Effects of Cutting, Pruning, and Grafting on the Expression of Age-Related Genes in Larix kaempferi
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. RNA Extraction and cDNA Synthesis
2.3. Sequence Analysis, Full-Length cDNA Cloning, and Annotation
2.4. Quantitative Reverse Transcription Polymerase Chain Reaction
3. Results and Discussion
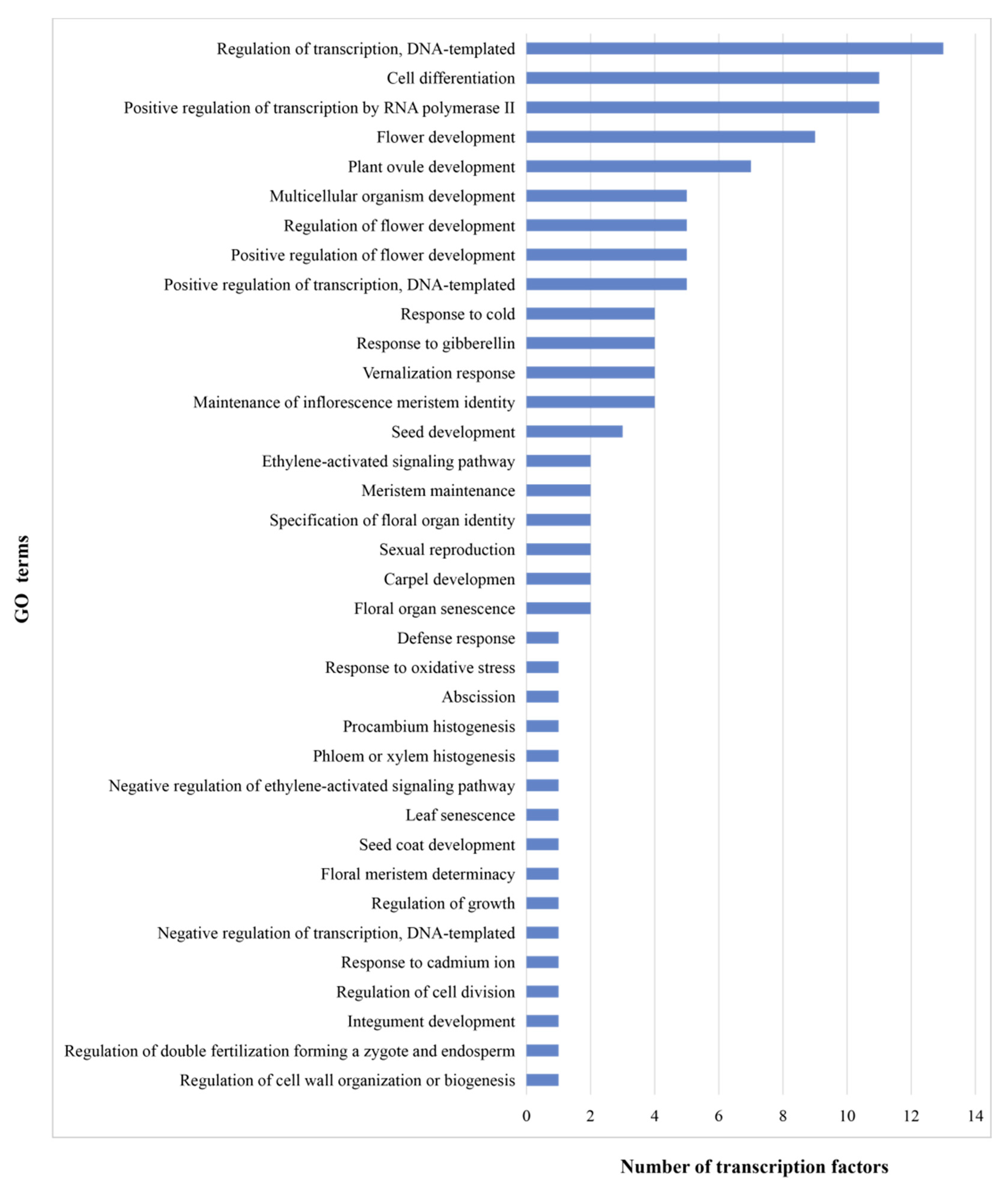
3.1. Cloning and Annotation of 20 Transcription Factors
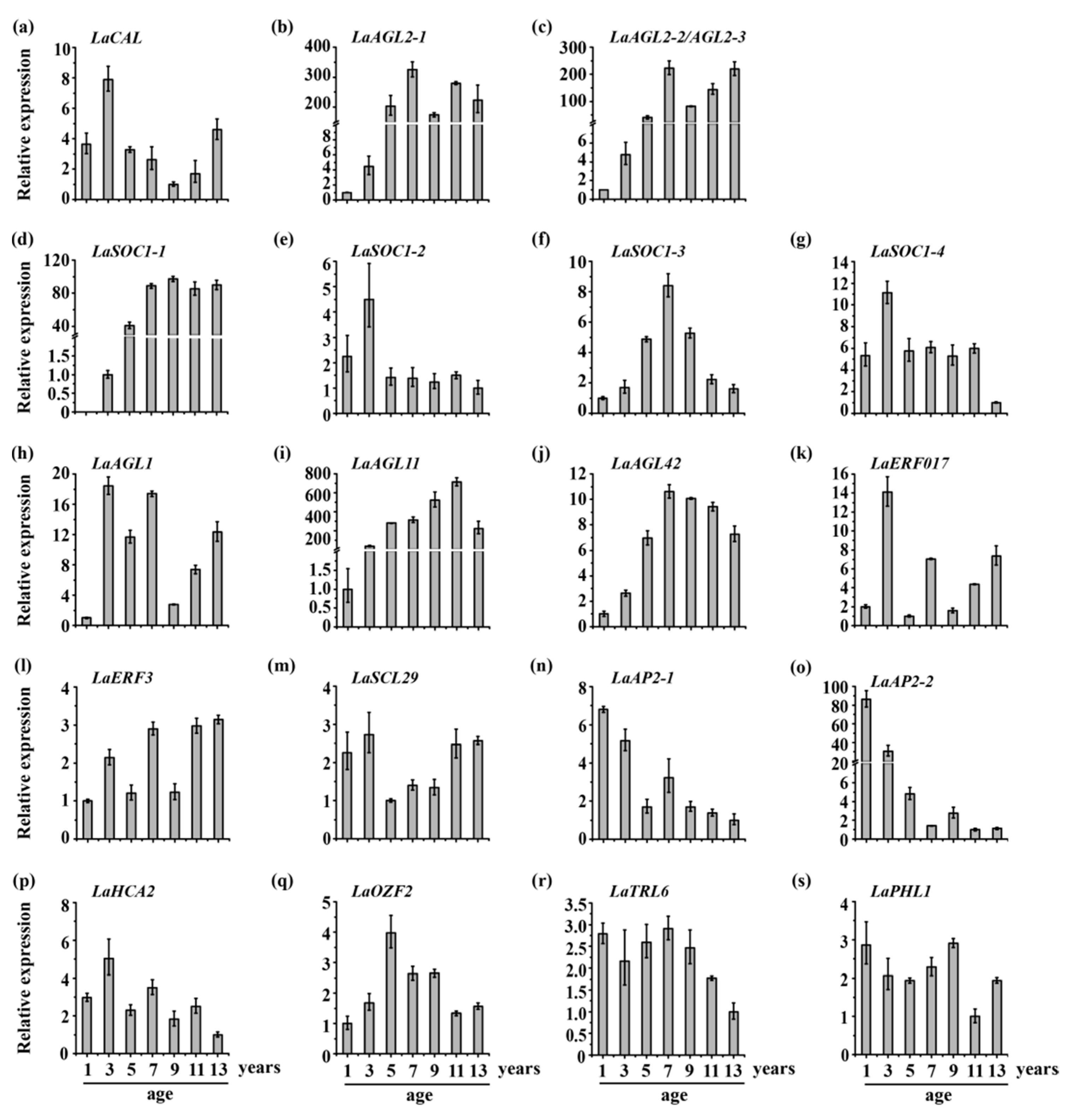
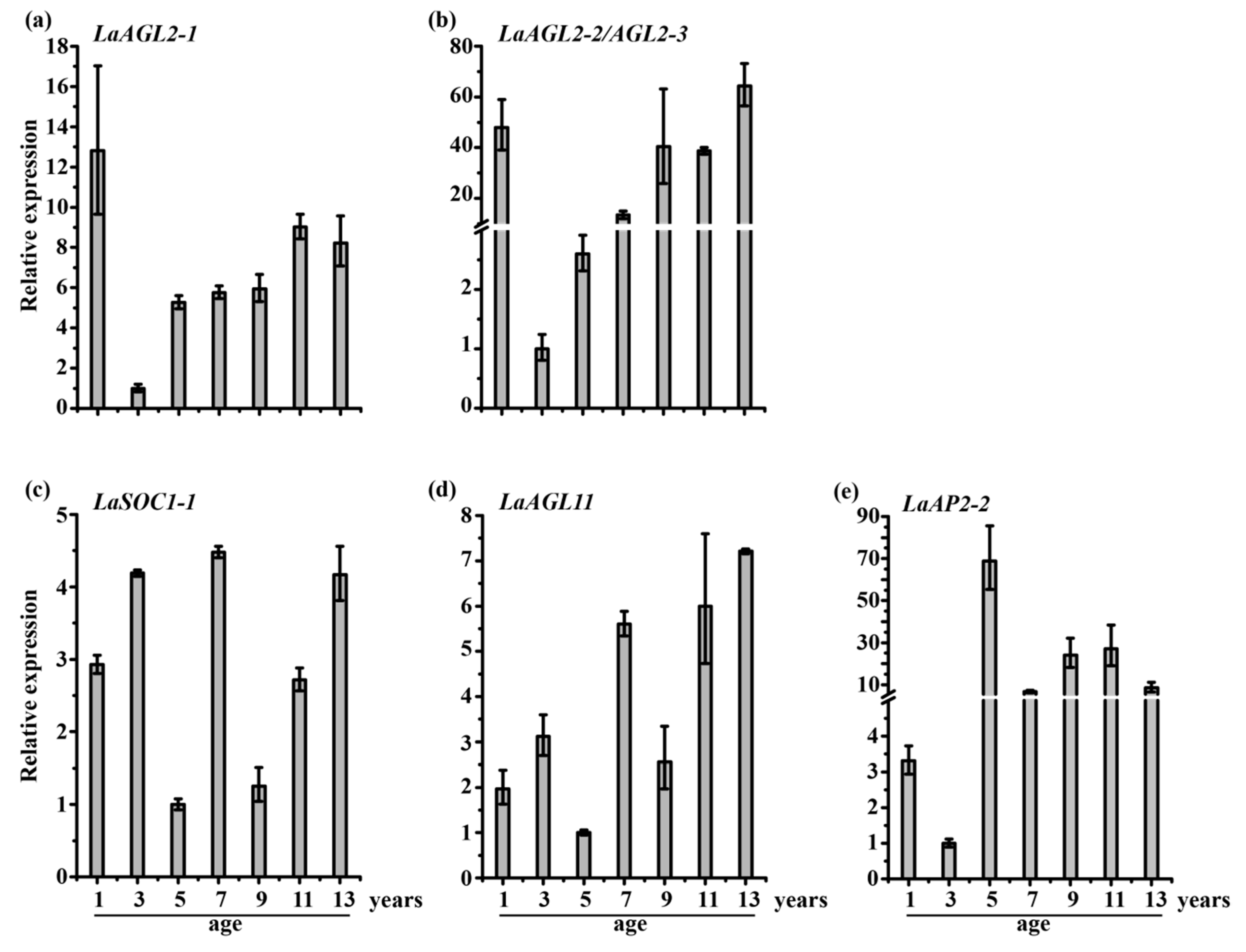
3.2. Expression Patterns of 20 Transcription Factors during Larch Tree Aging
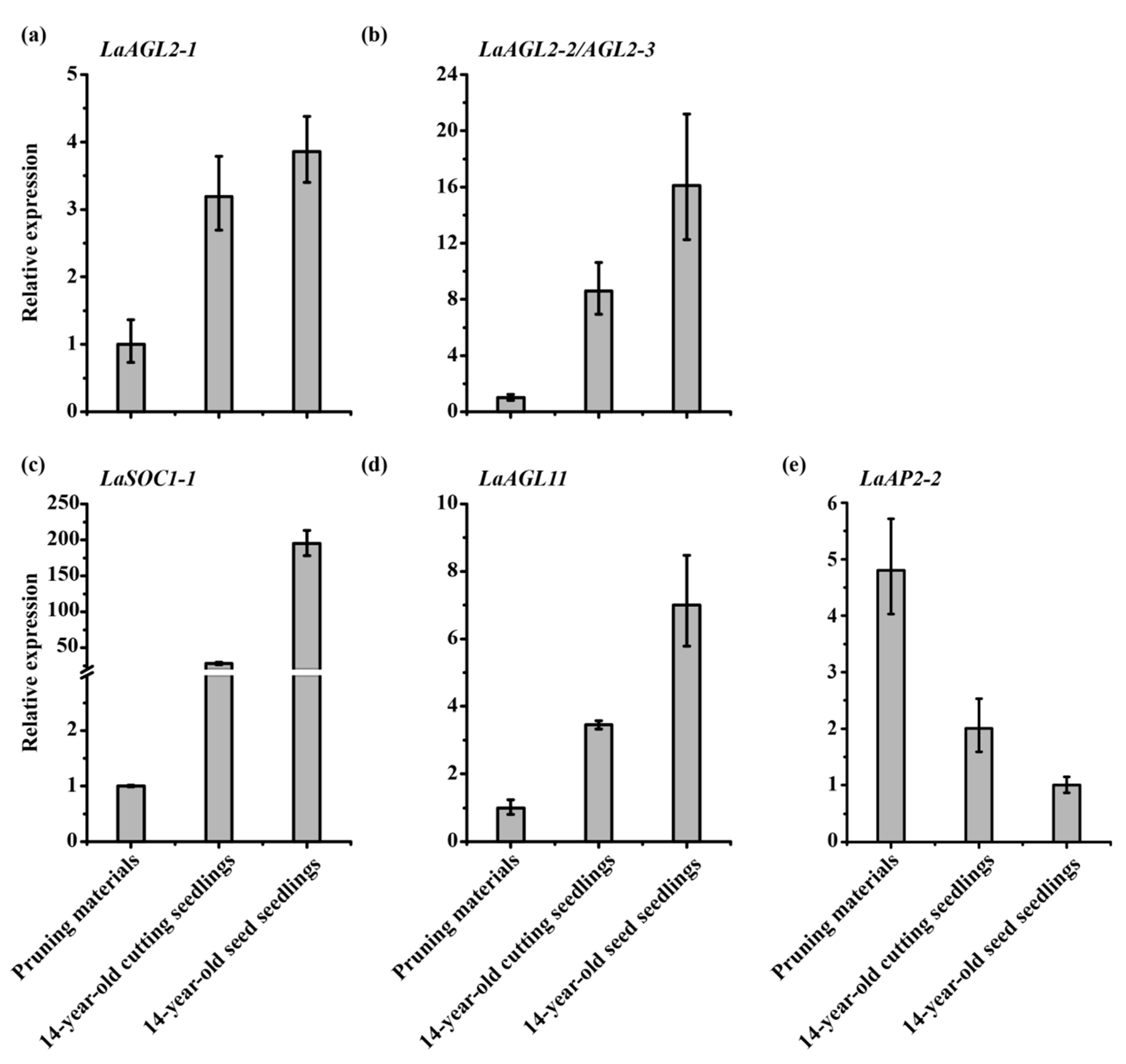
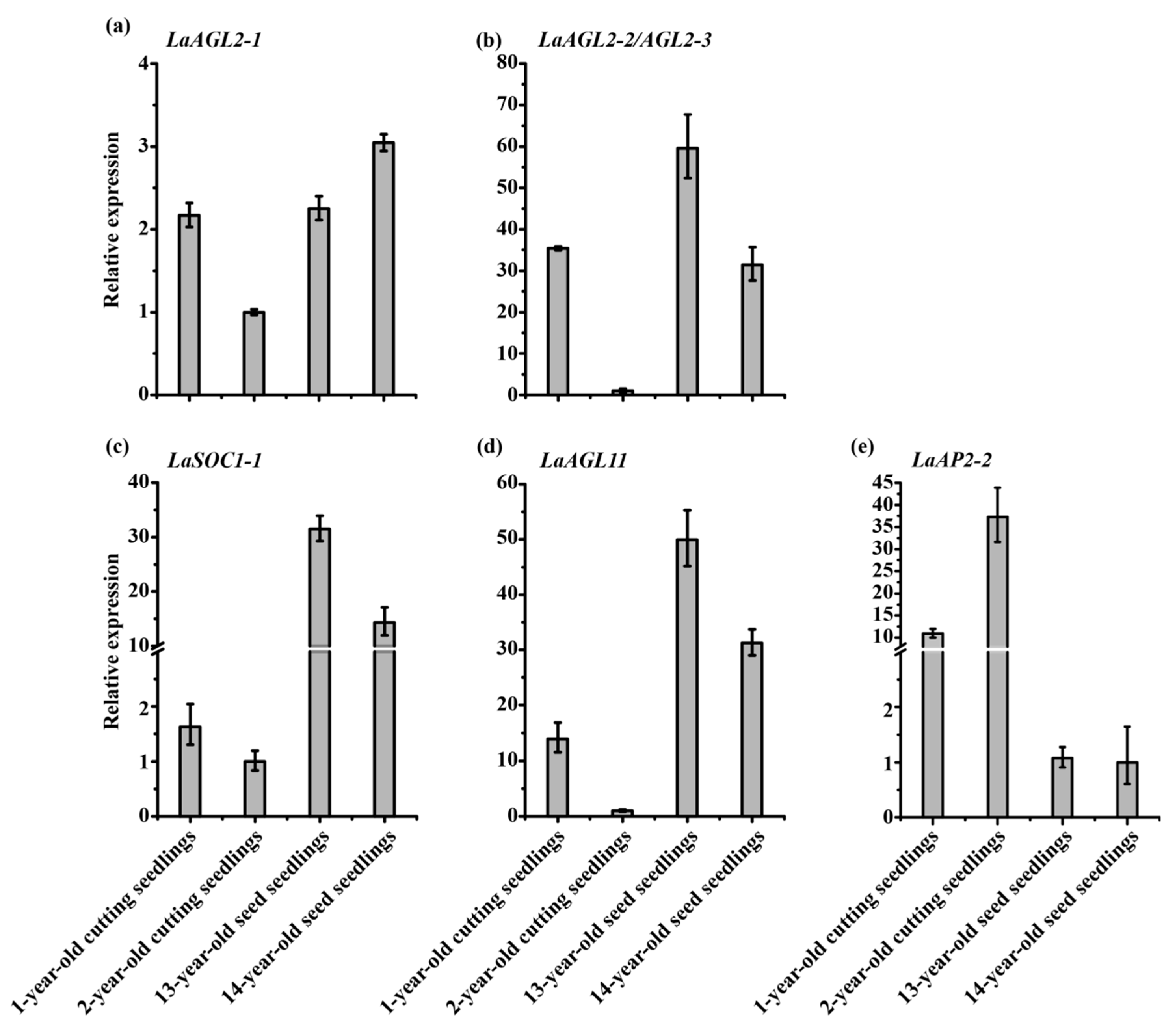
3.3. Effects of Pruning and Cutting on the Expression of Six Age-Related Genes
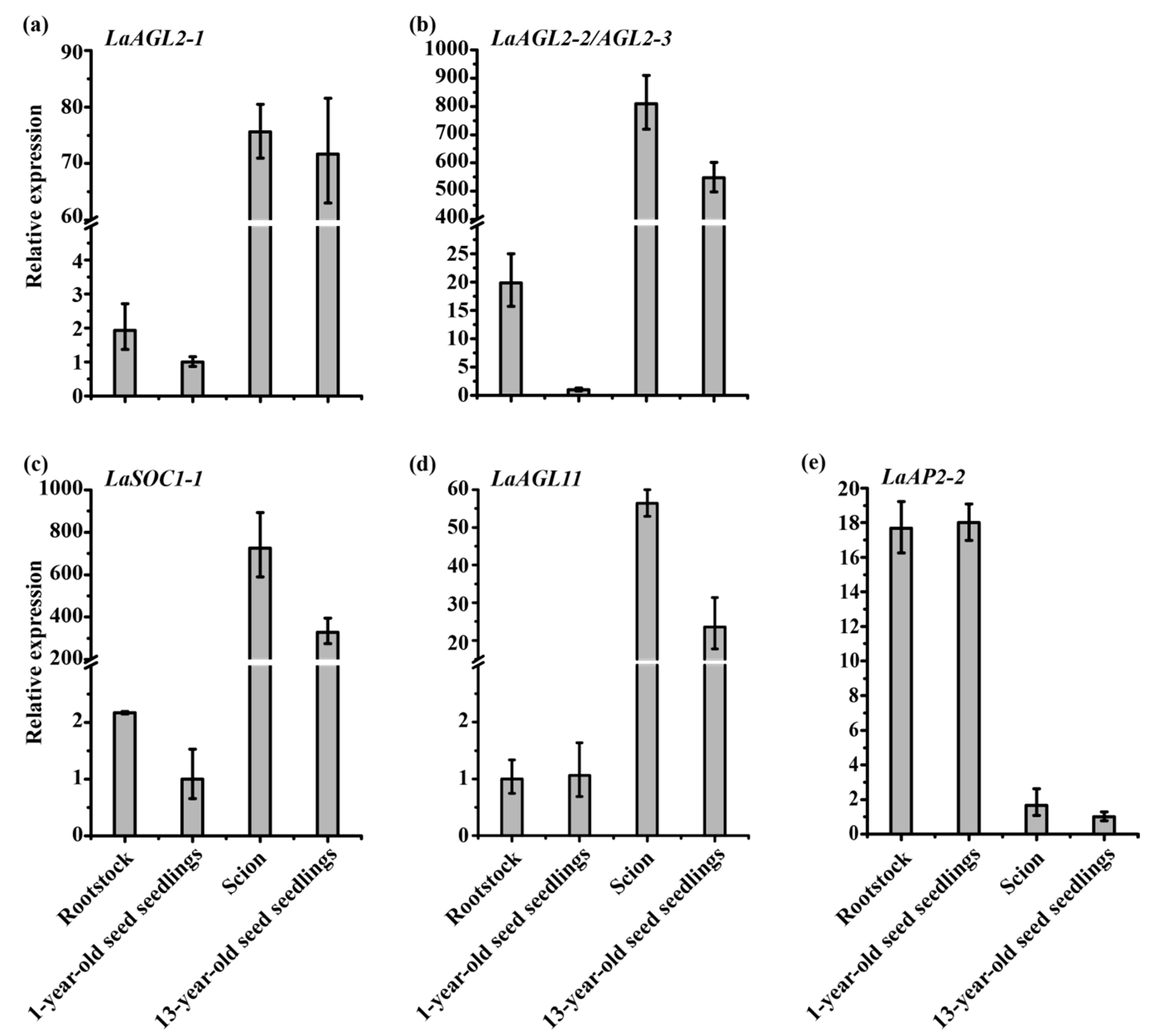
3.4. Expression Patterns of the Six Age-Related Genes after Grafting
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Díaz-Sala, C. Direct reprogramming of adult somatic cells toward adventitious root formation in forest tree species: The effect of the juvenile–adult transition. Front. Plant Sci. 2014, 5, 310. [Google Scholar] [CrossRef] [PubMed]
- Massoumi, M.; Krens, F.A.; Visser, R.G.F.; de Klerk, G.J.M. Azacytidine and miR156 promote rooting in adult but not in juvenile Arabidopsis tissues. J. Plant Physiol. 2017, 208, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Hand, P.; Besford, R.T.; Richardson, C.M.; Peppitt, S.D. Antibodies to phase related proteins in juvenile and mature Prunus avium. Plant Growth Regul. 1996, 20, 25–29. [Google Scholar] [CrossRef]
