Nano-Biomechanical Investigation of Phosphatidylserine-Mediated Ebola Viral Attachment via Human Gas6 and Axl
Abstract
1. Introduction
2. Materials and Methods
2.1. Protein Constructs and Ebola Viral-Like Particles
2.2. AFM Chip Functionalization, Substrate Preparation, and Biomolecule Immobilization
2.3. Single Molecule Force Spectroscopy
2.4. Statistical Analysis of Single Molecule Force Measurement Results
2.5. Microscale Thermophoresis Assay
3. Results
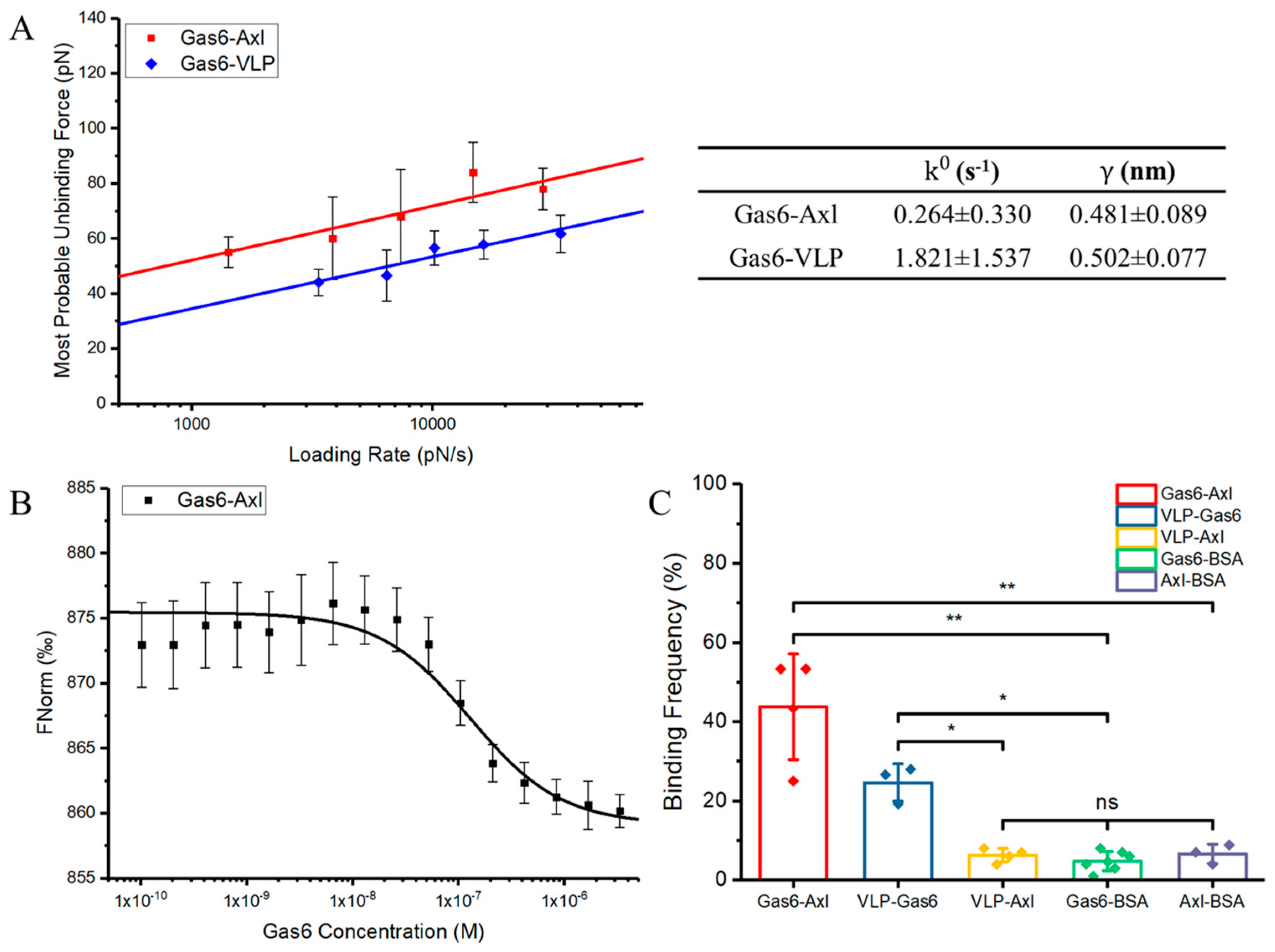
