Host Barriers Limit Viral Spread in a Spillover Host: A Study of Deformed Wing Virus in the Bumblebee Bombus terrestris
Abstract
1. Introduction
2. Materials and Methods
2.1. Source of Bees
2.2. Experimental Procedure
2.3. Preparation of Inoculum
2.4. Oral Inoculation
2.5. Inoculation by Injection
2.6. Sample Collection over Time
2.7. Collection of Haemolymph Samples
2.8. Collection of Faecal Samples
2.9. Viral RNA Extraction
2.10. Detection of Virus by One-Step Real-Time PCR
2.11. Data Analysis
3. Results
3.1. Abundance of DWV-B in Host Faeces Following Oral Inoculation
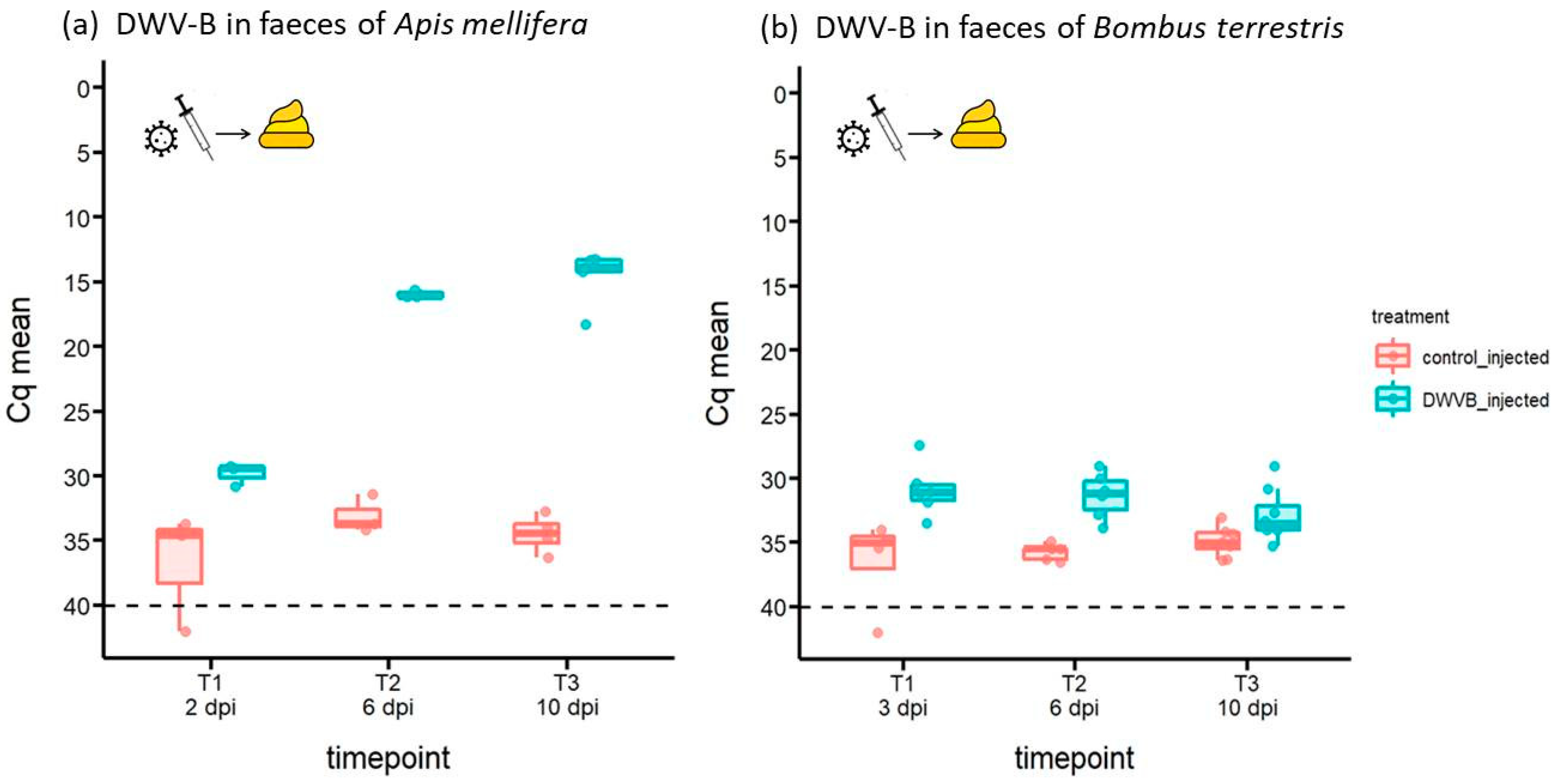
3.2. Abundance of DWV-B in Host Faeces Following Inoculation by Injection
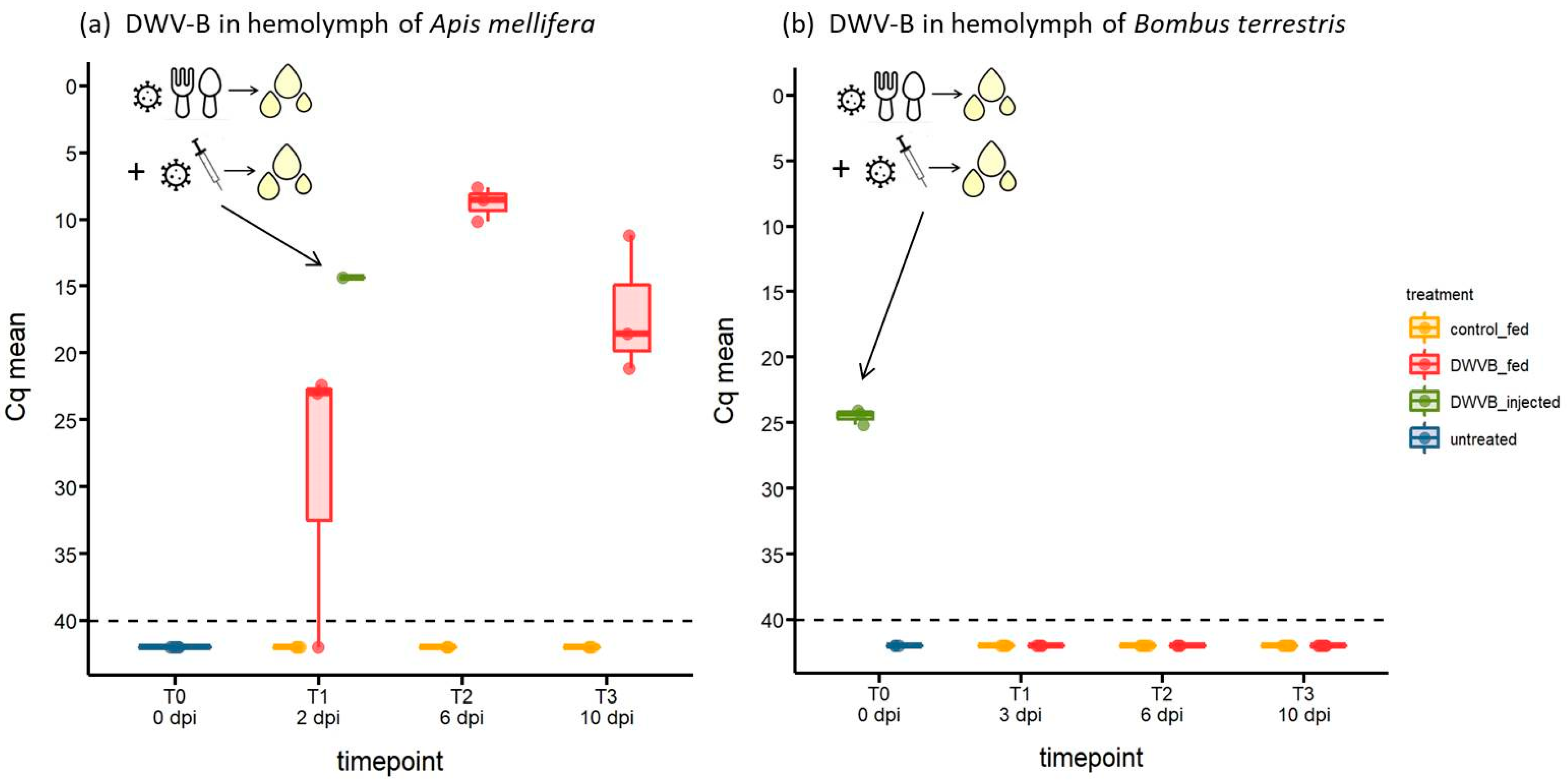
3.3. Abundance of DWV-B in Host Haemolymph Following Oral Inoculation
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Daszak, P.; Cunningham, A.A.; Hyatt, A.D. Emerging infectious diseases of wildlife- threats to biodiversity and human health. Science 2000, 287, 443–449. [Google Scholar] [CrossRef]
- Delwart, E. Animal virus discovery: Improving animal health, understanding zoonoses, and opportunities for vaccine development. Curr. Opin. Virol. 2012, 2, 344–352. [Google Scholar] [CrossRef] [PubMed]
- Holmes, E.C. The ecology of viral emergence. Annu. Rev. Virol. 2022, 9, 173–192. [Google Scholar] [CrossRef]
- Shi, M.; Lin, X.D.; Chen, X.; Tian, J.H.; Chen, L.J.; Li, K.; Wang, W.; Eden, J.S.; Shen, J.J.; Liu, L.; et al. The evolutionary history of vertebrate RNA viruses. Nature 2018, 556, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.E.; Patel, N.G.; Levy, M.A.; Storeygard, A.; Balk, D.; Gittleman, J.L.; Daszak, P. Global trends in emerging infectious diseases. Nature 2008, 451, 990–993. [Google Scholar] [CrossRef]
- Wilfert, L.; Long, G.; Leggett, H.C.; Schmid-Hempel, P.; Butlin, R.; Martin, S.J.M.; Boots, M. Deformed wing virus is a recent global epidemic in honeybees driven by Varroa mites. Science 2016, 351, 594–597. [Google Scholar] [CrossRef]
- Grozinger, C.M.; Flenniken, M.L. Bee viruses: Ecology, pathogenicity, and impacts. Annu. Rev. Entomol. 2019, 64, 205–226. [Google Scholar] [CrossRef] [PubMed]
