Cascading from SARS-CoV-2 to Parkinson’s Disease through Protein-Protein Interactions
Abstract
1. Introduction
2. Antecedents
- Several viruses have been associated with Parkinsonism [30], which include RNA viruses of the families of Bornaviridae, Orthomyxoviridae, Paramyxoviridae, Picornavirisae, Retroviridae, and Flaviviridae. After the 1918 pandemic influenza outbreak caused by H1N1 influenza virus, there were several cases of postencephalic Parkinsonism [31]. Recently, Jang et al. [32] have reported that a highly pathogenic H5N1 influenza virus can induce Parkisonian pathology in mice.
- The cytokine storm hypothesis is not able to explain the extrapulmonary damages produced by SARS-CoV-2 infection as the median levels of IL-6 in patients with severe COVID-19 are 10- to 200-fold smaller (see Table 1) than those observed in patients with hyperinflammatory phenotype of acute respiratory distress syndrome (ARDS) (see also the comments in References [33,34]):
| Severe COVID-19 | Population | IL-6 Level pg/mL | ||
|---|---|---|---|---|
| [35] | 84 | 7 (6–11) | ||
| [36] | 54 | 11 (8–14) | ||
| [37] | 286 | 25 (10–55) | ||
| [38] | 237 | 26 (11–69) | ||
| [39] | 85 | 64 (31–165) | ||
| [40] | 17 * | 64 (25.6–111.9) | ||
| ARDS | “hypoinflamatory” | hyperinflamatory | ||
| pop. | IL-6 level pg/mL | pop. | IL-6 level pg/mL | |
| [41] | 638 | 86 (34–216) | 246 | 578 (181–2621) |
| [42] | 386 | 154 (67–344) | 135 | 1525 (584–3802) |
| [43] | 451 | 282 (111–600) | 269 | 1618 (517–3205) |
- SARS-CoV-2 vRNA has been detected using PCR techniques in different parts of the brain [44,45]. While the number of copies per mL of RNA from homogenized organs and tissues in 11 patients deceased from COVID-19 ranged 114.8–19,498 for different parts of the respiratory system, it ranged only 2.3–4.9 for different parts of the nervous system [44].
- It has been recently determined by Philippen et al. [46] that SARS-CoV-2 infection causes brain inflammation in the macaque model. Post-mortem analysis demonstrated infiltration of T-cells and activated microglia in the brain. Viral RNA was detected in brain tissues from one animal. The authors observed Lewy bodies in brains of all rhesus macaques. In humans, Lewy body formation is an indication for the development of Parkinson’s disease.
- Wölfel et al. [47] reported infectious virus readily isolated from samples derived from the throat or lung of COVID-19 patients, but not from stool samples—in spite of high concentrations of virus RNA. Blood and urine samples never yielded virus. Therefore, the identification of vRNA does not necessarily indicates viral tropism. There is huge evidence that RNA, including viral one, can be delivered to mammalian cells by means of extracellular vesicles, such as exosomes [48].
- It is known today that exosomes [49] facilitate the spread of viruses improving virus infection pathogenesis [50,51,52]. For instance, proteins and noncoding RNA from HIV are known to be transported by exosomes to (i) increase susceptibility to infection, (ii) influence virus budding and spread, and (iii) increase neuropathogenesis. Viral RNA and proteins from Zika virus are transported by exosomes to improve viral spread to neighboring cells. Virus spread is also known to be facilitated by exosomes in EV-A71, Rabies virus, and HCV. The use of exosomes containing viral proteins and/or viral RNA is also know to evade the immune system in the case of EBV, KSHV, HSV1, HTLV-1, and avian influenza (H5N1).
- Ramakrishnaiah et al. [53] found experimentally that purified exosomes isolated from HCV-infected human cells contained full-length viral RNA and proteins, which were capable of transmitting infection to human cells. They also shown that exosomes-transmitted infection was resistant to antibody neutralization.
- V’kovski et al. [57] have shown recently that SARS-CoV-2 replicated to higher titers when infections were performed at 33 °C rather than 37 °C. The reverse is found for SARS-CoV. They also found that SARS-CoV-2 triggered a pronounced antiviral and pro-inflammatory response earlier and more strongly induced at 37 °C than at 33 °C. These temperatures correspond to the ones of the upper (33 °C) and lower (37 °C) respiratory tract.
- The temperature of healthy brain is slightly higher than 37 °C (ventral striatum 37.6 °C; dorsal striatum 37.2 °C; cerebellum 37.3 °C) [58], (frontal lobe (37.2 ± 0.6 °C) and thalamus (37.7 ± 0.6 °C)) [59], which may indicate a limited capacity for SARS-CoV-2 for reproducing in this organ where it also would trigger higher immunological response.
- Post-translational modification of host proteins is a key strategy of viral pathogens to modulate host factors critical for infection, which are essential for viruses’ replication, propagation, and evasion from host immune responses [60].
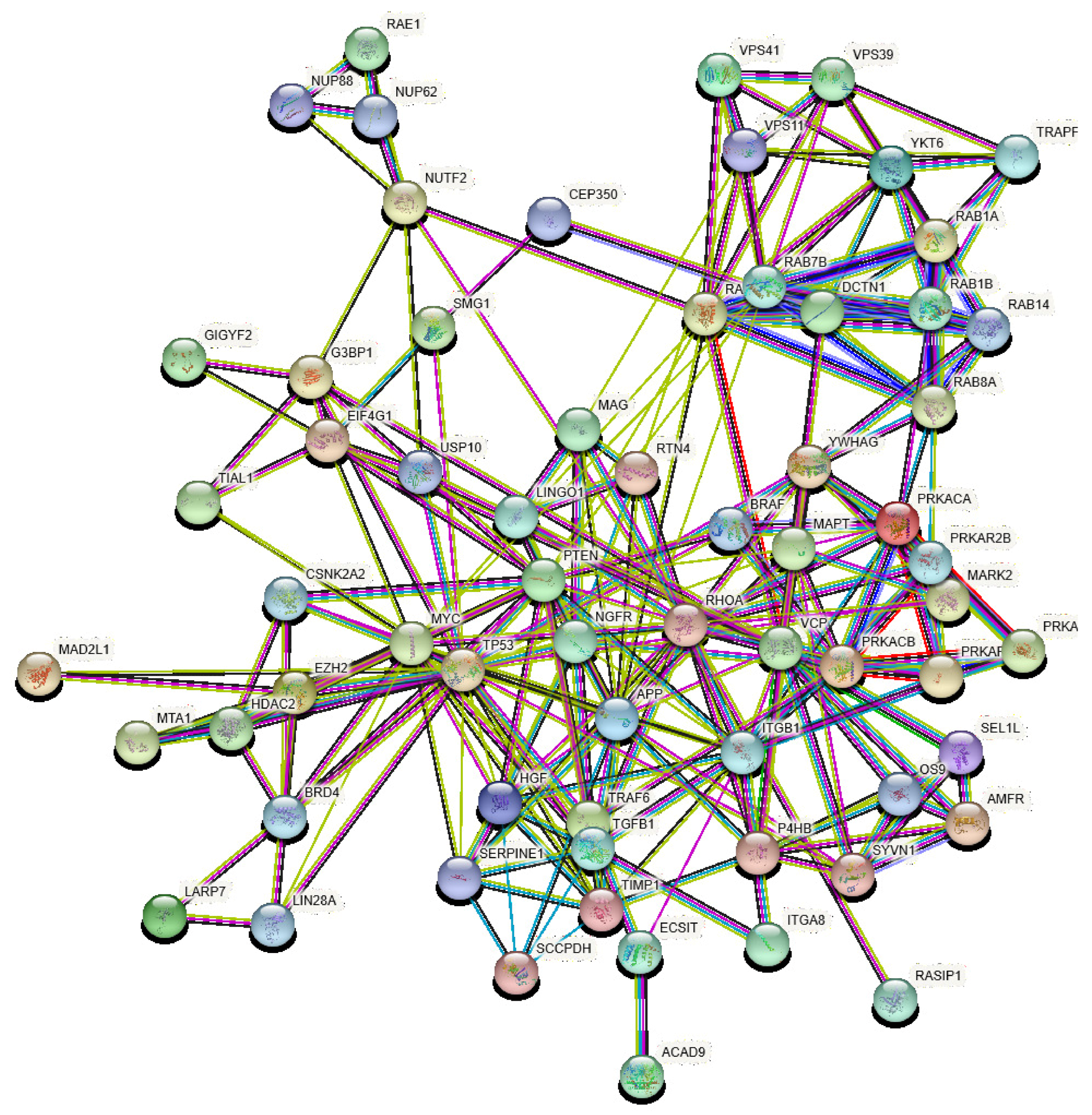
- It is today understood that the spread of perturbations across the subcellular networks, such as protein-protein interaction networks, is one of the major causes of diseases [61,62]. Such perturbations can be either of topological nature, e.g., deletion of nodes (proteins) or edges (interactions), or dynamical, i.e., the propagation of changes in the concentrations of given proteins in the cell.