- Wendling, I.; Trueman, S.J.; Xavier, A. Maturation and related aspects in clonal forestry—part II: Reinvigoration, rejuvenation and juvenility maintenance. New For. 2014, 45, 473–486. [Google Scholar] [CrossRef]
- Moon, H.K.; Yi, J.S. Cutting propagation of Quercus acutissima clones after rejuvenation through serial grafting. Ann. Sci. For. 1993, 50, 314–318. [Google Scholar] [CrossRef]
- Wang, J.; Jiang, L.; Wu, R. Plant grafting: How genetic exchange promotes vascular reconnection. New Phytol. 2017, 214, 56–65. [Google Scholar] [CrossRef]
- Majada, J.; Martínez-Alonso, C.; Feito, I.; Kidelman, A.; Aranda, I.; Alía, R. Mini-cuttings: An effective technique for the propagation of Pinus pinaster Ait. New For. 2011, 41, 399–412. [Google Scholar] [CrossRef]
- Jung, J.H.; Ju, Y.; Seo, P.J.; Lee, J.H.; Park, C.M. The SOC1-SPL module integrates photoperiod and gibberellic acid signals to control flowering time in Arabidopsis. Plant J. 2012, 69, 577–588. [Google Scholar] [CrossRef]
- Murai, K.; Miyamae, M.; Kato, H.; Takumi, S.; Ogihara, Y. WAP1, a Wheat APETALA1 homolog, plays a central role in the phase transition from vegetative to reproductive growth. Plant Cell Physiol. 2003, 44, 1255–1265. [Google Scholar] [CrossRef]
- Preston, J.C.; Hileman, L.C. Functional evolution in the plant SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) gene family. Front. Plant Sci. 2013, 4, 80. [Google Scholar] [CrossRef]
