Genetic Consequences of Hybridization in Relict Isolated Trees Pinus sylvestris and the Pinus mugo Complex
Abstract
1. Introduction
2. Materials and Methods
2.1. Material
2.2. DNA Extraction, Amplification and Marker Genotyping
2.3. Diversity and Differentiation Measures
2.4. Biometrics
2.5. Statistical Analyses of Biometric Data
3. Results
3.1. Genetic Diversity and Differentiation
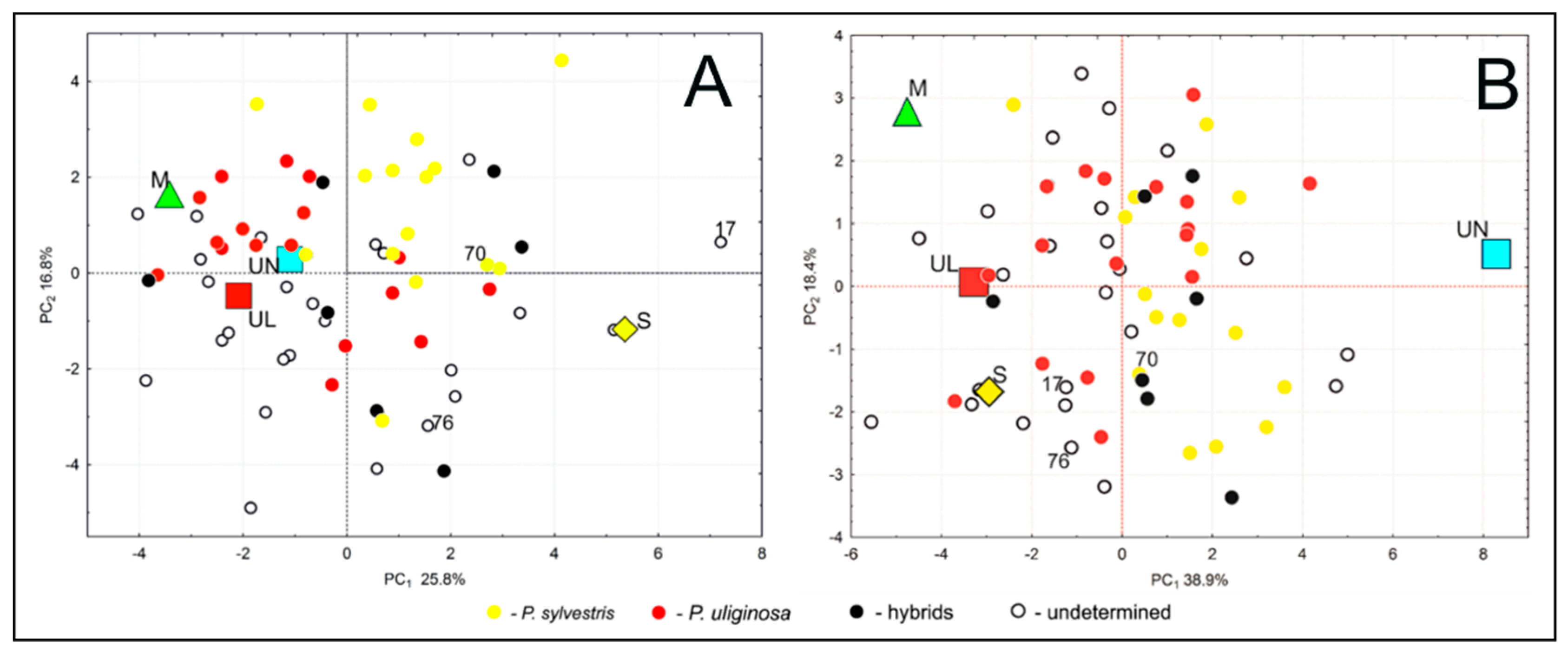
3.2. Phenotypic Variability and Differentiation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Rieseberg, L.H.; Ellstrand, N.C.; Arnold, M. What can molecular and morphological markers tell us about plant hybridization? Crit. Rev. Plant Sci. 1993, 12, 213–241. [Google Scholar] [CrossRef]
- Arnold, M.L. Natural Hybridization and Evolution; Oxford University Press: Oxford, UK, 1997. [Google Scholar]
- Arnold, M.L. Evolution through Genetic Exchange; Oxford University Press: Oxford, UK, 2007; pp. 1–252. [Google Scholar] [CrossRef]
- Mallet, J. Hybridization, ecological races and the nature of species: Empirical evidence for the ease of speciation. Philos. Trans. R Soc. B Biol. Sci. 2008, 363, 2971–2986. [Google Scholar] [CrossRef] [PubMed]
- Soltis, P.S.; Soltis, D.E. The role of hybridization in plant speciation. Annu. Rev. Plant Biol. 2009, 60, 561–588. [Google Scholar] [CrossRef] [PubMed]
- Abbott, R.; Albach, D.; Ansell, S.; Arntzen, J.W.; Baird, S.J.E.; Bierne, N.; Boughman, J.; Brelsford, A.; Buerkle, C.A.; Buggs, R.; et al. Hybridization and speciation. J. Evol. Biol. 2013, 26, 229–246. [Google Scholar] [CrossRef]
- Boratyński, A.; Boratyńska, K.; Lewandowski, A.; Gołąb, Z.; Kiciński, P. Evidence of the possibility of natural reciprocal crosses between Pinus sylvestris and P. uliginosa based on the phenology of reproductive organs. Flora 2003, 198, 377–388. [Google Scholar] [CrossRef]
- Stacy, E.A.; Paritosh, B.; Johnson, M.A.; Price, D.K. Incipient ecological speciation between successional varieties of a dominant tree involves intrinsic postzygotic isolating barriers. Ecol. Evol. 2017, 7, 2501–2512. [Google Scholar] [CrossRef]
- Gernandt, D.S.; López, G.G.; García, S.O.; Liston, A. Phylogeny and classification of Pinus. Taxon 2005, 54, 29–42. [Google Scholar] [CrossRef]
