Immunology of Bats and Their Viruses: Challenges and Opportunities
Abstract
:1. Introduction
| Virus | Disease | Reservoir Host |
|---|---|---|
| Rabies virus and other lyssaviruses [1,32] | Rabies | Many bat species, world-wide distribution |
| Marburg virus [33] | Marburg virus disease | Egyptian fruit bat (Rousettus aegyptiacus) |
| Ebolaviruses [34] | Ebola virus disease | Hammer-headed bat (Hypsignathus monstrosus), Franquet’s epauletted fruit bat (Epomops franqueti), little collared fruit bat (Myonycteris torquata) 1 |
| SARS-CoV [26] | Severe acute respiratory syndrome | Chinese horseshoe bat (Rhinolophus spp.) 2 |
| MERS-CoV [35] | Middle East respiratory syndrome | Egyptian tomb bat (Taphozous perforatus) 2 |
| Nipah and Hendra viruses [36,37] | Encephalitis | Certain flying foxes (Pteropus spp.) |
| Sosuga virus [38] | Severe Acute Febrile Disease | Rousettus spp. 1 |
2. Viral Diseases of Bats
2.1. Rabies
2.2. Tacaribe Virus
2.3. Lloviu Virus
3. Bats as Reservoir Hosts of Important Human Pathogens
4. Bat Immune Responses to Viruses
4.1. Pattern Recognition Receptors
4.2. Interferons
4.2.1. Type I IFN
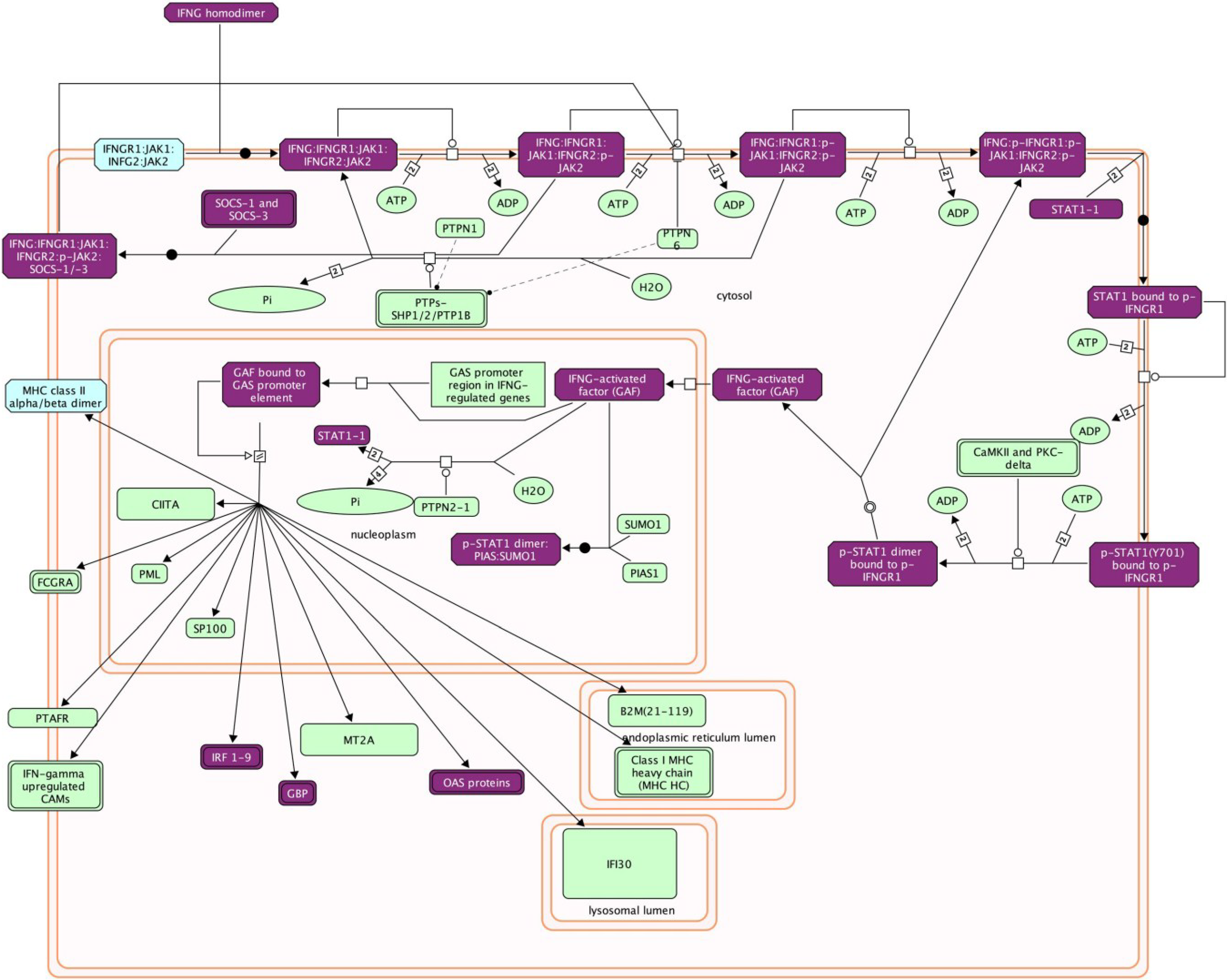
4.2.2. Type II IFN
4.2.3. Type III IFN
4.3. Responses of Innate Immune Cells and Lymphocytes
5. Approaches for Studying Bat Immune Responses
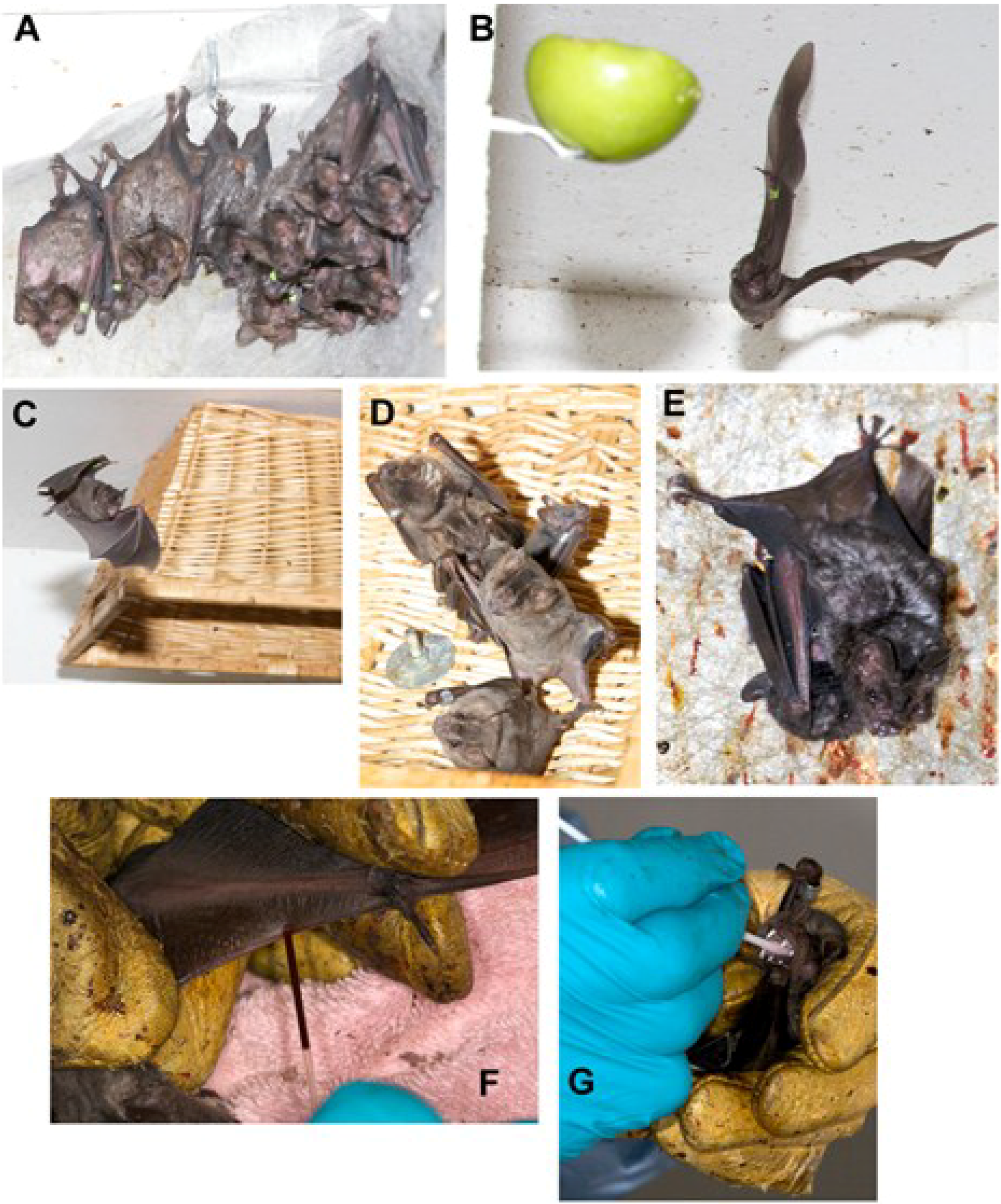
5.1. Husbandry

5.2. RNA-Seq
5.3. Real-Time PCR
5.4. Metabolomics
5.5. Immune Cell Culture
6. Conclusions
Acknowledgments
Conflicts of Interest
References
- Calisher, C.H.; Childs, J.E.; Field, H.E.; Holmes, K.V.; Schountz, T. Bats: Important reservoir hosts of emerging viruses. Clin. Microbiol. Rev. 2006, 19, 531–545. [Google Scholar] [CrossRef] [PubMed]
- Papenfuss, A.T.; Baker, M.L.; Feng, Z.P.; Tachedjian, M.; Crameri, G.; Cowled, C.; Ng, J.; Janardhana, V.; Field, H.E.; Wang, L.F. The immune gene repertoire of an important viral reservoir, the Australian black flying fox. BMC Genomics 2012, 13, e261. [Google Scholar] [CrossRef]
- Shaw, T.I.; Srivastava, A.; Chou, W.C.; Liu, L.; Hawkinson, A.; Glenn, T.C.; Adams, R.; Schountz, T. Transcriptome sequencing and annotation for the Jamaican fruit bat (Artibeus jamaicensis). PLOS ONE 2012, 7, e48472. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Cowled, C.; Shi, Z.; Huang, Z.; Bishop-Lilly, K.A.; Fang, X.; Wynne, J.W.; Xiong, Z.; Baker, M.L.; Zhao, W.; et al. Comparative analysis of bat genomes provides insight into the evolution of flight and immunity. Science 2013, 339, 456–460. [Google Scholar] [CrossRef] [PubMed]
- Calisher, C.H.; Holmes, K.V.; Dominguez, S.R.; Schountz, T.; Cryan, P. Bats prove to be rich reservoirs for emerging viruses. Microbe 2008, 3, 521–528. [Google Scholar]
- Omatsu, T.; Watanabe, S.; Akashi, H.; Yoshikawa, Y. Biological characters of bats in relation to natural reservoir of emerging viruses. Comp. Immunol. Microbiol. Infect. Dis. 2007, 30, 357–374. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.-F. Bats and Viruses: A brief review. Virologica Sinica 2009, 24, 93–99. [Google Scholar] [CrossRef]
- Wibbelt, G.; Moore, M.S.; Schountz, T.; Voigt, C.C. Emerging diseases in chiroptera: Why bats? Biol. Lett. 2010, 6, 438–440. [Google Scholar] [CrossRef] [PubMed]
- Blehert, D.S.; Hicks, A.C.; Behr, M.; Meteyer, C.U.; Berlowski-Zier, B.M.; Buckles, E.L.; Coleman, J.T.; Darling, S.R.; Gargas, A.; Niver, R.; et al. Bat white-nose syndrome: An emerging fungal pathogen? Science 2009, 323, 227. [Google Scholar] [CrossRef] [PubMed]