3.1. Gas6 Binds to Axl with Higher Mechanical Strength and Binding Affinities Compared to PS on Ebola Viral-Like Particles
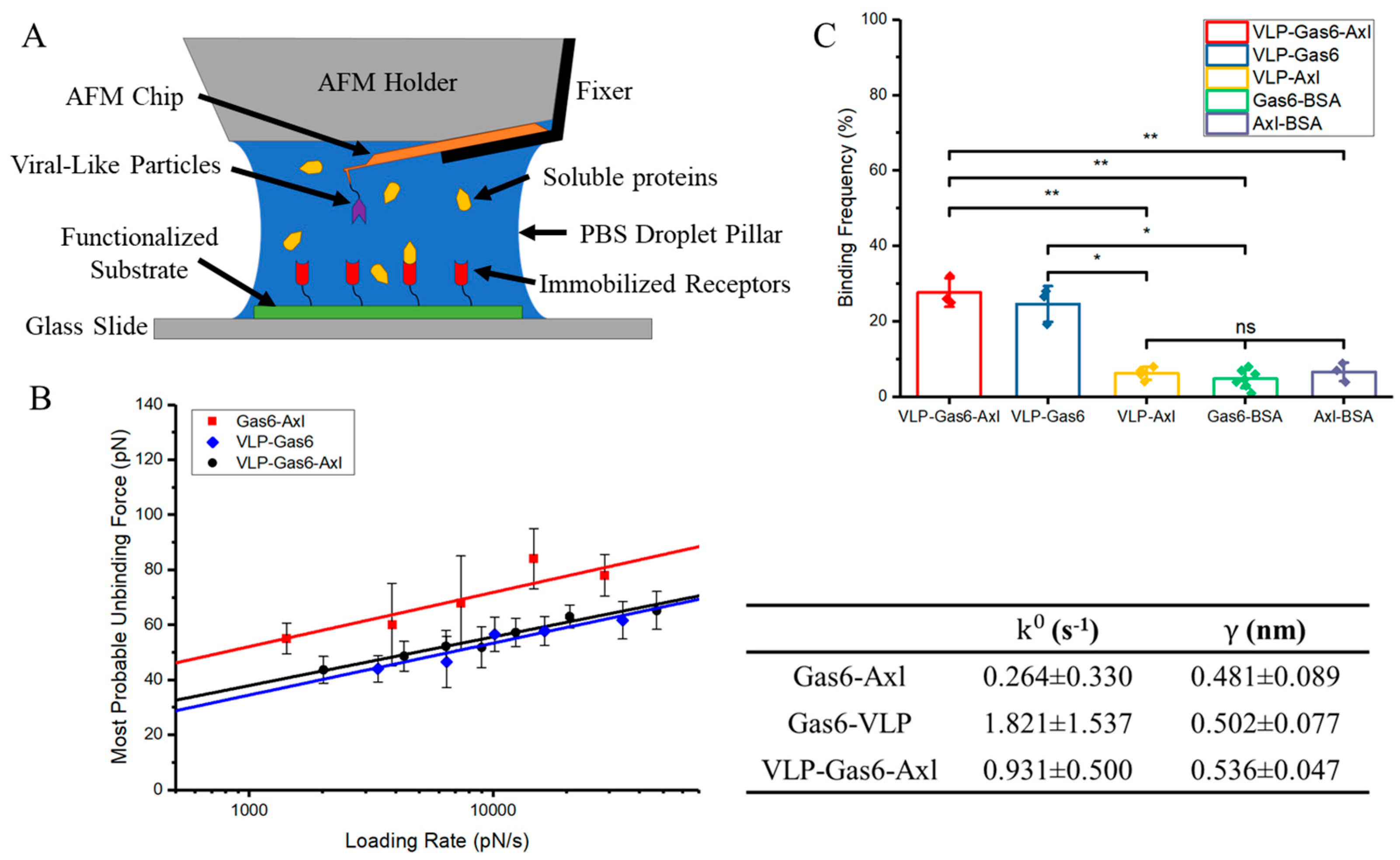
3.2. Characterization of Mechanical Strength and Binding Affinity Among Ebola VLP, Soluble Gas6, and Axl
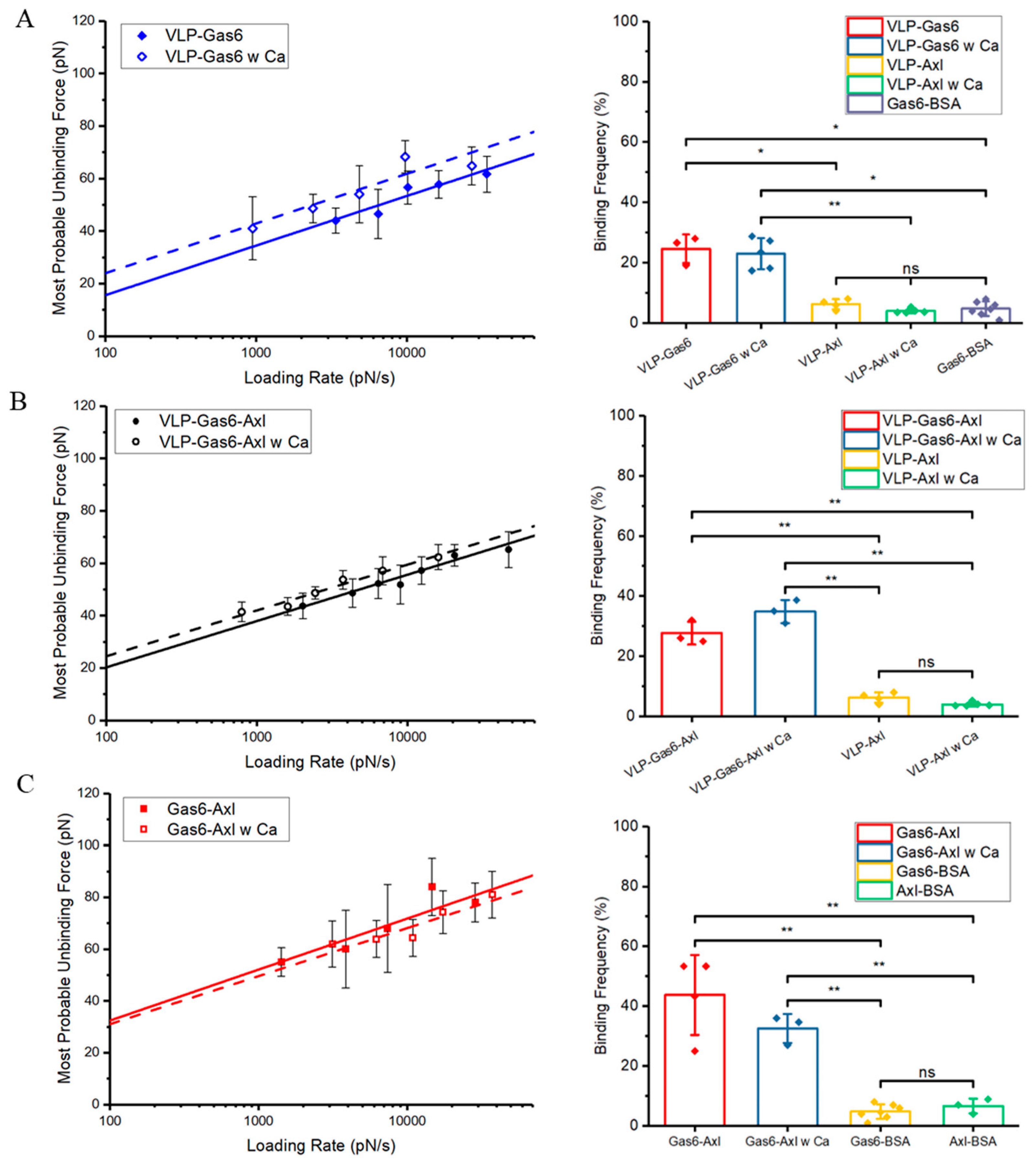
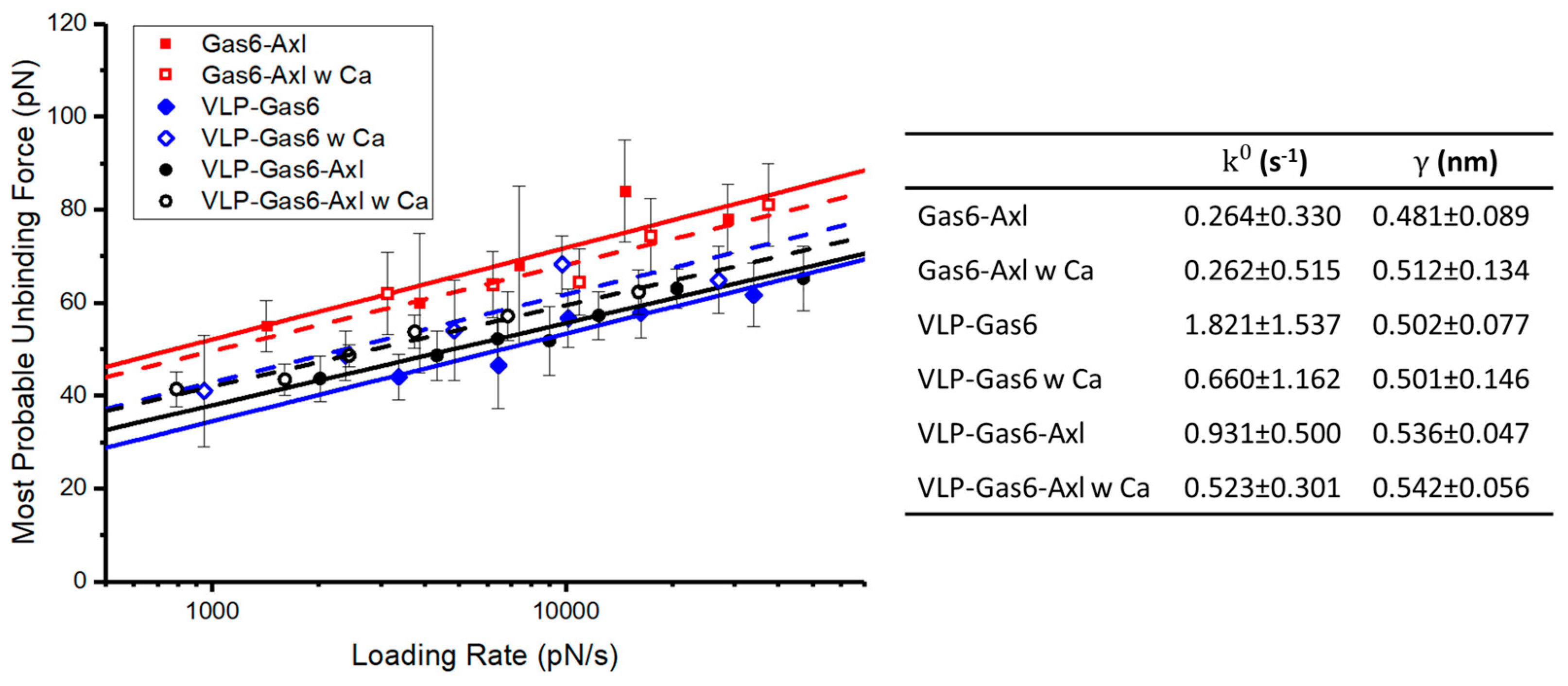
3.3. Calcium Ions Significantly Strengthen the Mechanical Strength and Binding Affinity of PS-Gas6 Bound Complex
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- WHO. Ebola Virus Disease. 2023. Available online: https://www.who.int/news-room/fact-sheets/detail/ebola-virus-disease (accessed on 29 February 2024).
- Hunt, C.L.; Lennemann, N.J.; Maury, W. Filovirus entry: A novelty in the viral fusion world. Viruses 2012, 4, 258–275. [Google Scholar] [CrossRef] [PubMed]
- Meyers, L.; Frawley, T.; Goss, S.; Kang, C. Ebola virus outbreak 2014: Clinical review for emergency physicians. Ann. Emerg. Med. 2015, 65, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Biedenkopf, N.; Bukreyev, A.; Chandran, K.; Di Paola, N.; Formenty, P.B.; Griffiths, A.; Hume, A.J.; Mühlberger, E.; Netesov, S.V.; Palacios, G. Renaming of genera Ebolavirus and Marburgvirus to Orthoebolavirus and Orthomarburgvirus, respectively, and introduction of binomial species names within family Filoviridae. Arch. Virol. 2023, 168, 220. [Google Scholar] [CrossRef] [PubMed]
- CDC. Ebola Disease Outbreak Locations. 2024. Available online: https://www.cdc.gov/ebola/outbreaks/index.html (accessed on 15 August 2024).