3. Materials and Methods
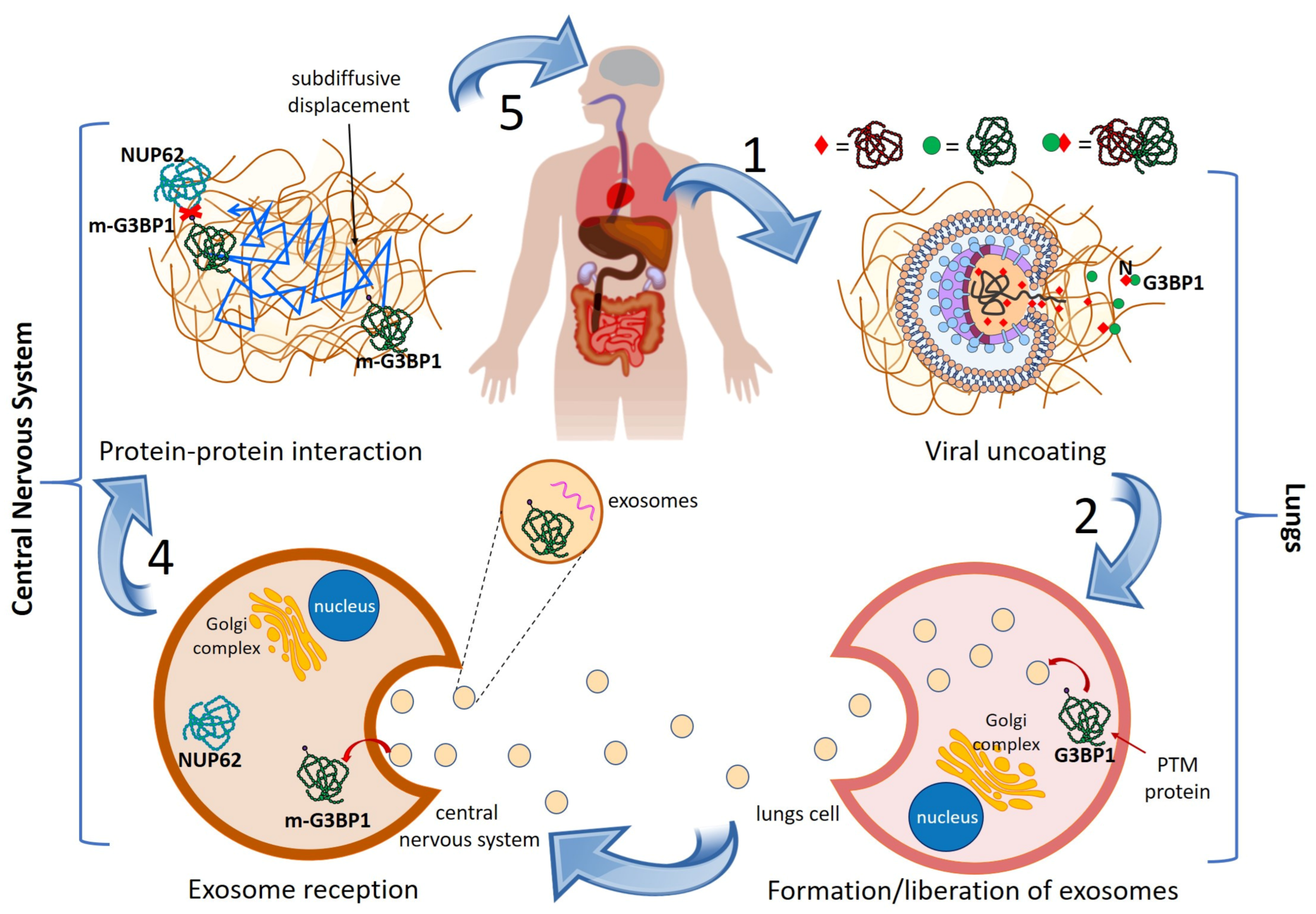
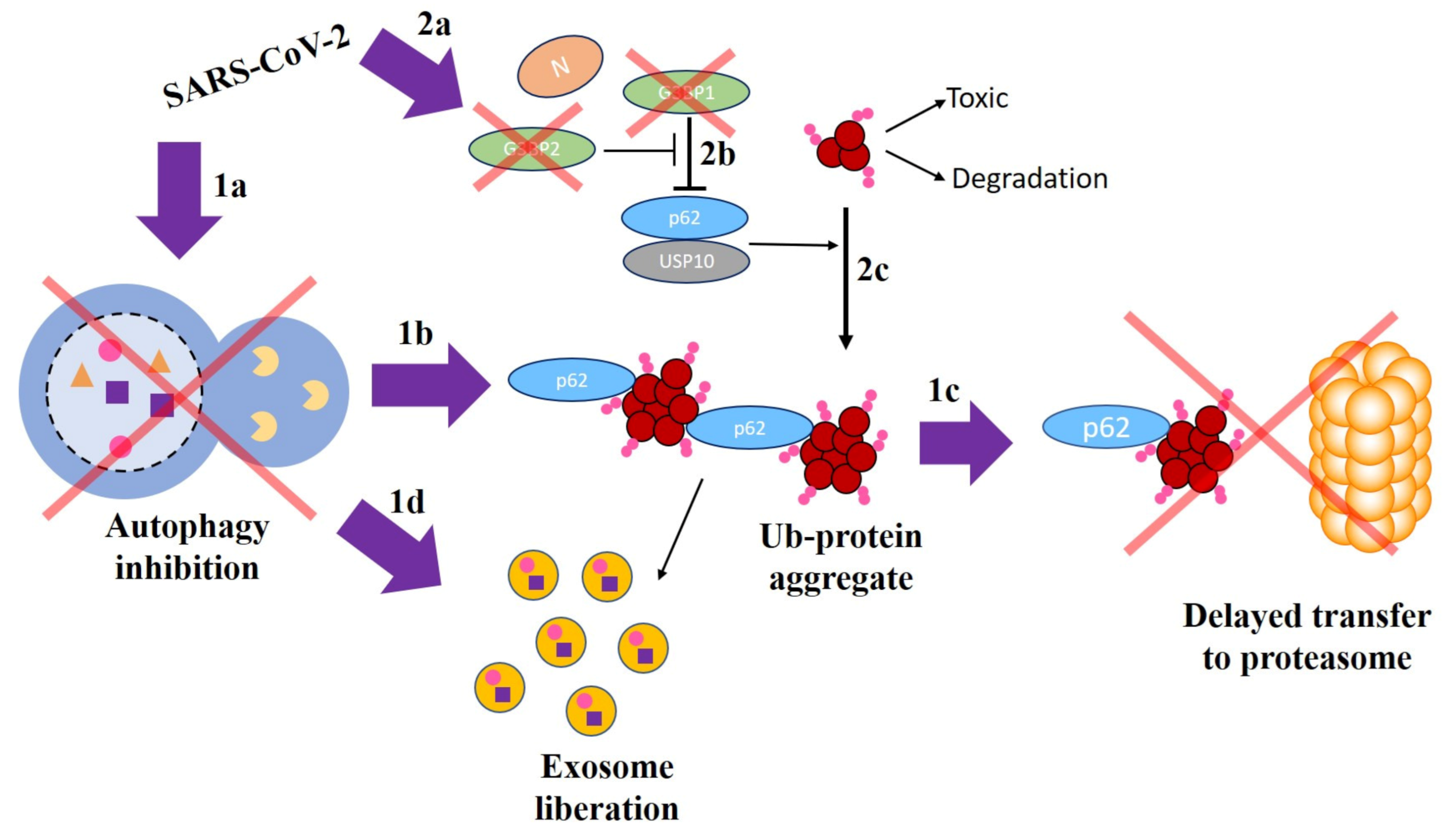
3.1. Cascading Mechanism
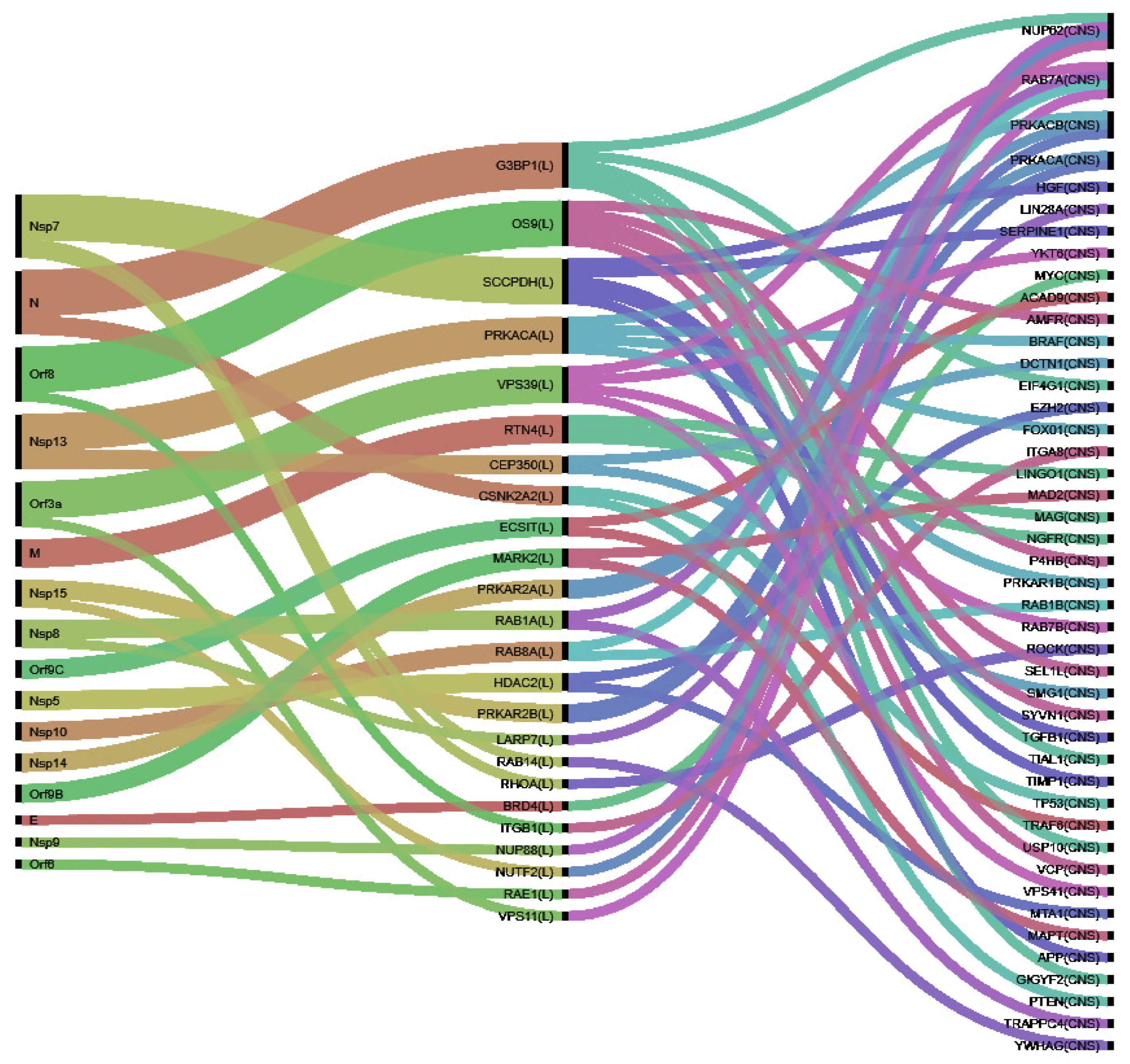
3.2. Identification of VP and Their Perturbators
4. Results
5. Discussion
6. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Gorbalenya, A.; Baker, S.; Baric, R.; de Groot, R.; Drosten, C.; Gulyaeva, A.; Haagmans, B.; Lauber, C.; Leontovich, A.; Neuman, B.; et al. Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 2020, 3–4. [Google Scholar]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.M.; Wang, W.; Song, Z.G.; Hu, Y.; Tao, Z.W.; Tian, J.H.; Pei, Y.Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Estrada, E. COVID-19 and SARS-CoV-2. Modeling the present, looking at the future. Phys. Rep. 2020, 869, 1–51. [Google Scholar] [CrossRef]
- Gupta, A.; Madhavan, M.V.; Sehgal, K.; Nair, N.; Mahajan, S.; Sehrawat, T.S.; Bikdeli, B.; Ahluwalia, N.; Ausiello, J.C.; Wan, E.Y.; et al. Extrapulmonary manifestations of COVID-19. Nat. Med. 2020, 26, 1017–1032. [Google Scholar] [CrossRef]
- Prasanna, P.L.; Abilash, V. Coronaviruses pathogenesis, comorbidities and multi-organ damage–A review. Life Sci. 2020. [Google Scholar] [CrossRef]
- Robba, C.; Battaglini, D.; Pelosi, P.; Rocco, P.R. Multiple organ dysfunction in SARS-CoV-2: MODS-CoV-2. Expert Rev. Respir. Med. 2020, 14, 865–868. [Google Scholar] [CrossRef]
- Jiang, F.; Deng, L.; Zhang, L.; Cai, Y.; Cheung, C.W.; Xia, Z. Review of the clinical characteristics of coronavirus disease 2019 (COVID-19). J. Gen. Intern. Med. 2020, 35, 545–1549. [Google Scholar] [CrossRef]
- Zhou, M.; Zhang, X.; Qu, J. Coronavirus disease 2019 (COVID-19): A clinical update. Front. Med. 2020, 14, 126–135. [Google Scholar] [CrossRef]
- Abdel-Mannan, O.; Eyre, M.; Löbel, U.; Bamford, A.; Eltze, C.; Hameed, B.; Hemingway, C.; Hacohen, Y. Neurologic and radiographic findings associated with COVID-19 infection in children. JAMA Neurol. 2020, 77, 1440–1445. [Google Scholar] [CrossRef]
- Pinzon, R.T.; Wijaya, V.O.; Buana, R.B.; Al Jody, A.; Nunsio, P.N. Neurologic Characteristics in Coronavirus Disease 2019 (COVID-19): A Systematic Review and Meta-Analysis. Front. Neurol. 2020, 11, 565. [Google Scholar] [CrossRef] [PubMed]
- Meppiel, E.; Peiffer-Smadja, N.; Maury, A.; Bekri, I.; Delorme, C.; Desestret, V.; Gorza, L.; Hautecloque-Raysz, G.; Landre, S.; Lannuzel, A.; et al. Neurologic manifestations associated with COVID-19: A multicentre registry. Clin. Microbiol. Infect. 2020, 27, 458–466. [Google Scholar] [CrossRef] [PubMed]
- Verkhratsky, A.; Li, Q.; Melino, S.; Melino, G.; Shi, Y. Can COVID-19 pandemic boost the epidemic of neurodegenerative diseases? Biol. Direct 2020, 15, 1–8. [Google Scholar] [CrossRef]
- Faber, I.; Brandao, P.R.; Menegatti, F.; de Carvalho Bispo, D.D.; Maluf, F.B.; Cardoso, F. Coronavirus disease 2019 and Parkinsonism: A non-post-encephalitic case. Mov. Disord. 2020, 35, 1721–1722. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.E.; Eichel, R.; Steiner-Birmanns, B.; Janah, A.; Ioshpa, M.; Bar-Shalom, R.; Paul, J.J.; Gaber, H.; Skrahina, V.; Bornstein, N.M.; et al. A case of probable Parkinson’s disease after SARS-CoV-2 infection. Lancet Neurol. 2020, 19, 804–805. [Google Scholar] [CrossRef]
- Méndez-Guerrero, A.; Laespada-García, M.I.; Gómez-Grande, A.; Ruiz-Ortiz, M.; Blanco-Palmero, V.A.; Azcarate-Diaz, F.J.; Rábano-Suárez, P.; Álvarez-Torres, E.; Pérez, D.V.; Rodríguez-Montalbán, R.; et al. Acute hypokinetic-rigid syndrome following SARS-CoV-2 infection. Neurology 2020, 95, e2109–e2118. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Latapi, P.; Fearon, C.; Fasano, A.; Lang, A.E. Parkinson’s disease and COVID-19: Do we need to be more patient? Mov. Disord. 2021, 36, 277. [Google Scholar] [CrossRef] [PubMed]
- Outeiro, T.F.; Krisko, A. Reply to: “Parkinson’s Disease and COVID-19: Do We Need to Be More Patient?”. Mov. Disord. 2021, 36, 278. [Google Scholar] [CrossRef]
- Méndez-Guerrero, A.; Blanco-Palmero, V.A.; Laespada-García, M.I.; Azcárate-Díaz, F.J.; de la Aleja, J.G. Author response: Acute hypokinetic-rigid syndrome following SARS-CoV-2 infection. Neurology 2020, 96. [Google Scholar] [CrossRef]
