.S.; Chueh, L.L. Morbilliviral infection in a pygmy sperm whale (Kogia breviceps) from Taiwanese waters. Vet. Microbiol. 2006, 116, 69–76. [Google Scholar]
- NOAA. 2013–2014 Bottlenose Dolphin Unusual Mortality Event in the Mid-Atlantic. 2014. Available online: http://www.nmfs.noaa.gov/pr/health/mmume/midatldolphins2013.html (accessed on 28 August 2014).
- Nielsen, O.; Smith, G.; Weingartl, H.; Lair, S.; Measures, L. Use of a SLAM transfected Vero cell line to isolate and characterize marine mammal morbilliviruses using an experimental ferret model. J. Wildl. Dis. 2008, 44, 600–611. [Google Scholar]
- Orvell, C.; Blixenkrone-Möller, M.; Svansson, V.; Have, P. Immunological relationships between phocid and canine distemper virus studied with monoclonal antibodies. J. Gen. Virol. 1990, 71, 2085–2092. [Google Scholar] [CrossRef] [PubMed]
- Saliki, J.T.; Lehenbauer, T.W. Monoclonal antibody-based competitive enzyme-linked immunosorbent assay for detection of morbillivirus antibody in marine mammal sera. J. Clin. Microbiol. 2001, 39, 1877–1881. [Google Scholar] [CrossRef] [PubMed]
- Reidarson, T.H.; McBain, J.; House, C.; King, D.P.; Stott, J.L.; Krafft, A.; Taubenberger, J.K.; Heyning, J.; Lipscomb, T.P. Morbillivirus infection in stranded common dolphins from the Pacific Ocean. J. Wildl. Dis. 1998, 34, 771–776. [Google Scholar]
- Lindmark, R.; Thorén-Tolling, K.; Sjöquist, J. Binding of immunoglobulins to protein A and immunoglobulin levels in mammalian sera. J. Immunol. Methods. 1983, 62, 1–13. [Google Scholar] [CrossRef]
- Grant, R.J.; Kelley, K.L.; Maruniak, J.E.; Garcia-Maruniak, A.; Barrett, T.; Manire, C.A.; Romero, C.H. Expression from baculovirus and serological reactivity of the nucleocapsid protein of dolphin morbillivirus. Vet. Microbiol. 2010, 143, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, O.; Stewart, R.E.; Measures, L.; Duignan, P.; House, C. A morbillivirus antibody survey of Atlantic walrus, narwhal and beluga in Canada. J. Wildl. Dis. 2000, 36, 508–517. [Google Scholar] [CrossRef]
- Duignan, P.J.; Nielsen, O.; House, C.; Kovacs, K.M.; Duffy, N.; Early, G.; Sadove, S.; St Aubin, D.J.; Rima, B.K.; Geraci, J.R. Epizootiology of morbillivirus infection in harp, hooded, and ringed seals from the Canadian Arctic and western Atlantic. J. Wildl. Dis. 1997, 33, 7–19. [Google Scholar] [CrossRef] [PubMed]
- Krafft, A.; Lichy, J.H.; Lipscomb, T.P.; Klaunberg, B.A.; Kennedy, S.; Taubenberger, J.K. Postmortem diagnosis of morbillivirus infection in bottlenose dolphins (Tursiops truncatus) in the Atlantic and Gulf of Mexico epizootics by polymerase chain reaction-based assay. J. Wildl. Dis. 1995, 31, 410–415. [Google Scholar] [CrossRef] [PubMed]
- Rubio-Guerri, C.; Melero, M.; Rivera-Arroyo, B.; Bellière, E.N.; Crespo, J.L.; García-Párraga, D.; Esperón, F.; Sánchez-Vizcaíno, J.M. Simultaneous diagnosis of Cetacean morbillivirus infection in dolphins stranded in the Spanish Mediterranean sea in 2011 using a novel Universal Probe Library (UPL) RT-PCR assay. Vet. Microbiol. 2013, 165, 109–114. [Google Scholar]
