Transcriptome Analysis for Genes Associated with Small Ruminant Lentiviruses Infection in Goats of Carpathian Breed
Abstract
1. Introduction
2. Methods
2.1. Animals and Blood Sample Collection
2.2. Proviral Load Quantification
2.3. Transcriptome Sequencing and Data Analysis
2.4. GO Enrichment and Pathways Analysis
2.5. Quantitative Polymerase Chain Reaction (qPCR) Analysis
2.6. Statistical Analysis
3. Results
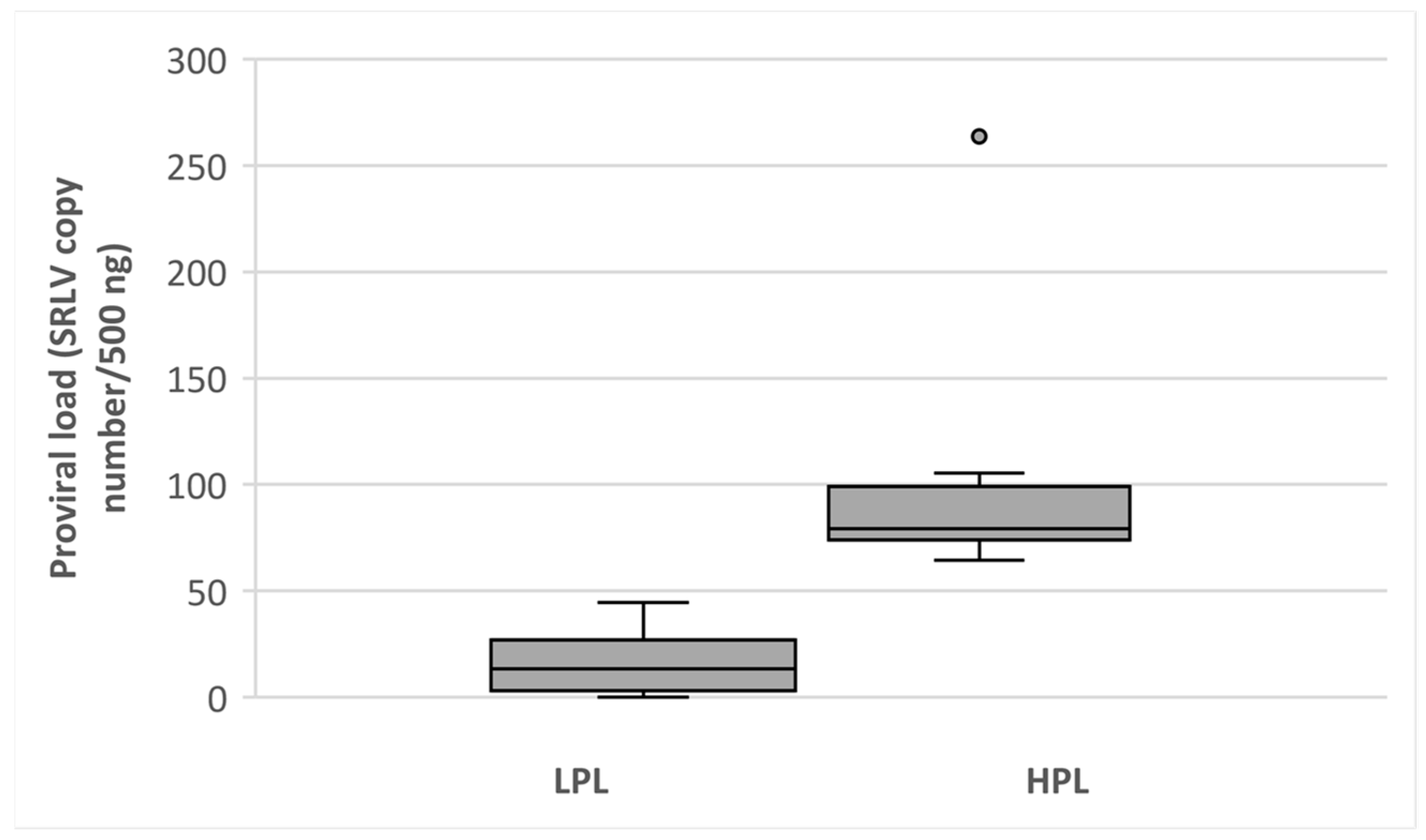
3.1. Classification of Goats on HPL and LPL
3.2. RNA-seq Data
3.2.1. Transcriptome Quantification
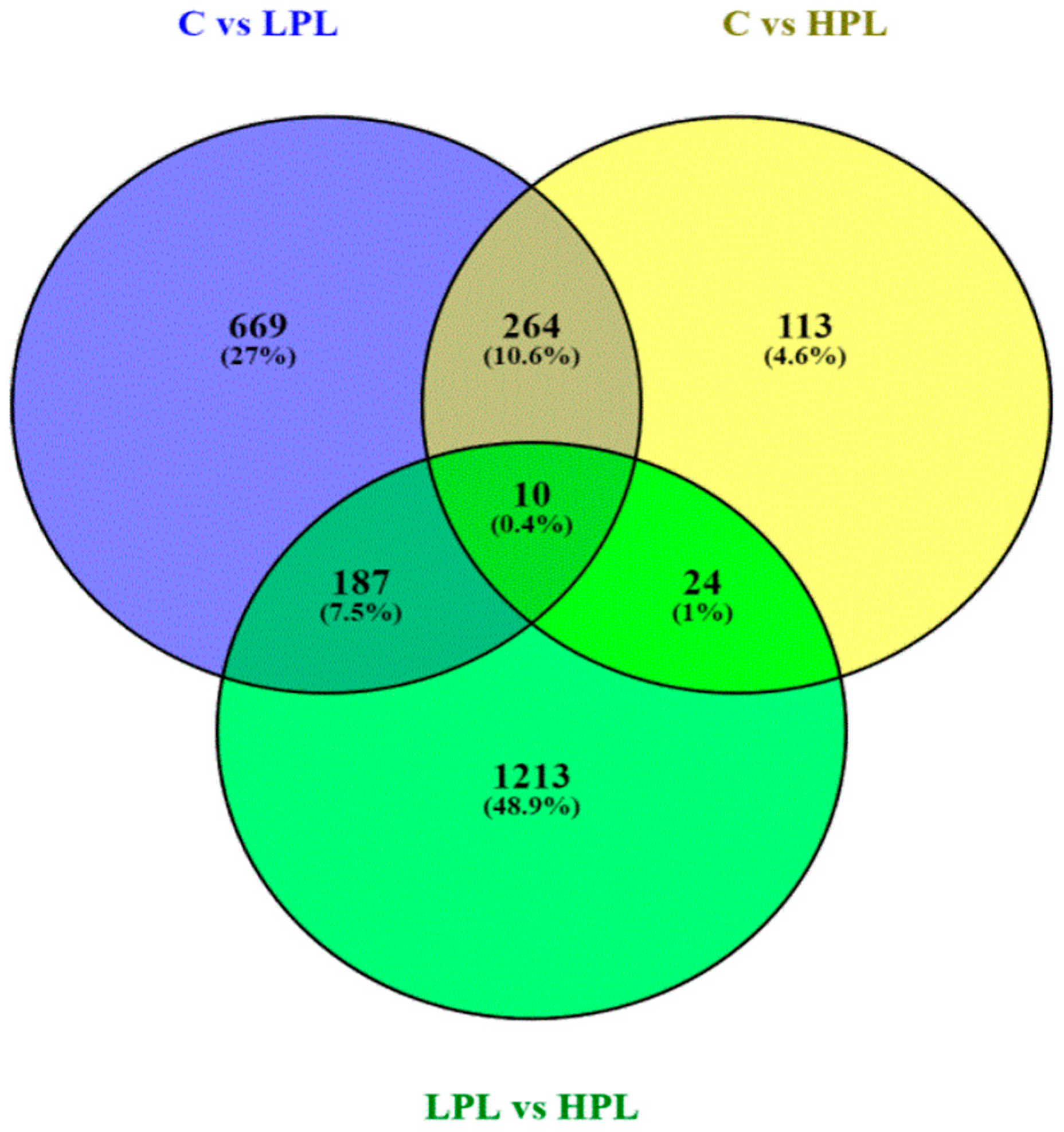
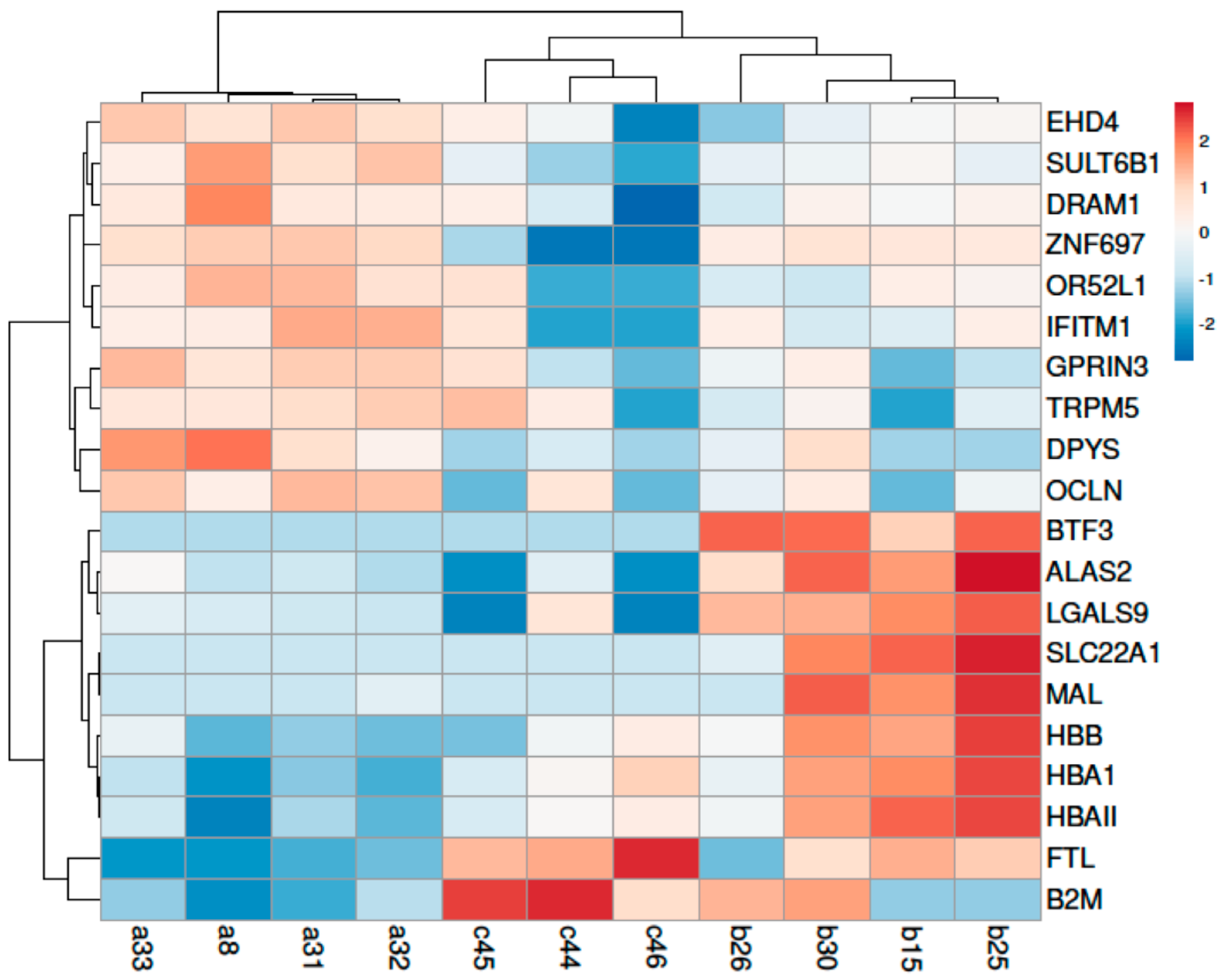
3.2.2. DEGs Analysis
3.3. Gene Ontology and Pathways Annotation
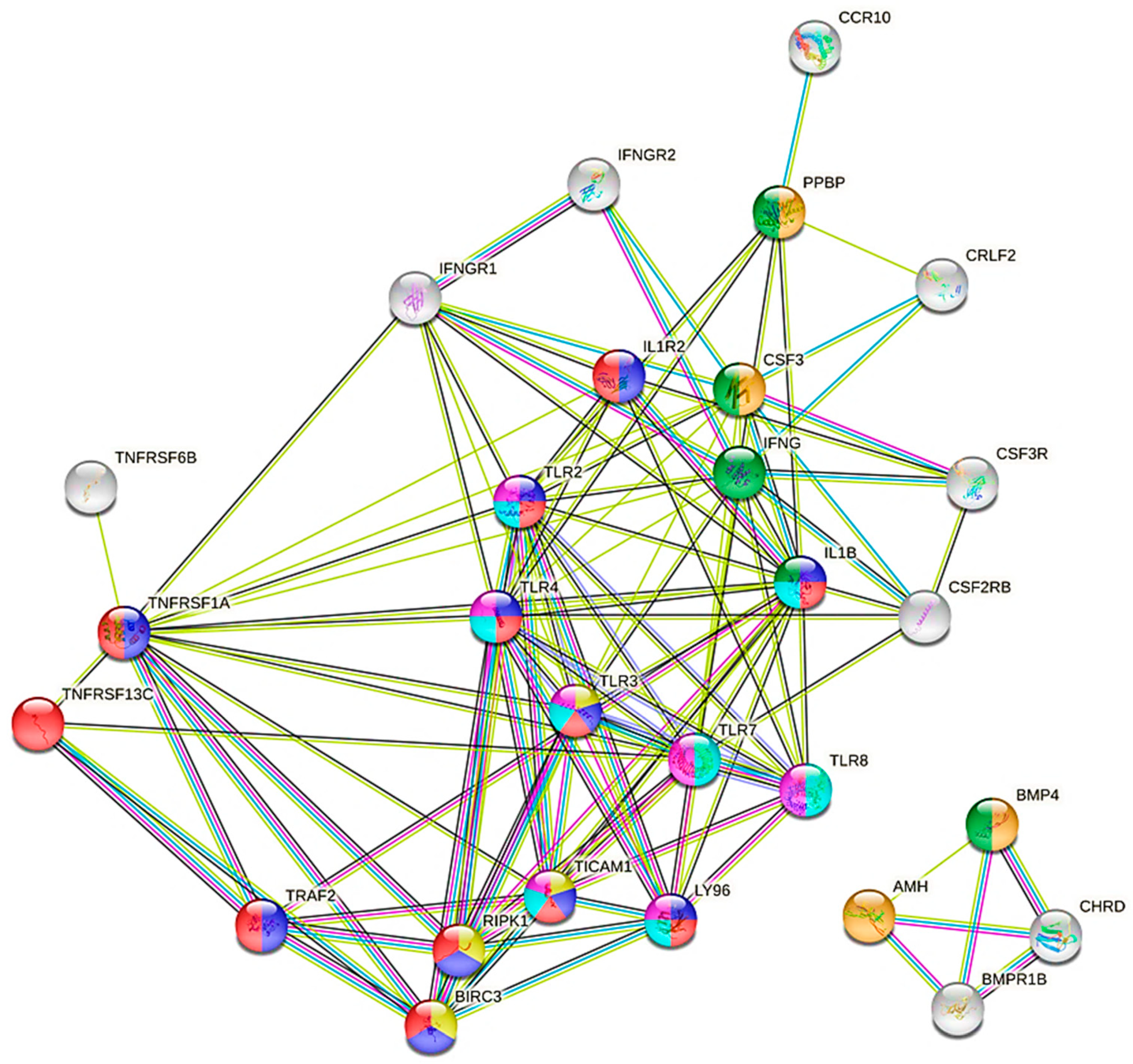
3.3.1. DEGs Detected between Control Group and Groups with Proviral Load
3.3.2. DEGs Detected between LPL and HPL Groups
3.4. qPCR Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Minguijón, E.; Reina, R.; Pérez, M.; Polledo, L.; Villoria, M.; Ramírez, H.; Leginagoikoa, I.; Badiola, J.J.; García-Marín, J.F.; de Andrés, D.; et al. Small ruminant lentivirus infections and diseases. Vet. Microbiol. 2015, 181, 75–89. [Google Scholar] [CrossRef] [PubMed]
- Blacklaws, B.A. Small ruminant lentiviruses: Immunopathogenesis of visna-maedi and caprine arthritis and encephalitis virus. Comp. Immunol. Microbiol. Infect. Dis. 2012, 35, 259–269. [Google Scholar] [CrossRef] [PubMed]
- Peterhans, E.; Greenland, T.; Badiola, J.; Harkiss, G.; Bertoni, G.; Amorena, B.; Eliaszewicz, M.; Juste, R.A.; Krassnig, R.; Lafont, J.P.; et al. Routes of transmission and consequences of small ruminant lentiviruses (SRLVs) infection and eradication schemes. Vet. Res. 2004, 35, 257–274. [Google Scholar] [CrossRef]
- Martínez-Navalón, B.; Peris, C.; Gómez, E.A.; Peris, B.; Roche, M.L.; Caballero, C.; Goyena, E.; Berriatua, E. Quantitative estimation of the impact of caprine arthritis encephalitis virus infection on milk production by dairy goats. Vet. J. 2013, 197, 311–317. [Google Scholar] [CrossRef]
- Scheer-Czechowski, P.; Vogt, H.R.; Tontis, A.; Peterhans, E.; Zanoni, R. Pilotprojekt zur Sanierung der Maedi-Visna bei Walliser Schwarznasenschafen (Pilot project for eradicating maedi-visna in Walliser blacknose sheep). Schweiz. Arch. Tierheilkd. 2000, 142, 155–164. [Google Scholar] [PubMed]
- Kalogianni, A.I.; Bossis, I.; Ekateriniadou, L.V.; Gelasakis, A.I. Etiology, Epizootiology and Control of Maedi-Visna in Dairy Sheep: A Review. Animals 2020, 10, 616. [Google Scholar] [CrossRef]
- Tavella, A.; Bettini, A.; Ceol, M.; Zambotto, P.; Stifter, E.; Kusstatscher, N.; Lombardi, R.; Nardeli, S.; Beato, M.S.; Capello, K.; et al. Achievements of an eradication programme against caprine arthritis encephalitis virus in South Tyrol, Italy. Vet. Rec. 2018, 182, 51. [Google Scholar] [CrossRef]