- Carlsbecker, A.; Tandre, K.; Johanson, U.; Englund, M.; Engström, P. The MADS-box gene DAL1 is a potential mediator of the juvenile-to-adult transition in Norway spruce (Picea abies). Plant J. 2004, 40, 546–557. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Park, M.Y.; Wang, L.J.; Koo, Y.; Chen, X.Y.; Weigel, D.; Poethig, R.S. MiRNA control of vegetative phase change in trees. PLoS Genet. 2011, 7, e1002012. [Google Scholar] [CrossRef] [PubMed]
- Curaba, J.; Talbot, M.; Li, Z.; Helliwell, C. Over-expression of microRNA171 affects phase transitions and floral meristem determinancy in barley. BMC Plant Biol. 2013, 13, 6. [Google Scholar] [CrossRef]
- Sgamma, T.; Jackson, A.; Muleo, R.; Thomas, B.; Massiah, A. TEMPRANILLO is a regulator of juvenility in plants. Sci. Rep. 2014, 4, 3704. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, R.; Wang, C.T.; Ma, C.; Shevchenko, O.; Dye, S.J.; Puzey, J.R.; Etherington, E.; Sheng, X.; Meilan, R.; Strauss, S.H.; et al. Populus CEN/TFL1 regulates first onset of flowering, axillary meristem identity and dormancy release in Populus. Plant J. 2010, 62, 674–688. [Google Scholar] [CrossRef]
- Mimida, N.; Kotoda, N.; Ueda, T.; Igarashi, M.; Hatsuyama, Y.; Iwanami, H.; Moriya, S.; Abe, K. Four TFL1/CEN -Like genes on distinct linkage groups show different expression patterns to regulate vegetative and reproductive development in Apple (Malus × domestica Borkh.). Plant Cell Physiol. 2009, 50, 394–412. [Google Scholar] [CrossRef]