- Rubio-Guerri, C.; Melero, M.; Esperón, F.; Bellière, E.N.; Arbelo, M.; Crespo, J.L.; Sierra, E.; García-Párraga, D.; Sánchez-Vizcaíno, J.M. Unusual striped dolphin mass mortality episode related to cetacean morbillivirus in the Spanish Mediterranean sea. BMC Vet. Res. 2013, 9, e106. [Google Scholar] [CrossRef]
- Ludlow, M.; McQuaid, S.; Milner, D.; de Swart, R.L.; Duprex, W.P. Pathological consequences of systemic measles virus infection. J. Pathol. 2015, 235, 253–265. [Google Scholar] [CrossRef] [PubMed]
- De Swart, R.L.; Ludlow, M.; de Witte, L.; Yanagi, Y.; van Amerongen, G.; McQuaid, S.; Yuksel, S.; Geijtenbeek, T.B.; Duprex, W.P.; Osterhaus, A.D. Predominant infection of CD150+ lymphocytes and dendritic cells during measles virus infection of macaques. PLoS Pathog. 2007, 3, e178. [Google Scholar] [CrossRef] [PubMed]
- Appel, M.P.G. Pathogenesis of canine distemper. Am. J. Vet. Res. 1969, 30, 1167–1182. [Google Scholar] [PubMed]
- Lemon, K.; de Vries, R.D.; Mesman, A.W.; McQuaid, S.; van Amerongen, G.; Yuksel, S.; Ludlow, M.; Rennick, L.J.; Kuiken, T.; Rima, B.K.; et al. Early target cells of measles virus after aerosol infection of non-human primates. PLoS Pathog. 2011, 7, e1001263. [Google Scholar] [CrossRef] [PubMed]
- Schulman, F.Y.; Lipscomb, T.P.; Moffett, D.; Krafft, A.E.; Lichy, J.H.; Tsai, M.M.; Taubenberger, J.K.; Kennedy, S. Histologic, immunohistochemical, and polymerase chain reaction studies of bottlenose dolphins from the 1987–1988 United States Atlantic coast epizootic. Vet. Pathol. 1997, 34, 288–295. [Google Scholar] [CrossRef] [PubMed]
- Fernández, A.; Esperón, F.; Herraéz, P.; Espinosa de los Monteros, A.; Clavel, C.; Bernabé, A.; Sanchez-Vizcaino, M.; Verborgh, P.; DeStephanis, R.; Toledano, F.; et al. Morbillivirus and pilot whale deaths, Mediterranean Sea. Emerg. Infect. Dis. 2008, 14, 792–794. [Google Scholar] [CrossRef] [PubMed]
- Mazzariol, S.; Marcer, F.; Mignone, W.; Serracca, L.; Goria, M.; Marsili, L.; di Guardo, G.; Casalone, C. Dolphin Morbillivirus and Toxoplasma gondii coinfection in a Mediterranean fin whale (Balaenoptera physalus). BMC Vet. Res. 2012, 8, e20. [Google Scholar] [CrossRef]
- Stone, B.M.; Blyde, D.J.; Saliki, J.T.; Morton, J.M. Morbillivirus infection in live stranded, injured, trapped, and captive cetaceans in southeastern Queensland and northern New South Wales, Australia. J. Wildl. Dis. 2012, 48, 47–55. [Google Scholar] [CrossRef]
- Soto, S.; Gonzalez, B.; Willoughby, K.; Maley, M.; Olivera, A.; Kennedy, S.; Marco, A.; Domingo, M. Systemic herpesvirus and morbillivirus co-infection in a striped dolphin (Stenella coeruleoalba). J. Comp. Pathol. 2012, 146, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Taubenberger, J.K.; Tsai, M.; Krafft, A.E.; Lichy, J.H.; Reid, A.H.; Schulman, F.Y.; Lipscomb, T.P. Two morbilliviruses implicated in bottlenose dolphin epizootics. Emerg. Infect. Dis. 1996, 2, 213–216. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.H.; Kouyos, R.D.; Adams, R.J.; Grenfell, B.T.; Griffin, D.E. Prolonged persistence of measles virus RNA is characteristic of primary infection dynamics. Proc. Natl. Acad. Sci. USA 2012, 109, 14989–14994. [Google Scholar] [CrossRef] [PubMed]