- National Center for Infectious Diseases (U.S.). Ebola Hemorrhagic Fever: Known Cases and Outbreaks of Ebola Hemorrhagic Fever, in Chronological Order [last updated 12 October 2011]. 2011. Available online: https://stacks.cdc.gov/view/cdc/22147 (accessed on 29 February 2024).
- Regules, J.A.; Beigel, J.H.; Paolino, K.M.; Voell, J.; Castellano, A.R.; Hu, Z.; Muñoz, P.; Moon, J.E.; Ruck, R.C.; Bennett, J.W. A recombinant vesicular stomatitis virus Ebola vaccine. N. Engl. J. Med. 2017, 376, 330–341. [Google Scholar] [CrossRef]
- Feldmann, H.; Sprecher, A.; Geisbert, T.W. Ebola. N. Engl. J. Med. 2020, 382, 1832–1842. [Google Scholar] [CrossRef]
- Feldmann, H. Ebola—A growing threat? N. Engl. J. Med. 2014, 371, 1375–1378. [Google Scholar] [CrossRef]
- Takada, A. Filovirus tropism: Cellular molecules for viral entry. Front. Microbiol. 2012, 3, 34. [Google Scholar] [CrossRef]
- De Clercq, E. Ebola virus (EBOV) infection: Therapeutic strategies. Biochem. Pharmacol. 2015, 93, 1–10. [Google Scholar] [CrossRef]
- Nanbo, A.; Imai, M.; Watanabe, S.; Noda, T.; Takahashi, K.; Neumann, G.; Halfmann, P.; Kawaoka, Y. Ebolavirus is internalized into host cells via macropinocytosis in a viral glycoprotein-dependent manner. PLoS Pathog. 2010, 6, e1001121. [Google Scholar] [CrossRef]
- Noda, T.; Ebihara, H.; Muramoto, Y.; Fujii, K.; Takada, A.; Sagara, H.; Kim, J.H.; Kida, H.; Feldmann, H.; Kawaoka, Y. Assembly and budding of Ebolavirus. PLoS Pathog. 2006, 2, e99. [Google Scholar] [CrossRef]
- Bharat, T.A.M.; Riches, J.D.; Kolesnikova, L.; Welsch, S.; Krähling, V.; Davey, N.; Parsy, M.-L.; Becker, S.; Briggs, J.A.G. Cryo-electron tomography of Marburg virus particles and their morphogenesis within infected cells. PLoS Biol. 2011, 9, e1001196. [Google Scholar] [CrossRef] [PubMed]
- Geisbert, T.W.; Hensley, L.E. Ebola virus: New insights into disease aetiopathology and possible therapeutic interventions. Expert Rev. Mol. Med. 2004, 6, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Bohan, D.; Van Ert, H.; Ruggio, N.; Rogers, K.J.; Badreddine, M.; Aguilar Briseno, J.A.; Elliff, J.M.; Rojas Chavez, R.A.; Gao, B.; Stokowy, T.; et al. Phosphatidylserine receptors enhance SARS-CoV-2 infection. PLoS Pathog. 2021, 17, e1009743. [Google Scholar] [CrossRef] [PubMed]
- Ghosh Roy, S. TAM receptors: A phosphatidylserine receptor family and its implications in viral infections. Int. Rev. Cell Mol. Biol. 2020, 357, 81–122. [Google Scholar] [CrossRef] [PubMed]
- Kirui, J.; Abidine, Y.; Lenman, A.; Islam, K.; Gwon, Y.D.; Lasswitz, L.; Evander, M.; Bally, M.; Gerold, G. The Phosphatidylserine Receptor TIM-1 Enhances Authentic Chikungunya Virus Cell Entry. Cells 2021, 10, 1828. [Google Scholar] [CrossRef]
- Brunton, B.; Rogers, K.; Phillips, E.K.; Brouillette, R.B.; Bouls, R.; Butler, N.S.; Maury, W. TIM-1 serves as a receptor for Ebola virus in vivo, enhancing viremia and pathogenesis. PLoS Neglected Trop. Dis. 2019, 13, e0006983. [Google Scholar] [CrossRef]
- Younan, P.; Iampietro, M.; Nishida, A.; Ramanathan, P.; Santos, R.I.; Dutta, M.; Lubaki, N.M.; Koup, R.A.; Katze, M.G.; Bukreyev, A. Ebola virus binding to Tim-1 on T lymphocytes induces a cytokine storm. MBio. 2017, 8, e00845-17. [Google Scholar] [CrossRef]
