Haploids in Conifer Species: Characterization and Chromosomal Integrity of a Maritime Pine Cell Line
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Chromosome Integrity Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Uddenberg, D.; Akhter, S.; Ramachandran, P.; Sundström, J.F.; Carlsbecker, A. Sequenced genomes and rapidly emerging technologies pave the way for conifer evolutionary developmental biology. Front. Plant Sci. 2015, 6, 970. [Google Scholar] [CrossRef] [PubMed]
- Birol, I.; Raymond, A.; Jackman, S.D.; Pleasance, S.; Coope, R.; Taylor, G.A.; Yuen, M.M.S.; Keeling, C.I.; Brand, D.; Vandervalk, B.P.; et al. Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shot gun sequencing data. Bioinformatics 2013, 29, 1492–1497. [Google Scholar] [CrossRef] [PubMed]
- Nystedt, B.; Street, N.R.; Wetterbom, A.; Zuccolo, A.; Lin, Y.C.; Scofield, D.G.; Vezzi, F.; Delhomme, N.; Giacomello, S.; Alexeyenko, A.; et al. The Norway spruce genome sequence and conifer genome evolution. Nature 2013, 497, 579–584. [Google Scholar] [CrossRef] [PubMed]
- Neale, D.B.; Wegrzyn, J.L.; Stevens, K.A.; Zimin, A.V.; Puiu, D.; Crepeau, M.W.; Cardeno, C.; Koriabine, M.; Holtz-Morris, A.E.; Liechty, J.D.; et al. Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies. Genome Biol. 2014, 15, R59. [Google Scholar] [CrossRef] [PubMed]
- Zimin, A.; Stevens, K.A.; Crepeau, M.W.; Holtz-Morris, A.; Koriabine, M.; Marçais, G.; Puiu, D.; Roberts, M.; Wegrzyn, J.L.; de Jong, P.J.; et al. Sequencing and assembly of the 22-gb loblolly pine genome. Genetics 2014, 196, 875–889. [Google Scholar] [CrossRef] [PubMed]
- Rohr, R. State of development of technology for haploid production from gymnosperm microgametophyte and megagametophyte culture. Propag. Ornam. Plants 2004, 4, 3–18. [Google Scholar]
- Baldursson, S.; Nørgaard, J.; Krogstrup, P. Factors influencing haploid callus initiation and proliferation in megagametophyte cultures of Sitka spruce (Picea sitchensis). Silvae Gene 1993, 42, 79–86. [Google Scholar]
- Simola, L.K.; Honkanen, J. Organogenesis and fine structure in megagametophyte callus lines of Picea abies. Physiol. Plant 1983, 59, 551–561. [Google Scholar] [CrossRef]
- Simola, L.K.; Santanen, A. Improvement of nutrient medium for growth and embryogenesis of megagametophyte and embryo callus lines of Picea abies. Physiol. Plant 1990, 80, 27–35. [Google Scholar] [CrossRef]
- Von Aderkas, P.; Bonga, J.M.; Nagmani, R. Promotion of embryogenesis in cultured megagametophytes of Larix decidua. Can. J. For. Res. 1987, 7, 293–296. [Google Scholar]
- Von Aderkas, P.; Bonga, J.M. Morphological definition of phenocritical period for initiation of haploid embryogenic tissues from explants of Larix decidua. In Somatic Cell Genetics of Woody Plants; Ajuha, M.R., Ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1988; pp. 29–38. [Google Scholar]
- Von Aderkas, P.; Klimaszewska, K.; Bonga, J.M. Diploid and haploid embryogenesis in Larix leptolepis L. decidua, and their reciprocal hybrids. Can. J. For. Res. 1990, 20, 9–14. [Google Scholar] [CrossRef]
- Von Aderkas, P.; Pattanavibool, R.; Hristoforoglu, K.; Ma, Y. Embryogenesis and genetic stability in long term megagametophyte derived cultures of larch. Plant Cell Tissue Organ Cult. 2003, 75, 27–34. [Google Scholar] [CrossRef]
- Von Aderkas, P. Embryogenesis from protoplasts of haploid European larch. Can. J. For. Res. 1992, 22, 396–402. [Google Scholar] [CrossRef]
- Von Aderkas, P.; Bonga, J.M. Plants from haploid tissue of Larix decidua. Theor. Appl. Genet. 1993, 87, 225–228. [Google Scholar] [CrossRef] [PubMed]
