The First Complete Genome Sequence of a Novel Tetrastichus brontispae RNA Virus-1 (TbRV-1)
Abstract
1. Introduction
2. Materials and Methods
2.1. Insect Collection
2.2. Transcriptome of the Tetrastichus brontispae Abdomen
2.3. Viral Metagenomics Analysis
2.4. Virus Genome Sequencing
2.5. MicroRNA Library Construction and Sequencing
2.6. Isolation of Virus RNA
2.7. Tissue Distribution and Developmental Expression Patterns of TbRV-1
3. Results
3.1. Five Types of RNA Virus Present in the T. brontispae Transcriptome
3.2. The Genome Organization of TbRV-1
3.3. Detection of TbRV-1 siRNAs
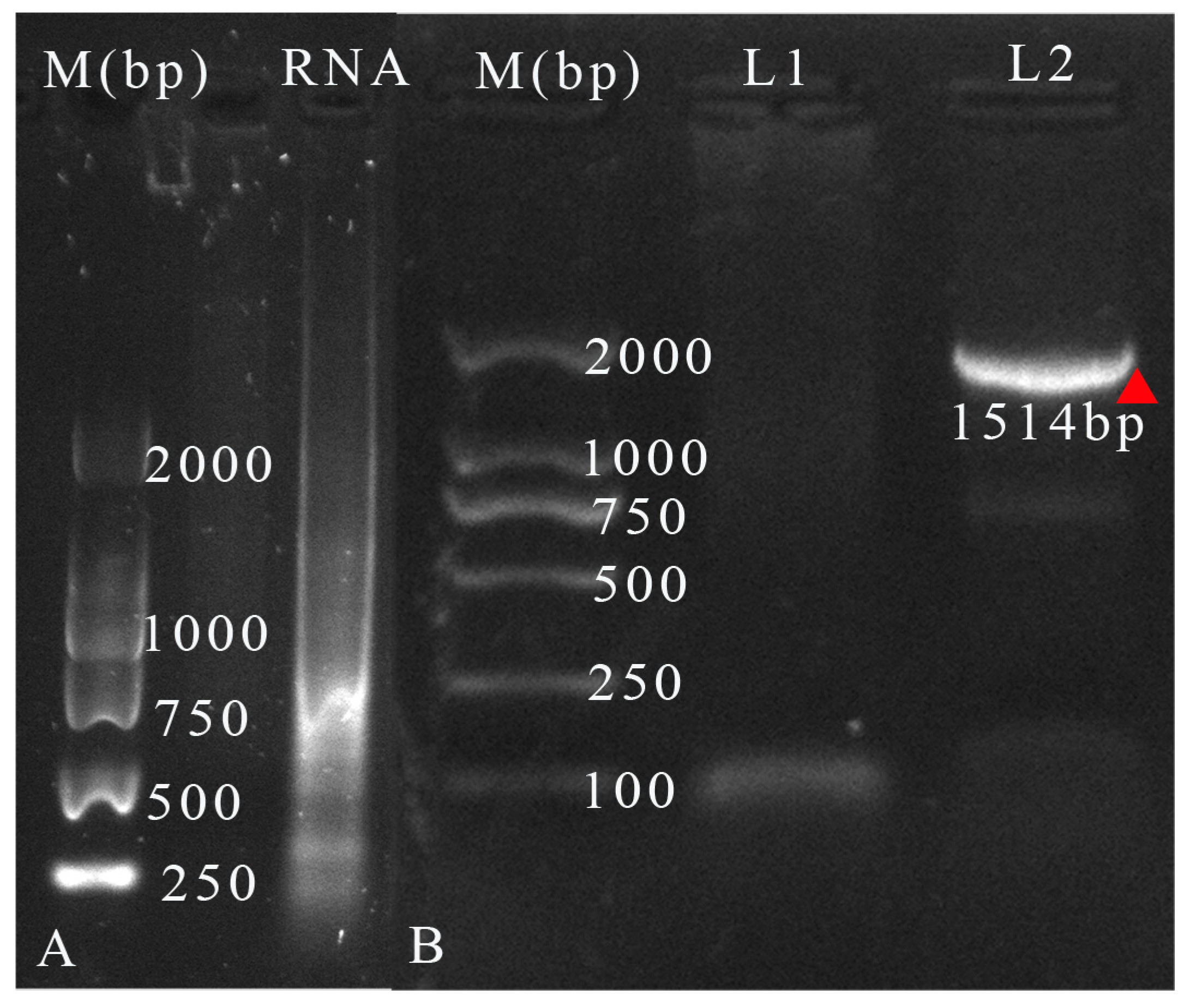
3.4. Identification of Virus RNA from the Isolated Mixtures
3.5. The Quantification of Virus RNA Copy Numbers
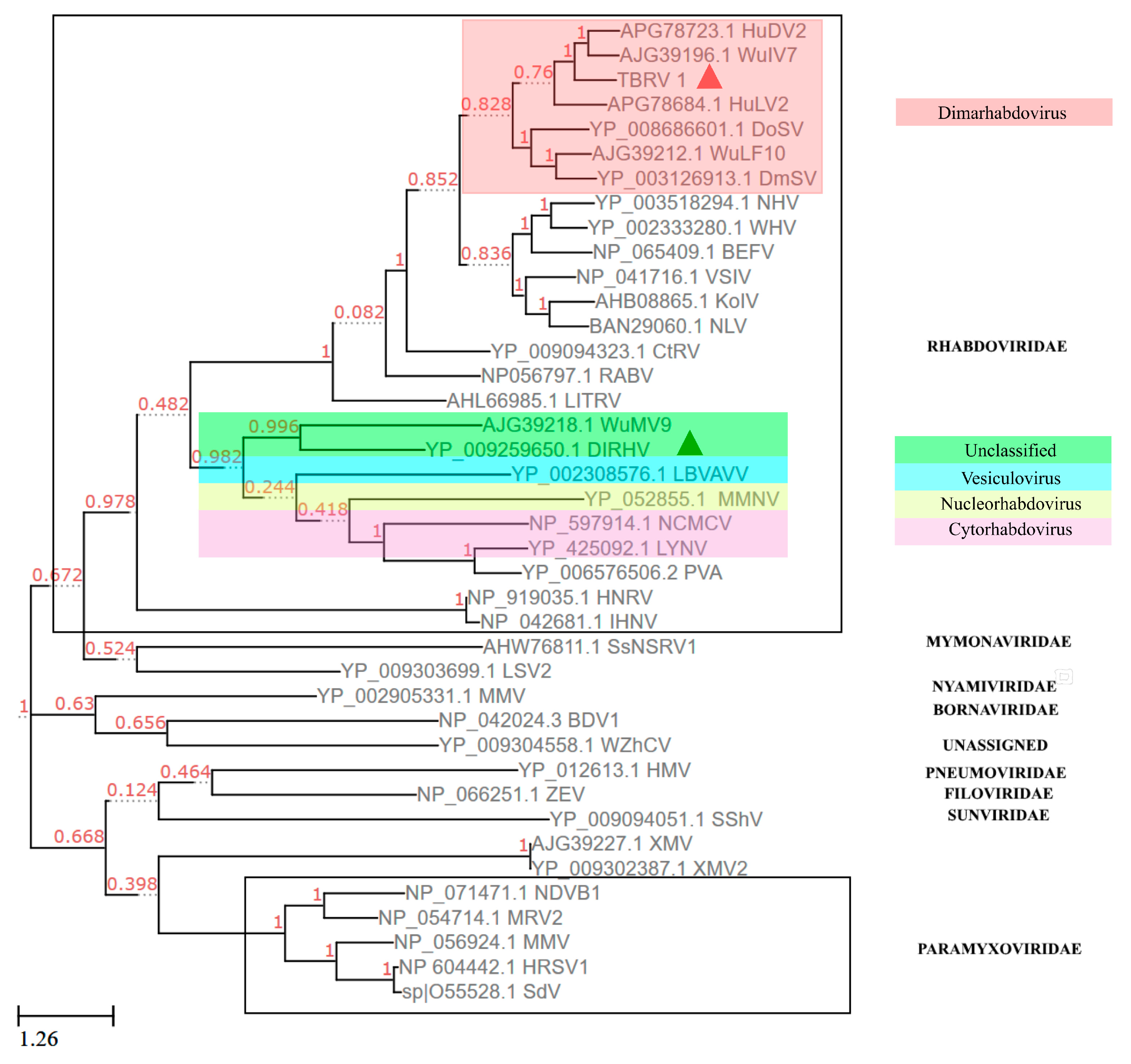
3.6. Phylogenetic Analysis Using the Protein Sequence of RdRP
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Walker, P.; Blasdell, K.; Calisher, C.H.; Dietzgen, R.G.; Kondo, H.; Kurath, G.; Longdon, B.; Stone, D.M.; Tesh, R.B.; Tordo, N.; et al. ICTV Report Consortium. ICTV Virus Taxonomy Profile: Rhabdoviridae. J. Gen. Virol. 2018, 99, 447–448. [Google Scholar] [CrossRef] [PubMed]
- Hogenhout, S.A.; Redinbaugh, M.G.; Ammar, E.-D. Plant and animal rhabdovirus host range: A bug’s view. Trends Microbiol. 2003, 11, 264–271. [Google Scholar] [CrossRef]
- Longdon, B.; Murray, G.G.R.; Palmer, W.J.; Day, J.P.; Parker, D.J.; Welch, J.J.; Obbard, D.J.; Jiggins, F.M. The evolution, diversity, and host associations of rhabdoviruses. Virus Evol. 2015, 1, e014. [Google Scholar] [CrossRef]
- International Committee Taxonomy of Viruses. Available online: http://www.ictvonline.org/virus-Taxonomy.asp (accessed on 12 March 2019).
- Hosang, M.L.A.; Alouw, J.C.; Novarianto, H. Biological control of Brontispa longissima (Gestro) in Indonesia. In Report of the Expert Consultation on Coconut Beetle Outbreak in APPPC Member Countries; RAP Publication: Manado, Indonesia, 2004. [Google Scholar]
- Kuzmin, I.V.; Novella, I.S.; Dietzgen, R.G.; Padhi, A.; Rupprecht, C.E. The rhabdoviruses: Biodiversity, phylogenetics, and evolution. Infect. Genet. Evol. 2009, 9, 541–553. [Google Scholar] [CrossRef] [PubMed]