- Frick, W.F.; Pollock, J.F.; Hicks, A.C.; Langwig, K.E.; Reynolds, D.S.; Turner, G.G.; Butchkoski, C.M.; Kunz, T.H. An emerging disease causes regional population collapse of a common North American bat species. Science 2010, 329, 679–682. [Google Scholar] [CrossRef] [PubMed]
- Gargas, A.; Trest, M.T.; Christensen, M.; Volk, T.J.; Blehert, D.S. Geomyces destructans sp. nov. associated with bat white-nose syndrome. Mycotaxon. 2009, 108, 147–154. [Google Scholar]
- Bai, Y.; Kosoy, M.; Recuenco, S.; Alvarez, D.; Moran, D.; Turmelle, A.; Ellison, J.; Garcia, D.L.; Estevez, A.; Lindblade, K.; et al. Bartonella spp. in bats, Guatemala. Emerg. Infect. Dis. 2011, 17, 1269–1272. [Google Scholar] [CrossRef] [PubMed]
- Billeter, S.A.; Hayman, D.T.; Peel, A.J.; Baker, K.; Wood, J.L.; Cunningham, A.; Suu-Ire, R.; Dittmar, K.; Kosoy, M.Y. Bartonella species in bat flies (Diptera: Nycteribiidae) from western Africa. Parasitology 2012, 139, 324–329. [Google Scholar] [CrossRef] [PubMed]
- Kosoy, M.; Bai, Y.; Lynch, T.; Kuzmin, I.V.; Niezgoda, M.; Franka, R.; Agwanda, B.; Breiman, R.F.; Rupprecht, C.E. Bartonella spp. in bats, Kenya. Emerg. Infect. Dis. 2010, 16, 1875–1881. [Google Scholar] [CrossRef] [PubMed]
- Morse, S.F.; Olival, K.J.; Kosoy, M.; Billeter, S.; Patterson, B.D.; Dick, C.W.; Dittmar, K. Global distribution and genetic diversity of Bartonella in bat flies (Hippoboscoidea, Streblidae, Nycteribiidae). Infect. Genet. Evol. J. Mol. Epidemiol. Evol. Genet. Infect. Dis. 2012, 12, 1717–1723. [Google Scholar] [CrossRef]
- Reeves, W.K.; Streicker, D.G.; Loftis, A.D.; Dasch, G.A. Serologic survey of Eptesicus fuscus from Georgia, U.S.A. for Rickettsia and Borrelia and laboratory transmission of a Rickettsia by bat ticks. J. Vector Ecol. 2006, 31, 386–389. [Google Scholar] [CrossRef] [PubMed]
- Muhldorfer, K.; Speck, S.; Kurth, A.; Lesnik, R.; Freuling, C.; Muller, T.; Kramer-Schadt, S.; Wibbelt, G. Diseases and causes of death in European bats: Dynamics in disease susceptibility and infection rates. PLOS ONE 2011, 6, e29773. [Google Scholar] [CrossRef] [PubMed]
- Muhldorfer, K.; Speck, S.; Wibbelt, G. Diseases in free-ranging bats from Germany. BMC Vet. Res. 2011, 7, e61. [Google Scholar] [CrossRef]
- Muhldorfer, K.; Wibbelt, G.; Haensel, J.; Riehm, J.; Speck, S. Yersinia species isolated from bats, Germany. Emerg. Infect. Dis. 2010, 16, 578–580. [Google Scholar] [CrossRef] [PubMed]
- Greer, D.L.; McMurray, D.N. Pathogenesis of experimental histoplasmosis in the bat, Artibeus lituratus. Am. J. Trop. Med. Hyg. 1981, 30, 653–659. [Google Scholar] [PubMed]
- Breed, A.C.; Field, H.E.; Smith, C.S.; Edmonston, J.; Meers, J. Bats Without Borders: Long-distance movements and implications for disease risk management. EcoHealth 2010, 7, 204–212. [Google Scholar] [CrossRef] [PubMed]
- Barr, J.; Smith, C.; Smith, I.; de Jong, C.; Todd, S.; Melville, D.; Broos, A.; Crameri, S.; Haining, J.; Marsh, G.; et al. Isolation of multiple novel paramyxoviruses from pteropid bat urine. J. Gener. Virol. 2014. [Google Scholar] [CrossRef]
- Carrington, C.V.; Foster, J.E.; Zhu, H.C.; Zhang, J.X.; Smith, G.J.; Thompson, N.; Auguste, A.J.; Ramkissoon, V.; Adesiyun, A.A.; Guan, Y. Detection and phylogenetic analysis of group 1 coronaviruses in South American bats. Emerg. Infect. Dis. 2008, 14, 1890–1893. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.K.; Poon, L.L.; Guan, Y.; Peiris, J.S. Novel astroviruses in insectivorous bats. J. Virol. 2008, 82, 9107–9114. [Google Scholar] [CrossRef] [PubMed]