- Von Aderkas, P.; Anderson, P. Aneuploidy and polyploidization in haploid tissue cultures of Larix decidua. Physiol. Plant 1993, 88, 73–77. [Google Scholar] [CrossRef]
- Fang, K.; Zhang, L.; Lin, J. A rapid, efficient method for the mass production of pollen protoplast from Pinus bungeana Zucc. ex Endl and Picea wilsonii Mast. Flora 2006, 201, 74–80. [Google Scholar] [CrossRef]
- Krutovsky, K.V.; Tretyakova, I.N.; Oreshkova, N.V.; Pak, M.E.; Kvitko, O.V.; Vaganov, E.A. Somaclonal variation of haploid in vitro tissue culture obtained from Siberian larch (Larix sibirica Ledeb.) megagametophytes for whole genome de novo sequencing. In Vitro Cell. Dev. Biol.-Plant 2014, 50, 655–664. [Google Scholar] [CrossRef]
- Arrillaga, I.; Guevara, M.A.; Muñoz-Bertomeu, J.; Lázaro-Gimeno, D.; Sáez-Laguna, E.; Díaz, L.M.; Torralba, L.; Mendoza-Poudereux, I.; Segura, J.; Cervera, M.T. Selection of haploid cell lines from megagametophyte cultures of maritime pine as a DNA source for massive sequencing of the species. Plant Cell Tissue Organ Cult. 2014, 118, 147–155. [Google Scholar] [CrossRef]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bioassay with tobacco tissue cultures. Physiol. Plant 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Lelu-Walter, M.A.; Bernier-Cardou, M.; Klimaszewska, K. Simplified and improved somatic embriogenesis for clonal propagation or Pinus pinaster (Ait.). Plant Cell Rep. 2006, 25, 767–776. [Google Scholar] [CrossRef] [PubMed]
- Larkin, P.J.; Scowcroft, W.R. Somaclonal variation—A novel source of variability from cell cultures for plant improvement. Theor. Appl. Genet. 1981, 60, 197–214. [Google Scholar] [CrossRef] [PubMed]
- Tulecke, W. Somatic embryogenesis in woody perennials. In Cell and Tissue Culture in Forestry; Bonga, J.M., Durzan, D.J., Eds.; Martinus Nijhoff: Dordrecht, Germany, 1987; Volume 2, pp. 61–91. [Google Scholar]
- Bhojwani, S.S.; Dantu, P.K. Somaclonal variation. In Plant Tissue Culture: An Introductory Text; Springer: New Delhi, Indian, 2013; pp. 141–154. [Google Scholar]
- Burg, K.; Helmersson, A.; Bozhkov, P.; von Arnold, S. Developmental and genetic variation in nuclear microsatellite stability during somatic embryogenesis in pine. J. Exp. Bot. 2007, 58, 687–698. [Google Scholar] [CrossRef] [PubMed]
- Leva, A.; Petruchelli, R. Monitoring of cultivar identity in micropropagated olive plants using RAPD and ISSR markers. Biol. Plantarum. 2012, 56, 373–376. [Google Scholar] [CrossRef]
- Miguel, C.; Marum, L. An epigenetic view of plant cells cultured in vitro: Somaclonal variation and beyond. J. Exp. Bot. 2011, 62, 3713–3725. [Google Scholar] [CrossRef] [PubMed]
- De Miguel, M.; Cabezas, J.A.; de María, N.; Sánchez-Gómez, D.; Guevara, M.Á.; Vélez, M.D.; Sáez-Laguna, E.; Díaz, L.M.; Mancha, J.A.; Barbero, M.C.; et al. Genetic control of functional traits related to photosynthesis and water use efficiency in Pinus pinaster Ait. drought response: Integration of genome annotation, allele association and QTL detection for candidate gene identification. BMC Genom. 2014, 15, 464. [Google Scholar] [CrossRef]
- MacKay, J.; Dean, J.F.D.; Plomion, C.; Peterson, D.G.; Cánovas, F.M.; Pavy, N.; Ingvarsson, P.K.; Savolainen, O.; Guevara, M.A.; Fluch, S.; et al. Towards decoding the conifer giga-genome. Plant Mol. Biol. 2012, 80, 555–569. [Google Scholar] [CrossRef] [PubMed]
- Promoting a Functional and Comparative Understanding of the Conifer Genome Implementing Applied Aspects for More Productive and Adapted Forests (ProCoGen). Available online: http://www.procogen.eu/ (accessed on 10 November 2016).