- Longdon, B.; Jiggins, F.M. Vertically transmitted viral endosymbionts of insects: Do sigma viruses walk alone? Proc. Biol. Sci. 2012, 279, 3889–3898. [Google Scholar] [CrossRef] [PubMed]
- L‘héritier, P.H. Sensitivity to CO2 in Drosophila a review. Heredity 1948, 2, 325–348. [Google Scholar] [CrossRef]
- Strand, M.R.; Burke, G.R. Polydnaviruses from discovery to current insights. Virology 2015, 479–480, 393–402. [Google Scholar] [CrossRef]
- Gundersen-Rindala, D.; Dupuy, C.; Huguet, E.; Drezen, J.-M. Parasitoid polydnaviruses: Evolution, pathology and applications. Biocontrol Sci. Technol. 2013, 23, 1–61. [Google Scholar] [CrossRef]
- Madanagopal, N.; Kim, Y. Parasitism by Cotesia glomerata induces immunosuppression of Pieris rapae: Effects of ovarian protein and polydnavirus. J. Asia-Pacif. Entomol. 2006, 9, 339–346. [Google Scholar] [CrossRef]
- Furihata, S.; Tanaka, K.; Ryuda, M.; Ochiai, M.; Matsumoto, H.; Csikos, G.; Hayakawa, Y. Immunoevasive protein (IEP)-containing surface layer covering polydnavirus particles is essential for viral infection. J. Invertebr. Pathol. 2014, 115, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Fang, Q.; Wang, B.; Yan, Z.; Hong, J.; Bao, Y.; Kuhn, J.H.; Werren, J.H.; Song, Q.; Ye, G. A novel negative-stranded RNA virus mediates sex ratio in its parasitoid host. PLoS Pathog. 2016, 13, e1006201. [Google Scholar] [CrossRef]
- Lawrence, P.O.; Matos, L.F. Transmission of the Diachasmimorpha longicaudata rhabdovirus (DlRhV) to wasp offspring: An ultrastructural analysis. J. Insect Physiol. 2005, 51, 235–241. [Google Scholar] [CrossRef] [PubMed]
- Simmondsa, T.J.; Carrillo, D.; Burke, G.R. Characterization of a venom gland-associated rhabdovirus in the parasitoid wasp Diachasmimorpha longicaudata. J. Insect Physiol. 2016, 91–92, 48–55. [Google Scholar] [CrossRef]
- Chen, Q.; Peng, Z.; Xu, C.; Tang, C.; Lu, B.; Jin, Q.; Wen, H.; Wan, F. Biological assessment of Tetrastichus brontispae, a pupal parasitoid of coconut leaf beetle Brontispa longissima. Biocontrol Sci. Technol. 2010, 20, 283–295. [Google Scholar] [CrossRef]
