Soil Microbial Communities in Natural and Managed Cloud Montane Forests
Abstract
:1. Introduction
2. Materials and Methods
2.1. Site Description
2.2. Biochemical Assays
2.3. Statistical Analysis
3. Results
3.1. Soil Characteristics and Microbial Biomass
3.2. PLFA Analyses
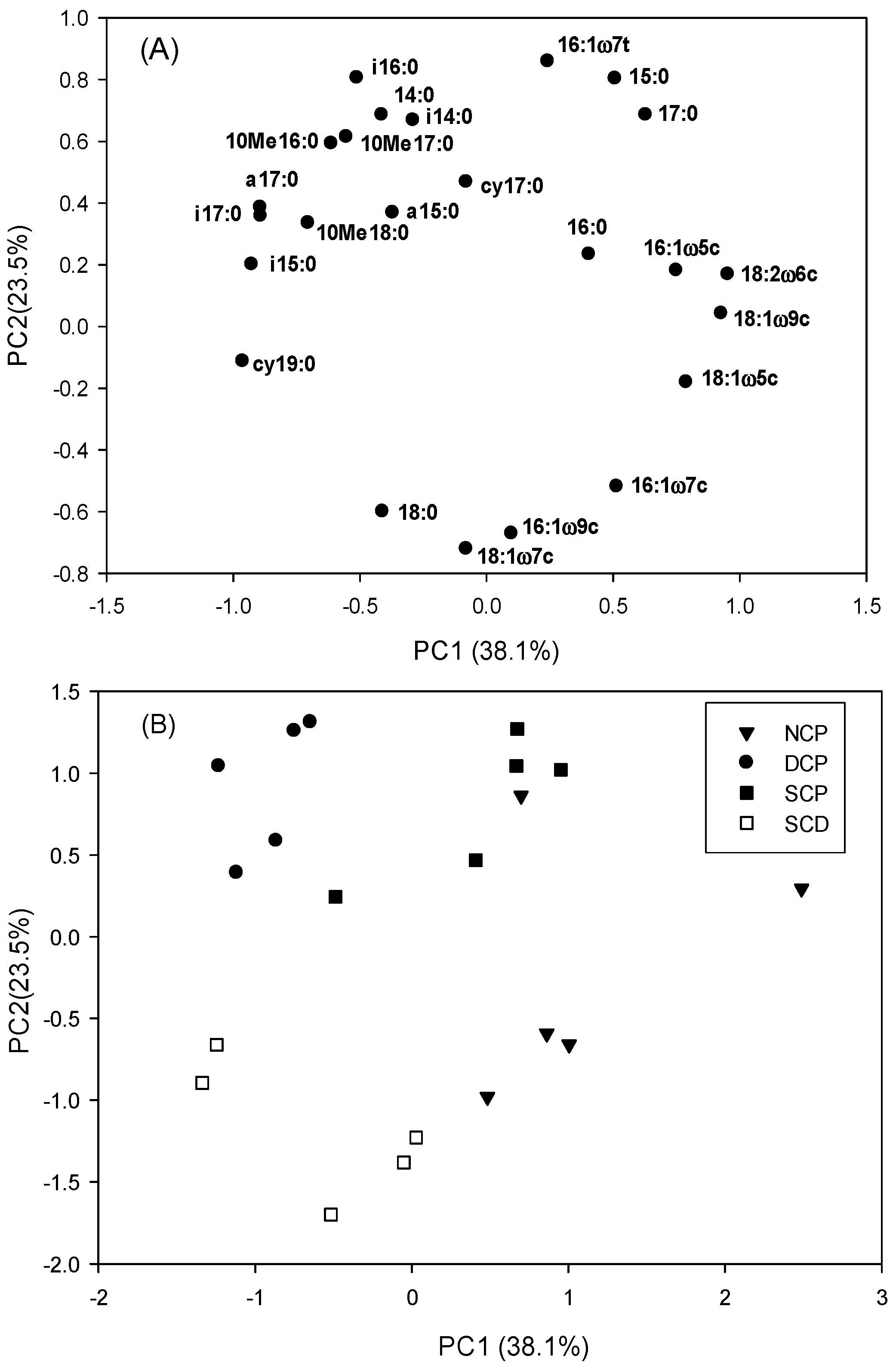
3.3. Microbial Community Patterns by Forest Management
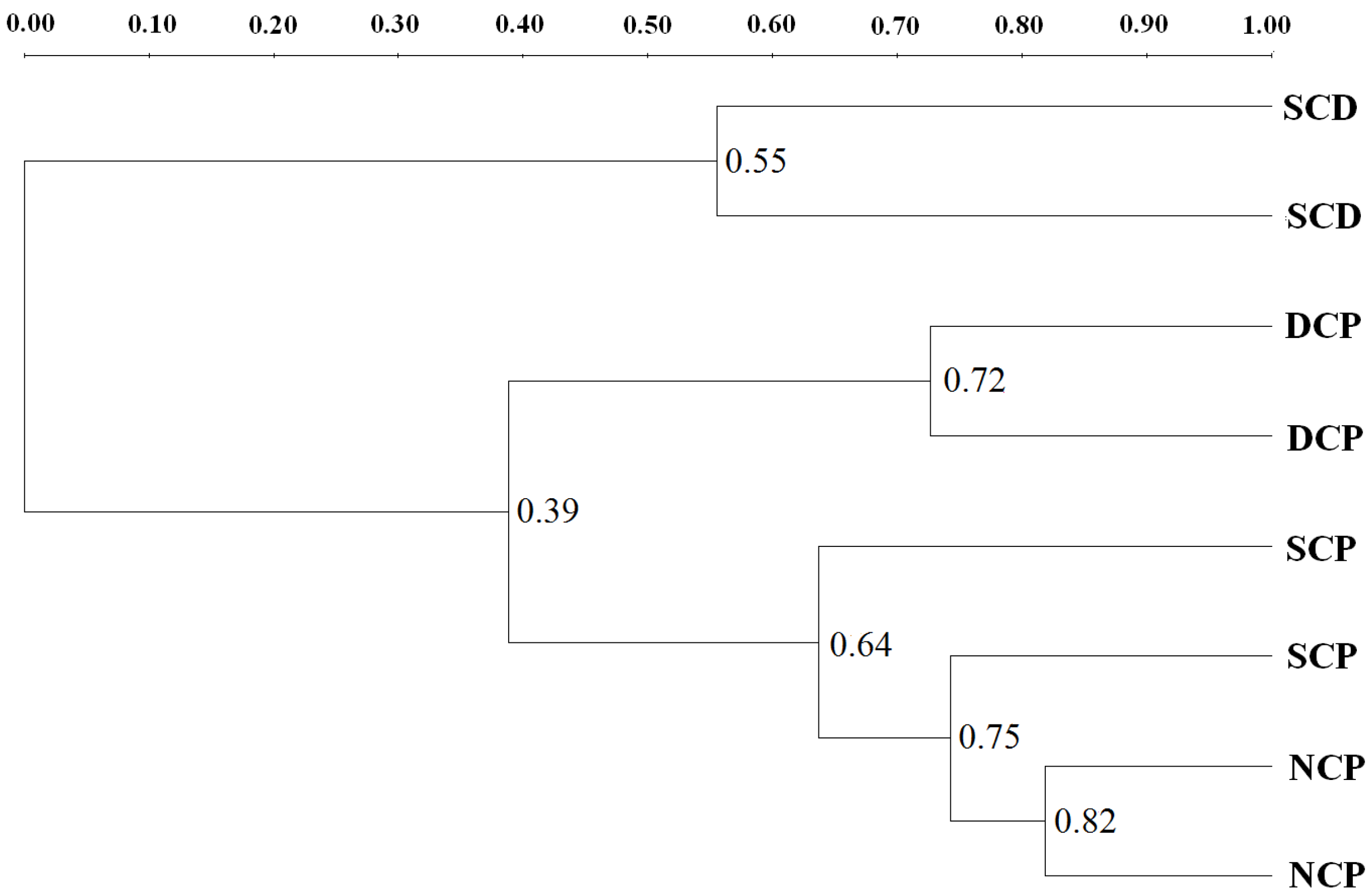
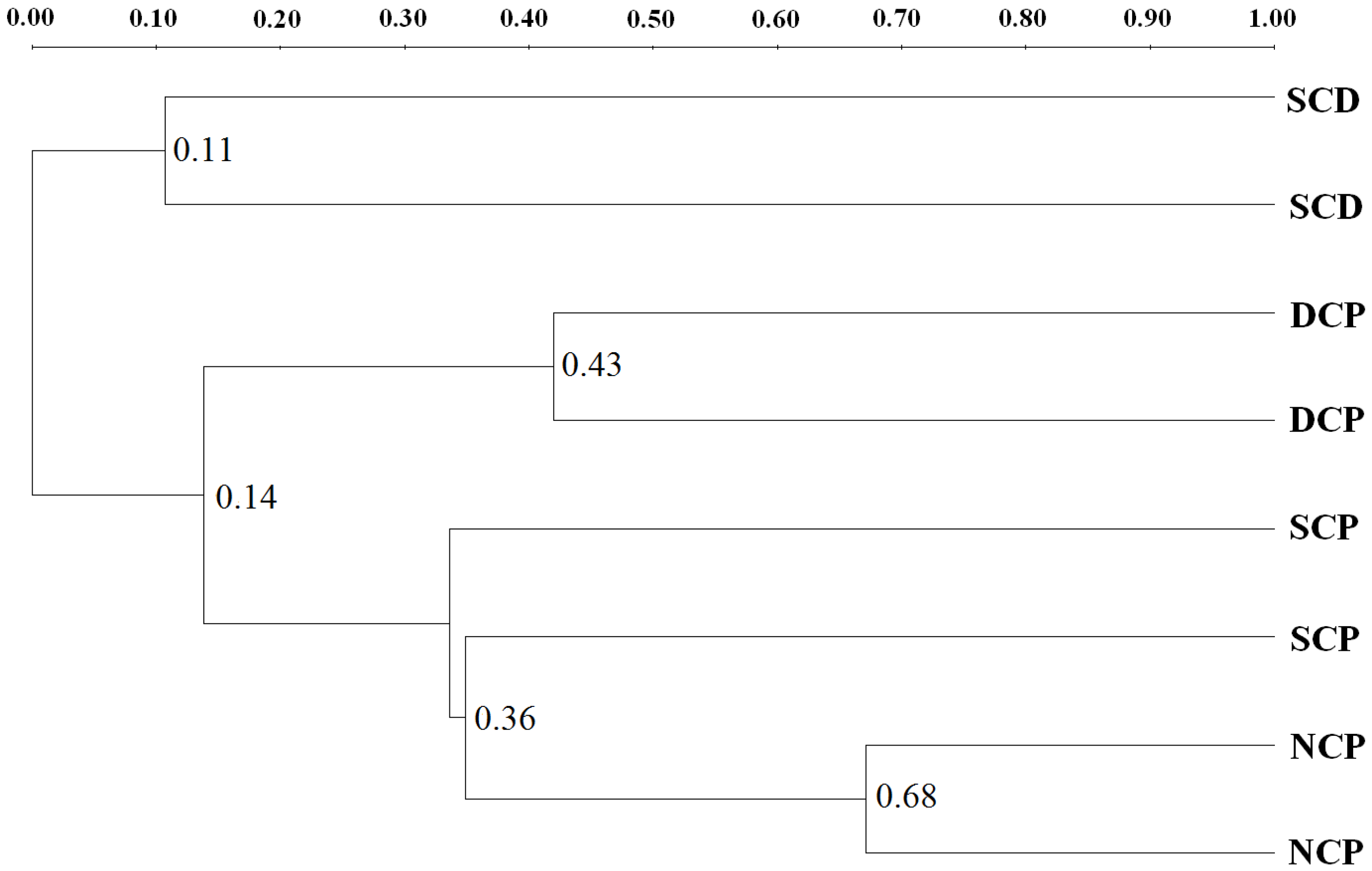
3.4. DGGE Analysis
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nannipieri, P.; Ascher, J.; Ceccherini, M.T.; Landi, L.; Pietramellara, G.; Renella, G. Microbial diversity and soil functions. Eur. J. Soil Sci. 2003, 54, 655–670. [Google Scholar] [CrossRef]
- Ushio, M.; Kitayama, K.; Balser, T.C. Tree species-mediated spatial patchiness of the composition of microbial community and physicochemical properties in the topsoils of a tropical montane forest. Soil Biol. Biochem. 2010, 42, 1588–1595. [Google Scholar] [CrossRef]
- Lin, Y.T.; Jangid, K.; Whitman, W.B.; Coleman, D.C.; Chiu, C.Y. Soil bacterial communities in native and regenerated perhumid montane forests. Appl. Soil Ecol. 2011, 47, 111–118. [Google Scholar] [CrossRef]
- Lucas-Borja, M.E.; Candel, D.; Jindo, K.; Moreno, J.L.; Andres, M.; Bastida, F. Soil microbial community structure and activity in monospecific and mixed forest stands, under Mediterranean humid conditions. Plant Soil 2012, 354, 359–370. [Google Scholar] [CrossRef]
- Chang, E.H.; Chen, T.H.; Tian, G.; Hsu, C.K.; Chiu, C.Y. Effect of 40 and 80 Years of conifer regrowth on soil microbial activities and community structure in subtropical low mountain forests. Forests 2016, 7, 244. [Google Scholar] [CrossRef]