- Martin, S.J.; Brettell, L.E. Deformed wing virus in honeybees and other insects. Annu. Rev. Virol. 2019, 6, 49–69. [Google Scholar] [CrossRef]
- Alger, S.A.; Burnham, P.A.; Boncristiani, H.F.; Brody, A.K. RNA virus spillover from managed honeybees (Apis Mellifera) to wild bumblebees (Bombus Spp.). PLoS ONE 2019, 14, e0217822. [Google Scholar] [CrossRef]
- Fürst, M.A.; McMahon, D.P.; Osborne, J.L.; Paxton, R.J.; Brown, M.J.F. Disease associations between honeybees and bumblebees as a threat to wild pollinators. Nature 2014, 506, 364–366. [Google Scholar] [CrossRef]
- Manley, R.; Temperton, B.; Doyle, T.; Gates, D.; Hedges, S.; Boots, M.; Wilfert, L. Knock-on community impacts of a novel vector: Spillover of emerging DWV-B from varroa-infested honeybees to wild bumblebees. Ecol. Lett. 2019, 22, 1306–1315. [Google Scholar] [CrossRef] [PubMed]
- McMahon, D.P.; Fürst, M.A.; Caspar, J.; Theodorou, P.; Brown, M.J.F.; Paxton, R.J. A sting in the spit: Widespread cross-infection of multiple RNA viruses across wild and managed bees. J. Anim. Ecol. 2015, 84, 615–624. [Google Scholar] [CrossRef] [PubMed]
- Radzevičiūtė, R.; Theodorou, P.; Husemann, M.; Japoshvili, G.; Kirkitadze, G.; Zhusupbaeva, A.; Paxton, R.J. Replication of honey bee-associated RNA viruses across multiple bee species in apple orchards of Georgia, Germany and Kyrgyzstan. J. Invertebr. Pathol. 2017, 146, 14–23. [Google Scholar] [CrossRef]
- Plowright, R.K.; Parrish, C.R.; McCallum, H.; Hudson, P.J.; Ko, A.I.; Graham, A.L.; Lloyd-Smith, J.O. Pathways to zoonotic spillover. Nat. Rev. Microbiol. 2017, 15, 502–510. [Google Scholar] [CrossRef]
- Becker, D.J.; Washburne, A.D.; Faust, C.L.; Pulliam, J.R.C.; Mordecai, E.A.; Lloyd-Smith, J.O.; Plowright, R.K. Dynamic and integrative approaches to understanding pathogen spillover. Philos. Trans. R. Soc. B Biol. Sci. 2019, 374, 20190014. [Google Scholar] [CrossRef]
- Parrish, C.R.; Holmes, E.C.; Morens, D.M.; Park, E.; Burke, D.S.; Calisher, C.H.; Laughlin, C.A.; Saif, L.J.; Daszak, P. Cross-species virus transmission and the emergence of new epidemic diseases. Microbiol. Mol. Biol. Rev. 2008, 72, 457–470. [Google Scholar] [CrossRef] [PubMed]