- Brundin, P.; Nath, A.; Beckham, J.D. Is COVID-19 a perfect storm for Parkinson’s disease? Trends Neurosci. 2020, 43, 931–933. [Google Scholar] [CrossRef]
- Sulzer, D.; Antonini, A.; Leta, V.; Nordvig, A.; Smeyne, R.J.; Goldman, J.E.; Al-Dalahmah, O.; Zecca, L.; Sette, A.; Bubacco, L.; et al. COVID-19 and possible links with Parkinson’s disease and parkinsonism: From bench to bedside. NPJ Park. Dis. 2020, 6, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Akilli, N.B.; Yosunkaya, A. Part of the Covid19 puzzle: Acute parkinsonism. Am. J. Emerg. Med. 2021. In Press. [Google Scholar] [CrossRef] [PubMed]
- Matschke, J.; Lütgehetmann, M.; Hagel, C.; Sperhake, J.P.; Schröder, A.S.; Edler, C.; Mushumba, H.; Fitzek, A.; Allweiss, L.; Dandri, M.; et al. Neuropathology of patients with COVID-19 in Germany: A post-mortem case series. Lancet Neurol. 2020, 19, 919–929. [Google Scholar] [CrossRef]
- Leisman, D.E.; Ronner, L.; Pinotti, R.; Taylor, M.D.; Sinha, P.; Calfee, C.S.; Hirayama, A.V.; Mastroiani, F.; Turtle, C.J.; Harhay, M.O.; et al. Cytokine elevation in severe and critical COVID-19: A rapid systematic review, meta-analysis, and comparison with other inflammatory syndromes. Lancet Respir. Med. 2020, 8, 1233–1244. [Google Scholar] [CrossRef]
- Aziz, M.; Fatima, R.; Assaly, R. Elevated interleukin-6 and severe COVID-19: A meta-analysis. J. Med Virol. 2020. [Google Scholar] [CrossRef]
- Leisman, D.E.; Deutschman, C.S.; Legrand, M. Facing COVID-19 in the ICU: Vascular dysfunction, thrombosis, and dysregulated inflammation. Intensive Care Med. 2020, 46, 1105–1108. [Google Scholar] [CrossRef]
- Sinha, P.; Matthay, M.A.; Calfee, C.S. Is a “cytokine storm” relevant to COVID-19? JAMA Intern. Med. 2020. [Google Scholar] [CrossRef]
- Sinha, P.; Delucchi, K.L.; McAuley, D.F.; O’Kane, C.M.; Matthay, M.A.; Calfee, C.S. Development and validation of parsimonious algorithms to classify acute respiratory distress syndrome phenotypes: A secondary analysis of randomised controlled trials. Lancet Respir. Med. 2020, 8, 247–257. [Google Scholar] [CrossRef]
- Merello, M.; Bhatia, K.P.; Obeso, J.A. SARS-CoV-2 and the risk of Parkinson’s disease: Facts and fantasy. Lancet Neurol. 2020, 20, 94–95. [Google Scholar] [CrossRef]
- Jang, H.; Boltz, D.A.; Webster, R.G.; Smeyne, R.J. Viral parkinsonism. Biochim. Et Biophys. Acta (BBA)-Mol. Basis Dis. 2009, 1792, 714–721. [Google Scholar] [CrossRef]
- Henry, J.; Smeyne, R.J.; Jang, H.; Miller, B.; Okun, M.S. Parkinsonism and neurological manifestations of influenza throughout the 20th and 21st centuries. Park. Relat. Disord. 2010, 16, 566–571. [Google Scholar] [CrossRef]
- Jang, H.; Boltz, D.; Sturm-Ramirez, K.; Shepherd, K.R.; Jiao, Y.; Webster, R.; Smeyne, R.J. Highly pathogenic H5N1 influenza virus can enter the central nervous system and induce neuroinflammation and neurodegeneration. Proc. Natl. Acad. Sci. USA 2009, 106, 14063–14068. [Google Scholar] [CrossRef]
- Hedrick, T.L.; Murray, B.P.; Hagan, R.S.; Mock, J.R. COVID-19: Clean up on IL-6. Am. J. Respir. Cell Mol. Biol. 2020, 63, 541–543. [Google Scholar] [CrossRef]
- Campochiaro, C.; Dagna, L. The conundrum of interleukin-6 blockade in COVID-19. Lancet Rheumatol. 2020, 2, e579–e580. [Google Scholar] [CrossRef]
- Wu, C.; Chen, X.; Cai, Y.; Zhou, X.; Xu, S.; Huang, H.; Zhang, L.; Zhou, X.; Du, C.; Zhang, Y.; et al. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern. Med. 2020, 180, 934–943. [Google Scholar] [CrossRef]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Qin, C.; Zhou, L.; Hu, Z.; Zhang, S.; Yang, S.; Tao, Y.; Xie, C.; Ma, K.; Shang, K.; Wang, W.; et al. Dysregulation of immune response in patients with COVID-19 in Wuhan, China. Clin. Infect. Dis. 2020, 71, 762–768. [Google Scholar] [CrossRef]
- Cummings, M.J.; Baldwin, M.R.; Abrams, D.; Jacobson, S.D.; Meyer, B.J.; Balough, E.M.; Aaron, J.G.; Claassen, J.; Rabbani, L.E.; Hastie, J.; et al. Epidemiology, clinical course, and outcomes of critically ill adults with COVID-19 in New York City: A prospective cohort study. Lancet 2020, 395, 1763–1770. [Google Scholar] [CrossRef]