- Reina, R.; Berriatua, E.; Luján, L.; Juste, R.; Sánchez, A.; de Andrés, D.; Amorena, B. Prevention strategies against small ruminant lentiviruses: An update. Vet. J. 2009, 182, 31–37. [Google Scholar] [CrossRef]
- Cirone, F.; Maggiolino, A.; Cirilli, M.; Sposato, A.; De Palo, P.; Ciappetta, G.; Pratelli, A. Small ruminant lentiviruses in goats in southern Italy: Serological evidence, risk factors and implementation of control programs. Vet. Microbiol. 2019, 228, 143–146. [Google Scholar] [CrossRef]
- Gjerset, B.; Rimstad, E.; Teige, J.; Soetaert, K.; Jonassen, C.M. Impact of natural sheep-goat transmission on detection and control of small ruminant lentivirus group C infections. Vet. Microbiol. 2009, 135, 231–238. [Google Scholar] [CrossRef]
- Herrmann-Hoesing, L.M.; White, S.N.; Mousel, M.R.; Lewis, G.S.; Knowles, D.P. Ovine progressive pneumonia provirus levels associate with breed and Ovar-DRB1. Immunogenetics 2008, 60, 749–758. [Google Scholar] [CrossRef]
- Larruskain, A.; Bernales, I.; Luján, L.; de Andrés, D.; Amorena, B.; Jugo, B.M. Expression analysis of 13 ovine immune response candidate genes in Visna/Maedi disease progression. Comp. Immunol. Microbiol. Infect. Dis. 2013, 36, 405–413. [Google Scholar] [CrossRef]
- White, S.N.; Knowles, D.P. Expanding possibilities for intervention against small ruminant lentiviruses through genetic marker-assisted selective breeding. Viruses 2013, 5, 1466–1499. [Google Scholar] [CrossRef]
- Cecchi, F.; Dadousis, C.; Bozzi, R.; Fratini, F.; Russo, C.; Bandecchi, P.; Cantile, C.; Mazzei, M. Genome scan for the possibility of identifying candidate resistance genes for goat lentiviral infections in the Italian Garfagnina goat breed. Trop. Anim. Health Prod. 2019, 51, 729–733. [Google Scholar] [CrossRef]
- White, S.N.; Mousel, M.R.; Herrmann-Hoesing, L.M.; Reynolds, J.O.; Leymaster, K.A.; Neibergs, H.L.; Lewis, G.S.; Knowles, D.P. Genome-wide association identifies multiple genomic regions associated with susceptibility to and control of ovine lentivirus. PLoS ONE 2012, 7, e47829. [Google Scholar] [CrossRef]
- Alshanbari, F.A.; Mousel, M.R.; Reynolds, J.O.; Herrmann-Hoesing, L.M.; Highland, M.A.; Lewis, G.S.; White, S.N. Mutations in Ovis aries TMEM154 are associated with lower small ruminant lentivirus proviral concentration in one sheep flock. Anim. Genet. 2014, 45, 565–571. [Google Scholar] [CrossRef]
- Colussi, S.; Desiato, R.; Beltramo, C.; Peletto, S.; Modesto, P.; Maniaci, M.G.; Campia, V.; Quasso, A.; Rosati, S.; Bertolotti, L.; et al. A single nucleotide variant in the promoter region of the CCR5 gene increases susceptibility to arthritis encephalitis virus in goats. BMC Vet. Res. 2019, 15, 230. [Google Scholar] [CrossRef] [PubMed]
- Molaee, V.; Eltanany, M.; Lühken, G. First survey on association of TMEM154 and CCR5 variants with serological maedi-visna status of sheep in German flocks. Vet. Res. 2018, 49, 36. [Google Scholar] [CrossRef]