- Nguyen, H.T.; Oo, T.T.; Ichiki, R.T.; Takano, S.; Murata, M.; Takasu, K.; Konishi, K.; Tunkumthong, S.; Chomphookhiaw, N.; Nakamura, S. Parasitisation of Tetrastichus brontispae (Hymenoptera: Eulophidae), a biological control agent of the coconut hispine beetle Brontispa longissima (Coleoptera: Chrysomelidae). Biocontrol Sci. Technol. 2012, 22, 955–968. [Google Scholar] [CrossRef]
- Tang, B.Z.; Xu, L.; Hou, Y.M. Effects of rearing conditions on the parasitism of Tetrastichus brontispae on its pupal host Octodonta nipae. BioControl 2014, 59, 647–657. [Google Scholar] [CrossRef]
- Liu, K.; Fu, B.L.; Lin, J.R.; Fu, Y.G.; Peng, Z.Q.; Jin, Q.A.; Tang, L.D. Parasitism performance of Tetrastichus brontispae Ferriere over the coconut hispine beetle, Brontispa longissima (Gestro). Neotrop. Entomol. 2016, 45, 420–426. [Google Scholar] [CrossRef] [PubMed]
- Meng, E.; Tang, B.; Hou, Y.; Chen, X.; Chen, J.; Yu, X. Altered immune function of Octodonta nipae (Maulik) to its pupal endoparasitoid, Tetrastichus brontispae Ferrière. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2016, 198, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Rotheram, S. Immune surface of eggs of a parasitic insect. Nature 1967, 214, 700. [Google Scholar] [CrossRef] [PubMed]
- Feddersen, I.; Sander, K.; Schmidt, O. Virus-like particles with host protein-like antigenic determinants protect an insect parasitoid from encapsulation. Experientia 1986, 42, 1278–1281. [Google Scholar] [CrossRef]
- Shi, M.; Lin, X.; Tian, J.; Chen, L.; Chen, X.; Li, C.; Qin, X.; Li, J.; Cao, J.; Eden, J.S.; et al. Redefining the invertebrate RNA virosphere. Nature 2016, 540, 539–543. [Google Scholar] [CrossRef] [PubMed]
- Tang, B.Z.; Chen, J.; Hou, Y.M.; Meng, E. Transcriptome immune analysis of the invasive beetle Octodonta nipae (Maulik) (Coleoptera: Chrysomelidae) parasitized by Tetrastichus brontispae Ferrière (Hymenoptera: Eulophidae). PLoS ONE 2014, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Reuter, G.; Boros, A.; Pal, J.; Kapusinszky, B.; Delwart, E.; Pankovics, P. Detection and genome analysis of a novel (dima)rhabdovirus (Riverside virus) from Ochlerotatus sp. mosquitoes in Central Europe. Infect. Genet. Evol. 2016, 39, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Kasuya, A.; Thornton, J.M. Three-dimensional structure analysis of PROSITE patterns. J. Mol. Biol. 1999, 286, 1673–1691. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Wu, Q.; Luo, Y.; Lu, R.; Lau, N.; Lai, E.C.; Li, W.X.; Ding, S.W. Virus discovery by deep sequencing and assembly of virus-derived small silencing RNAs. Proc. Natl. Acad. Sci. USA 2010, 107, 1606–1611. [Google Scholar] [CrossRef]
- Zerbino, D.R.; Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 18, 821–829. [Google Scholar] [CrossRef]
- Suzuki, M.; Tanaka, T. Virus-like particles in venom of Meteorus pulchricornis induce host hemocyte apoptosis. J. Insect Physiol. 2006, 52, 602–613. [Google Scholar] [CrossRef]
- Sylvester, E.S.; Richardson, J. Aphid-borne Rhabdoviruses—Relationships with Their Vectors. In Advances in Disease Vector Research; Harris, K.F., Ed.; Springer: New York, NY, USA, 1992; Volume 9. [Google Scholar]
- Walker, P.J.; Firth, C.; Widen, S.G.; Blasdell, K.R.; Guzman, H.; Wood, T.G.; Paradkar, P.N.; Holmes, E.C.; Tesh, R.B.; Vasilakis, N. Evolution of genome size and complexity in the rhabdoviridae. PLoS Pathog. 2015, 11, e1004664. [Google Scholar] [CrossRef]
- Gao, F.; Shen, J.; Liao, F.; Cai, W.; Lin, S.; Yang, H.; Chen, S. The first complete genome sequence of narcissus latent virus from Narcissus. Arch. Virol. 2018, 163, 1383–1386. [Google Scholar] [CrossRef] [PubMed]
- Barr, J.N.; Whelan, S.P.J.; Wertz, G.W. Transcriptional control of the RNA-dependent RNA polymerase of vesicular stomatitis virus. Biochim. Biophys. Acta 2002, 1577, 337–353. [Google Scholar] [CrossRef]
- Leyrat, C.; Ribeiro, E.A.; Gérard, F.C.; Ivanov, I.; Ruigrok, R.W.; Jamin, M. Structure, interactions with host cell and functions of rhabdovirus phosphoprotein. Future Virol. 2011, 6, 465–481. [Google Scholar] [CrossRef]
- Cook, S.; Chung, B.Y.; Bass, D.; Moureau, G.; Tang, S.; McAlister, E.; Culverwell, C.L.; Glucksman, E.; Wang, H.; Brown, T.D.; et al. Novel virus discovery and genome reconstruction from field RNA samples reveals highly divergent viruses in dipteran hosts. PLoS ONE 2013, 8, e80720. [Google Scholar] [CrossRef] [PubMed]
- Li, C.X.; Shi, M.; Tian, J.H.; Lin, X.D.; Kang, Y.J.; Chen, L.J.; Qin, X.C.; Xu, J.; Holmes, E.C.; Zhang, Y.Z. Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. eLife 2015, 4, e05378. [Google Scholar] [CrossRef]