- Mo, P.; Xing, Y.; Xiao, Y.; Deng, L.; Zhao, Q.; Wang, H.; Xiong, Y.; Cheng, Z.; Gao, S.; Liang, K.; et al. Clinical characteristics of refractory COVID-19 pneumonia in Wuhan, China. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Chen, X.; Zhao, B.; Qu, Y.; Chen, Y.; Xiong, J.; Feng, Y.; Men, D.; Huang, Q.; Liu, Y.; Yang, B.; et al. Detectable serum SARS-CoV-2 viral load (RNAaemia) is closely correlated with drastically elevated interleukin 6 (IL-6) level in critically ill COVID-19 patients. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Famous, K.R.; Delucchi, K.; Ware, L.B.; Kangelaris, K.N.; Liu, K.D.; Thompson, B.T.; Calfee, C.S. Acute respiratory distress syndrome subphenotypes respond differently to randomized fluid management strategy. Am. J. Respir. Crit. Care Med. 2017, 195, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Calfee, C.S.; Delucchi, K.; Parsons, P.E.; Thompson, B.T.; Ware, L.B.; Matthay, M.A.; Nhlbi Ards Network. Subphenotypes in acute respiratory distress syndrome: Latent class analysis of data from two randomised controlled trials. Lancet Respir. Med. 2014, 2, 611–620. [Google Scholar] [CrossRef]
- Sinha, P.; Delucchi, K.L.; Thompson, B.T.; McAuley, D.F.; Matthay, M.A.; Calfee, C.S. Latent class analysis of ARDS subphenotypes: A secondary analysis of the statins for acutely injured lungs from sepsis (SAILS) study. Intensive Care Med. 2018, 44, 1859–1869. [Google Scholar] [CrossRef] [PubMed]
- Deinhardt-Emmer, S.; Wittschieber, D.; Sanft, J.; Kleemann, S.; Elschner, S.; Haupt, K.F.; Vau, V.; Häring, C.; Rödel, J.; Henke, A.; et al. Early postmortem mapping of SARS-CoV-2 RNA in patients with COVID-19 and the correlation with tissue damage. Elife 2021, 10, e60361. [Google Scholar] [CrossRef]
- Liu, J.M.; Tan, B.H.; Wu, S.; Gui, Y.; Suo, J.L.; Li, Y.C. Evidence of central nervous system infection and neuroinvasive routes, as well as neurological involvement, in the lethality of SARS-CoV-2 infection. J. Med. Virol. 2021, 93, 1304–1313. [Google Scholar] [CrossRef]
- Philippens, I.H.; Boszormenyi, K.P.; Wubben, J.A.; Fagrouch, Z.C.; van Driel, N.; Mayenburg, A.Q.; Lozovagia, D.; Roos, E.; Schurink, B.; Bugiani, M.; et al. SARS-CoV-2 causes brain inflammation and induces Lewy body formation in macaques. bioRxiv 2021. [Google Scholar] [CrossRef]
- Wölfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Müller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef]
- O’Brien, K.; Breyne, K.; Ughetto, S.; Laurent, L.C.; Breakefield, X.O. RNA delivery by extracellular vesicles in mammalian cells and its applications. Nat. Rev. Mol. Cell Biol. 2020, 21, 585–606. [Google Scholar] [CrossRef]
- Pegtel, D.M.; Gould, S.J. Exosomes. Annu. Rev. Biochem. 2019, 88, 487–514. [Google Scholar] [CrossRef]
- Schorey, J.S.; Cheng, Y.; Singh, P.P.; Smith, V.L. Exosomes and other extracellular vesicles in host–pathogen interactions. EMBO Rep. 2015, 16, 24–43. [Google Scholar] [CrossRef]
- Alenquer, M.; Amorim, M.J. Exosome biogenesis, regulation, and function in viral infection. Viruses 2015, 7, 5066–5083. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.R.; Kashanchi, F.; Jacobson, S. Exosomes in viral disease. Neurotherapeutics 2016, 13, 535–546. [Google Scholar] [CrossRef]
- Ramakrishnaiah, V.; Thumann, C.; Fofana, I.; Habersetzer, F.; Pan, Q.; de Ruiter, P.E.; Willemsen, R.; Demmers, J.A.; Raj, V.S.; Jenster, G.; et al. Exosome-mediated transmission of hepatitis C virus between human hepatoma Huh7. 5 cells. Proc. Natl. Acad. Sci. USA 2013, 110, 13109–13113. [Google Scholar] [CrossRef] [PubMed]
- Estrada, E. Protein-driven mechanism of multiorgan damage in COVID-19. Med. Drug Discov. 2020, 8, 100069. [Google Scholar] [CrossRef] [PubMed]
- Hassanpour, M.; Rezaie, J.; Nouri, M.; Panahi, Y. The role of extracellular vesicles in COVID-19 virus infection. Infect. Genet. Evol. 2020, 85, 104422. [Google Scholar] [CrossRef] [PubMed]
- Patil, M.; Singh, S.; Henderson, J.; Krishnamurthy, P. Mechanisms of COVID-19-induced cardiovascular disease: Is sepsis or exosome the missing link? J. Cell. Physiol. 2021, 236, 3366–3382. [Google Scholar] [CrossRef] [PubMed]
- V’kovski, P.; Gultom, M.; Kelly, J.N.; Steiner, S.; Russeil, J.; Mangeat, B.; Cora, E.; Pezoldt, J.; Holwerda, M.; Kratzel, A.; et al. Disparate temperature-dependent virus–host dynamics for SARS-CoV-2 and SARS-CoV in the human respiratory epithelium. PLoS Biol. 2021, 19, e3001158. [Google Scholar] [CrossRef] [PubMed]