- Morizono, K.; Chen, I.S. Role of phosphatidylserine receptors in enveloped virus infection. J. Virol. 2014, 88, 4275–4290. [Google Scholar] [CrossRef]
- Carnec, X.; Meertens, L.; Dejarnac, O.; Perera-Lecoin, M.; Hafirassou, M.L.; Kitaura, J.; Ramdasi, R.; Schwartz, O.; Amara, A. The phosphatidylserine and phosphatidylethanolamine receptor CD300a binds dengue virus and enhances infection. J. Virol. 2016, 90, 92–102. [Google Scholar] [CrossRef]
- Linger, R.M.; Keating, A.K.; Earp, H.S.; Graham, D.K. Taking aim at Mer and Axl receptor tyrosine kinases as novel therapeutic targets in solid tumors. Expert Opin. Ther. Targets 2010, 14, 1073–1090. [Google Scholar] [CrossRef]
- Moller-Tank, S.; Maury, W. Phosphatidylserine receptors: Enhancers of enveloped virus entry and infection. Virology 2014, 468, 565–580. [Google Scholar] [CrossRef] [PubMed]
- Linger, R.M.; Keating, A.K.; Earp, H.S.; Graham, D.K. TAM receptor tyrosine kinases: Biologic functions, signaling, and potential therapeutic targeting in human cancer. Adv. Cancer Res. 2008, 100, 35–83. [Google Scholar] [PubMed]
- Mark, M.R.; Chen, J.; Hammonds, R.G.; Sadick, M.; Godowsk, P.J. Characterization of Gas6, a member of the superfamily of G domain-containing proteins, as a ligand for Rse and Axl. J. Biol. Chem. 1996, 271, 9785–9789. [Google Scholar] [CrossRef] [PubMed]
- Tie, J.K.; Stafford, D.W. Structural and functional insights into enzymes of the vitamin K cycle. J. Thromb. Haemost. 2016, 14, 236–247. [Google Scholar] [CrossRef] [PubMed]
- Brindley, M.A.; Hunt, C.L.; Kondratowicz, A.S.; Bowman, J.; Sinn, P.L.; McCray Jr, P.B.; Quinn, K.; Weller, M.L.; Chiorini, J.A.; Maury, W. Tyrosine kinase receptor Axl enhances entry of Zaire ebolavirus without direct interactions with the viral glycoprotein. Virology 2011, 415, 83–94. [Google Scholar] [CrossRef]
- Miller, E.H.; Chandran, K. Filovirus entry into cells–new insights. Curr. Opin. Virol. 2012, 2, 206–214. [Google Scholar] [CrossRef]
- Iida, M.; McDaniel, N.K.; Kostecki, K.L.; Welke, N.B.; Kranjac, C.A.; Liu, P.; Longhurst, C.; Bruce, J.Y.; Hong, S.; Salgia, R. AXL regulates neuregulin1 expression leading to cetuximab resistance in head and neck cancer. BMC Cancer 2022, 22, 447. [Google Scholar] [CrossRef]
- Puddu, A.; Ravera, S.; Panfoli, I.; Bertola, N.; Maggi, D. High Glucose Impairs Expression and Activation of MerTK in ARPE-19 Cells. Int. J. Mol. Sci. 2022, 23, 1144. [Google Scholar] [CrossRef]
- Sun, L.-W.; Kao, S.-H.; Yang, S.-F.; Jhang, S.-W.; Lin, Y.-C.; Chen, C.-M.; Hsieh, Y.-H. Corosolic acid attenuates the invasiveness of glioblastoma cells by promoting CHIP-mediated AXL degradation and inhibiting GAS6/AXL/JAK Axis. Cells 2021, 10, 2919. [Google Scholar] [CrossRef]
- Yeh, L.H.P.; Bajpai, R.K.; Sun, G.Y. Membrane lipid metabolism and phospholipase activity in insect Spodoptera frugiperda 9 ovarian cells. Lipids 1997, 32, 481–487. [Google Scholar] [CrossRef]
- Ilinykh, P.A.; Santos, R.I.; Gunn, B.M.; Kuzmina, N.A.; Shen, X.; Huang, K.; Gilchuk, P.; Flyak, A.I.; Younan, P.; Alter, G. Asymmetric antiviral effects of ebolavirus antibodies targeting glycoprotein stem and glycan cap. PLoS Pathog. 2018, 14, e1007204. [Google Scholar] [CrossRef] [PubMed]