- Domingo, M.; Vilafranca, M.; Visa, J.; Prats, N.; Trudgett, A.; Visser, I. Evidence for chronic morbillivirus infection in the Mediterranean striped dolphin (Stenella coeruleoalba). Vet. Microbiol. 1995, 44, 229–239. [Google Scholar] [CrossRef] [PubMed]
- Garg, R.K. Subacute sclerosing panencephalitis. J. Neurol. 2008, 255, 1861–1871. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, J.; Issacson, R.S.; Koppel, B.S. Subacute sclerosing panencephalitis: An update. Dev. Med. Child Neurol. 2010, 52, 901–917. [Google Scholar] [CrossRef]
- Headley, S.A.; Amude, A.M.; Alfieri, A.F.; Bracarense, A.P.; Alfieri, A.A.; Summers, B.A. Molecular detection of Canine distemper virus and the immunohistochemical characterization of the neurologic lesions in naturally occurring old dog encephalitis. J. Vet. Diagn. Invest. 2009, 21, 588–597. [Google Scholar] [CrossRef] [PubMed]
- Vandevelde, M.; Kristensen, B.; Braund, K.G.; Greene, C.E.; Swango, L.J.; Hoerlein, B.F. Chronic canine distemper virus encephalitis in mature dogs. Vet. Pathol. 1980, 17, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Griffin, D.E.; Lin, W.H.; Pan, C.H. Measles virus, immune control, and persistence. FEMS Microbiol. Rev. 2012, 36, 649–662. [Google Scholar] [CrossRef] [PubMed]
- Vandevelde, M.; Zurbriggen, A. The neurobiology of canine distemper virus infection. Vet. Microb. 1995, 44, 271–280. [Google Scholar] [CrossRef]
- Axthelm, M.K.; Krakowka, S. Experimental old dog encephalitis (ODE) in a gnotobiotic dog. Vet. Pathol. 1998, 35, 527–534. [Google Scholar] [CrossRef] [PubMed]
- Maxie, M.G.; Youssef, S. Nervous system. In Jubb, Kennedy, and Palmer’s Pathology of Domestic Animals; Maxie, M.G., Ed.; Saunders/Elsevier: Philadelphia, PA, USA, 2007; Volume 1, pp. 405–408. [Google Scholar]
- Di Guardo, G. Morbillivirus-host interaction: Lessons from aquatic mammals. Front. Microbiol. 2012, 3, e431. [Google Scholar] [CrossRef]
- Gomez de Segura, A.; Crespo, E.A.; Pedraza, S.N.; Hammond, P.S.; Raga, J.A. Abundance of small cetaceans in the waters of the central Spanish Mediterranean. Mar. Biol. 2006, 150, 149–160. [Google Scholar] [CrossRef]
- Schönberger, K.; Ludwig, M.S.; Wildner, M.; Weissbrich, B. Epidemiology of subacute sclerosing panencephalitis (SSPE) in Germany from 2003 to 2009: A risk estimation. PLoS One 2013, 8, e68909. [Google Scholar] [CrossRef] [PubMed]
- Uchida, K.; Muranaka, M.; Horii, Y.; Murakami, N.; Yamaguchi, R.; Tateyama, S. Non-purulent meningoencephalomyelitis of a Pacific striped dolphin (Lagenorhynchus obliquidens). The first evidence of morbillivirus infection in a dolphin at the Pacific Ocean around Japan. J. Vet. Med. Sci. 1999, 61, 159–162. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, S. Morbillivirus infections in aquatic mammals. J. Comp. Pathol. 1998, 119, 201–225. [Google Scholar] [CrossRef] [PubMed]