- Adler, L.S.; Irwin, R.E.; McArt, S.H.; Vannette, R.L. Floral traits affecting the transmission of beneficial and pathogenic pollinator-associated microbes. Curr. Opin. Insect Sci. 2021, 44, 1–7. [Google Scholar] [CrossRef]
- Burnham, P.A.; Alger, S.A.; Case, B.; Boncristiani, H.; Hébert-Dufresne, L.; Brody, A.K. Flowers as dirty doorknobs: Deformed wing virus transmitted between Apis mellifera and Bombus impatiens through shared flowers. J. Appl. Ecol. 2021, 58, 2065–2074. [Google Scholar] [CrossRef]
- Graystock, P.; Goulson, D.; Hughes, W.O.H. Parasites in bloom: Flowers aid dispersal and transmission of pollinator parasites within and between bee species. Proc. R. Soc. B Biol. Sci. 2015, 282, 20151371. [Google Scholar] [CrossRef]
- Traynor, K.S.; Mondet, F.; De Miranda, J.R.; Techer, M.; Kowallik, V.; Oddie, M.A.Y.; Chantawannakul, P.; Mcafee, A. Varroa destructor: A complex parasite, crippling honey bees worldwide. Trends Parasitol. 2020, 36, 592–606. [Google Scholar] [CrossRef]
- Paxton, R.J.; Schäfer, M.O.; Nazzi, F.; Zanni, V.; Annoscia, D.; Marroni, F.; Bigot, D.; Laws-Quinn, E.R.; Panziera, D.; Jenkins, C.; et al. Epidemiology of a major honey bee pathogen, deformed wing virus: Worldwide replacement of genotype A by genotype B. Int. J. Parasitol. Parasites Wildl. 2022, 18, 157–171. [Google Scholar] [CrossRef]
- You, E.; Moon, K.H.; Kim, H.; Kim, Y.H. Detection of deformed wing virus in the greenhouse for possible horizontal transmission of virus in honey bee colony. Entomol. Res. 2023, 53, 558–565. [Google Scholar] [CrossRef]
- Potts, S.G.; Imperatriz-Fonseca, V.; Ngo, H.T.; Aizen, M.A.; Biesmeijer, J.C.; Breeze, T.D.; Dicks, L.V.; Garibaldi, L.A.; Hill, R.; Settele, J.; et al. Safeguarding pollinators and their values to human well-being. Nature 2016, 540, 220–229. [Google Scholar] [CrossRef]