- Hashmi, G.; Huettel, R.; Meyer, R.; Krusberg, L.; Hammerschlag, F. RAPD analysis of somaclonal variants derived from embryo callus cultures of peach. Plant Cell Rep. 1997, 16, 624–627. [Google Scholar] [CrossRef]
- Renau-Morata, B.; Arrillaga, I.; Segura, J. In vitro storage of cedar shoot cultures under minimal growth conditions. Plant Cell Rep. 2006, 25, 636–642. [Google Scholar] [CrossRef] [PubMed]
- Orbovic, V.; Calovic, M.; Viloria, Z.; Nielsen, B.; Gmitter, F.; Castle, W.; Grosser, J. Analysis of genetic variability in various tissue culture-derived lemon plant populations using RAPD and flow cytometry. Euphytica 2008, 161, 329–335. [Google Scholar] [CrossRef]
- Carvalho, L.C.; Goulao, L.; Oliveira, C.; Gonçalves, J.C.; Amancio, S. RAPD assessment for identification of clonal identity and genetic stability of in vitro propagated chestnut hybrids. Plant Cell Tissue Organ Cult. 2004, 77, 23–27. [Google Scholar] [CrossRef]
- Prado, M.J.; Herrera, M.T.; Vázquez, R.A.; Romo, S.; González, M.V. Micropropagation of two selected male kiwifruit and analysis of genetic variation with AFLP markers. HortSci 2005, 40, 740–746. [Google Scholar]
- Popescu, C.F.; Flak, A.; Glimelius, K. Application of AFLPs to characterize somaclonal variation in anther-derived grapevines. Vitis 2002, 41, 177–182. [Google Scholar]
- Xiangqian, Li.; Mingliang, Xu.; Schuyler, S.; Korban, S. DNA methylation profiles differ between field- and in vitro grown leaves of apple. J. Plant Physiol. 2002, 159, 1229–1234. [Google Scholar]
- Etienne, H.; Bertrand, B. Somaclonal variation in Coffea arabica: Effects of genotype and embryogenic cell suspension age on frequency and phenotype of variants. Tree Physiol. 2003, 23, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Lopes, T.; Pinto, G.; Loureiro, J.; Costa, A.; Santos, C. Determination of genetic stability in long-term somatic embryogenic cultures and derived plantlets of cork oak using microsatellite markers. Tree Physiol. 2006, 26, 1145–1152. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.; Rajora, O. Microsatellite DNA somaclonal variation in micropropagated trembling aspen (Populus tremuloides). Plant Cell Rep. 2001, 20, 531–536. [Google Scholar]