- Ludlow, M.; de Vries, R.D.; Lemon, K.; McQuaid, S.; Millar, E.; van Amerongen, G.; Yüksel, S.; Verburgh, R.J.; Osterhaus, A.D.ME.; de Swart, R.L.; et al. Infection of lymphoid tissues in the macaque upper respiratory tract contributes to the emergence of transmissible measles virus. J. Gen. Virol. 2013, 94, 1933–1944. [Google Scholar] [CrossRef] [PubMed]
- Appel, M.J.; Shek, W.R.; Summers, B.A. Lymphocyte-mediated immune cytotoxicity in dogs infected with virulent canine distemper virus. Infect. Immun. 1982, 37, 592–600. [Google Scholar] [PubMed]
- Griffin, D.E.; Ward, B.J.; Esolen, L.M. Pathogenesis of measles virus infection: An hypothesis for altered immune responses. J. Infect. Dis. 1994, 170 (Suppl 1), S24–S31. [Google Scholar] [CrossRef] [PubMed]
- Heaney, J.; Barrett, T.; Cosby, S.L. Inhibition of in vitro leukocyte proliferation by morbilliviruses. J. Virol. 2002, 76, 3579–3584. [Google Scholar] [CrossRef] [PubMed]
- Schlender, J.; Schnorr, J.J.; Spielhoffer, P.; Cathomen, T.; Cattaneo, R.; Billeter, M.A.; ter Meulen, V.; Schneider-Schaulies, S. Interaction of measles virus glycoproteins with the surface of uninfected peripheral blood lymphocytes induces immunosuppression in vitro. Proc. Natl. Acad. Sci. USA 1996, 93, 13194–13199. [Google Scholar] [CrossRef] [PubMed]
- Black, F. Epidemiology of Paramyxoviridae. In The Paramyxoviruses; Kingsburry, D.W., Ed.; Plenum Press: New York, NY, USA, 1991; pp. 509–536. [Google Scholar]
- Van Bressem, M.-F.; van Waerebeek, K.; Raga, J.A. A review of virus infections of cetaceans and the potential impact of morbilliviruses, poxviruses and papillomaviruses on host population dynamics. Dis. Aquat. Organ. 1999, 38, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Rijks, J.M.; Osterhaus, A.D.M.E.; Kuiken, T.; Frölich, K. Morbillivirus Infections. In Infectious Diseases of Wild Mammals and Birds in Europe; Gavier-Widén, D., Duff, J.P., Meredith, A., Eds.; Wiley-Blackwell: Oxford, UK, 2012. [Google Scholar]
- Van Bressem, M.F.; Raga, J.A. Viruses of Cetaceans. In Studies in Viral Ecology; Hurst, C., Ed.; John Wiley & Sons: Hoboken, NJ, USA, 2011; pp. 309–332. [Google Scholar]
- Birkun, A., Jr.; Kuiken, T.; Krivokhizhin, S.; Haines, D.M.; Osterhaus, A.D.M.E.; van de Bildt, M.W.; Joiris, C.R.; Siebert, U. Epizootic of morbilliviral disease in common dolphins (Delphinus delphis ponticus) from the Black sea. Vet. Rec. 1999, 144, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Chiba, M.E.; Saito, M.; Suzuki, N.; Honda, Y.; Yaegashi, N. Measles infection in pregnancy. J. Infect. 2003, 47, 40–44. [Google Scholar] [CrossRef] [PubMed]
- Giusti, D.; Burette, J.; Nguyen, Y.; Lévêque, N.; Graesslin, O.; Andreoletti, L. Virological diagnosis and management of two cases of congenital measles. J. Med. Virol. 2013, 85, 2136–2138. [Google Scholar]