- Dominguez, S.R.; O'Shea, T.J.; Oko, L.M.; Holmes, K.V. Detection of group 1 coronaviruses in bats in North America. Emerg. Infect. Dis. 2007, 13, 1295–1300. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.; Woo, P.C.; Li, K.S.; Huang, Y.; Tsoi, H.W.; Wong, B.H.; Wong, S.S.; Leung, S.Y.; Chan, K.H.; Yuen, K.Y. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc. Natl. Acad. Sci. USA 2005, 102, 14040–14045. [Google Scholar] [CrossRef] [PubMed]
- Quan, P.L.; Firth, C.; Conte, J.M.; Williams, S.H.; Zambrana-Torrelio, C.M.; Anthony, S.J.; Ellison, J.A.; Gilbert, A.T.; Kuzmin, I.V.; Niezgoda, M.; et al. Bats are a major natural reservoir for hepaciviruses and pegiviruses. Proc. Natl. Acad. Sci. USA 2013, 110, 8194–8199. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Li, M.; Li, L.; Monagin, C.; Chmura, A.A.; Schneider, B.S.; Epstein, J.H.; Mei, X.; Shi, Z.; Daszak, P.; et al. Evidence for retrovirus and paramyxovirus infection of multiple bat species in China. Viruses 2014, 6, 2138–2154. [Google Scholar] [CrossRef] [PubMed]
- Yanagihara, R.; Gu, S.H.; Arai, S.; Kang, H.J.; Song, J.W. Hantaviruses: Rediscovery and new beginnings. Virus Res. 2014, 187, 6–14. [Google Scholar] [CrossRef] [PubMed]
- Pulliam, J.R.; Epstein, J.H.; Dushoff, J.; Rahman, S.A.; Bunning, M.; Jamaluddin, A.A.; Hyatt, A.D.; Field, H.E.; Dobson, A.P.; Daszak, P. Agricultural intensification, priming for persistence and the emergence of Nipah virus: A lethal bat-borne zoonosis. J. R. Soc. Interface/ R. Soc. 2012, 9, 89–101. [Google Scholar] [CrossRef]
- Wood, J.L.; Leach, M.; Waldman, L.; Macgregor, H.; Fooks, A.R.; Jones, K.E.; Restif, O.; Dechmann, D.; Hayman, D.T.; Baker, K.S.; et al. A framework for the study of zoonotic disease emergence and its drivers: Spillover of bat pathogens as a case study. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 2012, 367, 2881–2892. [Google Scholar] [CrossRef]
- Rupprecht, C.E.; Smith, J.S.; Fekadu, M.; Childs, J.E. The ascension of wildlife rabies: A cause for public health concern or intervention? Emerg. Infect. Dis. 1995, 1, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Towner, J.S.; Pourrut, X.; Albarino, C.G.; Nkogue, C.N.; Bird, B.H.; Grard, G.; Ksiazek, T.G.; Gonzalez, J.P.; Nichol, S.T.; Leroy, E.M. Marburg virus infection detected in a common African bat. PLOS ONE 2007, 2, e764. [Google Scholar] [PubMed]
- Leroy, E.M.; Kumulungui, B.; Pourrut, X.; Rouquet, P.; Hassanin, A.; Yaba, P.; Delicat, A.; Paweska, J.T.; Gonzalez, J.P.; Swanepoel, R. Fruit bats as reservoirs of Ebola virus. Nature 2005, 438, 575–576. [Google Scholar] [CrossRef] [PubMed]
- Memish, Z.A.; Mishra, N.; Olival, K.J.; Fagbo, S.F.; Kapoor, V.; Epstein, J.H.; Alhakeem, R.; Durosinloun, A.; Al Asmari, M.; Islam, A.; et al. Middle East respiratory syndrome coronavirus in bats, Saudi Arabia. Emerg. Infect. Dis. 2013, 19, 1819–1823. [Google Scholar] [CrossRef] [PubMed]
- Chua, K.B.; Koh, C.L.; Hooi, P.S.; Wee, K.F.; Khong, J.H.; Chua, B.H.; Chan, Y.P.; Lim, M.E.; Lam, S.K. Isolation of Nipah virus from Malaysian Island flying-foxes. Microbes Infect. 2002, 4, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Halpin, K.; Young, P.L.; Field, H.E.; Mackenzie, J.S. Isolation of Hendra virus from pteropid bats: A natural reservoir of Hendra virus. J. Gener. Virol. 2000, 81, 1927–1932. [Google Scholar]