- Passarelli, A.L. Barriers to success: How Baculoviruses establish efficient systemic infections. Virology 2011, 411, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Caccia, S.; Casartelli, M.; Tettamanti, G. The amazing complexity of insect midgut cells: Types, peculiarities, and functions. Cell Tissue Res. 2019, 377, 505–525. [Google Scholar] [CrossRef] [PubMed]
- Möckel, N.; Gisder, S.; Genersch, E. Horizontal transmission of deformed wing virus: Pathological consequences in adult bees (Apis mellifera) depend on the transmission route. J. Gen. Virol. 2011, 92, 370–377. [Google Scholar] [CrossRef] [PubMed]
- Streicher, T.; Tragust, S.; Paxton, R.J. Experimental viral spillover can harm Bombus terrestris workers under field conditions. Ecol. Entomol. 2022, 48, 81–89. [Google Scholar] [CrossRef]
- Tehel, A.; Streicher, T.; Tragust, S. Experimental infection of bumblebees with honeybee-associated viruses: No direct fitness costs but potential future threats to novel wild bee hosts. R. Soc. Open Sci. 2020, 7, 200480. [Google Scholar] [CrossRef] [PubMed]
- Tehel, A.; Streicher, T.; Tragust, S.; Paxton, R.J. Experimental cross species transmission of a major viral pathogen in bees is predominantly from honeybees to bumblebees. Proc. R. Soc. B Biol. Sci. 2022, 289, 20212255. [Google Scholar] [CrossRef]
- Wasik, B.R.; De Wit, E.; Munster, V.; Lloyd-Smith, J.O.; Martinez-Sobrido, L.; Parrish, C.R. Onward transmission of viruses: How do viruses emerge to cause epidemics after spillover? Philos. Trans. R. Soc. B Biol. Sci. 2019, 374, 20190017. [Google Scholar] [CrossRef]
- Gusachenko, O.N.; Woodford, L.; Cumming, K.B. Evidence for and against deformed wing virus spillover from honey bees to bumble bees: A reverse genetic analysis. Sci. Rep. 2020, 10, 16847. [Google Scholar] [CrossRef]
- Singh, R.; Levitt, A.L.; Rajotte, E.G.; Holmes, E.C.; Ostiguy, N.; Vanengelsdorp, D.; Lipkin, W.I.; Depamphilis, C.W.; Toth, A.L.; Cox-Foster, D.L. RNA viruses in hymenopteran pollinators: Evidence of inter-taxa virus transmission via pollen and potential impact on non-Apis hymenopteran species. PLoS ONE 2010, 5, e14357. [Google Scholar] [CrossRef] [PubMed]