- Riener, C.K.; Stroh, C.M.; Ebner, A.; Klampfl, C.; Gall, A.A.; Romanin, C.; Lyubchenko, Y.L.; Hinterdorfer, P.; Gruber, H.J. Simple test system for single molecule recognition force microscopy. Anal. Chim. Acta 2003, 479, 59–75. [Google Scholar] [CrossRef]
- Rankl, C.; Kienberger, F.; Wildling, L.; Wruss, J.; Gruber, H.J.; Blaas, D.; Hinterdorfer, P. Multiple receptors involved in human rhinovirus attachment to live cells. Proc. Natl. Acad. Sci. USA 2008, 105, 17778–17783. [Google Scholar] [CrossRef] [PubMed]
- Ebner, A.; Wildling, L.; Zhu, R.; Rankl, C.; Haselgrübler, T.; Hinterdorfer, P.; Gruber, H.J. Functionalization of probe tips and supports for single-molecule recognition force microscopy. In STM and AFM Studies on (Bio)Molecular Systems: Unravelling the Nanoworld; Springer: Berlin/Heidelberg, Germany, 2008; pp. 29–76. [Google Scholar]
- Ebner, A.; Hinterdorfer, P.; Gruber, H.J. Comparison of different aminofunctionalization strategies for attachment of single antibodies to AFM cantilevers. Ultramicroscopy 2007, 107, 922–927. [Google Scholar] [CrossRef] [PubMed]
- Cao, W.; Dong, C.; Kim, S.; Hou, D.; Tai, W.; Du, L.; Im, W.; Zhang, X.F. Biomechanical characterization of SARS-CoV-2 spike RBD and human ACE2 protein-protein interaction. Biophys. J. 2021, 120, 1011–1019. [Google Scholar] [CrossRef]
- Chesla, S.E.; Selvaraj, P.; Zhu, C. Measuring two-dimensional receptor-ligand binding kinetics by micropipette. Biophys. J. 1998, 75, 1553–1572. [Google Scholar] [CrossRef]
- Evans, E. Probing the relation between force—Lifetime—And chemistry in single molecular bonds. Annu. Rev. Biophys. Biomol. Struct. 2001, 30, 105–128. [Google Scholar] [CrossRef]
- Levy, R.; Maaloum, M. Measuring the spring constant of atomic force microscope cantilevers: Thermal fluctuations and other methods. Nanotechnology 2001, 13, 33. [Google Scholar] [CrossRef]
- Franz, C.M.; Taubenberger, A.; Puech, P.-H.; Muller, D.J. Studying integrin-mediated cell adhesion at the single-molecule level using AFM force spectroscopy. Sci. STKE 2007, 2007, pl5. [Google Scholar] [CrossRef]
- Kim, S.; Liu, Y.; Ziarnik, M.; Seo, S.; Cao, Y.; Zhang, X.F.; Im, W. Binding of human ACE2 and RBD of omicron enhanced by unique interaction patterns among SARS-CoV-2 variants of concern. J. Comput. Chem. 2023, 44, 594–601. [Google Scholar] [CrossRef]
- Hou, D.; Cao, W.; Kim, S.; Cui, X.; Ziarnik, M.; Im, W.; Zhang, X.F. Biophysical investigation of interactions between SARS-CoV-2 spike protein and neuropilin-1. Protein Sci. 2023, 32, e4773. [Google Scholar] [CrossRef] [PubMed]
- Nydegger, D.T.; Pujol-Giménez, J.; Kandasamy, P.; Vogt, B.; Hediger, M.A. Applications of the Microscale Thermophoresis Binding Assay in COVID-19 Research. Viruses 2023, 15, 1432. [Google Scholar] [CrossRef] [PubMed]
- Magnez, R.; Bailly, C.; Thuru, X. Microscale thermophoresis as a tool to study protein interactions and their implication in human diseases. Int. J. Mol. Sci. 2022, 23, 7672. [Google Scholar] [CrossRef] [PubMed]
- Vann, K.R.; Acharya, A.; Jang, S.M.; Lachance, C.; Zandian, M.; Holt, T.A.; Smith, A.L.; Pandey, K.; Durden, D.L.; El-Gamal, D. Binding of the SARS-CoV-2 envelope E protein to human BRD4 is essential for infection. Structure 2022, 30, 1224–1232.e1225. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Wang, S.; Zong, Z.; Pan, T.; Liu, S.; Mao, W.; Huang, H.; Yan, X.; Yang, B.; He, X. TRIM28-mediated nucleocapsid protein SUMOylation enhances SARS-CoV-2 virulence. Nat. Commun. 2024, 15, 244. [Google Scholar] [CrossRef]
