1. Introduction
Influenza is an acute respiratory infectious disease caused by influenza virus infection. Influenza viruses belong to the family Orthomyxoviridae; there are four main types (influenza A, influenza B, influenza C, and influenza D) [
1]. Influenza A and B viruses, which are the most common causes of seasonal and pandemic influenza in humans, have always been a major threat to human health [
2]. IAV harbors a genome comprising eight single-stranded negative-sense RNA segments that encode at least 14 proteins [
3]. Based on the properties of the hemagglutinin (HA) and neuraminidase (NA) surface proteins, IAVs can be divided into 18 different HA subtypes (H1–H18) and 11 different NA subtypes (N1–N11) [
4].
The smallest nonstructural protein (NS) segment encodes two kinds of viral proteins (NS1 and NS2/NEP). The length of NS1 protein varies from 215–237 amino acids, depending on the strain of virus [
5]. NS1 can be divided into two functional domains: the N-terminal RNA binding domain (RBD) and the C-terminal effect domain (ED), which interact with a variety of host factors. The RBD rotates to form a six-helix homodimer that binds to double-stranded RNA (dsRNA). The ED of most NS1 proteins contains nuclear output signals [
6]. Therefore, the NS1 protein plays important roles not only in the cytoplasm but also in the nucleus [
5]. NS1 is important for multiple viral functions, including temporal adjustment of viral RNA synthesis, control of viral mRNA splicing, adjustment of virus particle morphogenesis, and suppression of host apoptotic responses through activation of phosphoinositide 3-kinase (PI3K) [
7]. The NS1 protein plays a crucial role in regulating host antiviral immune responses through various mechanisms. For example, NS1 increases virulence and/or pathogenicity in the host by acting as an interferon (IFN) antagonist, thereby enabling the virus to escape from interferon-mediated responses and replicate efficiently [
8]. The RBD binds to both single-stranded RNA (ssRNA) and dsRNA, thereby blocking their recognition by retinoic acid-induced gene RIG-I and inhibiting expression of antiviral cytokines interferon-α/β [
9]. NS1 also inhibits the 3′ terminal processing of host genes by binding to CPSF30 to block processing of cellular mRNAs [
10].
However, the NS1 protein of the 2009 pandemic H1N1 IAV (pH1N1) virus lost the ability to bind CPSF30, so it cannot block expression of host genes, although studies show that this ability can be restored by introducing amino acid changes R108K, E125D, and G189D [
11,
12,
13]. Interestingly, recombinant pH1N1 with these alternative expression NS1 can inhibit expression of host genes and is more effective than the 2009 pandemic H1N1 IAV (pH1N1) in combating host innate immune responses [
14]. However, in vivo studies show that the NS1 mutant pH1N1 virus grows to a lower titer in the lungs of mice and has slightly lower pathogenicity than the WT virus [
12,
15]. There are vaccines against influenza viruses; however, these viruses mutate constantly, leading to increased virulence via mutation of viral proteins such as NS1. Since 2009, multiple amino acid sense mutations in the NS1 protein of pandemic H1N1 influenza virus have been reported, including the I123V and N205S mutations [
12].