- Fernández-Ocaña, A.; García-López, M.C.; Jiménez-Ruiz, J.; Saniger, L.; Macías, D.; Navarro, F.; Oya, R.; Belaj, A.; Rosa, R.; Corpas, F.J.; et al. Identification of a gene involved in the juvenile-to-adult transition (JAT) in cultivated olive trees. Tree Genet. Genomes 2010, 6, 891–903. [Google Scholar] [CrossRef]
- Li, A.; Zhou, Y.; Jin, C.; Song, W.; Chen, C.; Wang, C. LaAP2L1, a heterosis-associated AP2/EREBP transcription factor of Larix, increases organ size and final biomass by affecting cell proliferation in Arabidopsis. Plant Cell Physiol. 2013, 54, 1822–1836. [Google Scholar] [CrossRef]
- Nilsson, L.; Carlsbecker, A.; Sundås-Larsson, A.; Vahala, T. APETALA2 like genes from Picea abies show functional similarities to their Arabidopsis homologues. Planta 2007, 225, 589–602. [Google Scholar] [CrossRef]
- Xiang, W.B.; Li, W.F.; Zhang, S.G.; Qi, L.W. Transcriptome-wide analysis to dissect the transcription factors orchestrating the phase change from vegetative to reproductive development in Larix kaempferi. Tree Genet. Genomes 2019, 15, 68. [Google Scholar] [CrossRef]
- Ahsan, M.U.; Hayward, A.; Irihimovitch, V.; Fletcher, S.; Tanurdzic, M.; Pocock, A.; Beveridge, C.A.; Mitter, N. Juvenility and vegetative phase transition in tropical/subtropical tree crops. Front. Plant Sci. 2019, 10, 729. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q.; Bu, D.; Zhang, J.; Wu, Y.; Pei, D. The transcriptome landscape of walnut interspecies hybrid (Juglans hindsii × Juglans regia) and regulation of cambial activity in relation to grafting. Front. Genet. 