- Griffiths, C.D.; Bilawchuk, L.M.; McDonough, J.E.; Jamieson, K.C.; Elawar, F.; Cen, Y.; Duan, W.; Lin, C.; Song, H.; Casanova, J.-L. IGF1R is an entry receptor for respiratory syncytial virus. Nature 2020, 583, 615–619. [Google Scholar] [CrossRef]
- Lemke, G. Phosphatidylserine is the signal for TAM receptors and their ligands. Trends Biochem. Sci. 2017, 42, 738–748. [Google Scholar] [CrossRef]
- Dransfield, I.; Zagórska, A.; Lew, E.; Michail, K.; Lemke, G. Mer receptor tyrosine kinase mediates both tethering and phagocytosis of apoptotic cells. Cell Death Dis. 2015, 6, e1646. [Google Scholar] [CrossRef]
- Huang, M.; Rigby, A.C.; Morelli, X.; Grant, M.A.; Huang, G.; Furie, B.; Seaton, B.; Furie, B.C. Structural basis of membrane binding by Gla domains of vitamin K–dependent proteins. Nat. Struct. Mol. Biol. 2003, 10, 751–756. [Google Scholar] [CrossRef]
- Bhattacharyya, S.; Zagórska, A.; Lew, E.D.; Shrestha, B.; Rothlin, C.V.; Naughton, J.; Diamond, M.S.; Lemke, G.; Young, J.A. Enveloped viruses disable innate immune responses in dendritic cells by direct activation of TAM receptors. Cell Host Microbe 2013, 14, 136–147. [Google Scholar] [CrossRef]
- Lew, E.D.; Oh, J.; Burrola, P.G.; Lax, I.; Zagórska, A.; Través, P.G.; Schlessinger, J.; Lemke, G. Differential TAM receptor–ligand–phospholipid interactions delimit differential TAM bioactivities. eLife 2014, 3, e03385. [Google Scholar] [CrossRef] [PubMed]
- Morizono, K.; Xie, Y.; Olafsen, T.; Lee, B.; Dasgupta, A.; Wu, A.M.; Chen, I.S. The soluble serum protein Gas6 bridges virion envelope phosphatidylserine to the TAM receptor tyrosine kinase Axl to mediate viral entry. Cell Host Microbe 2011, 9, 286–298. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Richard, A.S.; Jackson, C.B.; Ojha, A.; Choe, H. Phosphatidylethanolamine and phosphatidylserine synergize to enhance GAS6/AXL-mediated virus infection and efferocytosis. J. Virol. 2020, 95, e02079-20. [Google Scholar] [CrossRef] [PubMed]
- Hunt, C.L.; Kolokoltsov, A.A.; Davey, R.A.; Maury, W. The Tyro3 receptor kinase Axl enhances macropinocytosis of Zaire ebolavirus. J. Virol. 2011, 85, 334–347. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hou, D.; Mu, Q.; Chen, W.; Cao, W.; Zhang, X.F. Nano-Biomechanical Investigation of Phosphatidylserine-Mediated Ebola Viral Attachment via Human Gas6 and Axl. Viruses 2024, 16, 1700. https://doi.org/10.3390/v16111700
Hou D, Mu Q, Chen W, Cao W, Zhang XF. Nano-Biomechanical Investigation of Phosphatidylserine-Mediated Ebola Viral Attachment via Human Gas6 and Axl. Viruses. 2024; 16(11):1700. https://doi.org/10.3390/v16111700
Chicago/Turabian StyleHou, Decheng, Qian Mu, Weixuan Chen, Wenpeng Cao, and Xiaohui Frank Zhang. 2024. "Nano-Biomechanical Investigation of Phosphatidylserine-Mediated Ebola Viral Attachment via Human Gas6 and Axl" Viruses 16, no. 11: 1700. https://doi.org/10.3390/v16111700
APA StyleHou, D., Mu, Q., Chen, W., Cao, W., & Zhang, X. F. (2024). Nano-Biomechanical Investigation of Phosphatidylserine-Mediated Ebola Viral Attachment via Human Gas6 and Axl. Viruses, 16(11), 1700. https://doi.org/10.3390/v16111700