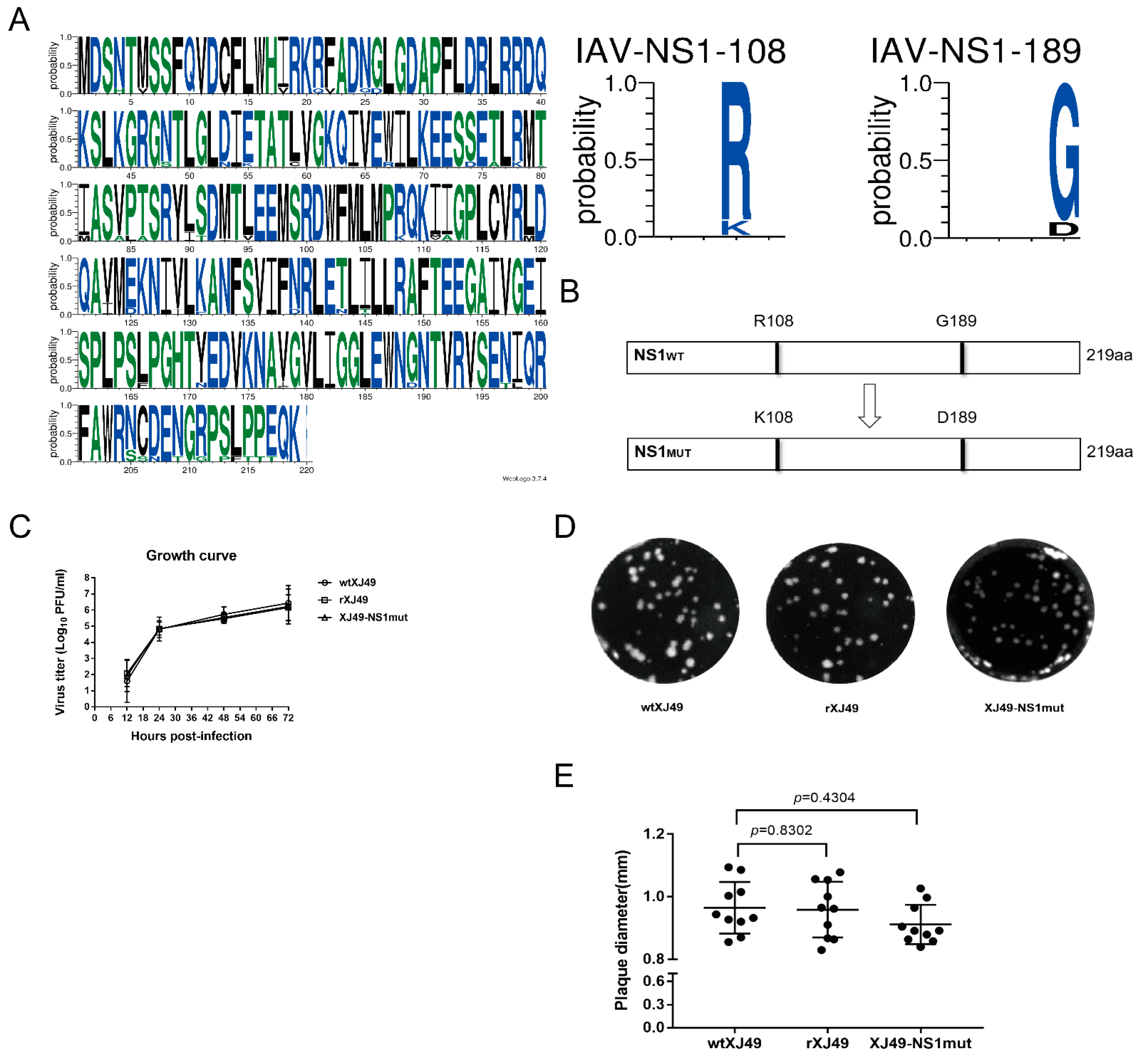
Here, we sequenced a seasonal 2009 H1N1 influenza virus isolated in 2018 and confirmed the presence of mutations I123V and N205S in important functional sites. However, 108R and 189G were conserved when compared with the early pandemic strain A/California/07/2009. To increase our understanding of the antagonistic mechanisms underlying interactions between influenza virus and host innate immunity, we used a reverse genetics approach to introduce a double mutation (R108K/G189D) into the NS1 protein of the seasonal 2009 H1N1 IAV, rescued the mutant virus, and analyzed its biological characteristics. We found that the mutated NS1 protein harboring R108K/G189D exhibited systematic and selective inhibition of cytokine responses beyond the complexity of host–influenza NS1 protein interactions.
2. Materials and Methods
2.1. Cell Culture and Virus
MDCK (Madin–Darby canine kidney) cells (ATCC, CCL-34), HEK293T (Human embryonic kidney) cells (ATCC, CRL-3216), and A549 (Human lung epithelial carcinoma) cells (ATCC, CCL-185) were cultured at 37 °C/5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM, ThermoFisher, New York, NY, USA; 11965092) supplemented with 10% heat-inactivated fetal bovine serum (Gibco) and 1% PSG (100 U/mL penicillin, 100 μg/mL streptomycin). IAV A/Urumqi/XJ49/2018 (H1N1) (XJ49) was isolated from throat swabs obtained from patients in Urumqi in 2018 (GenBank Accession no.: MN549338–MN549345).