- Molaee, V.; Otarod, V.; Abdollahi, D.; Lühken, G. Lentivirus Susceptibility in Iranian and German Sheep Assessed by Determination of TMEM154 E35K. Animals 2019, 9, 685. [Google Scholar] [CrossRef] [PubMed]
- Jarczak, J.; Kaba, J.; Reczyńska, D.; Bagnicka, E. Impaired Expression of Cytokines as a Result of Viral Infections with an Emphasis on Small Ruminant Lentivirus Infection in Goats. Viruses 2016, 8, 186. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.; Ma, F.; Quinn, M.; Xiang, S.H. Genome-Wide Search for Host Association Factors during Ovine Progressive Pneumonia Virus Infection. PLoS ONE 2016, 11, e0150344. [Google Scholar] [CrossRef]
- Ramírez, H.; Echeverría, I.; Benito, A.A.; Glaria, I.; Benavides, J.; Pérez, V.; de Andrés, D.; Reina, R. Accurate Diagnosis of Small Ruminant Lentivirus Infection Is Needed for Selection of Resistant Sheep through TMEM154 E35K Genotyping. Pathogens 2021, 10, 83. [Google Scholar] [CrossRef]
- Jáuregui, P.; Crespo, H.; Glaria, I.; Luján, L.; Contreras, A.; Rosati, S.; de Andrés, D.; Amorena, B.; Towers, G.J.; Reina, R. Ovine TRIM5α can restrict visna/maedi virus. J. Virol. 2012, 86, 9504–9509. [Google Scholar] [CrossRef]
- White, S.N.; Mousel, M.R.; Reynolds, J.O.; Lewis, G.S.; Herrmann-Hoesing, L.M. Common promoter deletion is associated with 3.9-fold differential transcription of ovine CCR5 and reduced proviral level of ovine progressive pneumonia virus. Anim. Genet. 2009, 40, 583–589. [Google Scholar] [CrossRef]
- Bowles, D.; Carson, A.; Isaac, P. Genetic Distinctiveness of the Herdwick Sheep Breed and Two Other Locally Adapted Hill Breeds of the UK. PLoS ONE 2014, 9, e87823. [Google Scholar] [CrossRef] [PubMed]
- Heaton, M.P.; Clawson, M.L.; Chitko-Mckown, C.G.; Leymaster, K.A.; Smith, T.P.; Harhay, G.P.; White, S.N.; Herrmann-Hoesing, L.M.; Mousel, M.R.; Lewis, G.S.; et al. Reduced lentivirus susceptibility in sheep with TMEM154 mutations. PLoS Genet. 2012, 8, e1002467. [Google Scholar] [CrossRef] [PubMed]
- Heaton, M.P.; Kalbfleisch, T.S.; Petrik, D.T.; Simpson, B.; Kijas, J.W.; Clawson, M.L.; Chitko-McKown, C.G.; Harhay, G.P.; Leymaster, K.A.; International Sheep Genomics Consortium. Genetic testing for TMEM154 mutations associated with lentivirus susceptibility in sheep. PLoS ONE 2013, 8, e55490. [Google Scholar] [CrossRef]
- Clawson, M.L.; Redden, R.; Schuller, G.; Heaton, M.P.; Workman, A.; Chitko-McKown, C.G.; Smith, T.P.; Leymaster, K.A. Genetic subgroup of small ruminant lentiviruses that infects sheep homozygous for TMEM154 frameshift deletion mutation A4Δ53. Vet. Res. 2015, 46, 22. [Google Scholar] [CrossRef] [PubMed]
- Sider, L.H.; Heaton, M.P.; Chitko-McKown, C.G.; Harhay, G.P.; Smith, T.P.; Leymaster, K.A.; Laegreid, W.W.; Clawson, M.L. Small ruminant lentivirus genetic subgroups associate with sheep TMEM154 genotypes. Vet. Res. 2013, 44, 64. [Google Scholar] [CrossRef]