- Kang, Y.-J.; Zhang, Y.-Z. Taxonomy and evolution of Rhabdoviruses. Chin. J. Vector Biol. Control 2014, 25, 374–377. [Google Scholar]
- Longdon, B.; Obbard, D.J.; Jiggins, F.M. Sigma viruses from three species of Drosophila form a major new clade in the rhabdovirus phylogeny. Proc. Biol. Sci. 2010, 277, 35–44. [Google Scholar] [CrossRef]
- Liu, S.; Vijayendran, D.; Bonning, B.C. Next generation sequencing technologies for insect virus discovery. Viruses 2011, 3, 1849–1869. [Google Scholar] [CrossRef]
- Simmonds, P. Methods for virus classification and the challenge of incorporating metagenomic sequence data. J. Gen. Virol. 2015, 96, 1193–1206. [Google Scholar] [CrossRef]
- Simmonds, P.; Adams, M.J.; Benko, M.; Breitbart, M.; Brister, J.R.; Carstens, E.B.; Davison, A.J.; Delwart, E.; Gorbalenya, A.E.; Harrach, B.; et al. Consensus statement: Virus taxonomy in the age of metagenomics. Nat. Rev. Microbiol. 2017, 15, 161–168. [Google Scholar] [CrossRef]
- Burke, G.R.; Strand, M.R. Polydnaviruses of parasitic wasps domestication of viruses to act as gene delivery vectors. Insects 2012, 3, 91–119. [Google Scholar] [CrossRef]
- Lawrence, P.O. Morphogenesis and cytopathic effects of the Diachasmimorpha longicaudata entomopoxvirus in host haemocytes. J. Insect Physiol. 2005, 51, 221–233. [Google Scholar] [CrossRef]
- Barratt, B.I.; Murney, R.; Easingwood, R.; Ward, V.K. Virus-like particles in the ovaries of Microctonus aethiopoides Loan (Hymenoptera: Braconidae): Comparison of biotypes from Morocco and Europe. J. Invertebr. Pathol. 2006, 91, 13–18. [Google Scholar] [CrossRef]
- Varaldi, J.; Ravallec, M.; Labrosse, C.; Lopez-Ferber, M.; Bouletreau, M.; Fleury, F. Artifical transfer and morphological description of virus particles associated with superparasitism behaviour in a parasitoid wasp. J. Insect Physiol. 2006, 52, 1202–1212. [Google Scholar] [CrossRef]
- Suzuki, M.; Miura, K.; Tanaka, T. Effects of the virus-like particles of a braconid endoparasitoid, Meteorus pulchricornis on hemocytes and hematopoietic organs of its noctuid host Pseudaletia separata. Appl. Entomol. Zool. 2009, 44, 115–125. [Google Scholar] [CrossRef]