2.2. Sequence Analysis
According to their lineage/subtype, as well as the year, 2155 seasonally circulating NS1 sequences were downloaded from the influenza virus database of National Center for Biotechnology Information randomly (NCBI,
https://www.ncbi.nlm.nih.gov/FLU, accessed on 10 November 2020). We used Molecular Evolutionary Genetic Analysis (MEGA) software v6.0 and Weblogo (
http://weblogo.threeplusone.com/, accessed on 15 March 2020) for sequence alignment and mutation site analysis.
2.3. Generation of Recombinant Viruses by Reverse Genetics
The pHW2000 plasmid was kindly provided by Professor Robert G. Webster (Department of Pathology, University of Tennessee). The rXJ49 H1N1 virus was generated using an eight-plasmid system based on pHW2000 by transfection into human HEK293T cells. To obtain pHW2000 plasmids containing eight segments of IAV XJ49, viral RNA was extracted from infected cells and cDNA synthesized by reverse transcription using Uni12 primers [
16]. The cDNA encoding the XJ49 viral genome was amplified using segment-specific primers and then cloned into the pHW2000 using BsmB I restriction enzymes, as described previously [
16]. The R108K and G189D mutations were inserted into the NS1 gene using the Q5 site-directed mutagenesis kit (NEB; E0554S) and the following primers: NS1-R108K_F: 5’-CTTATGCCTAaGCAAAAGATAATAG-3’; NS1-R108K_R: 5’-CATGAACCAGTCTCGTGAC-3’; NS1-G189D_F: 5’-AAGTGGAATGaTAACACGGTTC-3’; and NS1-G189D_R: 5’-AAGTCCTCCGATGAGGAC-3’). The plasmid encoding the NS1 protein of XJ49 was substituted with the mutated NS1 plasmid to obtain the rXJ49-NS1mut virus. XJ49 and rXJ49-NS1mut were passaged in MDCK cells. Correct insertion of the viral segments into pHW2000 and presence of the R108K and G189D mutations were confirmed by sequencing.
2.4. Virus Growth Kinetics
To access virus growth kinetics in vitro, triplicate wells containing confluent monolayers of MDCK cells (5 × 105 cells/well, 6-well plates) were infected with viruses at a MOI of 0.01. After 1 h of virus adsorption at 37 °C, cells were overlaid with DMEM containing TPCK-treated trypsin (2 μg/mL) and then incubated at 37 °C. At 0, 12, 24, 48, 72, and 96 h post-infection (hpi), 500 μL cell supernatant was collected from each well, and the viral titer was determined in a plaque assay.
2.5. Plaque Assay
Confluent monolayers of MDCK cells (105 cells/well in a 12-well plate) were infected with virus at 37 °C. After 1 h of virus adsorption, cells were overlaid with DMEM containing 1% Low Melting Point agarose and TPCK-treated trypsin (2 μg/mL) and then incubated at 37 °C for 3 days. At 3 dpi, cells were fixed for 15 min at room temperature with 4% paraformaldehyde. The agarose overlay was removed and stained for 10 min with 1% crystal violet in methanol. The number of plaques in each well was counted.
2.6. Western Blot Analysis
Samples of transfected cells were prepared in buffer containing 100 mM Tris-HCl (pH 6.8), 4% SDS, 20% glycerol, 0.2% bromophenol blue, and 20% mercaptoethanol. The samples were heated for 15 min at 100 °C and loaded onto an SDS–PAGE gel. Proteins were transferred to a PVDF membrane, blocked with PBST buffer containing 5% bovine serum albumin (BSA)/0.1% Tween-20, and then probed at room temperature for 2 h with anti-NS1 polyclonal antibody (Gene Tex; GTX125990). An anti-GAPDH monoclonal antibody (Abcam; ab8245) was used as a loading control. A horseradish peroxidase (HRP) secondary antibody (ZSJQ-BIO; ZB2305/ZB2301) was used to detect primary antibody binding. Protein bands were visualized using SuperSignal West Femto’s highest sensitivity chemiluminescence substrate kit (Thermo Science, New York, NY, USA; 34096).