- Hanacek, P.; Havel, L.; Truksa, M. Characterisation and determination of genetic stability of early somatic embryo culture clones of Norway spruce using RAPD markers. Biologia 2002, 57, 517–521. [Google Scholar]
- Harvengt, L.; Trontin, J.F.; Reymond, I.; Canlet, F.; Pâques, M. Molecular evidence of true-to-type propagation of a 3-year-old Norway spruce through somatic embryogenesis. Planta 2001, 213, 828–832. [Google Scholar] [CrossRef] [PubMed]
- Helmersson, A.; von Arnold, S.; Burg, K.; Bozhkov, P.V. High stability of nuclear microsatellite loci during the early stages of somatic embryogenesis in Norway spruce. Tree Physiol. 2004, 24, 1181–1186. [Google Scholar] [CrossRef] [PubMed]
- Marum, L.; Loureiro, J.; Rodriguez, E.; Oliveira, M.; Miguel, C. Flow cytometric and morphological analyses of Pinus pinaster somatic embryogenesis. Biotech 2009, 143, 288–295. [Google Scholar] [CrossRef] [PubMed]
- Marum, L.; Rocheta, M.; Maroco, J.; Oliveira, M.M.; Miguel, C. Analysis of genetic stability at SSR loci during somatic embryogenesis in maritime pine (Pinus pinaster). Plant Cell Rep. 2009, 28, 673–682. [Google Scholar] [CrossRef] [PubMed]
- Mariette, S.; Changné, D.; Decroocq, S.; Vendramin, G.; Lalanne, C.; Madur, D.; Plomion, C. Microsatellite markers for Pinus pinaster Ait. Ann. For. Sci. 2001, 58, 203–206. [Google Scholar] [CrossRef]
- Chagné, D.; Brown, G.R.; Lalanne, C.; Madur, D.; Pot, D.; Plomion, C. Comparative genome and QTL mapping between maritime and loblolly pines. Mol. Breed. 2003, 12, 185–195. [Google Scholar] [CrossRef]
- Chancerel, E.; Lepoittevin, C.; Le Provost, G.; Lin, Y.C.; Jaramillo-Correa, J.P.; Eckert, A.J.; Wegrzyn, J.L.; Zelenika, D.; Boland, A.; Frigerio, J.M.; et al. Development and implementation of a highly-multiplexed SNP array for genetic mapping in maritime pine and comparative mapping with loblolly pine. BMC Genom. 2011, 12, 368. [Google Scholar] [CrossRef] [PubMed]