- Ehrenberg, S.; Lewkowski, O.; Erler, S. Dyeing but not dying: Colourful dyes as a non-lethal method of food labelling for in vitro-reared honey bee (Apis mellifera) larvae. J. Insect Physiol. 2019, 113, 1–8. [Google Scholar] [CrossRef]
- McMahon, D.P.; Natsopoulou, M.E.; Doublet, V.; Fürst, M.; Weging, S.; Brown, M.J.F.; Gogol-Döring, A.; Paxton, R.J. Elevated virulence of an emerging viral genotype as a driver of honeybee loss. Proc. R. Soc. B 2016, 283, 20160811. [Google Scholar] [CrossRef]
- Huang, S.; Li, J.; Zhang, Y.; Li, Z.; Evans, J.D.; Rose, R.; Gilligan, T.M.; LeBrun, A.; He, N.; Zheng, T.; et al. A novel method for the detection and diagnosis of virus infections in honey bees. J. Virol. Methods 2021, 293, 114163. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Wang, H.; Meeus, I.; Piot, N.; Smagghe, G. Systemic israeli acute paralysis virus (IAPV) infection in bumblebees (Bombus terrestris) through feeding and injection. J. Invertebr. Pathol. 2018, 151, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Cilia, G.; Zavatta, L.; Ranalli, R.; Nanetti, A.; Bortolotti, L. Replicative deformed wing virus found in the head of adults from symptomatic commercial bumblebee (Bombus terrestris) colonies. Animals 2021, 8, 117. [Google Scholar] [CrossRef] [PubMed]
- Lipa, J.J.; Triggiani, O. Apicystis gen nov and Apicystis bombi (Liu, Macfarlane & Pengelly) comb nov (Protozoa: Neogregarinida), a cosmopolitan parasite of Bombus and Apis (Hymenoptera: Apidae). Apidologie 1996, 27, 29–34. [Google Scholar] [CrossRef]
- Armitage, S.A.; Genersch, E.; McMahon, D.P.; Rafaluk-Mohr, C.; Rolff, J. Tripartite interactions: How immunity, microbiota and pathogens interact and affect pathogen virulence evolution. Curr. Opin. Insect Sci. 2022, 50, 100871. [Google Scholar] [CrossRef]
- Li, J.; Peng, W.; Wu, J.; Strange, J.P.; Boncristiani, H.; Chen, Y. Cross-species infection of deformed wing virus poses a new threat to pollinator conservation. J. Econ. Entomol. 2011, 104, 732–739. [Google Scholar] [CrossRef]
- Fievet, J.; Tentcheva, D.; Gauthier, L.; De Miranda, J.; Cousserans, F.; Colin, M.E.; Bergoin, M. Localization of deformed wing virus infection in queen and drone Apis mellifera L. Virol. J. 2006, 3, 16. [Google Scholar] [CrossRef] [PubMed]