- Di Guardo, G.; Cocumelli, C.; Scholl, F.; di Francesco, C.E.; Speranza, R.; Pennelli, M.; Eleni, C. Morbilliviral encephalitis in a striped dolphin Stenella coeruleoalba calf from Italy. Dis. Aquat. Organ. 2011, 95, 247–251. [Google Scholar] [CrossRef] [PubMed]
- West, K.L.; Levine, G.; Jacob, J.; Jensen, B.; Sanchez, S.; Colegrove, K.; Rotstein, D. Coinfection and vertical transmission of Brucella and Morbillivirus in a neonatal sperm whale (Physeter macrocephalus) in Hawaii, USA. J. Wildl. Dis. 2014. [Google Scholar] [CrossRef]
- Almberg, E.S.; Cross, P.C.; Smith, D.W. Persistence of canine distemper virus in the Greater Yellowstone ecosystem’s carnivore community. Ecol. Appl. 2010, 7, 2058–2074. [Google Scholar] [CrossRef]
- Dobson, A.; Holdo, R.M.; Holt, R.D. Rinderpes. In Encyclopedia of Biological Invasions; Simberloff, D., Rejmánek, M., Eds.; University of California Press: Berkeley, CA, USA, 1991; pp. 597–604. [Google Scholar]
- Griffin, D.E.; Bellini, W.J. Measles virus. In Fields Virology, 3rd ed.; Fields, B.N., Knipe, D.M., Howley, P.M., Chanock, R.M., Melnick, J.I., Monath, T.P., Roizman, B., Straus, S.E., Eds.; Lippincott-Raven Publishers: Philadelphia, PA, USA, 1996; pp. 1267–1312. [Google Scholar]
- Wohlsein, P.; Puff, C.; Kreutzer, M.; Siebert, U.; Baumgärtner, W. Distemper in a dolphin. Emerg. Infect. Dis. 2007, 13, 1959–1961. [Google Scholar] [CrossRef] [PubMed]
- Jauniaux, T.; Charlier, G.; Desmecht, M.; Haelters, J.; Jacques, T.; Losson, B.; van Gompel, J.; Tavernier, J.; Coignoul, F. Pathological findings in two fin whales (Balaenoptera physalus) with evidence of morbillivirus infection. J. Comp. Pathol. 2000, 123, 198–201. [Google Scholar]
- Aguilar, A.; Raga, J.A. The striped dolphin epizootic in the Mediterranean Sea. Ambio 1993, 22, 524–528. [Google Scholar]
- Tsur, L.; Goffman, O.; Yakobsen, B.; Moffet, D.; Kennedy, S. Morbillivirus infection in a bottlenose dolphin (Tursiops truncatus) from the Mediterranean Sea. Eur. J. Vet. Pathol. 1997, 2, 83–85. [Google Scholar]
- Hersh, S.L.; Odell, D.K.; Asper, E.D. Bottlenose dolphin mortality patterns in the Indian/Banana river system in Florida. In The Bottlenose Dolphin; Leatherwood, S.P., Reeves, R.R., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 155–164. [Google Scholar]
- Update on the dolphin morbillivirus outbreak and the 2013–2014 U.S.Mid-Atlantic bottlenose dolphin (Tursiops truncatus) unusual mortality event. Available online: https://events.iwc.int/index.php/scientific/SC65B/paper/viewFile/759/757/SC-65b-E03.pdf (accessed on 15 August 2014).
- Tomo, I.; Bastianello, S.; Woolford, L.; Kemper, C.; Dickson, C.; Kelly, D.; Wang, J.; Bingham, J. Pathology of dolphins during an unusual mortality event in South Australia, 2013. In Proceedings of the 20 Biennial Conference on the Biology of Marine Mammals, Dunedin, New Zealand, 9–13 December 2013.