- Kiyatkin, E.A. Brain temperature homeostasis: Physiological fluctuations and pathological shifts. Front. Biosci. A J. Virtual Libr. 2010, 15, 73. [Google Scholar] [CrossRef]
- Corbett, R.; Laptook, A.; Weatherall, P. Noninvasive measurements of human brain temperature using volume-localized proton magnetic resonance spectroscopy. J. Cereb. Blood Flow Metab. 1997, 17, 363–369. [Google Scholar] [CrossRef]
- Ribet, D.; Cossart, P. Pathogen-mediated posttranslational modifications: A re-emerging field. Cell 2010, 143, 694–702. [Google Scholar] [CrossRef]
- Santolini, M.; Barabási, A.L. Predicting perturbation patterns from the topology of biological networks. Proc. Natl. Acad. Sci. USA 2018, 115, E6375–E6383. [Google Scholar] [CrossRef]
- Barabási, A.L.; Gulbahce, N.; Loscalzo, J. Network medicine: A network-based approach to human disease. Nat. Rev. Genet. 2011, 12, 56–68. [Google Scholar] [CrossRef]
- Estrada, E. Fractional diffusion on the human proteome as an alternative to the multi-organ damage of SARS-CoV-2. Chaos Interdiscip. J. Nonlinear Sci. 2020, 30, 081104. [Google Scholar] [CrossRef]
- Del Sol, A.; Balling, R.; Hood, L.; Galas, D. Diseases as network perturbations. Curr. Opin. Biotechnol. 2010, 21, 566–571. [Google Scholar] [CrossRef]
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 2020, 583, 459–468. [Google Scholar] [CrossRef]
- Bouhaddou, M.; Memon, D.; Meyer, B.; White, K.M.; Rezelj, V.V.; Marrero, M.C.; Polacco, B.J.; Melnyk, J.E.; Ulferts, S.; Kaake, R.M.; et al. The global phosphorylation landscape of SARS-CoV-2 infection. Cell 2020, 182, 685–712. [Google Scholar] [CrossRef]
- Tkach, M.; Théry, C. Communication by extracellular vesicles: Where we are and where we need to go. Cell 2016, 164, 1226–1232. [Google Scholar] [CrossRef]
- Lunati, A.; Lesage, S.; Brice, A. The genetic landscape of Parkinson’s disease. Rev. Neurol. 2018, 174, 628–643. [Google Scholar] [CrossRef]
- Mann, M.; Jensen, O.N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 2003, 21, 255–261. [Google Scholar] [CrossRef]
- Xu, H.; Wang, Y.; Lin, S.; Deng, W.; Peng, D.; Cui, Q.; Xue, Y. PTMD: A database of human disease-associated post-translational modifications. Genom. Proteom. Bioinform. 2018, 16, 244–251. [Google Scholar] [CrossRef]
- Howitt, J.; Hill, A.F. Exosomes in the pathology of neurodegenerative diseases. J. Biol. Chem. 2016, 291, 26589–26597. [Google Scholar] [CrossRef] [PubMed]
- Soria, F.N.; Pampliega, O.; Bourdenx, M.; Meissner, W.G.; Bezard, E.; Dehay, B. Exosomes, an unmasked culprit in neurodegenerative diseases. Front. Neurosci. 2017, 11, 26. [Google Scholar] [CrossRef] [PubMed]
- Han, C.; Xiong, N.; Guo, X.; Huang, J.; Ma, K.; Liu, L.; Xia, Y.; Shen, Y.; Li, J.; Jiang, H.; et al. Exosomes from patients with Parkinson’s disease are pathological in mice. J. Mol. Med. 2019, 97, 1329–1344. [Google Scholar] [CrossRef]
- Stuendl, A.; Kunadt, M.; Kruse, N.; Bartels, C.; Moebius, W.; Danzer, K.M.; Mollenhauer, B.; Schneider, A. Induction of α-synuclein aggregate formation by CSF exosomes from patients with Parkinson’s disease and dementia with Lewy bodies. Brain 2016, 139, 481–494. [Google Scholar] [CrossRef] [PubMed]
- Kamerkar, S.; LeBleu, V.S.; Sugimoto, H.; Yang, S.; Ruivo, C.F.; Melo, S.A.; Lee, J.J.; Kalluri, R. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 2017, 546, 498–503. [Google Scholar] [CrossRef] [PubMed]
- Rual, J.F.; Venkatesan, K.; Hao, T.; Hirozane-Kishikawa, T.; Dricot, A.; Li, N.; Berriz, G.F.; Gibbons, F.D.; Dreze, M.; Ayivi-Guedehoussou, N.; et al. Towards a proteome-scale map of the human protein–protein interaction network. Nature 2005, 437, 1173–1178. [Google Scholar] [CrossRef] [PubMed]
- Uhlén, M.; Fagerberg, L.; Hallström, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, Å.; Kampf, C.; Sjöstedt, E.; Asplund, A.; et al. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef] [PubMed]
- Gysi, D.M.; Valle, Í.D.; Zitnik, M.; Ameli, A.; Gan, X.; Varol, O.; Sanchez, H.; Baron, R.M.; Ghiassian, D.; Loscalzo, J.; et al. Network medicine framework for identifying drug repurposing opportunities for COVID-19. arXiv 2020, arXiv:2004.07229. [Google Scholar]