- Gisder, S.; Möckel, N.; Eisenhardt, D.; Genersch, E. In vivo evolution of viral virulence: Switching of deformed wing virus between hosts results in virulence changes and sequence shifts. Environ. Microbiol. 2018, 20, 4612–4628. [Google Scholar] [CrossRef] [PubMed]
- Gusachenko, O.N.; Woodford, L.; Balbirnie-Cumming, K.; Campbell, E.M.; Christie, C.R.; Bowman, A.S.; Evans, D.J. Green bees: Reverse genetic analysis of deformed wing virus transmission, replication, and tropism. Viruses 2020, 12, 532. [Google Scholar] [CrossRef] [PubMed]
- Kuiken, T.; Holmes, E.C.; McCauley, J.; Rimmelzwaan, G.F.; Williams, C.S.; Grenfell, B.T. Host species barriers to influenza virus infections. Science. 2006, 312, 394–397. [Google Scholar] [CrossRef]
- Lunn, T.J.; Restif, O.; Peel, A.J.; Munster, V.J.; De Wit, E.; Sokolow, S.; Van Doremalen, N.; Hudson, P.; McCallum, H. Dose-response and transmission: The nexus between reservoir hosts, environment and recipient hosts. Philos. Trans. R. Soc. B Biol. Sci. 2019, 374, 20190016. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Streicher, T.; Brinker, P.; Tragust, S.; Paxton, R.J. Host Barriers Limit Viral Spread in a Spillover Host: A Study of Deformed Wing Virus in the Bumblebee Bombus terrestris. Viruses 2024, 16, 607. https://doi.org/10.3390/v16040607
Streicher T, Brinker P, Tragust S, Paxton RJ. Host Barriers Limit Viral Spread in a Spillover Host: A Study of Deformed Wing Virus in the Bumblebee Bombus terrestris. Viruses. 2024; 16(4):607. https://doi.org/10.3390/v16040607
Chicago/Turabian StyleStreicher, Tabea, Pina Brinker, Simon Tragust, and Robert J. Paxton. 2024. "Host Barriers Limit Viral Spread in a Spillover Host: A Study of Deformed Wing Virus in the Bumblebee Bombus terrestris" Viruses 16, no. 4: 607. https://doi.org/10.3390/v16040607
APA StyleStreicher, T., Brinker, P., Tragust, S., & Paxton, R. J. (2024). Host Barriers Limit Viral Spread in a Spillover Host: A Study of Deformed Wing Virus in the Bumblebee Bombus terrestris. Viruses, 16(4), 607. https://doi.org/10.3390/v16040607