- Kennedy, S.; Kuiken, T.; Ross, H.M.; McAliskey, M.; Moffett, D.; McNiven, M.; Carole, M. Morbillivirus infection in two common porpoises (Phocoena phocoena) from the coasts of England and Scotland. Vet. Rec. 1992, 31, 286–290. [Google Scholar] [CrossRef]
- Jepson, P.D.; Baker, J.R.; Kuiken, T.; Simpson, V.R.; Kennedy, S.; Bennett, P.M. Pulmonary pathology of harbour porpoises (Phocoena phocoena) stranded in England and Wales between 1990 and 1996. Vet. Rec. 2000, 46, 721–728. [Google Scholar] [CrossRef]
- Siebert, U.; Wünschmann, A.; Weiss, R.; Frank, H.; Benke, H.; Frese, K. Post-mortem findings in harbour porpoises (Phocoena phocoena) from the German North and Baltic Seas. J. Comp. Pathol. 2001, 124, 102–114. [Google Scholar]
- Jauniaux, T.; Petitjean, D.; Brenez, C.; Borrens, M.; Brosens, L.; Haelters, J.; Tavernier, T.; Coignoul, F. Post-mortem findings and causes of death of harbour porpoises (Phocoena phocoena) stranded from 1990 to 2000 along the coastlines of Belgium and Northern France. J. Comp. Pathol. 2002, 126, 243–253. [Google Scholar] [CrossRef] [PubMed]
- Hammond, P.S.; Berggren, P.; Benke, H.; Borchers, D.L.; Collet, A.; Heide-Jorgensen, M.P.; Heimlich, S.; Hiby, A.R.; Leopold, M.F.; Øien, N. Abundance of harbour porpoise and other cetaceans in the North Sea and adjacent waters. J. Appl. Ecol. 2002, 39, 361–376. [Google Scholar] [CrossRef]
- Forcada, J.; Aguilar, A.; Hammond, P.S.; Pastor, X.; Aguilar, R. Distribution and numbers of striped dolphins in the western Mediterranean sea after the 1990 epizootic outbreak. Mar. Mamm. Sci. 1994, 10, 137–150. [Google Scholar] [CrossRef]
- Garibaldi, F.; Mignone, W.; Caroggio, P.; Ballardini, M.; Podestà, M.; Bozzetta, E.; Casalone, C.; Marsilio, F.; di Francesco, C.E.; Proietto, U.; et al. Serological evidence of Morbillivirus infection in striped dolphins (Stenella coeruleoalba) found stranded on the Ligurian Sea coast of Italy. In Proceedings of the 22th European Cetacean Society Conference, Egmond aan Zee, The Netherlands, 10–12 March 2008; pp. 192–193.
- Wierucka, K.; Verborgh, P.; Meade, R.; Colmant, L.; Gauffier, P.; Esteban, R.; de Stephanis, R.; Cañadas, A. Effects of a morbillivirus epizootic on long-finned pilot whales Globicephala melas in Spanish Mediterranean waters. Mar. Ecol. Prog. Ser. 2014, 502, 1–10. [Google Scholar] [CrossRef]
- Gomez De Segura, A.; Hammond, P.S.; Cañadas, A.; Raga, J.A. Comparing cetacean abundance estimates derived from spatial models and design-based line transect methods. Mar. Ecol. Prog. Ser 2007, 329, 289–299. [Google Scholar] [CrossRef]
- Soto, S. Morbillivirus Infection in Mediterranean Striped Dolphins (Stenella coeruleoalba) during the 2007 Epidemic and the Post-Epidemic Years. Ph.D. Thesis, Universidad Autonoma de Barcelona, Barcelona, Spain, September 2014. [Google Scholar]
- Aguilar, A.; Borrell, A. Abnormally high polychlorinated biphenyl levels in striped dolphins (Stenella coeruleoalba) affected by the 1990–1992 Mediterranean epizootic. Sci. Total Environ. 1994, 154, 237–247. [Google Scholar] [CrossRef] [PubMed]
- Aznar, F.J.; Perdiguero, D.; Pérez del Olmo, A.; Repullés, A.; Agustí, C.; Raga, J.A. Changes in epizoic crustacean infestations during cetacean die-offs: The mass mortality of Mediterranean striped dolphins Stenella coeruleoalba revisited. Dis. Aquat. Org. 2005, 67, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Fossi, M.C.; Casini, S.; Marsili, L. Potential toxicological hazard due to endocrine-disrupting chemicals on Mediterranean top predators: State of art, gender differences and methodological tools. Environ. Res. 2007, 104, 174–182. [Google Scholar] [CrossRef] [PubMed]