- Albarino, C.G.; Foltzer, M.; Towner, J.S.; Rowe, L.A.; Campbell, S.; Jaramillo, C.M.; Bird, B.H.; Reeder, D.M.; Vodzak, M.E.; Rota, P.; et al. Novel paramyxovirus associated with severe acute febrile disease, South Sudan and Uganda, 2012. Emerg. Infect. Dis. 2014, 20, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Alcami, A.; Koszinowski, U.H. Viral mechanisms of immune evasion. Trends in Microbiology 2000, 8, 410–418. [Google Scholar] [CrossRef] [PubMed]
- Grandvaux, N.; tenOever, B.R.; Servant, M.J.; Hiscott, J. The interferon antiviral response: From viral invasion to evasion. Curr. Opin. Infect. Dis. 2002, 15, 259–267. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.J.; Horvath, C.M. Host evasion by emerging paramyxoviruses: Hendra virus and Nipah virus V proteins inhibit interferon signaling. Viral Immunol. 2004, 17, 210–219. [Google Scholar] [CrossRef] [PubMed]
- Shaw, M.L. Henipaviruses Employ a Multifaceted approach to evade the antiviral interferon response. Viruses 2009, 1, 1190–1203. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.J.; Cruz, C.D.; Horvath, C.M. Identification of the nuclear export signal and STAT-binding domains of the Nipah virus V protein reveals mechanisms underlying interferon evasion. J. Virol. 2004, 78, 5358–5367. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.J.; Parisien, J.P.; Horvath, C.M. Nipah virus V protein evades alpha and gamma interferons by preventing STAT1 and STAT2 activation and nuclear accumulation. J. Virol. 2002, 76, 11476–11483. [Google Scholar] [CrossRef] [PubMed]
- Shaw, M.L.; Garcia-Sastre, A.; Palese, P.; Basler, C.F. Nipah virus V and W proteins have a common STAT1-binding domain yet inhibit STAT1 activation from the cytoplasmic and nuclear compartments, respectively. J. Virol. 2004, 78, 5633–5641. [Google Scholar] [CrossRef] [PubMed]
- Schountz, T.; Quackenbush, S.; Rovnak, J.; Haddock, E.; Black, W.C., IV; Feldmann, H.; Prescott, J. Differential lymphocyte and antibody responses in deer mice infected with Sin Nombre hantavirus or Andes hantavirus. J. Virol. 2014, 88, 8319–8331. [Google Scholar] [CrossRef] [PubMed]
- Halpin, K.; Hyatt, A.D.; Fogarty, R.; Middleton, D.; Bingham, J.; Epstein, J.H.; Rahman, S.A.; Hughes, T.; Smith, C.; Field, H.E.; et al. Pteropid bats are confirmed as the reservoir hosts of henipaviruses: A comprehensive experimental study of virus transmission. Am. J. Trop. Med. Hyg. 2011, 85, 946–951. [Google Scholar] [CrossRef] [PubMed]
- Amman, B.R.; Jones, M.E.; Sealy, T.K.; Uebelhoer, L.S.; Schuh, A.J.; Bird, B.H.; Coleman-McCray, J.D.; Martin, B.E.; Nichol, S.T.; Towner, J.S. Oral shedding of Marburg virus in experimentally infected Egyptian fruit bats (Rousettus Aegyptiacus). J. Wildl. Dis. 2014. [Google Scholar] [CrossRef]
- Paweska, J.T.; Jansen van Vuren, P.; Masumu, J.; Leman, P.A.; Grobbelaar, A.A.; Birkhead, M.; Clift, S.; Swanepoel, R.; Kemp, A. Virological and serological findings in Rousettus aegyptiacus experimentally inoculated with vero cells-adapted hogan strain of Marburg virus. PLOS ONE 2012, 7, e45479. [Google Scholar] [CrossRef] [PubMed]
- Luis, A.D.; Hayman, D.T.; O'Shea, T.J.; Cryan, P.M.; Gilbert, A.T.; Pulliam, J.R.; Mills, J.N.; Timonin, M.E.; Willis, C.K.; Cunningham, A.A.; et al. A comparison of bats and rodents as reservoirs of zoonotic viruses: Are bats special? Proc. Biol. Sci./R. Soc. 2013, 280, e20122753. [Google Scholar] [CrossRef]