2019, 10, 577. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Lu, M.; Zhu, B.; Wang, R.; Wan, X.; Deng, W.W.; Zhang, Z.Z. Integrated transcriptomic and phytochemical analyses provide insights into characteristic metabolites variation in leaves of 1-year-old grafted tea (Camellia sinensis). Tree Genet. Genomes 2019, 15, 58. [Google Scholar] [CrossRef]
- Li, W.F.; Yang, W.H.; Zhang, S.G.; Han, S.Y.; Qi, L.W. Transcriptome analysis provides insights into wood formation during larch tree aging. Tree Genet. Genomes 2017, 13, 19. [Google Scholar] [CrossRef]
- Wang, X.S.; Hu, X.S.; Wang, Y.C.; Nie, J.; Liu, S.M. The effect of age and basal-cut shapes of cutting on rooting of Larix leptolepis. For. Res. 1993, 6, 627–632. [Google Scholar]
- Wang, J.H.; Sun, X.M.; Wang, X.S.; Xu, C.L.; Ding, B.; Wang, X.D. Effects of age, type of auxin and treatment concentration on rooting ability of Larix leptolepis. For. Res. 2006, 19, 102–108. [Google Scholar]
- Trueman, S.J. Clonal propagation and storage of subtropical pines in Queensland, Australia. S. Afr. For. J. 2006, 208, 49–52. [Google Scholar] [CrossRef]
- Rosier, C.L.; Frampton, J.; Goldfarb, B.; Wise, F.C.; Blazich, F.A. Stumping height, crown position, and age of parent tree influence rooting of stem cuttings of Fraser fir. HortScience 2005, 40, 771–777. [Google Scholar] [CrossRef]
- Aimers-Halliday, J.; Menzies, M.I.; Faulds, T.; Holden, D.G.; Low, C.B.; Dibley, M.J. Nursery systems to control maturation in Pinus radiata cuttings, comparing hedging and serial propagation. N. Z. J. For. Sci. 2003, 33, 135–155. [Google Scholar]