- Valsecchi, E.; Amos, W.; Raga, J.A.; Podestà, M.; Sherwin, W. The effects of inbreeding on mortality during a morbillivirus outbreak in the Mediterranean striped dolphin (Stenella coeruleoalba). Anim. Cons. 2004, 7, 139–146. [Google Scholar] [CrossRef]
- Casalone, C.; Mazzariol, S.; Pautasso, A.; di Guardo, G.; di Nocera, F.; Lucifora, G.; Ligios, C.; Franco, A.; Fichi, G.; Cocumelli, C.; et al. Cetacean strandings in Italy: An unusual mortality event along the Tyrrhenian Sea coast in 2013. Dis. Aquat. Org. 2014, 109, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Nortobartolo di Sciara, G.; Birkun, G. Conserving Whales, Porpoises and Dolphins in the Mediterranean and Black Seas: An ACCOBAMS Status Report; ACOBAMS: Monaco, Principality of Monaco, 2010; p. 212. [Google Scholar]
- McLellan, W.; Friedlaender, A.; Mead, J.; Potter, C.; Pabst, D.A. Analysing 25 years of bottlenose dolphin (Tursiops truncatus) strandings along the Atlantic coast of the USA: Do historic records support the coastal migratory stock hypothesis. J. Cet. Res. Manag. 2002, 4, 297–304. [Google Scholar]
- Rosel, P.E.; Hansen, L.; Hohn, A.A. Restricted dispersal in a continuously distributed marine species: Common bottlenose dolphins Tursiops truncatus in coastal waters of the western North Atlantic. Mol. Ecol. 2009, 18, 5030–5045. [Google Scholar] [CrossRef] [PubMed]
- Lipscomb, T.P.; Kennedy, S.; Moffett, D.; Krafft, A.; Klaunberg, B.A.; Lichy, J.H.; Regan, G.T.; Worthy, G.A.; Taubenberger, J.K. Morbilliviral epizootic in bottlenose dolphins of the Gulf of Mexico. J. Vet. Diagn. Invest. 1996, 8, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Worthy, A.J. Patterns of Bottlenose Dolphin, Tursiops truncatus, Strandings in Texas. In Characteristics and Causes of Texas Marine Strandings; Zimmennan, R., Ed.; NOAA Technical Report NMFS 143; U.S. Department Commerce: Galveston, TX, USA, 1998; p. 85. [Google Scholar]
- Cantor, M.; Wedekin, L.L.; Daura-Jorge, F.G.; Rossi-Santos, M.R.; Simoes-Lopes, P. Assessing population parameters and trends of Guiana dolphins (Sotalia guianensis): An eight-year mark-recapture study. Mar. Mamm. Sci. 2012, 28, 63–83. [Google Scholar] [CrossRef]
- Rossi-Santos, M.; Wedekin, L.L.; Sousa-Lima, R.S. Distribution and habitat use of small cetaceans off Abrolhos Bank, eastern Brazil. LAJAM 2006, 5, 23–28. [Google Scholar] [CrossRef]
- Martins, C.C.A.; Morete, M.E.; Engel, M.H.; Freitas, A.C.; Secchi, E.R.; Kinas, P.G. Aspects of habitat use patterns of humpback whales in the Abrolhos Bank, Brazil, breeding ground. Mem. Qld. Mus. 2001, 47, 563–570. [Google Scholar]
- Wedekin, L.L.; Freitas, A.; Engel, M.H.; Sazima, I. Rough-toothed dolphins (Steno bredanensis) catch diskfishes while interacting with humpback whales (Megaptera novaeangliae) off Abrolhos Bank breeding ground, southwest Atlantic. Aquat. Mamm. 2004, 30, 327–329. [Google Scholar] [CrossRef]
- Groch, K.R.; Carvalho, V.L.; Meirelles, A.C.O.; Gonzales-Viera, O.; Kolesnikovas, C.K.M.; Groch, K.R.; Zucca, D.; Fernández, A.; Catão-Dias, J.L. Morbillivirus in live stranded cetaceans from Brazilian coast: Immunohistochemical evidence. In Proceedings of the 20th Biennial Conference on the Biology of Marine Mammals, Dunedin, New Zealand, 9–13 December 2013; pp. 85–86.