- O’Shea, T.J.; Cryan, P.M.; Cunningham, A.A.; Fooks, A.R.; Hayman, D.T.; Luis, A.D.; Peel, A.J.; Plowright, R.K.; Wood, J.L. Bat flight and zoonotic viruses. Emerg. Infect. Dis. 2014, 20, 741–745. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.F.; Walker, P.J.; Poon, L.L. Mass extinctions, biodiversity and mitochondrial function: Are bats ‘special’ as reservoirs for emerging viruses? Curr. Opin. Virol. 2011, 1, 649–657. [Google Scholar] [CrossRef] [PubMed]
- Botten, J.; Mirowsky, K.; Kusewitt, D.; Bharadwaj, M.; Yee, J.; Ricci, R.; Feddersen, R.M.; Hjelle, B. Experimental infection model for Sin Nombre hantavirus in the deer mouse (Peromyscus maniculatus). Proc. Natl. Acad. Sci. USA 2000, 97, 10578–10583. [Google Scholar] [CrossRef] [PubMed]
- Botten, J.; Mirowsky, K.; Kusewitt, D.; Ye, C.; Gottlieb, K.; Prescott, J.; Hjelle, B. Persistent Sin Nombre virus infection in the deer mouse (Peromyscus maniculatus) model: Sites of replication and strand-specific expression. J. Virol. 2003, 77, 1540–1550. [Google Scholar] [CrossRef] [PubMed]
- Fulhorst, C.F.; Ksiazek, T.G.; Peters, C.J.; Tesh, R.B. Experimental infection of the cane mouse Zygodontomys brevicauda (family Muridae) with Guanarito virus (Arenaviridae), the etiologic agent of Venezuelan hemorrhagic fever. J. Infect. Dis. 1999, 180, 966–969. [Google Scholar] [CrossRef] [PubMed]
- Yanagihara, R.; Amyx, H.L.; Gajdusek, D.C. Experimental infection with Puumala virus, the etiologic agent of nephropathia epidemica, in bank voles (Clethrionomys glareolus). J. Virol. 1985, 55, 34–38. [Google Scholar] [PubMed]
- Easterbrook, J.D.; Zink, M.C.; Klein, S.L. Regulatory T cells enhance persistence of the zoonotic pathogen Seoul virus in its reservoir host. Proc. Natl. Acad. Sci. USA 2007, 104, 15502–15507. [Google Scholar] [CrossRef] [PubMed]
- Schountz, T.; Prescott, J.; Cogswell, A.C.; Oko, L.; Mirowsky-Garcia, K.; Galvez, A.P.; Hjelle, B. Regulatory T cell-like responses in deer mice persistently infected with Sin Nombre virus. Proc. Natl. Acad. Sci. USA 2007, 104, 15496–15501. [Google Scholar] [CrossRef] [PubMed]
- Calisher, C.H.; Ellison, J.A. The other rabies viruses: The emergence and importance of lyssaviruses from bats and other vertebrates. Travel Med. Infect. Dis. 2012, 10, 69–79. [Google Scholar] [CrossRef] [PubMed]
- Kuzmin, I.V.; Bozick, B.; Guagliardo, S.A.; Kunkel, R.; Shak, J.R.; Tong, S.; Rupprecht, C.E. Bats, emerging infectious diseases, and the rabies paradigm revisited. Emerg. Health Threats J. 2011, 4, e7159. [Google Scholar]
- Rupprecht, C.E.; Turmelle, A.; Kuzmin, I.V. A perspective on lyssavirus emergence and perpetuation. Curr. Opin. Virol. 2011, 1, 662–670. [Google Scholar] [CrossRef] [PubMed]
- Turmelle, A.S.; Jackson, F.R.; Green, D.; McCracken, G.F.; Rupprecht, C.E. Host immunity to repeated rabies virus infection in big brown bats. J. Gener. Virol. 2010, 91, 2360–2366. [Google Scholar] [CrossRef]
- Davis, A.D.; Jarvis, J.A.; Pouliott, C.E.; Morgan, S.M.; Rudd, R.J. Susceptibility and pathogenesis of little brown bats (Myotis lucifugus) to heterologous and homologous rabies viruses. J. Virol. 2013, 87, 9008–9015. [Google Scholar] [CrossRef] [PubMed]
- Downs, W.G.; Anderson, C.R.; Spence, L.; Aitken, T.H.G.; Greenhall, A.H. Tacaribe virus, a new agent isolated from Artibeus bats and mosquitos in Trinidad, West Indies. American J. Trop. Med. Hyg. 1963, 12, 640–646. [Google Scholar]
- Tikasingh, E.; Caribbean Epidemiology Centre, Port of Spain, Trinidad and Tobago. Personal communication, 2011.