- Renaulta, S.; Stasiaka, K.; Federicib, B.; Bigota, Y. Commensal and mutualistic relationships of reoviruses with their parasitoid wasp hosts. J. Insect Physiol. 2005, 51, 137–148. [Google Scholar] [CrossRef]
- Dheilly, N.M.; Maure, F.; Ravallec, M.; Galinier, R.; Doyon, J.; Duval, D.; Leger, L.; Volkoff, A.N.; Misse, D.; Nidelet, S.; et al. Who is the puppet master? Replication of a parasitic wasp-associated virus correlates with host behaviour manipulation. Proc. Biol. Sci. 2015, 282, 20142773. [Google Scholar] [CrossRef]


| Query Virus | Gene ID | Length (bp) | Subject Protein | Subject Virus | E Value | Taxonomy of Subject Virus | Coverage Range |
|---|---|---|---|---|---|---|---|
| Tetrastichus brontispae RNA virus-1 (TbRV-1) | Unigene0017563 | 1431 | Nucleoprotein | Oita rhabdovirus 296/1972 | 8.2 × 10−100 | Rhabdoviridae (negative sense single strand RNA viruses) | 1–1431 |
| Unigene0019568 | 1954 | Matrix protein | Tibtogaran virus | 9 × 10−6 | 1442–3395 | ||
| Unigene0023051 | 5167 | RdRp | Scophthalmus maximus Rhabdovirus | 0 | 3411–8578 | ||
| Unigene0014453 | 1037 | RdRp | 2 × 10−21 | 11,075–12,171 | |||
| Unigene0016900 | 339 | RdRp | 4 × 10−35 | 9036–9375 | |||
| Unigene0016901 | 404 | RdRp | 4 × 10−33 | 8656–9058 | |||
| Unigene0008666 | 895 | RdRp | 1 × 10−35 | 9744–10,603 | |||
| Unigene0005445 | 216 | RdRp | Grass carp rhabdovirus V76 | 9 × 10−13 | 10,581–10,796 | ||
| Tetrastichus brontispae RNA virus -2 (TbRV-2) | Unigene0000118 | 2396 | Polymerase PB1 | Quaranfil virus | 0 | Orthomyxoviridae (negative sense single strand RNA viruses, 6–8 segments) | / |
| Unigene0022762 | 2405 | Polymerase PB1 | 7 × 10−171 | ||||
| Unigeno0022160 | 1447 | Polymerase PA | 1 × 10−36 | ||||
| Unigene0018460 | 2416 | Polymerase PA | 6 × 10−39 | ||||
| Unigene0020769 | 2500 | Polymerase PB2 | 2 × 10−21 | ||||
| Unigene0000506 | 427 | Polymerase PB2 | 2 × 10−7 | ||||
| Unigene0021353 | 1762 | Hypothetical protein | 1 × 10−36 | ||||
| Unigene0021482 | 1772 | Hypothetical protein | 4 × 10−20 | ||||
| Unigene0016137 | 1656 | Hemagglutinin | 2 × 10−32 | ||||
| Unigene0020778 | 1614 | Hemagglutinin | 3 × 10−42 | ||||