- Keerthikumar, S.; Chisanga, D.; Ariyaratne, D.; Al Saffar, H.; Anand, S.; Zhao, K.; Samuel, M.; Pathan, M.; Jois, M.; Chilamkurti, N.; et al. ExoCarta: A web-based compendium of exosomal cargo. J. Mol. Biol. 2016, 428, 688–692. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2016, 45, gkw937. [Google Scholar] [CrossRef]
- Pinero, J.; Ramirez-Anguita, J.M.; Sauch-Pitarch, J.; Ronzano, F.; Centeno, E.; Sanz, F.; Furlong, L.I. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 2020, 48, D845–D855. [Google Scholar]
- Nagy, J.A.; Benjamin, L.; Zeng, H.; Dvorak, A.M.; Dvorak, H.F. Vascular permeability, vascular hyperpermeability and angiogenesis. Angiogenesis 2008, 11, 109–119. [Google Scholar] [CrossRef]
- Wen, H.; Zhan, L.; Chen, S.; Long, L.; Xu, E. Rab7 may be a novel therapeutic target for neurologic diseases as a key regulator in autophagy. J. Neurosci. Res. 2017, 95, 1993–2004. [Google Scholar] [CrossRef]
- Saridaki, T.; Nippold, M.; Dinter, E.; Roos, A.; Diederichs, L.; Fensky, L.; Schulz, J.B.; Falkenburger, B.H. FYCO1 mediates clearance of α-synuclein aggregates through a Rab7-dependent mechanism. J. Neurochem. 2018, 146, 474–492. [Google Scholar] [CrossRef]
- Miao, G.; Zhao, H.; Li, Y.; Ji, M.; Chen, Y.; Shi, Y.; Bi, Y.; Wang, P.; Zhang, H. ORF3a of the COVID-19 virus SARS-CoV-2 blocks HOPS complex-mediated assembly of the SNARE complex required for autolysosome formation. Dev. Cell 2020, 56, 427–442. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Camfield, R.; Gorski, S.M. The interplay between exosomes and autophagy—Partners in crime. J. Cell Sci. 2018, 131, jcs215210. [Google Scholar] [CrossRef]
- Gassen, N.C.; Papies, J.; Bajaj, T.; Dethloff, F.; Emanuel, J.; Weckmann, K.; Heinz, D.E.; Heinemann, N.; Lennarz, M.; Richter, A.; et al. Analysis of SARS-CoV-2-controlled autophagy reveals spermidine, MK-2206, and niclosamide as putative antiviral therapeutics. bioRxiv 2020. [Google Scholar] [CrossRef]
- Fecchi, K.; Anticoli, S.; Peruzzu, D.; Iessi, E.; Gagliardi, M.C.; Matarrese, P.; Ruggieri, A. Coronavirus Interplay with lipid rafts and autophagy unveils promising therapeutic targets. Front. Microbiol. 2020, 11, 1821. [Google Scholar] [CrossRef] [PubMed]
- Shojaei, S.; Suresh, M.; Klionsky, D.J.; Labouta, H.I.; Ghavami, S. Autophagy and SARS-CoV-2 infection: A possible smart targeting of the autophagy pathway. Virulence 2020. [Google Scholar] [CrossRef]
- Shin, W.H.; Park, J.H.; Chung, K.C. The central regulator p62 between ubiquitin proteasome system and autophagy and its role in the mitophagy and Parkinson’s disease. BMB Rep. 2020, 53, 56. [Google Scholar] [CrossRef]
- Anisimov, S.; Takahashi, M.; Kakihana, T.; Katsuragi, Y.; Kitaura, H.; Zhang, L.; Kakita, A.; Fujii, M. G3BP1 inhibits ubiquitinated protein aggregations induced by p62 and USP10. Sci. Rep. 2019, 9, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.K.; Mishra, S.; Rajput, C.; Ur Rasheed, M.S.; Patel, D.K.; Singh, M.P. Cypermethrin activates autophagosome formation albeit inhibits autophagy owing to poor lysosome quality: Relevance to Parkinson’s disease. Neurotox. Res. 2018, 33, 377–387. [Google Scholar] [CrossRef] [PubMed]
- Mamais, A.; Manzoni, C.; Nazish, I.; Arber, C.; Sonustun, B.; Wray, S.; Warner, T.T.; Cookson, M.R.; Lewis, P.A.; Bandopadhyay, R. Analysis of macroautophagy related proteins in G2019S LRRK2 Parkinson’s disease brains with Lewy body pathology. Brain Res. 2018, 1701, 75–84. [Google Scholar] [CrossRef]
- Nogalska, A.; Terracciano, C.; D’Agostino, C.; Engel, W.K.; Askanas, V. p62/SQSTM1 is overexpressed and prominently accumulated in inclusions of sporadic inclusion-body myositis muscle fibers, and can help differentiating it from polymyositis and dermatomyositis. Acta Neuropathol. 2009, 118, 407–413. [Google Scholar] [CrossRef]
- Nabeel-Shah, S.; Lee, H.; Ahmed, N.; Marcon, E.; Farhangmehr, S.; Pu, S.; Burke, G.L.; Ashraf, K.; Wei, H.; Zhong, G.; et al. SARS-CoV-2 Nucleocapsid protein attenuates stress granule formation and alters gene expression via direct interaction with host mRNAs. bioRxiv 2020. [Google Scholar] [CrossRef]
- McBride, R.; Van Zyl, M.; Fielding, B.C. The coronavirus nucleocapsid is a multifunctional protein. Viruses 2014, 6, 2991–3018. [Google Scholar] [CrossRef]