- Cogswell-Hawkinson, A.; Bowen, R.; James, S.; Gardiner, D.; Calisher, C.H.; Adams, R.; Schountz, T. Tacaribe virus causes fatal infection of an ostensible reservoir host, the Jamaican fruit bat. J. Virol. 2012, 86, 5791–5799. [Google Scholar] [CrossRef] [PubMed]
- Abraham, J.; Corbett, K.D.; Farzan, M.; Choe, H.; Harrison, S.C. Structural basis for receptor recognition by New World hemorrhagic fever arenaviruses. Nat. Struct. Mol. Biol. 2010, 17, 438–444. [Google Scholar] [CrossRef] [PubMed]
- Charrel, R.N.; Feldmann, H.; Fulhorst, C.F.; Khelifa, R.; de Chesse, R.; de Lamballerie, X. Phylogeny of New World arenaviruses based on the complete coding sequences of the small genomic segment identified an evolutionary lineage produced by intrasegmental recombination. Biochem. Biophys. Res. Commun. 2002, 296, 1118–1124. [Google Scholar] [CrossRef] [PubMed]
- Ortega, J.; Castro-Arellano, I. Mammalian Species: Artibeus jamaicensis. Am. Soc. Mammal. 2001, 662, 1–9. [Google Scholar] [CrossRef]
- Gard, G.; Australian Animal Health Laboratory, East Geelong, Victoria, Australia. Personal communication, 2009.
- Negredo, A.; Palacios, G.; Vazquez-Moron, S.; Gonzalez, F.; Dopazo, H.; Molero, F.; Juste, J.; Quetglas, J.; Savji, N.; de la Cruz Martinez, M.; et al. Discovery of an ebolavirus-like filovirus in europe. PLOS Pathogens 2011, 7, e1002304. [Google Scholar] [CrossRef] [PubMed]
- Thompson, N.N.; Auguste, A.J.; Travassos da Rosa, A.P.; Carrington, C.V.; Blitvich, B.J.; Chadee, D.D.; Tesh, R.B.; Weaver, S.C.; Adesiyun, A.A. Seroepidemiology of selected alphaviruses and flaviviruses in bats in Trinidad. Zoonoses Public Health 2014. [Google Scholar]
- Xiao, J.; Li, J.; Hu, G.; Chen, Z.; Wu, Y.; Chen, Y.; Chen, Z.; Liao, Y.; Zhou, J.; Ke, X.; et al. Isolation and phylogenetic characterization of bat astroviruses in southern China. Arch. Virol. 2011, 156, 1415–1423. [Google Scholar] [CrossRef] [PubMed]
- Badrane, H.; Tordo, N. Host switching in lyssavirus history from the Chiroptera to the Carnivora orders. J. Virol. 2001, 75, 8096–8104. [Google Scholar] [CrossRef] [PubMed]
- Holmes, E.C.; Woelk, C.H.; Kassis, R.; Bourhy, H. Genetic constraints and the adaptive evolution of rabies virus in nature. Virology 2002, 292, 247–257. [Google Scholar] [CrossRef] [PubMed]
- Krysko, D.V.; Agostinis, P.; Krysko, O.; Garg, A.D.; Bachert, C.; Lambrecht, B.N.; Vandenabeele, P. Emerging role of damage-associated molecular patterns derived from mitochondria in inflammation. Trends Immunol. 2011, 32, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Ching, S.; Zhang, H.; Belevych, N.; He, L.; Lai, W.; Pu, X.A.; Jaeger, L.B.; Chen, Q.; Quan, N. Endothelial-specific knockdown of interleukin-1 (IL-1) type 1 receptor differentially alters CNS responses to IL-1 depending on its route of administration. J. Neurosci.: Off. J. Soc. Neurosci. 2007, 27, 10476–10486. [Google Scholar] [CrossRef]
- Wilhelms, D.B.; Kirilov, M.; Mirrasekhian, E.; Eskilsson, A.; Kugelberg, U.O.; Klar, C.; Ridder, D.A.; Herschman, H.R.; Schwaninger, M.; Blomqvist, A.; et al. Deletion of prostaglandin e2 synthesizing enzymes in brain endothelial cells attenuates inflammatory Fever. J. Neurosci.: Off. J. Soc. Neurosci. 2014, 34, 11684–11690. [Google Scholar] [CrossRef]
- Baker, M.L.; Tachedjian, M.; Wang, L.F. Immunoglobulin heavy chain diversity in Pteropid bats: Evidence for a diverse and highly specific antigen binding repertoire. Immunogenetics 2010, 62, 173–184. [Google Scholar] [CrossRef] [PubMed]
- Cowled, C.; Baker, M.L.; Zhou, P.; Tachedjian, M.; Wang, L.F. Molecular characterisation of RIG-I-like helicases in the black flying fox, Pteropus alecto. Dev. Compar. Immunol. 2012, 36, 657–664. [Google Scholar] [CrossRef]
- Janardhana, V.; Tachedjian, M.; Crameri, G.; Cowled, C.; Wang, L.F.; Baker, M.L. Cloning, expression and antiviral activity of IFNgamma from the Australian fruit bat, Pteropus alecto. Dev. Compar. Immunol. 2012, 36, 610–618. [Google Scholar] [CrossRef]
- Virtue, E.R.; Marsh, G.A.; Baker, M.L.; Wang, L.F. Interferon production and signaling pathways are antagonized during henipavirus infection of fruit bat cell lines. PLOS ONE 2011, 6, e22488. [Google Scholar] [CrossRef] [PubMed]
- Wynne, J.W.; di Rubbo, A.; Shiell, B.J.; Beddome, G.; Cowled, C.; Peck, G.R.; Huang, J.; Grimley, S.L.; Baker, M.L.; Michalski, W.P. Purification and characterisation of immunoglobulins from the Australian black flying fox (Pteropus alecto) using anti-fab affinity chromatography reveals the low abundance of IgA. PLOS ONE 2013, 8, e52930. [Google Scholar] [CrossRef] [PubMed]
- Wynne, J.W.; Shiell, B.J.; Marsh, G.A.; Boyd, V.; Harper, J.A.; Heesom, K.; Monaghan, P.; Zhou, P.; Payne, J.; Klein, R.; et al. Proteomics informed by transcriptomics reveals Hendra virus sensitizes bat cells to TRAIL mediated apoptosis. Genome Biol. 2014, 15, e532. [Google Scholar] [CrossRef]