2.7. Pathogenicity Study in Mice
Five-week-old female C57BL/6N mice were purchased from Charles River (Beijing, China). All animal experiments were approved by the Ethical Committee of Beijing Institute of Microbiology and Epidemiology. To determine the MLD50 of influenza virus, four groups of mice (n = 5/group) were anesthetized with pentobarbital sodium (240 mg/kg body weight). One group was used as a negative control group (PBS-infected), another was used as a positive control group (CA07), and the remaining groups were infected with virus. Of these, one group was infected with XJ49-NS1mut virus (XJ49-NS1mut), and one group was infected with rXJ49 virus (rXJ49). The MLD50 of each virus was calculated using the method of Reed and Muench. Survival and body weight of each individual mouse was monitored daily for 15 days. Mice were sacrificed and scored dead when weight loss reached 25% of the baseline weight (i.e., the weight on the day of infection). Mice were sacrificed at 1, 3, and 6 dpi, and lungs were extracted and homogenized prior to measurement of virus titer. Virus titers were determined in a plaque assay in MDCK cells. Lungs were collected at 6 dpi and fixed with 4% neutral phosphate-buffered formalin. Fixed specimens were embedded in paraffin, sectioned, and stained with hematoxylin and eosin (H&E). For immunohistochemical staining, sections were probed with antibodies specific for the A/H1N1 HA protein or mouse IFN-β. Tissue sections were examined under a light microscope (PANNORAMIC DESK/MIDI/250/1000; 3DHISTECH (Budapest, Hungary)). Cytokine and chemokine expression in lung homogenates was measured using a ProcartaPlex Multiplex Immunoassay kit (Thermo Science, New York, NY, USA; MAN0016936).
2.8. Reporter System Assay
The 293T cells cultured in 24-well plates were transiently co-transfected with the reporter plasmid p125-luc encoding FF-Luc under the control of the IFN promoter, a plasmid pRL constitutively expressing Rluc under the control of the HSV–TK promoter, and the WT and mutant NS1 pHW2000-expressing plasmids (pHW2000-XJ49NS or pHW2000-NS1mut), using Lipofectamine 3000 reagent (ThermoFisher, New York, NY, USA; L3000-3000). The empty pHW2000 plasmid was used as a negative control, while poly (IC) was used as a positive control. At 24 h post-transfection, cells were harvested and lysed. Reporter activity was analyzed using a Dual-Luciferase Reporter Assay System kit (Promega, Beijing, China; E1910). All assays were performed independently and in triplicate.
2.9. qRT-PCR
Total RNA was extracted from cultured cells or lung homogenate using a PureLink viral RNA/DNA Mini kit (ThermoFisher, New York, NY, USA; 12280-050). Real-time PCR was performed using a One Step TB Green PrimeScript PLUS RT-PCR Kit (TaKaRa; RR096A) and a Light Cycler 480 II (Roche, Mannheim, Germany). The specific primers used to target human interferon and the amplification and quantitative real-time PCR (qRT-PCR) conditions can be provided upon request. Relative expression was calculated using the comparative ΔΔCT method. The results are expressed as the n-fold difference relative to expression of GADPH (2−ΔΔCT).
2.10. Measurement of the IFN-β Levels
Culture supernatants were collected from A549 cells and assayed using the human interferon enzyme-linked immunosorbent assay kit (Gene-Protein Link, Beijing, China; P09H193).
2.11. Statistical Analysis
Data are presented as means ± SD of at least three independent experiments. Differences between two groups were evaluated using Student’s t-test, and differences between three or more groups were evaluated using analysis of variance (ANOVA). Statistical significance was set at p < 0.05. (*, p < 0.05; **, p < 0.01).
4. Discussion
The pathogenesis of IAV depends on the interaction between virus proteins and the host immune system. To replicate successfully in the host, IAVs have evolved to evade host immune response by encoding proteins such as NS1, which regulate host responses.
Previous studies show that the IAV NS1 protein binds to CPSF30, which suppresses expression of host genes encoding IFN and ISGs, thereby counteracting innate immune responses [
17]. The NS1 protein of the 2009 pandemic H1N1 virus lost the ability to bind to CPSF30, so it does not block expression of these host genes [
11,
18]. Three amino acids (R108, E125, and G189) in the NS1 protein of the 2009 pandemic H1N1 virus are highly conserved, which is not the case for former seasonal influenza viruses and avian viruses. A previous study showed that reinstating R108K, E125D, and G189D rescued the ability to inhibit CPSF30 [
11].