- Larruskain, A.; Minguijón, E.; García-Etxebarria, K.; Moreno, B.; Arostegui, I.; Juste, R.A.; Jugo, B.M. MHC class II DRB1 gene polymorphism in the pathogenesis of Maedi-Visna and pulmonary adenocarcinoma viral diseases in sheep. Immunogenetics 2010, 62, 75–83. [Google Scholar] [CrossRef]
- Yaman, Y.; Keleş, M.; Aymaz, R.; Sevim, S.; Sezenler, T.; Önaldı, A.T.; Kaptan, C.; Baskurt, A.; Koncagül, S.; Öner, Y.; et al. Association of TMEM154 variants with visna/maedi virus infection in Turkish sheep. Small Rumin. Res. 2019, 177, 61–67. [Google Scholar] [CrossRef]
- Leymaster, K.A.; Chitko-McKown, C.G.; Clawson, M.L.; Harhay, G.P.; Heaton, M.P. Effects of TMEM154 haplotypes 1 and 3 on susceptibility to ovine progressive pneumonia virus following natural exposure in sheep. J. Anim. Sci. 2013, 91, 5114–5121. [Google Scholar] [CrossRef]
- Abendaño, N.; Esparza-Baquer, A.; Bernales, I.; Reina, R.; de Andrés, D.; Jugo, B.M. Gene Expression Profiling Reveals New Pathways and Genes Associated with Visna/Maedi Viral Disease. Animals 2021, 11, 1785. [Google Scholar] [CrossRef] [PubMed]
- Panei, C.J.; Takeshima, S.N.; Omori, T.; Nunoya, T.; Davis, W.C.; Ishizaki, H.; Matoba, K.; Aida, Y. Estimation of bovine leukemia virus (BLV) proviral load harbored by lymphocyte subpopulations in BLV-infected cattle at the subclinical stage of enzootic bovine leucosis using BLV-CoCoMo-qPCR. BMC Vet. Res. 2013, 9, 95. [Google Scholar] [CrossRef] [PubMed]
- Ravazzolo, A.P.; Nenci, C.; Vogt, H.R.; Waldvogel, A.; Obexer-Ruff, G.; Peterhans, E.; Bertoni, G. Viral load, organ distribution, histopathological lesions, and cytokine mRNA expression in goats infected with a molecular clone of the caprine arthritis encephalitis virus. Virology 2006, 350, 116–127. [Google Scholar] [CrossRef] [PubMed]
- Herrmann-Hoesing, L.M.; Noh, S.M.; White, S.N.; Snekvik, K.R.; Truscott, T.; Knowles, D.P. Peripheral ovine progressive pneumonia provirus levels correlate with and predict histological tissue lesion severity in naturally infected sheep. Clin. Vaccine Immunol. 2009, 16, 551–557. [Google Scholar] [CrossRef][Green Version]
- Crespo, H.; Bertolotti, L.; Proffiti, M.; Cascio, P.; Cerruti, F.; Acutis, P.L.; de Andrés, D.; Reina, R.; Rosati, S. Low proviral small ruminant lentivirus load as biomarker of natural restriction in goats. Vet. Microbiol. 2016, 192, 152–162. [Google Scholar] [CrossRef]
- Stonos, N.; Wootton, S.K.; Karrow, N. Immunogenetics of small ruminant lentiviral infections. Viruses 2014, 6, 3311–3333. [Google Scholar] [CrossRef]
- Olech, M.; Kuźmak, J. Molecular Characterization of Small Ruminant Lentiviruses of Subtype A5 Detected in Naturally Infected but Clinically Healthy Goats of Carpathian Breed. Pathogens 2020, 9, 992. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 22 June 2020).