- Zhou, P.; Cowled, C.; Marsh, G.A.; Shi, Z.; Wang, L.F.; Baker, M.L. Type III IFN receptor expression and functional characterisation in the pteropid bat, Pteropus alecto. PLOS ONE 2011, 6, e25385. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Cowled, C.; Wang, L.F.; Baker, M.L. Bat Mx1 and Oas1, but not Pkr are highly induced by bat interferon and viral infection. Dev. Compar. Immunol. 2013, 3-4, 240–247. [Google Scholar] [CrossRef]
- Baker, M.L.; Schountz, T.; Wang, L.F. Antiviral immune responses of bats: A review. Zoonoses Public Health 2013, 60, 104–116. [Google Scholar] [CrossRef] [PubMed]
- Cowled, C.; Stewart, C.R.; Likic, V.A.; Friedlander, M.R.; Tachedjian, M.; Jenkins, K.A.; Tizard, M.L.; Cottee, P.; Marsh, G.A.; Zhou, P.; et al. Characterisation of novel microRNAs in the Black flying fox (Pteropus alecto) by deep sequencing. BMC Genomics 2014, 15, e682. [Google Scholar] [CrossRef]
- Dennehy, J.J. What ecologists can tell virologists. Annu. Rev. Microbial. 2014, 68, 117–135. [Google Scholar] [CrossRef]
- Weitzman, M.D.; Weitzman, J.B. What’s the damage? The impact of pathogens on pathways that maintain host genome integrity. Cell Host Microbe 2014, 15, 283–294. [Google Scholar] [CrossRef] [PubMed]
- Takeda, K.; Kaisho, T.; Akira, S. Toll-like receptors. Annu. Rev. Immunol. 2003, 21, 335–376. [Google Scholar] [CrossRef] [PubMed]
- Medzhitov, R.; Janeway, C.A., Jr. Innate immunity: The virtues of a nonclonal system of recognition. Cell 1997, 91, 295–298. [Google Scholar] [CrossRef] [PubMed]
- Barbalat, R.; Ewald, S.E.; Mouchess, M.L.; Barton, G.M. Nucleic acid recognition by the innate immune system. Annu. Rev. Immunol. 2011, 29, 185–214. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Shaw, M.H.; Kim, Y.G.; Nunez, G. NOD-like receptors: Role in innate immunity and inflammatory disease. Annu. Rev. Pathol. 2009, 4, 365–398. [Google Scholar] [CrossRef] [PubMed]
- Cowled, C.; Baker, M.; Tachedjian, M.; Zhou, P.; Bulach, D.; Wang, L.F. Molecular characterisation of Toll-like receptors in the black flying fox Pteropus alecto. Dev. Compar. Immunol. 2011, 35, 7–18. [Google Scholar] [CrossRef]
- Iha, K.; Omatsu, T.; Watanabe, S.; Ueda, N.; Taniguchi, S.; Fujii, H.; Ishii, Y.; Kyuwa, S.; Akashi, H.; Yoshikawa, Y. Molecular cloning and expression analysis of bat toll-like receptors 3, 7 and 9. J. Vet. Med. Sci./Jpn. Soc. Vet. Sci. 2010, 72, 217–220. [Google Scholar] [CrossRef]
- Diaz, M.O.; Pomykala, H.M.; Bohlander, S.K.; Maltepe, E.; Malik, K.; Brownstein, B.; Olopade, O.I. Structure of the human type-I interferon gene cluster determined from a YAC clone contig. Genomics 1994, 22, 540–552. [Google Scholar] [CrossRef] [PubMed]
- Biesold, S.E.; Ritz, D.; Gloza-Rausch, F.; Wollny, R.; Drexler, J.F.; Corman, V.M.; Kalko, E.K.; Oppong, S.; Drosten, C.; Muller, M.A. Type I interferon reaction to viral infection in interferon-competent, immortalized cell lines from the African fruit bat Eidolon helvum. PLOS ONE 2011, 6, e28131. [Google Scholar] [CrossRef] [PubMed]
- Sheppard, P.; Kindsvogel, W.; Xu, W.; Henderson, K.; Schlutsmeyer, S.; Whitmore, T.E.; Kuestner, R.; Garrigues, U.; Birks, C.; Roraback, J.; et al. IL-28, IL-29 and their class II cytokine receptor IL-28R. Nat. Immunol. 2003, 4, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Cowled, C.; Todd, S.; Crameri, G.; Virtue, E.R.; Marsh, G.A.; Klein, R.; Shi, Z.; Wang, L.F.; Baker, M.L. Type III IFNs in pteropid bats: Differential expression patterns provide evidence for distinct roles in antiviral immunity. J. Immunol. 2011, 186, 3138–3147. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, A.K.; Chakravarty, A.K. Plaque forming cell assay for antibody secreting cells in the bat Pteropus giganteus. Indian J. Exp. Biol. 1983, 21, 5–7. [Google Scholar] [PubMed]
- Chakravarty, A.K.; Sarkar, S.K. Immunofluorescence analysis of immunoglobulin bearing lymphocytes in the Indian fruit bat: Pteropus giganteus. Lymphology 1994, 27, 97–104. [Google Scholar] [PubMed]
- Paul, B.N.; Chakravarty, A.K. In vitro analysis of delayed immune response in a bat, Pteropus giganteus: Process of con-A mediated activation. Dev. Compar. Immunol. 1986, 10, 55–67. [Google Scholar] [CrossRef]
- Sarkar, S.K.; Chakravarty, A.K. Analysis of immunocompetent cells in the bat, Pteropus giganteus: Isolation and scanning electron microscopic characterization. Dev. Compar. Immunol. 1991, 15, 423–430. [Google Scholar] [CrossRef]
- Bratsch, S.; Wertz, N.; Chaloner, K.; Kunz, T.H.; Butler, J.E. The little brown bat, M. lucifugus, displays a highly diverse V H, D H and J H repertoire but little evidence of somatic hypermutation. Dev. Compar. Immunol. 2011, 35, 421–430. [Google Scholar] [CrossRef]
- Butler, J.E.; Wertz, N.; Zhao, Y.; Zhang, S.; Bao, Y.; Bratsch, S.; Kunz, T.H.; Whitaker, J.O., Jr.; Schountz, T. The two suborders of chiropterans have the canonical heavy-chain immunoglobulin (Ig) gene repertoire of eutherian mammals. Dev. Compar. Immunol. 2011, 35, 273–284. [Google Scholar] [CrossRef]