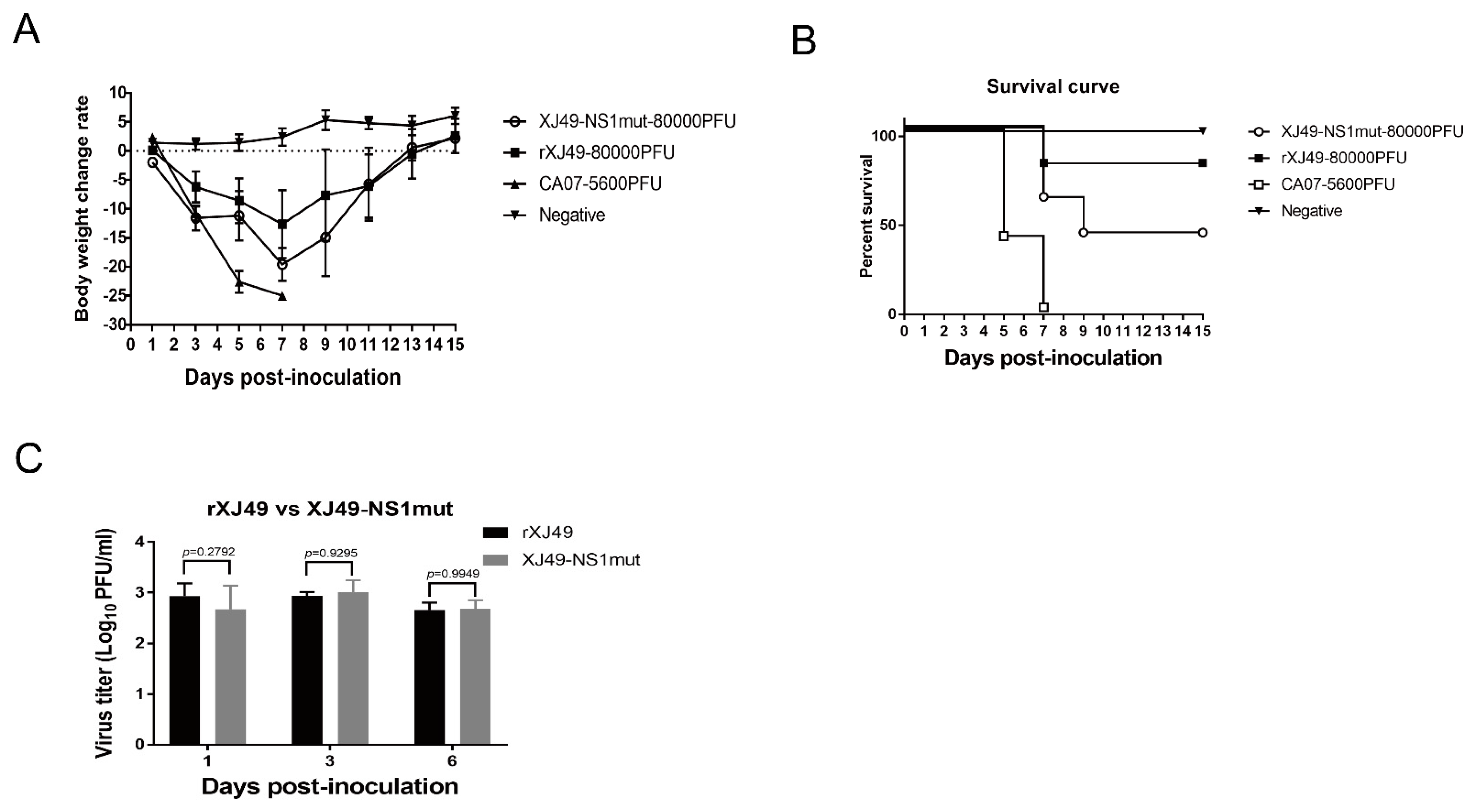
Here, we compared the sequences of 2155 strains of H1N1 virus identified since 2009 and found that, although some had R108K and G189D mutations in the NS1 protein, most retained R108 and G189. In 2018, we isolated a seasonal pandemic H1N1-origin influenza virus strain called A/Urumqi/XJ49/2018 from throat swabs of patients. We sequenced it and found that the NS1 protein contained the E125D mutation but retained R108 and G189. The isolate exhibited much lower morbidity in a mouse model than the early pandemic 2009 H1N1 isolate CA07. Thus, human H1N1 virus with triple or individual mutations at residues 108, 125, and 189 seemed not to be emerging constantly. Here, we used a H1N1 reverse genetics system to introduce mutations at sites 108 and 189 of the NS1 protein, hoping to obtain information about the molecular mechanism underlying how polymorphisms at these residues affect the interaction between NS1 and host factors. As expected, the rXJ49-NS1mut virus harboring the mutant NS1 protein replicated in MDCK cells in a manner similar to that of WT XJ49 virus but was more pathogenic in C57BL/6N mice. A previous study showed that a triple mutant NS1 virus based on the CA07 backbone showed less morbidity, whereas the mutant NS1 virus based on the XJ49 backbone exhibited higher morbidity. Nogales et al. revealed that inhibition of host gene expression depends on a strict balance between the NS1 and PA-X proteins. Compared with the pandemic CA07 virus, the seasonal XJ49 virus contained N204S, R221Q, L229S mutations in PA-X, which may be the reason for the different morbidity, even though it contained the same triple R108K, E125D, and G189D mutations [
18].