- Dodt, M.; Roehr, J.T.; Ahmed, R.; Dieterich, C. FLEXBAR-Flexible Barcode and Adapter Processing for Next-Generation Sequencing Platforms. Biology 2012, 1, 895–905. [Google Scholar] [CrossRef]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering splice junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Sahu, A.R.; Wani, S.A.; Saxena, S.; Rajak, K.K.; Chaudhary, D.; Sahoo, A.P.; Khanduri, A.; Pandey, A.; Mondal, P.; Malla, W.A.; et al. Selection and validation of suitable reference genes for qPCR gene expression analysis in goats and sheep under Peste des petits ruminants virus (PPRV), lineage IV infection. Sci. Rep. 2018, 8, 15969. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: http://www.R-project.org/ (accessed on 22 June 2020).
- Swaminathan, S.; Qiu, J.; Rupert, A.W.; Hu, Z.; Higgins, J.; Dewar, R.L.; Stevens, R.; Rehm, C.A.; Metcalf, J.A.; Sherman, B.T.; et al. Interleukin-15 (IL-15) Strongly Correlates with Increasing HIV-1 Viremia and Markers of Inflammation. PLoS ONE 2016, 11, e0167091. [Google Scholar] [CrossRef]
- Jarczak, J.; Słoniewska, D.; Kaba, J.; Bagnicka, E. The expression of cytokines in the milk somatic cells, blood leukocytes and serum of goats infected with small ruminant lentivirus. BMC Vet. Res. 2019, 15, 424. [Google Scholar] [CrossRef]
- Di Paolo, N.C.; Shayakhmetov, D.M. Interleukin 1α and the inflammatory process. Nat. Immunol. 2016, 17, 906–913. [Google Scholar] [CrossRef]
- Gabay, C.; Lamacchia, C.; Palmer, G. IL-1 pathways in inflammation and human diseases. Nat. Rev. Rheumatol 2010, 6, 232–241. [Google Scholar] [CrossRef]
- Zhang, Z.; Watt, N.J.; Hopkins, J.; Harkiss, G.; Woodall, C.J. Quantitative analysis of maedi-visna virus DNA load in peripheral blood monocytes and alveolar macrophages. J. Virol. Methods. 2000, 86, 13–20. [Google Scholar] [CrossRef]
- Mikula, I.; Bhide, M.; Pastorekova, S.; Mikula, I. Characterization of ovine TLR7 and TLR8 protein coding regions, detection of mutations and Maedi Visna virus infection. Vet. Immunol. Immunopathol. 2010, 138, 51–59. [Google Scholar] [CrossRef]
- Olech, M.; Ropka-Molik, K.; Szmatoła, T.; Piórkowska, K.; Kuźmak, J. Single Nucleotide Polymorphisms in Genes Encoding Toll-Like Receptors 7 and 8 and Their Association with Proviral Load of SRLVs in Goats of Polish Carpathian Breed. Animals 2021, 11, 1908. [Google Scholar] [CrossRef]
- Ge, Y.; Mansell, A.; Ussher, J.E.; Brooks, A.E.; Manning, K.; Wang, C.J.; Taylor, J.A. Rotavirus NSP4 Triggers Secretion of Proinflammatory Cytokines from Macrophages via Toll-Like Receptor 2. J. Virol. 2013, 87, 11160–11167. [Google Scholar] [CrossRef] [PubMed]
- Lester, S.N.; Li, K. Toll-like receptors in antiviral innate immunity. J. Mol. Biol. 2014, 426, 1246–1264. [Google Scholar] [CrossRef] [PubMed]
- Oliveira-Nascimento, L.; Massari, P.; Wetzler, L.M. The Role of TLR2 in Infection and Immunity. Front. Immunol. 2012, 3, 79. [Google Scholar] [CrossRef] [PubMed]
- Hernández, J.C.; Stevenson, M.; Latz, E.; Urcuqui-Inchima, S. HIV type 1 infection upregulates TLR2 and TLR4 expression and function in vivo and in vitro. AIDS Res. Hum. Retroviruses 2012, 28, 1313–1328. [Google Scholar] [CrossRef] [PubMed]
- Lechner, F.; Vogt, H.R.; Seow, H.F.; Bertoni, G.; Cheevers, W.P.; von Bodungen, U.; Zurbriggen, A.; Peterhans, E. Expression of cytokine mRNA in lentivirus-induced arthritis. Am. J. Pathol. 1997, 151, 1053–1065. [Google Scholar]