- Peer, K.R.; Greenwood, M.S. Maturation, topophysis and other factors in relation to rooting in Larix. Tree Physiol. 2001, 21, 267–272. [Google Scholar] [CrossRef] [PubMed]
- Wendling, I.; Warburton, P.M.; Trueman, S.J. Maturation in Corymbia torelliana × C. citriodora stock plants: Effects of pruning height on shoot production, adventitious rooting capacity, stem anatomy, and auxin and abscisic acid concentrations. Forests 2015, 6, 3763–3778. [Google Scholar] [CrossRef]
- Cameron, R.; Harrison-Murray, R.; Fordham, M.; Judd, H.; Ford, Y.; Marks, T.; Edmondson, R. Rooting cuttings of syringa vulgaris cv. charles joly and corylus avellana cv. aurea: The influence of stock plant pruning and shoot growth. Trees 2003, 17, 451–462. [Google Scholar] [CrossRef]
- Arnaud, Y.; Franclet, A.; Tranvan, H.; Jacques, M. Micropropagation and rejuvenation of Sequoia sempervirens (Lamb) Endl: A review. Ann. Sci. For. 1993, 50, 273–295. [Google Scholar] [CrossRef]
- Huang, L.C.; Chow, T.Y.; Tseng, T.C.; Kuo, C.I.; Liu, S.M.; Ngoh, M.G.; Murashige, T.; Huang, H.J. Association of mitochondrial plasmids with rejuvenation of the coastal redwood, Sequoia sempervirens (D. Don) Endl. Bot. Bull. Acad. Sin. 2003, 44, 25–30. [Google Scholar]
- Ewald, D.; Kretzschmar, U. The influence of micrografting in vitro on tissue culture behavior and vegetative propagation of old European larch trees. Plant Cell Tissue Organ Cult. 1996, 44, 249–252. [Google Scholar]