- Bodewes, R.; Morick, D.; van de Bildt, M.W.G.; Osinga, N.; Rubio Garcia, A.; Sanchez Contreras, G.J.; Smits, S.L.; Reperant, L.A.P.; Kuiken, T.; Osterhaus, A.D.M.E. 2014 Prevalence of phocine distemper virus specific antibodies: Bracing for the next seal epizootic in north-western Europe. Emerg. Microbes. Infect. 2013, 2, e3. [Google Scholar] [CrossRef]
- Prachayangprecha, S.; Schapendonk, C.M.; Koopmans, M.P.; Osterhaus, A.D.M.E.; Schürch, A.C.; Pas, S.D.; van der Eijk, A.A.; Poovorawan, Y.; Haagmans, B.L.; Smits, S.L. Exploring the potential of next-generation sequencing in detection of respiratory viruses. J. Clin. Microbiol. 2014, 52, 3722–3730. [Google Scholar] [CrossRef] [PubMed]
- Muniraju, M.; Munir, M.; Parthiban, A.R.; Banyard, A.C.; Bao, J.; Wang, Z.; Ayebazibwe, C.; Ayelet, G.; El Harrak, M.; Mahapatra, M.; et al. Molecular evolution of peste des petits ruminants virus. Emerg. Infect. Dis. 2014, 20, 2023–2033. [Google Scholar] [CrossRef] [PubMed]
- Drexler, J.F.; Corman, V.M.; Müller, M.A.; Maganga, G.D.; Vallo, P.; Binger, T.; Gloza-Rausch, F.; Cottontail, V.M.; Rasche, A.; Yordanov, S.; et al. Bats host major mammalian paramyxoviruses. Nat. Commun. 2012, 3, e796. [Google Scholar]
- Mazzariol, S.; Peletto, S.; Mondin, A.; Centelleghe, C.; di Guardo, G.; di Francesco, C.E.; Casalone, C.; Acutis, P.L. Dolphin morbillivirus infection in a captive harbor seal (Phoca vitulina). J. Clin. Microbiol. 2013, 51, 708–711. [Google Scholar] [CrossRef] [PubMed]
- Van de Bildt, M.W.; Martina, B.E.; Sidi, B.A.; Osterhaus, A.D.M.E. Morbillivirus infection in a bottlenose dolphin and a Mediterranean monk seal from the Atlantic coast of West Africa. Vet. Rec. 2001, 148, 210–211. [Google Scholar] [CrossRef] [PubMed]
- Grenfell, B.T.; Lonergan, M.E.; Harwood, J. Quantitative investigations of the epidemiology of phocine distemper virus (PDV) in European common seal populations. Sci. Total Environ. 1992, 115, 15–29. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Van Bressem, M.-F.; Duignan, P.J.; Banyard, A.; Barbieri, M.; Colegrove, K.M.; De Guise, S.; Di Guardo, G.; Dobson, A.; Domingo, M.; Fauquier, D.; et al. Cetacean Morbillivirus: Current Knowledge and Future Directions. Viruses 2014, 6, 5145-5181. https://doi.org/10.3390/v6125145
Van Bressem M-F, Duignan PJ, Banyard A, Barbieri M, Colegrove KM, De Guise S, Di Guardo G, Dobson A, Domingo M, Fauquier D, et al. Cetacean Morbillivirus: Current Knowledge and Future Directions. Viruses. 2014; 6(12):5145-5181. https://doi.org/10.3390/v6125145
Chicago/Turabian StyleVan Bressem, Marie-Françoise, Pádraig J. Duignan, Ashley Banyard, Michelle Barbieri, Kathleen M Colegrove, Sylvain De Guise, Giovanni Di Guardo, Andrew Dobson, Mariano Domingo, Deborah Fauquier, and et al. 2014. "Cetacean Morbillivirus: Current Knowledge and Future Directions" Viruses 6, no. 12: 5145-5181. https://doi.org/10.3390/v6125145
APA StyleVan Bressem, M.-F., Duignan, P. J., Banyard, A., Barbieri, M., Colegrove, K. M., De Guise, S., Di Guardo, G., Dobson, A., Domingo, M., Fauquier, D., Fernandez, A., Goldstein, T., Grenfell, B., Groch, K. R., Gulland, F., Jensen, B. A., Jepson, P. D., Hall, A., Kuiken, T., ... Wellehan, J. F. (2014). Cetacean Morbillivirus: Current Knowledge and Future Directions. Viruses, 6(12), 5145-5181. https://doi.org/10.3390/v6125145