- Lechner, F.; Machado, J.; Bertoni, G.; Seow, H.F.; Dobbelaere, D.A.; Peterhans, E. Caprine arthritis encephalitis virus dysregulates the expression of cytokines in macrophages. J. Virol. 1997, 71, 7488–7497. [Google Scholar] [CrossRef]
- Gómez-Lucía, E.; Collado, V.M.; Miró, G.; Doménech, A. Effect of type-I interferon on retroviruses. Viruses 2009, 1, 545–573. [Google Scholar] [CrossRef]
- White, S.N.; Mousel, M.R.; Reynolds, J.O.; Herrmann-Hoesing, L.M.; Knowles, D.P. Deletion variant near ZNF389 is associated with control of ovine lentivirus in multiple sheep flocks. Anim. Genet. 2014, 45, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Chen, G.; Lv, F.; Wang, X.; Ji, X.; Xu, Y.; Sun, J.; Wu, L.; Zheng, Y.T.; Gao, G. Zinc-finger antiviral protein inhibits HIV-1 infection by selectively targeting multiply spliced viral mRNAs for degradation. Proc. Natl. Acad. Sci. USA 2011, 108, 15834–15839. [Google Scholar] [CrossRef] [PubMed]
| GO | Accession Number | Number of Genes | FDR | Identified Genes |
|---|---|---|---|---|
| MyD88-dependent toll-like receptor signaling pathway | GO: 0002755 | 4 | 0.019 | TLR2, TLR4, TLR8, TLR7 |
| Response to growth factor | GO: 0070848 | 5 | 0.050 | HHIP, BMPR1B, ADAMTS3, PDGFRB, BMP6 |
| Regulation of cytokine secretion | GO: 0001817 | 3 | 0.050 | TLR2, TLR4, TLR8 |
| Inflammatory response | GO: 0006954 | 4 | 0.050 | TLR2, TLR4, TLR8, TLR7 |
| Transmembrane signaling receptor activity | GO: 0004888 | 3 | 0.010 | TLR2, TLR4, TLR7 |
| Innate immune response | GO: 0045087 | 3 | 0.010 | TLR2, TLR4, TLR8 |
| Extracellular region | GO: 0045087 | 6 | 0.010 | KITLG, BMP6, INHBB, INSL3, IGFBP1, IGFBP3 |
| Integral component of membrane | GO: 0016021 | 13 | 0.010 | ATP6, KITLG, ND3, ND4, BMPR1B, CALCRL, COX1, DGAT2, SLC11A1, SCD, TLR2, TLR8, TLR7 |
| Biological Pathways | Number of Genes Upregulated | Upregulated Genes | Number of Genes Downregulated | Downregulated Genes | FDR |
|---|---|---|---|---|---|
| Toll-like receptor signaling pathway | 6 | LY96, PIK3R5, TLR2, TLR4, TLR7, TLR8 | 1 | MAPK12 | 0.050 |
| Rheumatoid arthritis | 4 | ATP6V1A, TLR2, TLR4, TGFB2 | 0 | - | 0.010 |
| Ras signaling pathway | 5 | GAB2, EFNA4, KDR, PIK3R5, PDGFRB | 5 | KITLG, ANGPT2, PAK6, PLA2G1B, RGL1 | 0.010 |
| PI3K-Akt signaling pathway | 8 | CSF3R, EFNA4, GYS1, KDR, PIK3R5, PDGFRB, TLR2, TLR4 | 4 | KITLG, ANGPT2, COMP, COL1A1 | 0.010 |
| TNF signaling pathway | 3 | TNFRSF1A, PIK3R5, SOCS3 | 1 | MAPK12 | 0.010 |
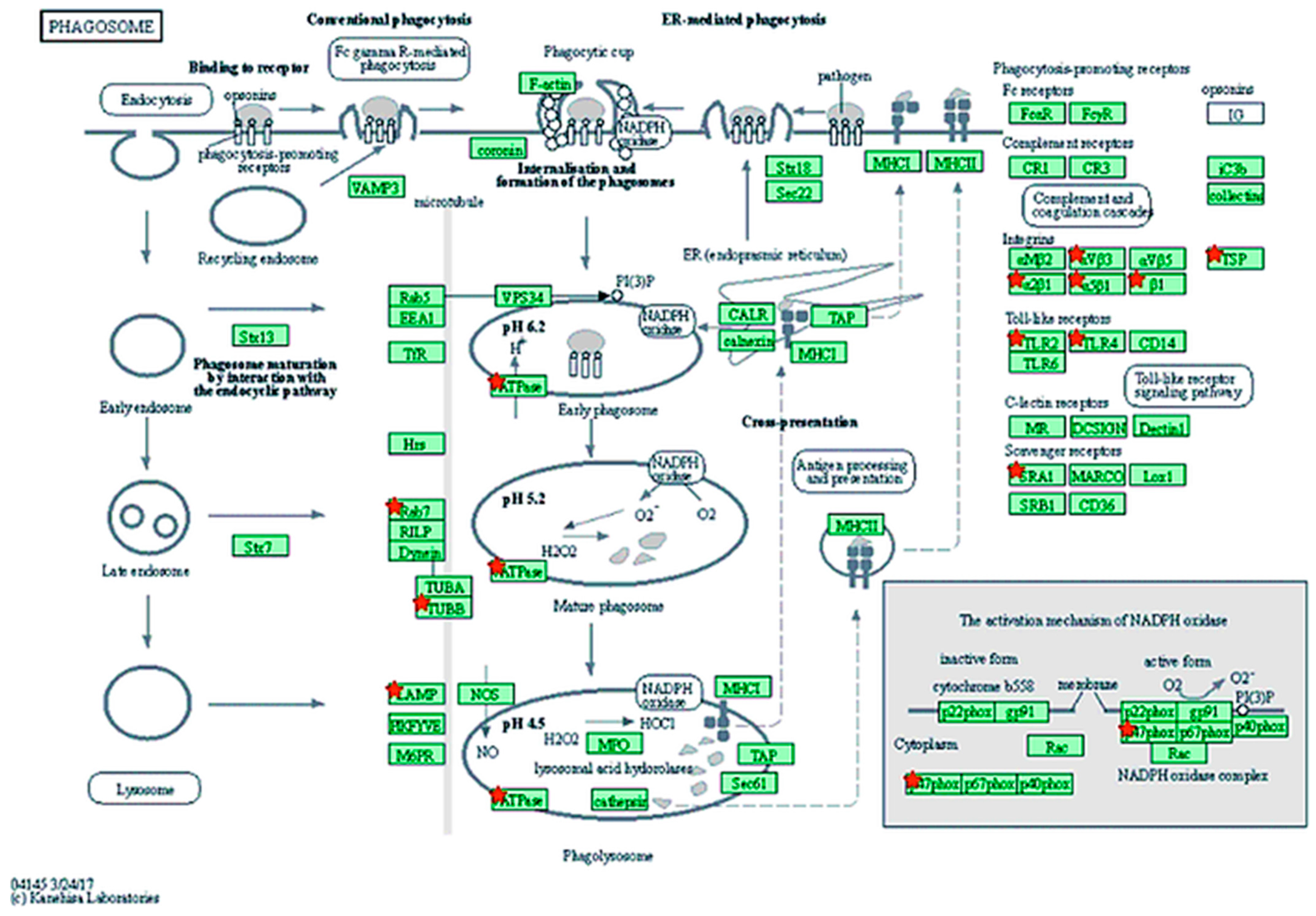
| Phagosome | 9 | ATP6V1A, RAB7B, LAMP2, MSR1, NCF1, TLR2, TLR4, TUBB1, TUBB4A | 1 | COMP | 0.020 |
| Cytokine-cytokine receptor interaction | 9 | TNFRSF1A, CSF2RB, CSF3R, CRLF2, IFNGR1, IFNGR2, IL1R2, PPBP, TGFB2 | 5 | CCR10, TNFRSF13C, TNFRSF6B, AMH, BMPR1B, | 0.004 |
| GO | Accession Number | Number of Genes | FDR | Identified Genes |
|---|---|---|---|---|
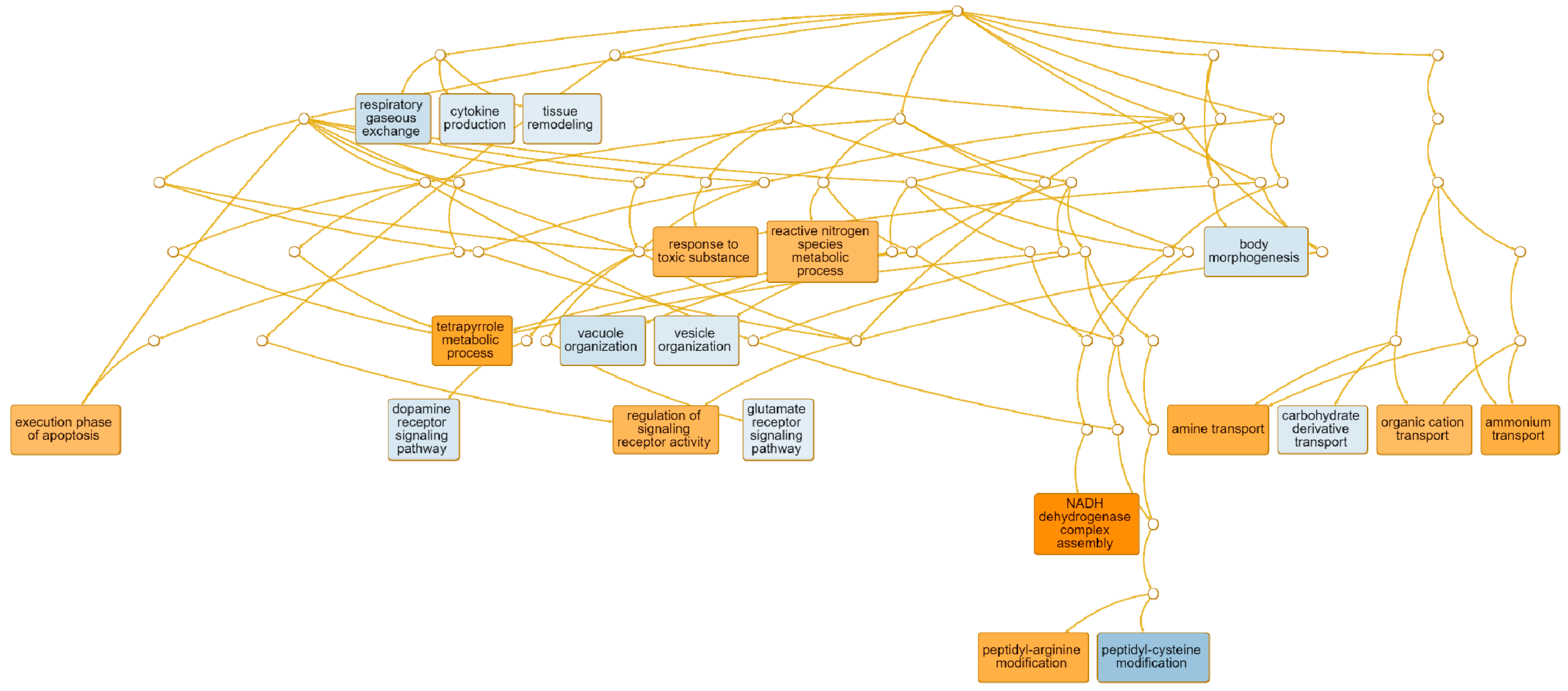
| execution phase of apoptosis | GO: 0097194 | 10 | 0.0240 | PTGIS, ENDOG, BOK, ERN2, SHARPIN, CIDEC, CXCR3, SIRT2, RPS3, BAX |
| tetrapyrrole metabolic process | GO: 0033013 | 9 | 0.00679 | ALAS2, ALAD, UROD, CYP1A2, UROS, HNF1A, MMAB, ATP5IF1, BLVRB |
| response to toxic substance | GO: 0009636 | 43 | 0.00691 | HBM, CHRNB2, LYPD1, GNRH1, MPO, CHRNA6, DRD3, CHRND, LTC4S, IL6 |
| regulation of signaling receptor activity | GO: 0010469 | 53 | <0.0000 | FGF16, CCL2, CGA, INHBE, CLEC11A, CXCL5, TNFSF11, AVP, LYPD1, FOXH1, GNRH1, NPY, CCL17, OXT, GHRH, NOG, EDA, CXCL9, MIF, RETN |
| peptidyl-arginine modification | GO: 0018195 | 10 | 0.01500 | ART1, PADI6, PADI3, PADI1, KRTCAP2, COPRS, PARK7, PRMT7, PRMT2, PRMT1 |