- Evans, V.C.; Barker, G.; Heesom, K.J.; Fan, J.; Bessant, C.; Matthews, D.A. De novo derivation of proteomes from transcriptomes for transcript and protein identification. Nat. Methods 2012, 9, 1207–1211. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Watanabe, S.; Yamane, D.; Ueda, N.; Iha, K.; Taniguchi, S.; Kato, K.; Tohya, Y.; Kyuwa, S.; Yoshikawa, Y.; et al. Functional analysis of Rousettus aegyptiacus “signal transducer and activator of transcription 1” (STAT1). Dev. Compar. Immunol. 2010, 34, 598–602. [Google Scholar] [CrossRef]
- Valmas, C.; Grosch, M.N.; Schumann, M.; Olejnik, J.; Martinez, O.; Best, S.M.; Krahling, V.; Basler, C.F.; Muhlberger, E. Marburg virus evades interferon responses by a mechanism distinct from ebola virus. PLOS Pathogens 2010, 6, e1000721. [Google Scholar] [CrossRef] [PubMed]
- Glenn, T.C. Field guide to next-generation DNA sequencers. Mol. Ecol. Resour. 2011, 11, 759–769. [Google Scholar] [CrossRef] [PubMed]
- Griseri, P.; Pages, G. Control of pro-angiogenic cytokine mRNA half-life in cancer: The role of AU-rich elements and associated proteins. J. Interferon Cytokine Res.: Off. J. Int. Soc. Interferon Cytokine Res. 2014, 34, 242–254. [Google Scholar] [CrossRef]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef] [PubMed]
- Schulz, M.H.; Zerbino, D.R.; Vingron, M.; Birney, E. Oases: Robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 2012, 28, 1086–1092. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, eR106. [Google Scholar] [CrossRef]
- Anders, S.; McCarthy, D.J.; Chen, Y.; Okoniewski, M.; Smyth, G.K.; Huber, W.; Robinson, M.D. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nat. Protocols 2013, 8, 1765–1786. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, e323. [Google Scholar] [CrossRef]
- Forster, S.C.; Finkel, A.M.; Gould, J.A.; Hertzog, P.J. RNA-eXpress annotates novel transcript features in RNA-seq data. Bioinformatics 2013, 29, 810–812. [Google Scholar] [CrossRef] [PubMed]
- Dennis, G., Jr.; Sherman, B.T.; Hosack, D.A.; Yang, J.; Gao, W.; Lane, H.C.; Lempicki, R.A. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003, 4, eP3. [Google Scholar] [CrossRef]
- Vastrik, I.; D’Eustachio, P.; Schmidt, E.; Gopinath, G.; Croft, D.; de Bono, B.; Gillespie, M.; Jassal, B.; Lewis, S.; Matthews, L.; et al. Reactome: A knowledge base of biologic pathways and processes. Genome Biol. 2007, 8, eR39. [Google Scholar] [CrossRef]
- Ligtenberg, W.P.; Bosnacki, D.; Hilbers, P.A. Reconn: A cytoscape plug-in for exploring and visualizing reactome. J. Bioinform. Comput. Biol. 2013, 11, e1350004. [Google Scholar] [CrossRef]
- Schountz, T.; Shaw, T.I.; Glenn, T.C.; Feldmann, H.; Prescott, J. Expression profiling of lymph node cells from deer mice infected with Andes virus. BMC Immunol. 2013, 14, e18. [Google Scholar] [CrossRef]
- Chukkapalli, V.; Heaton, N.S.; Randall, G. Lipids at the interface of virus-host interactions. Curr. Opin. Microbiol. 2012, 15, 512–518. [Google Scholar] [CrossRef] [PubMed]
- Heaton, N.S.; Randall, G. Multifaceted roles for lipids in viral infection. Trends Microbiol. 2011, 19, 368–375. [Google Scholar] [CrossRef] [PubMed]
- Pearce, E.L.; Pearce, E.J. Metabolic pathways in immune cell activation and quiescence. Immunity 2013, 38, 633–643. [Google Scholar] [CrossRef] [PubMed]
- Diamond, D.L.; Syder, A.J.; Jacobs, J.M.; Sorensen, C.M.; Walters, K.A.; Proll, S.C.; McDermott, J.E.; Gritsenko, M.A.; Zhang, Q.; Zhao, R.; et al. Temporal proteome and lipidome profiles reveal hepatitis C virus-associated reprogramming of hepatocellular metabolism and bioenergetics. PLOS Pathogens 2010, 6, e1000719. [Google Scholar] [CrossRef] [PubMed]
- Lutz, M.B.; Kukutsch, N.; Ogilvie, A.L.; Rossner, S.; Koch, F.; Romani, N.; Schuler, G. An advanced culture method for generating large quantities of highly pure dendritic cells from mouse bone marrow. J. Immunol. Methods 1999, 223, 77–92. [Google Scholar] [CrossRef] [PubMed]
- Davenport, B.J.; Willis, D.G.; Prescott, J.; Farrell, R.M.; Coons, T.A.; Schountz, T. Generation of competent bone marrow-derived antigen presenting cells from the deer mouse (Peromyscus maniculatus). BMC Immunol. 2004, 5, e23. [Google Scholar] [CrossRef]
- Higgins, D.G.; Sharp, P.M. CLUSTAL: A package for performing multiple sequence alignment on a microcomputer. Gene 1988, 73, 237–244. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schountz, T. Immunology of Bats and Their Viruses: Challenges and Opportunities. Viruses 2014, 6, 4880-4901. https://doi.org/10.3390/v6124880
Schountz T. Immunology of Bats and Their Viruses: Challenges and Opportunities. Viruses. 2014; 6(12):4880-4901. https://doi.org/10.3390/v6124880
Chicago/Turabian StyleSchountz, Tony. 2014. "Immunology of Bats and Their Viruses: Challenges and Opportunities" Viruses 6, no. 12: 4880-4901. https://doi.org/10.3390/v6124880
APA StyleSchountz, T. (2014). Immunology of Bats and Their Viruses: Challenges and Opportunities. Viruses, 6(12), 4880-4901. https://doi.org/10.3390/v6124880