- De Miguel, M.; de Maria, N.; Guevara, M.A.; Diaz, L.; Sáez-Laguna, E.; Sánchez-Gómez, D.; Chancerel, E.; Aranda, I.; Collada, C.; Plomion, C.; et al. Annotated genetic linkage maps of Pinus pinaster Ait. from a Central Spain population using microsatellite and gene based markers. BMC Genom. 2012, 13, 527. [Google Scholar] [CrossRef] [PubMed]
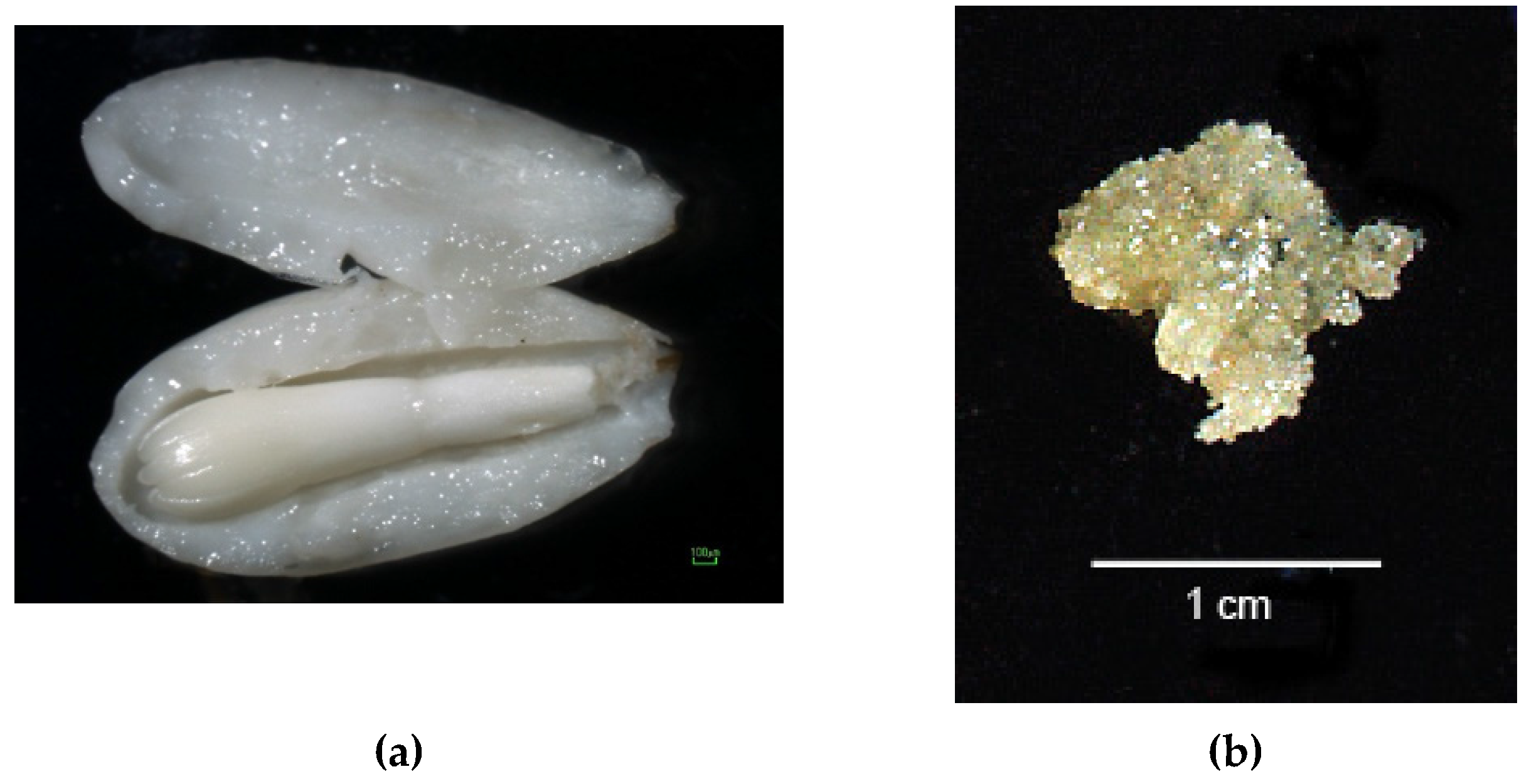

| Species | Haploid Tissue Obtained | References |
|---|---|---|
| Larix decidua | embryogenic cultures and plantlets | [10,11,12,13,14,15] |
| Larix leptolepis | embryogenic cultures | [12] |
| L. leptolepis x L. decidua reciprocal crosses | embryogenic cultures | [12] |
| Larix decidua | embryogenic tissue from protoplast | [14] |
| Larix sibirica | proliferating calli | [18] |
| Picea abies | organogenic cultures | [8] |
| Picea abies | embryogenic lines | [9] |
| Picea sitchensis | proliferating calli | [7] |
| Pinus pinaster | proliferating calli | [19] |
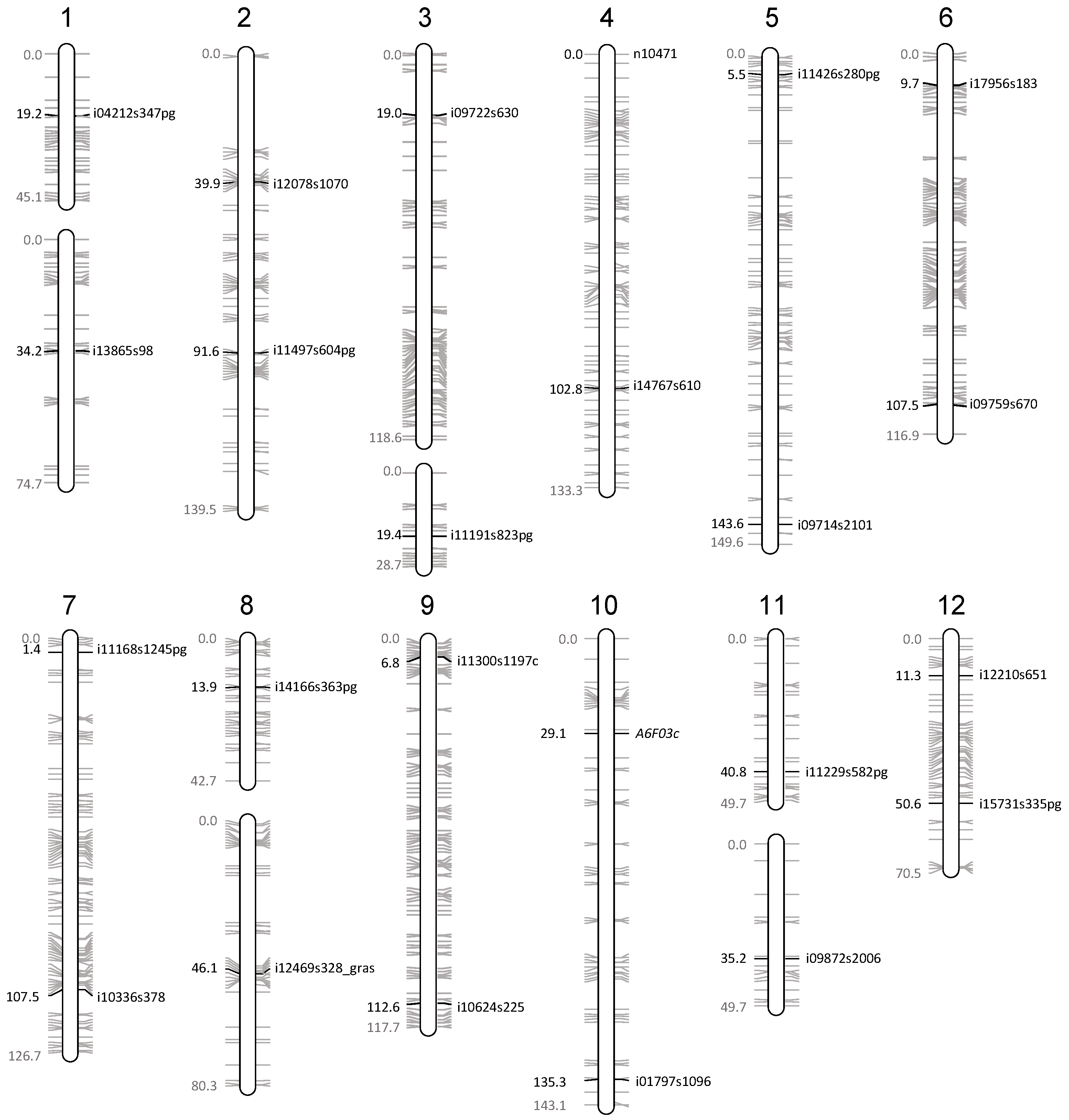
| Chromosome | Marker Name | Marker Type | Oria 6 Genetic Map | Genotyping | ||||
|---|---|---|---|---|---|---|---|---|
| Number | Arm | Position (cM) | LG | LG Size (cM) | Oria 6 | Callus State | ||
| 1 | upper | i04212s347pg | SNP | 19 | 1a | 45 | heteroz. | haploid |
| 1 | lower | i13865s98 | SNP | 34 | 1b | 75 | heteroz. | haploid |
| 2 | upper | i12078s1070 | SNP | 39 | 2 | 139 | heteroz. | haploid |
| 2 | lower | i11497s604 | SNP | 92 | 2 | heteroz. | haploid | |
| 3 | upper | i09722s630 | SNP | 19 | 3a | 119 | heteroz. | haploid |
| 3 | lower | i11191s823pg | SNP | 19 | 3b | 29 | heteroz. | haploid |
| 4 | upper | n10471 | SNP | 0 | 4 | 133 | heteroz. | haploid |
| 4 | lower | i14767s610 | SNP | 103 | 4 | heteroz. | haploid | |
| 5 | upper | i11426s280 | SNP | 5 | 5 | 150 | heteroz. | haploid |
| 5 | lower | i09714s2101 | SNP | 144 | 5 | heteroz. | haploid | |
| 6 | upper | i17956s183 | SNP | 10 | 6 | 117 | heteroz. | haploid |