- Grayston, S.J.; Prescott, C.E. Microbial communities in forest floors under four tree species in coastal British Columbia. Soil Biol. Biochem. 2005, 37, 1157–1167. [Google Scholar] [CrossRef]
- Yin, K.; Zhang, L.; Chen, D.; Tian, Y.; Zhang, F.; Wen, M.; Yuan, C. Understory herb layer exerts strong controls on soil microbial communities in subtropical plantations. Sci. Rep. 2016, 6, 27066. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.T.; Lin, C.P.; Chaw, S.M.; Whitman, W.B.; Coleman, D.C.; Chiu, C.Y. Bacterial community of very wet and acidic subalpine forest and fire-induced grassland soils. Plant Soil 2010, 332, 417–427. [Google Scholar] [CrossRef]
- Suzuki, C.; Nagaoka, K.; Shimada, A.; Takenaka, M. Bacterial communities are more dependent on soil type than fertilizer type, but the reverse is true for fungal communities. Soil Sci. Plant Nutr. 2009, 55, 80–90. [Google Scholar] [CrossRef]
- Vetrovsky, T.; Baldrian, P. Analysis of soil fungal communities by amplicon pyrosequencing: Current approaches to data analysis and the introduction of the pipeline SEED. Biol. Fert. Soils 2013, 49, 1027–1037. [Google Scholar] [CrossRef]
- Demoling, F.; Nilsson, L.O.; Bååth, E. Bacterial and fungal response to nitrogen fertilization in three coniferous forest soils. Soil Biol. Biochem. 2008, 40, 370–379. [Google Scholar] [CrossRef]
- Frostegård, Å.; Tunlid, A.; Bååth, E. Use and misuse of PLFA measurements in soils. Soil Biol. Biochem. 2011, 43, 1621–1625. [Google Scholar] [CrossRef]
- Vance, E.D.; Brookes, P.C.; Jenkinson, D.S. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 1987, 19, 703–707. [Google Scholar] [CrossRef]