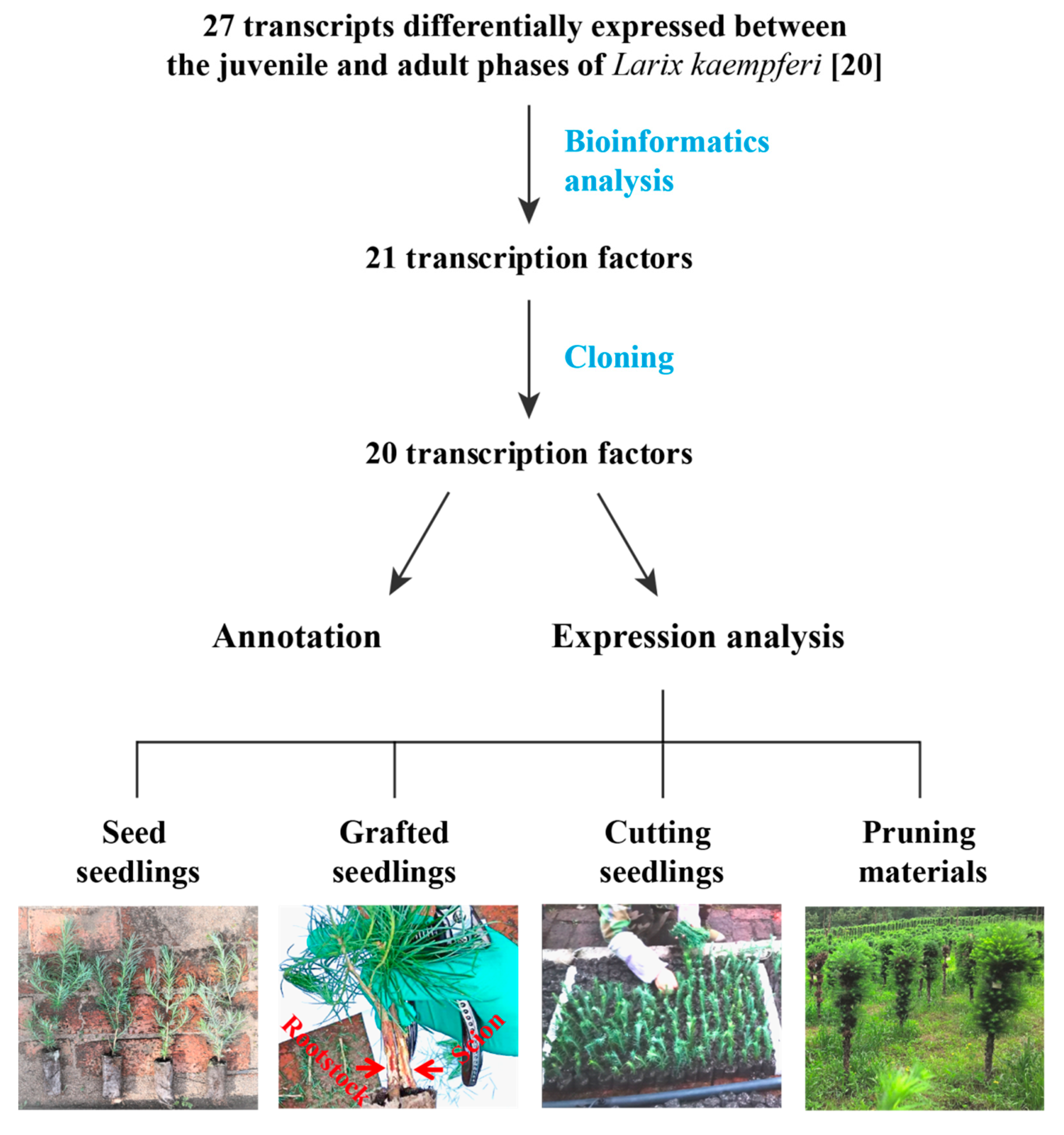
| Samples | Age | Other Information |
|---|---|---|
| Seed Seedlings | 1 year | These seedlings were grown from seeds |
| 3 years | ||
| 5 years | ||
| 7 years | ||
| 9 years | ||
| 11 years | ||
| 13 years | ||
| 14 years | ||
| Grafted Seedlings | 3 months | Rootstocks were from 1-year-old and scions from 12-year-old seed seedlings |
| Cutting Seedlings | 1 year | Cuttings were sampled from 12-year-old seed seedlings |
| 2 years | ||
| 3 years | - | |
| 5 years | - | |
| 7 years | - | |
| 9 years | - | |
| 11 years | - | |
| 13 years | - | |
| 14 years | Cuttings were sampled from 8-year-old seed seedlings | |
| Pruning Materials | 14 years | Cutting seedlings were pruned; they were propagated from cuttings from 8-year-old seed seedlings |
| Family | Name | Transcript ID | Accession Number |
|---|---|---|---|
| MADS-box | LaCAL | comp81209_c0_seq1 | MN790743 |
| LaAGL2-1 | comp125095_c0_seq5 | MN790744 | |
| comp126977_c0_seq13 | |||
| LaAGL2-2 | comp125095_c0_seq9 | MN790745 | |
| comp126977_c0_seq2 | |||
| comp126977_c0_seq5 | |||
| LaAGL2-3 | comp126977_c0_seq16 | MN790746 | |
| LaSOC1-1 | comp128412_c0_seq11 | MN790747 | |
| LaSOC1-2 | comp128412_c0_seq23 | MN790748 | |
| LaSOC1-3 | comp128412_c0_seq8 | MN790749 | |
| comp128471_c0_seq5 | |||
| comp128471_c0_seq24 | |||
| comp128471_c0_seq9 | |||
| LaSOC1-4 | comp129709_c0_seq16 | MN790750 | |
| NC | comp129709_c0_seq2 | ||
| LaAGL1 | comp129017_c0_seq15 | MN790751 | |