| Tetrastichus brontispae RNA virus 3 (TbRV-3) | Unigene000903 | 7027 | Polyprotein | Deformed wing virus | 1 × 10−69 | Iflaviridae | / |
| Unigene000905 | 2933 | Polyprotein | 4 × 10−101 | ||||
| Tetrastichus brontispae RNA virus -4 (TbRV-4) | Unigene0022228 | 2784 | RdRp | Saccharomyces 23S RNA Narnavirus | 5 × 10−18 | Narnaviridae (positive sense single strand RNA viruses) | / |
| Tetrastichus brontispae RNA virus -5 (TbRV-5) | Unigene0008109 | 260 | VP3 | Kadipiro virus | 5 × 10−8 | Reoviridae (double strand RNA viruses) | / |
| Unigene0002126 | 2047 | VP4 | 3 × 10−109 | ||||
| Unigene0011479 | 1396 | VP5 | 4 × 10−46 | ||||
| Unigene0004982 | 1092 | VP7 | 2 × 10−27 | ||||
| Unigene0005372 | 850 | VP12 | 3 × 10−22 | ||||
| Unigene0012923 | 483 | RdRp | Liaoning virus | 1 × 10−26 | |||
| Unigene0012924 | 204 | RdRp | 1 × 10−12 | ||||
| Unigene0013487 | 253 | VP1 | 4 × 10−18 | ||||
| Unigene0002892 | 281 | VP3 | 9 × 10−21 | ||||
| Unigene0010175 | 228 | VP2 | 1 × 10−17 | ||||
| Unigene0016608 | 1775 | VP2 | 2 × 10−88 | ||||
| Unigene0016692 | 426 | VP2 | 1 × 10−27 | ||||
| Unigene0016693 | 270 | VP2 | 1 × 10−16 | ||||
| Unigene0006368 | 206 | VP3 | 2 × 10−13 | ||||
| Unigene0005945 | 251 | VP3 | 3 × 10−17 | ||||
| Unigene0011375 | 856 | VP6 | 1 × 10−26 | ||||
| Unigene0008523 | 1035 | VP8 | 8 × 10−48 | ||||
| Unigene0014543 | 278 | RdRp | 3 × 10−14 | ||||
| Unigene0006593 | 250 | RdRp | 4 × 10−16 | ||||
| Unigene0007896 | 425 | Capping enzyme | 1 × 10–13 | ||||
| Unigene0001578 | 236 | VP1 | Banna virus | 2 × 10–12 |
| Virus ORF | Position (bp) | ORF Length (nt) | Protein Length | Protein Mass (kDa) | pI | Signal Peptide | No. of O-linked Glycosylation Sites | No. of Phosphorylation Sites | Top Blastx Match | E Value | Domain | E Value of Domain | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T | Y | S | ||||||||||||
| ORF I | 113–1393 | 1287 | 426 | 49.27 | 6.92 | NO | 9 | 16 | 9 | 26 | Putative nucleoprotein (Hubei dimarhabdovirus 2) | 1 × 10−100 | Rhabdo-ncap | 2.78 × 10−64 |
| ORF II | 1689–2600 | 912 | 303 | 34.79 | 5.50 | NO | 13 | 11 | 3 | 15 | ND | - | ND | - |
| ORF III | 2699–3355 | 657 | 218 | 25.97 | 6.91 | NO | 0 | 10 | 6 | 7 | Matrix protein (Hubei dimarhabdovirus 2) | 2 × 10−22 | Vesiculo_matrix super family | 2.49 × 10−4 |