- Amato, M.; Ladd, J.N. Assay for microbial biomass based on ninhydrin-reactive nitrogen in extracts of fumigated soils. Soil Biol. Biochem. 1988, 20, 107–114. [Google Scholar] [CrossRef]
- Frostegård, A.; Tunlid, A.; Bååth, E. Phospholipid fatty acid composition, biomass, and activity of microbial communities from two soil types exposed to different heavy metals. Appl. Environ. Microbiol. 1993, 59, 3605–3617. [Google Scholar] [PubMed]
- Chang, E.H.; Chen, T.H.; Tian, G.; Chiu, C.Y. The effect of altitudinal gradient on soil microbial community activity and structure in moso bamboo plantations. Appl. Soil Ecol. 2016, 98, 213–220. [Google Scholar] [CrossRef]
- Zogg, G.P.; Zak, D.R.; Ringleberg, D.B.; MacDonald, N.W.; Pregitzer, K.S.; White, D.C. Compositional and functional shifts in microbial communities due to soil warming. Soil Sci. Soc. Am. J. 1997, 61, 475–481. [Google Scholar] [CrossRef]
- Zelles, L. Fatty acid patterns of phospholipids and lipopolysaccharides in the characterisation of microbial communities in soil: A review. Biol. Fert. Soils 1999, 29, 111–129. [Google Scholar] [CrossRef]
- Nübel, U.; Engelen, B.; Felske, A.; Snaidr, J.; Wieshuber, A.; Amann, R.I.; Ludwig, W.; Backhaus, H. Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J. Bacteriol. 1996, 178, 5636–5643. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.W. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, J.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Williamson, J.R.; Neilsen, W.A. The influence of forest site on rate and extent of soil compaction and profile disturbance of skid trails during ground-based harvesting. Can. J. For. Res. 2000, 30, 1196–1205. [Google Scholar] [CrossRef]