- Liu, W.; Liu, L.; Kou, G.; Zheng, Y.; Ding, Y.; Ni, W.; Wang, Q.; Tan, L.; Wu, W.; Tang, S.; et al. Evaluation of nucleocapsid and spike protein-based enzyme-linked immunosorbent assays for detecting antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020, 58. [Google Scholar] [CrossRef]
- Chang, C.k.; Hou, M.H.; Chang, C.F.; Hsiao, C.D.; Huang, T.H. The SARS coronavirus nucleocapsid protein–forms and functions. Antivir. Res. 2014, 103, 39–50. [Google Scholar] [CrossRef]
- Ruiz-Martinez, J.; Krebs, C.E.; Makarov, V.; Gorostidi, A.; Martí-Massó, J.F.; Paisán-Ruiz, C. GIGYF2 mutation in late-onset Parkinson’s disease with cognitive impairment. J. Hum. Genet. 2015, 60, 637–640. [Google Scholar] [CrossRef]
- Al Yemni, E.; Monies, D.; Alkhairallah, T.; Bohlega, S.; Abouelhoda, M.; Magrashi, A.; Mustafa, A.; AlAbdulaziz, B.; Alhamed, M.; Baz, B.; et al. Integrated analysis of whole exome sequencing and copy number evaluation in Parkinson’s disease. Sci. Rep. 2019, 9, 1–9. [Google Scholar]
- Marchese, D.; Botta-Orfila, T.; Cirillo, D.; Rodriguez, J.A.; Livi, C.M.; Fernández-Santiago, R.; Ezquerra, M.; Martí, M.J.; Bechara, E.; Tartaglia, G.G.; et al. Discovering the 3’ UTR-mediated regulation of alpha-synuclein. Nucleic Acids Res. 2017, 45, 12888–12903. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Liu, Y.; Chen, T. Identification of key genes and pathways in Parkinson’s disease through integrated analysis. Mol. Med. Rep. 2017, 16, 3769–3776. [Google Scholar] [CrossRef] [PubMed]
- Chi, L.M.; Wang, L.P.; Jiao, D. Identification of differentially expressed genes and long noncoding RNAs associated with Parkinson’s disease. Parkinson’s Dis. 2019, 2019, 6078251. [Google Scholar] [CrossRef]
- Zeng, R.; Luo, D.X.; Li, H.P.; Zhang, Q.S.; Lei, S.S.; Chen, J.H. MicroRNA-135b alleviates MPP+-mediated Parkinson’s disease in in vitro model through suppressing FoxO1-induced NLRP3 inflammasome and pyroptosis. J. Clin. Neurosci. 2019, 65, 125–133. [Google Scholar] [CrossRef]
- Chang, M.Y.; Oh, B.; Choi, J.E.; Sulistio, Y.A.; Woo, H.J.; Jo, A.; Kim, J.; Kim, E.H.; Kim, S.W.; Hwang, J.; et al. LIN 28A loss of function is associated with Parkinson’s disease pathogenesis. EMBO J. 2019, 38, e101196. [Google Scholar] [CrossRef]
- Soyal, S.M.; Zara, G.; Ferger, B.; Felder, T.K.; Kwik, M.; Nofziger, C.; Dossena, S.; Schwienbacher, C.; Hicks, A.A.; Pramstaller, P.P.; et al. The PPARGC1A locus and CNS-specific PGC-1α isoforms are associated with Parkinson’s Disease. Neurobiol. Dis. 2019, 121, 34–46. [Google Scholar] [CrossRef]
- Sardoiwala, M.N.; Srivastava, A.K.; Kaundal, B.; Karmakar, S.; Choudhury, S.R. Recuperative effect of metformin loaded polydopamine nanoformulation promoting EZH2 mediated proteasomal degradation of phospho-α-synuclein in Parkinson’s disease model. Nanomed. Nanotechnol. Biol. Med. 2020, 24, 102088. [Google Scholar] [CrossRef]
- Arlehamn, C.S.L.; Pham, J.; Alcalay, R.N.; Frazier, A.; Shorr, E.; Carpenter, C.; Sidney, J.; Dhanwani, R.; Agin-Liebes, J.; Garretti, F.; et al. Widespread Tau-specific CD4 T cell reactivity in the general population. J. Immunol. 2019, 203, 84–92. [Google Scholar] [CrossRef]
- Craven, K.M.; Kochen, W.R.; Hernandez, C.M.; Flinn, J.M. Zinc exacerbates tau pathology in a tau mouse model. J. Alzheimer’s Dis. 2018, 64, 617–630. [Google Scholar] [CrossRef]
- Li, Y.I.; Wong, G.; Humphrey, J.; Raj, T. Prioritizing Parkinson’s disease genes using population-scale transcriptomic data. Nat. Commun. 2019, 10, 1–10. [Google Scholar] [CrossRef]
- Kumar, A.S.; Jagadeeshan, S.; Subramanian, A.; Chidambaram, S.B.; Surabhi, R.P.; Singhal, M.; Bhoopalan, H.; Sekar, S.; Pitani, R.S.; Duvuru, P.; et al. Molecular mechanism of regulation of MTA1 expression by granulocyte Colony-stimulating factor. J. Biol. Chem. 2016, 291, 12310–12321. [Google Scholar] [CrossRef] [PubMed]
- Uenaka, T.; Satake, W.; Cha, P.C.; Hayakawa, H.; Baba, K.; Jiang, S.; Kobayashi, K.; Kanagawa, M.; Okada, Y.; Mochizuki, H.; et al. In silico drug screening by using genome-wide association study data repurposed dabrafenib, an anti-melanoma drug, for Parkinson’s disease. Hum. Mol. Genet. 2018, 27, 3974–3985. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, H. ER stress and diseases. FEBS J. 2007, 274, 630–658. [Google Scholar] [CrossRef] [PubMed]
- Zinzula, L. Lost in deletion: The enigmatic ORF8 protein of SARS-CoV-2. Biochem. Biophys. Res. Commun. 2020, 538, 116–124. [Google Scholar] [CrossRef]