| 6 | lower | i09759s670 | SNP | 108 | 6 | heteroz. | haploid | |
| 7 | upper | i11168s1245 | SNP | 1 | 7 | 127 | heteroz. | haploid |
| 7 | lower | i10336s378 | SNP | 107 | 7 | heteroz. | haploid | |
| 8 | upper | i14166s363 | SNP | 14 | 8a | 43 | heteroz. | haploid |
| 8 | lower | 12469s328_gras | SNP | 46 | 8b | 80 | heteroz. | haploid |
| 9 | upper | i11300s1197 | SNP | 7 | 9 | 118 | heteroz. | haploid |
| 9 | lower | i10624s225 | SNP | 113 | 9 | heteroz. | haploid | |
| 10 | upper | A6F03 | SSR | 29 | 10 | 143 | heteroz. | haploid |
| 10 | lower | i01797s1096 | SNP | 135 | 10 | heteroz. | haploid | |
| 11 | upper | i13232s811 | SNP | 23 | 11a | 50 | heteroz. | haploid |
| 11 | lower | i09872s2006 | SNP | 35 | 11b | heteroz. | haploid | |
| 12 | upper | i12210s651 | SNP | 11 | 12 | 71 | heteroz. | haploid |
| 12 | lower | i15731s335 | SNP | 51 | 12 | heteroz. | haploid | |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cabezas, J.A.; Morcillo, M.; Vélez, M.D.; Díaz, L.; Segura, J.; Cervera, M.T.; Arrillaga, I. Haploids in Conifer Species: Characterization and Chromosomal Integrity of a Maritime Pine Cell Line. Forests 2016, 7, 274. https://doi.org/10.3390/f7110274
Cabezas JA, Morcillo M, Vélez MD, Díaz L, Segura J, Cervera MT, Arrillaga I. Haploids in Conifer Species: Characterization and Chromosomal Integrity of a Maritime Pine Cell Line. Forests. 2016; 7(11):274. https://doi.org/10.3390/f7110274
Chicago/Turabian StyleCabezas, José Antonio, Marian Morcillo, María Dolores Vélez, Luis Díaz, Juan Segura, María Teresa Cervera, and Isabel Arrillaga. 2016. "Haploids in Conifer Species: Characterization and Chromosomal Integrity of a Maritime Pine Cell Line" Forests 7, no. 11: 274. https://doi.org/10.3390/f7110274
APA StyleCabezas, J. A., Morcillo, M., Vélez, M. D., Díaz, L., Segura, J., Cervera, M. T., & Arrillaga, I. (2016). Haploids in Conifer Species: Characterization and Chromosomal Integrity of a Maritime Pine Cell Line. Forests, 7(11), 274. https://doi.org/10.3390/f7110274