| LaAGL11 | comp128471_c0_seq14 | MN790752 | |
| LaAGL42 | comp128471_c0_seq18 | MN790753 | |
| ERF | LaERF017 | comp124322_c0_seq3 | MN790754 |
| LaERF3 | comp129386_c0_seq9 | MN790755 | |
| GRAS | LaSCL29 | comp114072_c0_seq3 | MN790756 |
| AP2 | LaAP2-1 | comp128327_c0_seq16 | MN790757 |
| LaAP2-2 | comp122930_c0_seq2 | MN790758 | |
| DOF | LaHCA2 | comp120092_c0_seq3 | MN790759 |
| C3H | LaOZF2 | comp111742_c0_seq2 | MN790760 |
| MYB_Related | LaTRFL6 | comp128112_c0_seq2 | MN790761 |
| G2-Like | LaPHL1 | comp130729_c0_seq1 | MN790762 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Zang, Q.-L.; Qi, L.-W.; Han, S.-Y.; Li, W.-F. Effects of Cutting, Pruning, and Grafting on the Expression of Age-Related Genes in Larix kaempferi. Forests 2020, 11, 218. https://doi.org/10.3390/f11020218
Zhang Y, Zang Q-L, Qi L-W, Han S-Y, Li W-F. Effects of Cutting, Pruning, and Grafting on the Expression of Age-Related Genes in Larix kaempferi. Forests. 2020; 11(2):218. https://doi.org/10.3390/f11020218
Chicago/Turabian StyleZhang, Yao, Qiao-Lu Zang, Li-Wang Qi, Su-Ying Han, and Wan-Feng Li. 2020. "Effects of Cutting, Pruning, and Grafting on the Expression of Age-Related Genes in Larix kaempferi" Forests 11, no. 2: 218. https://doi.org/10.3390/f11020218
APA StyleZhang, Y., Zang, Q.-L., Qi, L.-W., Han, S.-Y., & Li, W.-F. (2020). Effects of Cutting, Pruning, and Grafting on the Expression of Age-Related Genes in Larix kaempferi. Forests, 11(2), 218. https://doi.org/10.3390/f11020218