Inhibition of interferon-induced cellular signals is very important for influenza infection, because production of cytokines such as type I interferon is the initial innate immune response that fights infection. The NS1 protein, which weakens the IFN response through a variety of mechanisms during influenza virus infection, is the most important IFN-antagonistic protein encoded by the virus [
19]. The protein is highly expressed in the cytoplasm and nucleus of infected cells and can interact with different components involved in the IFN response [
20]. During influenza virus infection, the host cell initiates a cascade of antiviral signals through pattern recognition receptor (PRR)-mediated recognition of pathogen-related molecular patterns (PAMPs). The main influenza virus PAMP is viral dsRNA with a triphosphate group at the 5’ end [
21]. The NS1 protein can use its RNA binding region to bind competitively to PAMPs and inhibit activation of the RIG–I signal pathway, thereby inhibiting transcription and translation of the IFN-β promoter. Here, we used a double luciferase assay to examine the regulatory effect of the NS1 protein on the IFN promoter; we found that WT NS1 did not affect IFN expression, whereas NS1 harboring R108K/G189D suppressed it.
The TRIM family is a superfamily with Ring-finger E3 ubiquitin ligase activity, which has more than 70 members in humans. TRIM family members are critical regulators of innate immune responses against microbial pathogens, including viruses [
22]. During influenza virus infection, NS1 inhibits oligomerization of TRIM25 by interacting with the E3 ubiquitin ligase domain to prevent ubiquitination of RIG-I K63, thereby blocking formation of the RIG-I-MAVS signal complex and effectively inhibiting production of IFN [
9]. Early studies show that CPSF30 recognizes the polyadenylation signal at the 3’ end of host mRNA during transcription, cleaves host RNA, and binds poly A polymerase to increase the tail of poly A; thus, NS1 binding to CPSF30 can inhibit polyadenosine acidification at the 3’ end of mRNA and block mRNA production in the nucleus, thereby inhibiting host cell gene expression [
23]. Here, we infected A549 cells with two viruses and found that expression of mRNA encoding TRIM25 and RIG-I in cells infected with XJ49-NS1mut was significantly lower than that in cells infected with rXJ49. Trim22 is a typical TRIM family protein that restricts replication of influenza viruses by targeting IAV nucleoproteins for degradation [
24]. Published studies reveal that recognition of viral DNA/RNA triggers type I interferon production dependent on IRF3 activation. Secreted IFN-α/β bind to IFNAR to activate the Stat1–Stat2 heterodimer and induce expression of TRIM22, thereby protecting the host against diverse viral infections. The mutant NS1 protein may inhibit expression of TRIM25 and TRIM22, block the RIG–I pathway, and inhibit expression of IFN. In addition, the levels of IL-28 and IL-29 in cells infected with rXJ49-NS1mut decreased significantly, indicating that the R108K/G189D mutation in the NS1 protein could indeed inhibit host immunity during the early stage of virus infection by suppressing activation of the JAK–Stat pathway.