- Ichikawa, T.; Yamaguchi, T.; Takahashi, T.; Asano, Y. Effects of the conversion of the forest management type from natural deciduous broad-leaved forests to artificial Japanese cypress (Chamaecyparis obtusa) and Japanese cedar (Cryptomeria japonica) forests on soil microbial flora and mineralization characteristics of organic carbon. Jpn. J. For. Environ. 2003, 45, 81–87. [Google Scholar]
- Mahmood, S.; Finlay, R.D.; Erland, S. Effects of repeated harvesting of forest residues on the ectomycorrhizal community in a Swedish spruce forest. New Phytol. 1999, 142, 577–585. [Google Scholar] [CrossRef]
- Zelles, L.; Bai, Q.Y.; Beck, T.; Beese, F. Signature fatty acids in phospholipids and lipopolysaccharides as indicators of microbial biomass and community structure in agricultural soils. Soil Biol. Biochem. 1992, 24, 317–332. [Google Scholar] [CrossRef]
- Bossio, D.A.; Scow, K.M.; Gunapala, N.; Graham, K.J. Determinants of soil microbial communities: Effects of agricultural management, season, and soil type on phospholipid fatty acid profiles. Microb. Ecol. 1998, 36, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Fierer, N.; Mark, A.B.; Robert, B.J. Toward an ecological classification of soil bacteria. Ecology 2007, 88, 1354–1364. [Google Scholar] [CrossRef] [PubMed]
- Kourtev, P.S.; Ehrenfeld, J.G.; Haggblom, M. Experimental analysis of the effect of exotic and native plant species on the structure and function of soil microbial communities. Soil Biol. Biochem. 2003, 35, 895–905. [Google Scholar] [CrossRef]