| amine transport | GO: 0015837 | 15 | 0.0060 | CHRNB2, CHRNA6, DRD3, ACE2, SNCG, SYT2, SLC22A16, SYT1, SLC18A1, DTNBP1 |
| organic cation transport | GO: 0015695 | 3 | 0.0290 | SLC22A1, SLC22A16, SLC18A3 |
| ammonium transport | GO: 0015696 | 15 | <0.0000 | RHAG, CHRNB2, CHRNA6, DRD3, SLC6A2, SNCG, SYT2, ADCYAP1, SLC22A16, SYT1 |
| NADH dehydrogenase complex assembly | GO: 0010257 | 21 | 0.0010 | NDUFA13, NDUFS5, NDUFS7, NDUFB9, NDUFA9, NDUFA8, BCS1L, NDUFB2, NDUFA5, NDUFA2 |
| GO | Accession Number | Number of Genes | FDR | Genes |
|---|---|---|---|---|
| peptidyl-cysteine modification | GO: 0018198 | 7 | 0.0020 | GOLGA7B, MAP6D1, TMX3, RAB3D, ZDHHC23, RAB6A, ZDHHC22 |
| cytokine production | GO: 0001816 | 95 | <0.000 | ADAMTS3, IL15, CMKLR1, NOD2, ABCA1, GDF2, CCR2, HFE, RSAD2, CHUK, ITK, LBP, ADCY7, IL6R, IL1A, RAB7B, IFIH1, C3AR1, LPL, CD274 |
| vacuole organization | GO: 0007033 | 34 | 0.0351 | ABCA1, PIP4K2B, UBXN2A, RALB, RAB7B, PPT1, GAA, ACP2, TBC1D14, AKTIP |
| vesicle organization | GO: 0016050 | 38 | 0.0300 | SEC16B, RAB8B, ABCA1, SAMD9, VPS39, RAB7B, DYSF, STX19, EXOC8, BCL2 |
| glutamate receptor signaling pathway | GO: 0007215 | 5 | 0.0600 | CACNG3, GRM2, CRHBP, HOMER2, PRNP |
| tissue remodeling | GO: 0048771 | 14 | 0.0600 | IHH, DCSTAMP, RASSF2, CCR2, DLL4, RAB3D, IL1A, EFNA2, NOTCH2, CLDN18 |
| Gene | Protein Name | FC | Adjpval | Protein Function |
|---|---|---|---|---|
| TLR4 | Toll-like receptor 4 | 1.49 | 0.02 | Acts via MYD88, TIRAP, and TRAF6, leading to NF-kappa-B activation, cytokine secretion, and the inflammatory response. |
| TLR2 | Toll-like receptor 2 | 1.60 | 0.03 | Related to mediating the innate immune response to bacterial lipoproteins or lipopeptides, related to cytokine secretion and the inflammatory response. |
| TLR6 | Toll-like receptor 6 | 1.66 | 0.04 | Acts via MYD88 and TRAF6, leading to NF-kappa-B activation, cytokine secretion, and the inflammatory response. |
| CHUK | Inhibitor of nuclear factor kappa-B kinase subunit alpha | 2.44 | 0.03 | Plays an essential role in the NF- kappa-B signaling pathway activated by multiple stimuli also by viral products. |
| CSF1R | Macrophage colony-stimulating factor 1 receptor | 1.80 | 0.03 | Controlling the proliferation and differentiation of hematopoietic precursor cells, especially mononuclear phagocytes, such as macrophages and monocytes. |
| IRF1 | Interferon regulatory factor 1 | 1.41 | 0.02 | Regulation of IFN and IFN-inducible genes, host response to viral and bacterial infections. |
| NRLP3 | NACHT, LRR, and PYD domain-containing protein 3 | Plays a crucial role in innate immunity and inflammation. | ||
| IFIH1 | Interferon-induced helicase C domain-containing protein 1 | 2.14 | 0.05 | Plays a major role in sensing viral infection and in the activation of a cascade of antiviral responses, including the induction of type I interferons and proinflammatory cytokines. |
| TBK1 | Serine/threonine-protein kinase TBK1 | 1.80 | 0.02 | Regulation of transcriptional activation of proinflammatory and antiviral genes including IFNA and IFNB. |
| CD14 | Monocyte differentiation antigen CD14 | 1.67 | 0.05 | Acts via MyD88, TIRAP, and TRAF6, leading to NF-kappa-B activation, cytokine secretion, and the inflammatory response. |
| MYD88 | Myeloid differentiation primary response protein MyD88 | 1.82 | 0.01 | Acts via toll-like receptor and IL-1 receptor signaling pathway in the innate immune response. |