- Wang, Q.; Jiao, F.; Zhang, P.; Yan, J.; Zhang, Z.; He, F.; Zhang, Q.; Lv, Z.; Peng, X.; Cai, H.; et al. CDK5-mediated phosphorylation-dependent ubiquitination and degradation of E3 ubiquitin ligases GP78 accelerates neuronal death in Parkinson’s disease. Mol. Neurobiol. 2018, 55, 3709–3717. [Google Scholar] [CrossRef]
- Ranjan, P.; Kumar, A. The Involvement of His50 during Protein Disulfide Isomerase Binding Is Essential for Inhibiting α-Syn Fibril Formation. Biochemistry 2016, 55, 2677–2680. [Google Scholar] [CrossRef]
- Omura, T.; Matsuda, H.; Nomura, L.; Imai, S.; Denda, M.; Nakagawa, S.; Yonezawa, A.; Nakagawa, T.; Yano, I.; Matsubara, K. Ubiquitin ligase HMG-CoA reductase degradation 1 (HRD1) prevents cell death in a cellular model of Parkinson’s disease. Biochem. Biophys. Res. Commun. 2018, 506, 516–521. [Google Scholar] [CrossRef]
- Al-Obeidi, E.; Al-Tahan, S.; Surampalli, A.; Goyal, N.; Wang, A.K.; Hermann, A.; Omizo, M.; Smith, C.; Mozaffar, T.; Kimonis, V. Genotype-phenotype study in patients with valosin-containing protein mutations associated with multisystem proteinopathy. Clin. Genet. 2018, 93, 119–125. [Google Scholar] [CrossRef]
- Santiago, J.A.; Potashkin, J.A. Integrative network analysis unveils convergent molecular pathways in Parkinson’s disease and diabetes. PLoS ONE 2013, 8, e83940. [Google Scholar] [CrossRef]
- Pan, H.; Zhao, Y.; Zhai, Z.; Zheng, J.; Zhou, Y.; Zhai, Q.; Cao, X.; Tian, J.; Zhao, L. Role of plasminogen activator inhibitor-1 in the diagnosis and prognosis of patients with Parkinson’s disease. Exp. Ther. Med. 2018, 15, 5517–5522. [Google Scholar] [CrossRef]
- Masuda, T.; Itoh, J.; Koide, T.; Tomidokoro, Y.; Takei, Y.; Ishii, K.; Tamaoka, A. Transforming growth factor-β1 in the cerebrospinal fluid of patients with distinct neurodegenerative diseases. J. Clin. Neurosci. 2017, 35, 47–49. [Google Scholar] [CrossRef][Green Version]
- Koike, H.; Ishida, A.; Shimamura, M.; Mizuno, S.; Nakamura, T.; Ogihara, T.; Kaneda, Y.; Morishita, R. Prevention of onset of Parkinson’s disease by in vivo gene transfer of human hepatocyte growth factor in rodent model: A model of gene therapy for Parkinson’s disease. Gene Ther. 2006, 13, 1639–1644. [Google Scholar] [CrossRef][Green Version]
- Chen, Y.C.; Wu, Y.R.; Mesri, M.; Chen, C.M. Associations of matrix metalloproteinase-9 and tissue inhibitory factor-1 polymorphisms with Parkinson disease in Taiwan. Medicine 2016, 95, e2672. [Google Scholar] [CrossRef]
| Protein | Biological Function | Protein Expression | RNA Expression (pTPM) |
|---|---|---|---|
| BRD4 | - | high | 29.6 |
| CEP350 | centrosome | high | 11.5 |
| CSNK2A2 | stress granules | - | 40.9 |
| ECSIT | respiratory electron transport | medium | 30.8 |
| G3BP1 | stress granules | medium | 41.0 |
| HDAC2 | - | high | 44.6 |
| ITGB1 | - | - | 317.6 |
| LARP7 | 7SK snRNP | - | 49.0 |
| MARK2 | MARK kinases | - | 18.5 |
| NUP88 | nuclear pore | medium | 11.1 |
| NUTF2 | - | low | 62.0 |
| OS9 | ER protein quality control | high | 179.9 |
| PRKACA | protein kinase A signaling | medium | 68.5 |
| PRKAR2A | protein kinase A signaling | high | 13.0 |
| PRKAR2B | protein kinase A signaling | low | 11.2 |
| RAB1A | Rab signaling | low | 167.3 |
| RAB14 | Rab signaling | medium | 55.6 |
| RAE1 | nuclear pore | - | 15.3 |
| RHOA | - | medium | 554.4 |
| RTN4 | ER morphology | - | 300.8 |
| SCCPDH | - | low | 20.3 |
| VPS11 | HOPS complex | medium | - |
| VPS39 | HOPS complex | - | 30.6 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Estrada, E. Cascading from SARS-CoV-2 to Parkinson’s Disease through Protein-Protein Interactions. Viruses 2021, 13, 897. https://doi.org/10.3390/v13050897
Estrada E. Cascading from SARS-CoV-2 to Parkinson’s Disease through Protein-Protein Interactions. Viruses. 2021; 13(5):897. https://doi.org/10.3390/v13050897
Chicago/Turabian StyleEstrada, Ernesto. 2021. "Cascading from SARS-CoV-2 to Parkinson’s Disease through Protein-Protein Interactions" Viruses 13, no. 5: 897. https://doi.org/10.3390/v13050897
APA StyleEstrada, E. (2021). Cascading from SARS-CoV-2 to Parkinson’s Disease through Protein-Protein Interactions. Viruses, 13(5), 897. https://doi.org/10.3390/v13050897