- Heipieper, H.J.; Meulenbeld, G.; VanOirschot, Q.; de Bont, J.A.M. Effect of environmental factors on the trans/cis ratio of unsaturated fatty acids in Pseudomonas putida S12. Appl. Environ. Microbiol. 1996, 62, 2773–2777. [Google Scholar] [PubMed]
- Sakai, H.; Inagaki, M.; Noguchi, K.; Sakata, T.; Yatskov, M.A.; Tanouchi, H.; Takahashi, M. Changes in soil organic carbon and nitrogen in an area of Andisol following afforestation with Japanese cedar and Hinoki cypress. Soil Sci. Plant Nutr. 2010, 56, 332–343. [Google Scholar] [CrossRef]
- Ollinger, S.V.; Smith, M.L.; Martin, M.E.; Hallett, R.A.; Goodale, C.L.; Aber, J.D. Regional variation in foliar chemistry and N cycling among forests of diverse history and composition. Ecology 2002, 83, 339–355. [Google Scholar]
- Zornoza, R.; Guerrero, C.; Mataix-Solera, J.; Scow, K.M.; Arcenegui, V.; Mataix-Beneyto, J. Changes in soil microbial community structure following the abandonment of agricultural terraces in mountainous areas of Eastern Spain. Appl. Soil Ecol. 2009, 42, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Burns, J.H.; Anacker, B.L.; Strauss, S.Y.; Burke, D.J. Soil microbial community variation correlates most strongly with plant species identity, followed by soil chemistry, spatial location and plant genus. AoB Plants 2015, 7, plv030. [Google Scholar] [CrossRef] [PubMed]
- Priha, O.; Grayston, S.J.; Pennanen, T.; Smolander, A. Microbial activities related to C and N cycling and microbial community structure in the rhizospheres of Pinus sylvestris, Picea abies and Betula pendula seedlings in an organic and mineral soil. Fems Microb. Ecol. 1999, 30, 187–199. [Google Scholar] [CrossRef]