| Biological Pathways | Number of Genes Upregulated | Upregulated Genes | Number of Genes Downregulated | Downregulated Genes | adjP * |
|---|---|---|---|---|---|
| B-cell receptor signaling pathway | 8 | LYN, CHUK, DAPP1, GRB2, PIK3CA, PIK3CB, PIK3AP1, PIK3R1 | 4 | NFKBIB, HRAS, CD79B, CD79A | 0.018 |
| Fc gamma R-mediated phagocytosis | 7 | CRKL, LYN, ARPC5, PIK3CA, PIK3CB, PIK3R1, PRKCD | 5 | DOCK2, LAT, PLPP2, PRKCG, RPS6KB2 | 0.034 |
| Apoptosis | 6 | FAS, CHUK, PIK3CA, PIK3CB, PIK3R1, TNFSF10 | 4 | AIFM1, ENDOG, IL3RA, NTRK1 | 0.034 |
| Natural killer cell-mediated cytotoxicity | 8 | FAS, GRB2, IFNAR1, PIK3CA, PIK3CB, PIK3R1, PTK2B, TNFSF10 | 4 | HRAS, HCST, LAT, PRKCG | 0.057 |
| T-cell receptor signaling pathway | 7 | CHUK, GRB2, MAPK14, PIK3CA, PIK3CB, PIK3R1, TEC | 3 | HRAS, NFKBIB, LAT | 0.036 |
| mTOR signaling pathway | 5 | EIF4E, PIK3CA, PIK3CB, PIK3R1, ULK2 | 2 | PRKCG, RPS6KB2 | 0.037 |
| FoxO signaling pathway | 13 | EP300, CHUK, CDKN1A, GADD45A, GRB2, MAPK14, PIK3CA, PIK3CB, PIK3R1, PRKAB1, STAT3, TGFBR2, TNFSF10 | 3 | HRAS, FOXO1, G6PC3 | 0.034 |
| TNF signaling pathway | 9 | FAS, TNFRSF1B, CHUK, IL15, MAPK14, NOD2, PIK3CA, PIK3CB, PIK3R1 | 0 | - | 0.044 |
| Toll-like receptor signaling pathway | 8 | TBK1, CHUK, IFNAR1, MAPK14, PIK3CA, PIK3CB, PIK3R1, TLR6 | 0 | - | 0.044 |
| Regulation of actin cytoskeleton | 10 | CRKL, GIT1, IQGAP1, ARPC5, CYFIP1, PXN, PIK3CA, PIK3CB, PIK3R1, PPP1CB | 4 | HRAS, FGFR2, PFN1, PPP1CA | 0.017 |
| HIF-1 signaling pathway | 8 | EP300, CUL2, CDKN1A, EIF4E, PIK3CA, PIK3CB, PIK3R1, STAT3 | 4 | TIMP1, FLT1, PRKCG, RPS6KB2 | 0.054 |
| Inflammatory mediator regulation of TRP channels | 8 | ADCY7, MAPK14, PIK3CA, PIK3CB, PIK3R1, PLA2G4A, PRKCD, PPP1CB | 4 | CALM3, NTRK1, PRKCG, PPP1CA | 0.050 |
| Signaling pathways regulating pluripotency of stem cells | 8 | JAK1, JAK2, GRB2, MAPK14, PIK3CA, PIK3CB, PIK3R1, STAT3 | 2 | HRAS, FGFR2 | 0.049 |
| Ras signaling pathway | 11 | RAP1B, RALB, TBK1, CSF1R, CHUK, GRB2, PIK3CA, PIK3CB, PIK3R1, PLA2G4A, RALBP1 | 6 | HRAS, CALM3, FGFR2, FLT1, LAT, PRKCG | 0.033 |
| Correlation Coefficient | ||
|---|---|---|
| Gene Symbol | qPCR vs. RNA-seq 1 | qPCR vs. Provirus Copy Number 2 |
| CCL2 | 0.850 | 0.778 *** |
| IL15 | 0.919 ** | −0.242 ns |
| CXCR3 | 0.794 * | 0.359 ns |
| MIF | −0.248 ns | −0.270 ns |
| NOD2 | 0.433 * | −0.470 * |
| CCR | 0.441 ns | 0.515 ** |
| BCL2 | −0.107 ns | 0.673 ** |
| CXCL5 | −0.232 ns | 0.759 *** |
| ITK | −0.589 ns | 0.478 * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Olech, M.; Ropka-Molik, K.; Szmatoła, T.; Piórkowska, K.; Kuźmak, J. Transcriptome Analysis for Genes Associated with Small Ruminant Lentiviruses Infection in Goats of Carpathian Breed. Viruses 2021, 13, 2054. https://doi.org/10.3390/v13102054
Olech M, Ropka-Molik K, Szmatoła T, Piórkowska K, Kuźmak J. Transcriptome Analysis for Genes Associated with Small Ruminant Lentiviruses Infection in Goats of Carpathian Breed. Viruses. 2021; 13(10):2054. https://doi.org/10.3390/v13102054
Chicago/Turabian StyleOlech, Monika, Katarzyna Ropka-Molik, Tomasz Szmatoła, Katarzyna Piórkowska, and Jacek Kuźmak. 2021. "Transcriptome Analysis for Genes Associated with Small Ruminant Lentiviruses Infection in Goats of Carpathian Breed" Viruses 13, no. 10: 2054. https://doi.org/10.3390/v13102054
APA StyleOlech, M., Ropka-Molik, K., Szmatoła, T., Piórkowska, K., & Kuźmak, J. (2021). Transcriptome Analysis for Genes Associated with Small Ruminant Lentiviruses Infection in Goats of Carpathian Breed. Viruses, 13(10), 2054. https://doi.org/10.3390/v13102054