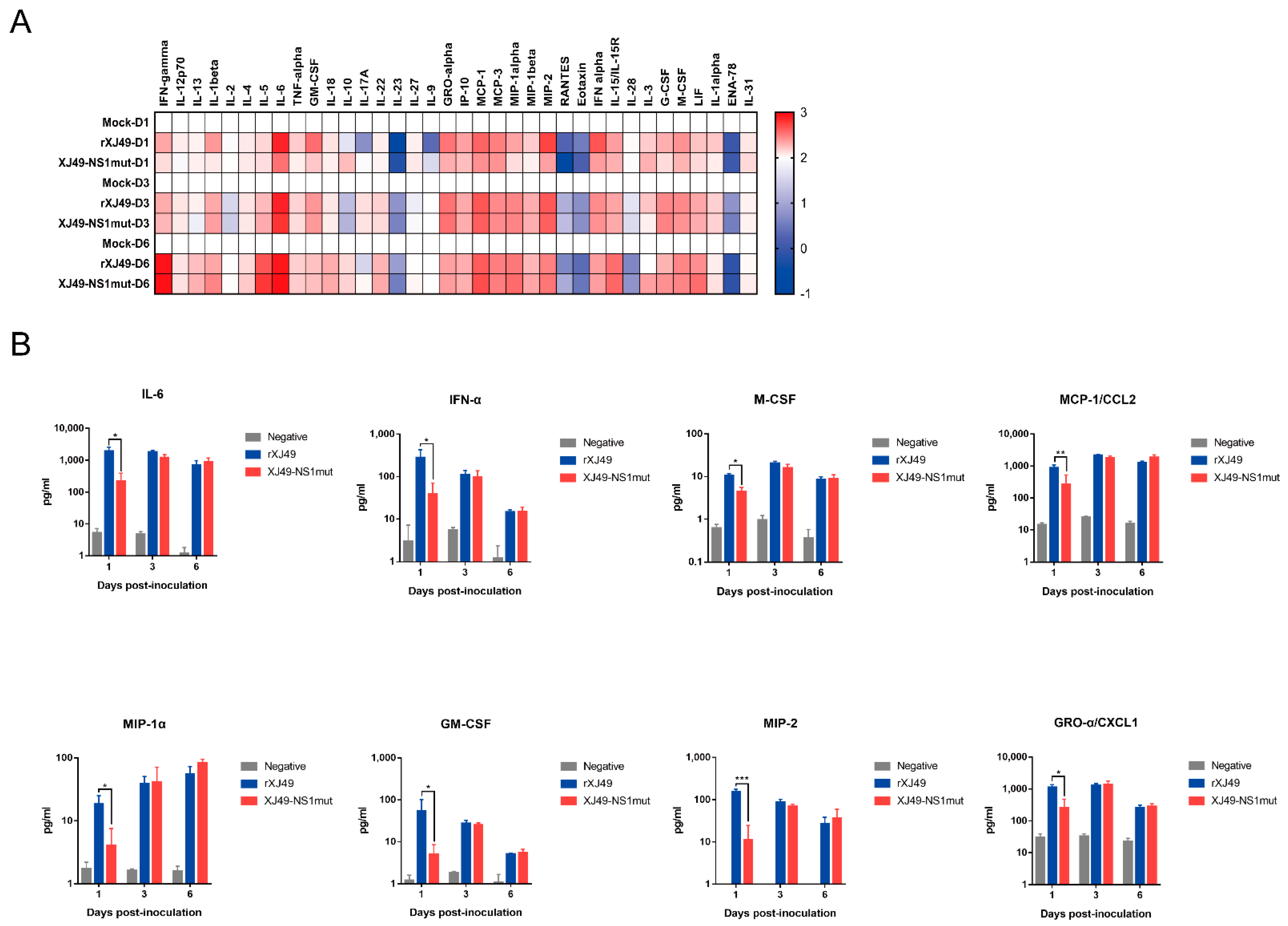
It seems that different cytokine waves appear as infection progresses. The first cytokines produced in the lungs after IAV infection are chemokines CCL5/RANTES, CCL2/MCP-1, and CXCL8/IL-8, as well as the main antiviral cytokines IFN-α, IFN-β, and IFN-γ. When the virus reaches the lower respiratory tract, it can infect activated alveolar macrophages and secrete higher levels of proinflammatory and antiviral cytokines [
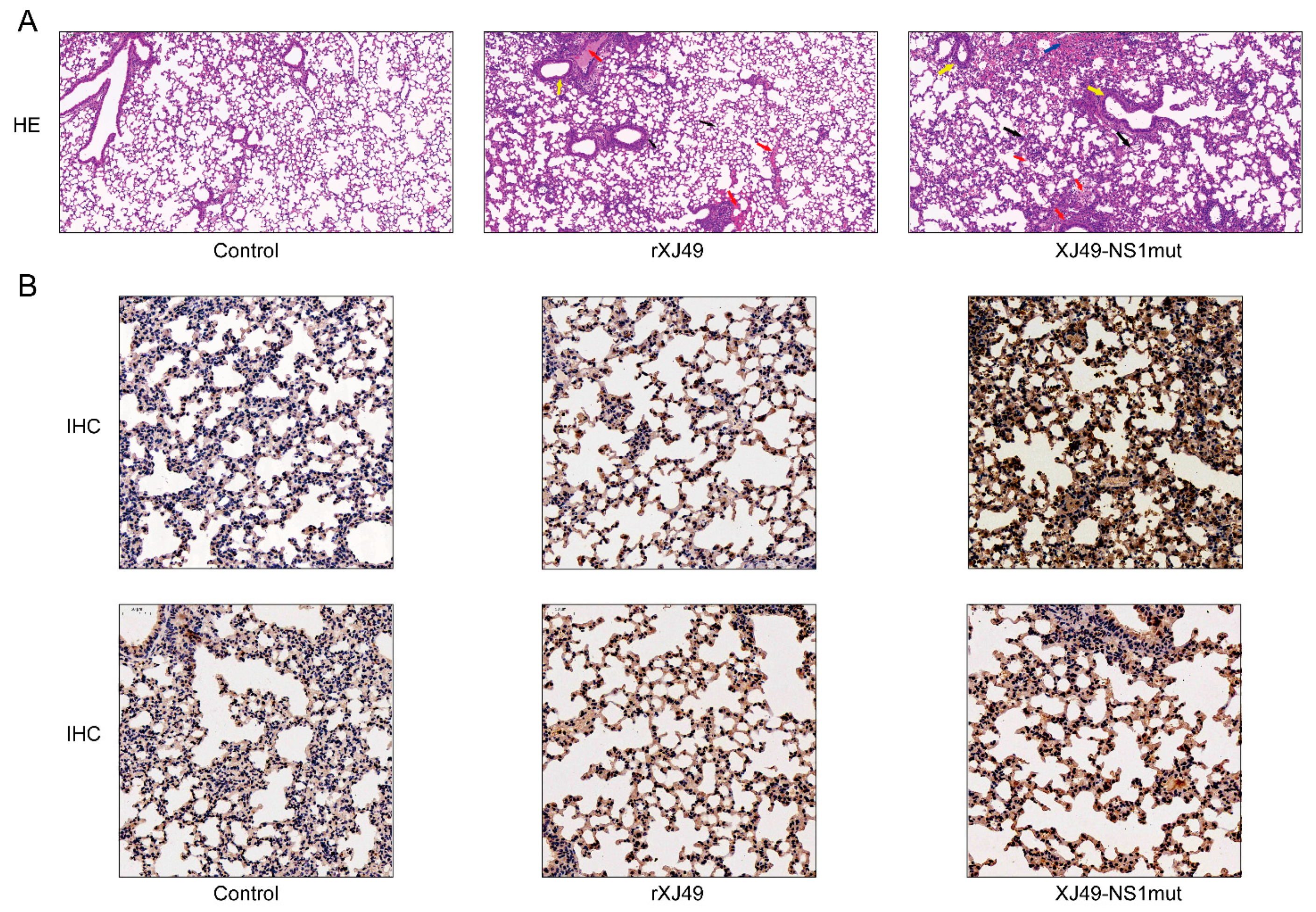
9]. In the lung homogenates of infected mice, we detected representative cytokines; indeed, expression of these cytokines in the lungs of mice infected with rXJ49-NS1mut and WT rXJ49 virus were significantly upregulated, especially those involved in inflammation (CCL2/MCP-1, CCL3/MIP-1α, CCL4/MIP-1β, and CCL7/MCP-3). Interestingly, we analyzed the pathology of mouse lungs and found that the pathological damage caused by rXJ49-NS1mut virus was more serious than that caused by WT rXJ49. It can be speculated that the R108K and G189D mutations in the NS1 protein enhance the innate immune response of the influenza virus against the host, resulting in the weakening of the host antiviral response, thus achieving the purpose of immune escape.