| Forest Type 1 | pH | Organic C (%) | Total N (%) | C/N | Cmic (mg·kg−1) | Nmic (mg·kg−1) |
|---|---|---|---|---|---|---|
| NCP | 3.5 a 2 | 48.6 a | 1.89 b | 23.1 a | 4931 a | 728 ab |
| DCP | 3.7 a | 44.3 a | 2.28 a | 18.9 b | 3601 b | 619 b |
| SCP | 3.6 a | 46.9 a | 1.97 b | 25.5 a | 4839 a | 845 a |
| SCD | 3.7 a | 19.7 b | 1.16 c | 16.5 b | 1542 c | 264 c |
| Forest Type 1 | Total PLFAs | Bacteria | Fungi | VAM Fungi | Actino-Bacteria | G+ | G− | G+/G− | 16:1ω7t/16:1ω7c |
|---|---|---|---|---|---|---|---|---|---|
| NCP | 550 a 2 | 240 ab | 110 a | 21.4 a | 34.7 b | 71.2 b | 57.7a | 1.24 c | 0.117 b |
| DCP | 511 ab | 268 a | 64.5 c | 15.1 b | 42.7 a | 97.3 a | 46.5 b | 2.12 a | 0.225 a |
| SCP | 445 c | 197 b | 80.5 b | 15.7 b | 32.6 b | 68.1 b | 39.4 b | 1.75 b | 0.280 a |
| SCD | 483 bc | 232 ab | 51.7 c | 10.8 c | 32.7 b | 71.2 b | 49.8 ab | 1.43 bc | 0.099 b |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, E.-H.; Tian, G.; Chiu, C.-Y. Soil Microbial Communities in Natural and Managed Cloud Montane Forests. Forests 2017, 8, 33. https://doi.org/10.3390/f8010033
Chang E-H, Tian G, Chiu C-Y. Soil Microbial Communities in Natural and Managed Cloud Montane Forests. Forests. 2017; 8(2):33. https://doi.org/10.3390/f8010033
Chicago/Turabian StyleChang, Ed-Haun, Guanglong Tian, and Chih-Yu Chiu. 2017. "Soil Microbial Communities in Natural and Managed Cloud Montane Forests" Forests 8, no. 2: 33. https://doi.org/10.3390/f8010033
APA StyleChang, E.-H., Tian, G., & Chiu, C.-Y. (2017). Soil Microbial Communities in Natural and Managed Cloud Montane Forests. Forests, 8(2), 33. https://doi.org/10.3390/f8010033