| ORF IV | 3761–5410 | 1650 | 549 | 63.28 | 7.45 | NO | 2 | 19 | 9 | 27 | Glycoprotein (Wuhan insect virus 7) | 2 × 10−61 | Rhabdo_glycop super family | 1.78 × 10−23 |
| ORF V | 5781–12,141 | 6360 | 2119 | 245.17 | 8.27 | NO | 10 | 67 | 38 | 113 | RdRp (Hubei dimarhabdovirus 2) | 0 | Mononeg_RNA-pol super family; Methyltrans_Mon super family; Mononeg_mRNAcap super family; | 01.15 × 10−132; 4.61 × 10−50 |
| Library | Small Interfering RNA Sequence | Identity | Coverage Range (nt) | Domain |
|---|---|---|---|---|
| TbBL1 | CAGGTTTTTTATTATTTTATCCTCTTGATTTGTGTCTCTAAC | 100% | 11,037–11,078 | RdRP |
| TbBL2 | TTGGAGTGTTGTTAGTGCATTAGCCATAGAAAGGGAATCCAA | 100% | 7790–7831 | RdRP |
| TbBL3 | TTTAAGTTTAAATAAAATATTAACCTGATGATCTGGAGCTCC | 100% | 3122–3163 | Matrix |
| Sex | Tissues | Copies/ng Total RNA |
|---|---|---|
| Female wasps | Head and thorax | 690 ± 189 a |
| Ovary | 706 ± 120 a | |
| Venom apparatus | 446 ± 130 a | |
| Remaining abdomen | 376 ± 6 a | |
| Male wasps | Whole body | 376 ± 119 a |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Meng, E.; Tang, B.; Sanchez-Garcia, F.J.; Qiao, T.; Fu, L.; Wang, Y.; Hou, Y.-M.; Wu, J.-L.; Chen, Z.-M. The First Complete Genome Sequence of a Novel Tetrastichus brontispae RNA Virus-1 (TbRV-1). Viruses 2019, 11, 257. https://doi.org/10.3390/v11030257
Meng E, Tang B, Sanchez-Garcia FJ, Qiao T, Fu L, Wang Y, Hou Y-M, Wu J-L, Chen Z-M. The First Complete Genome Sequence of a Novel Tetrastichus brontispae RNA Virus-1 (TbRV-1). Viruses. 2019; 11(3):257. https://doi.org/10.3390/v11030257
Chicago/Turabian StyleMeng, E, Baozhen Tang, Francisco Javier Sanchez-Garcia, Ting Qiao, Lang Fu, Yu Wang, You-Ming Hou, Jiang-Lin Wu, and Zhi-Ming Chen. 2019. "The First Complete Genome Sequence of a Novel Tetrastichus brontispae RNA Virus-1 (TbRV-1)" Viruses 11, no. 3: 257. https://doi.org/10.3390/v11030257
APA StyleMeng, E., Tang, B., Sanchez-Garcia, F. J., Qiao, T., Fu, L., Wang, Y., Hou, Y.-M., Wu, J.-L., & Chen, Z.-M. (2019). The First Complete Genome Sequence of a Novel Tetrastichus brontispae RNA Virus-1 (TbRV-1). Viruses, 11(3), 257. https://doi.org/10.3390/v11030257