- Kormuťák, A.; Ostrolucká, M.; Vooková, B.; Preťová, A.; Fečková, M. Artificial hybridization of Pinus sylvestris L. and Pinus mugo Turra. Acta. Biol. Crac. Ser. Bot. 2005, 47, 129–134. [Google Scholar]
- Delgado, P.; Salas-Lizana, R.; Vazquez-Lobo, A.; Wegier, A.; Anzidei, M.; Alvarez-Buylla, E.R.; Vendramin, G.G.; Pinero, D. Introgressive hybridization in Pinus montezumae Lamb and Pinus pseudostrobus Lindl. (Pinaceae): Morphological and molecular (cpSSR) evidence. Int. J. Plant Sci. 2007, 168, 861–875. [Google Scholar] [CrossRef]
- Krajmerová, D.; Paule, L.; Zhelev, P.; Voleková, M.; Evtimov, I.; Gagov, V.; Gömöry, D. Natural hybridization in eastern-Mediterranean firs: The case of Abies borisii-regis. Plant Biosyst. 2016, 150, 1189–1199. [Google Scholar] [CrossRef]
- Wachowiak, W.; Palme, A.E.; Savolainen, O. Speciation history of three closely related pines Pinus mugo (T.), P. uliginosa (N.) and P. sylvestris (L.). Mol. Ecol. 2011, 20, 1729–1743. [Google Scholar] [CrossRef] [PubMed]
- Wachowiak, W.; Zaborowska, J.; Labiszak, B.; Perry, A.; Zucca, G.M.; Gonzalez-Martinez, S.C.; Cavers, S. Molecular signatures of divergence and selection in closely related pine taxa. Tree Genet. Genomes 2018, 14. [Google Scholar] [CrossRef] [PubMed]
- Christensen, K.l. Taxonomic revision of the Pinus mugo complex and P. rhaetica (P. mugo sylvestris) (Pinaceae). Nord. J. Bot. 1987, 7, 383–408. [Google Scholar] [CrossRef]
- Businský, R.; Kirschner, J. Pinus mugo and P. uncinata as parents of hybrids a taxonomic and nomenclatural survey. Phyton 2010, 50, 27–57. [Google Scholar]
- Staszkiewicz, J.; Tyszkiewicz, M. Variability of the natural hybrids of Pinus sylvestris L. x Pinus mugo Turra (= P. x rotundata Link) in south-western Poland and in some selected localities of Bohemia and Moravia. Fragm. Flor. Geobot. 1972, 18, 173–191. [Google Scholar]
- Bobowicz, M.A. Differentiation of Pinus sylvestris L. and Pinus mugo Turra, pines from Bór na Czerwonem and from Zieleniec in traits of one- and two-year-old cones. Bull. Soc. Amis. Sci. Lett. Pozn. Ser. D Sci. Biol. 1988, 26, 99–108. [Google Scholar]
- Neet-Sarqueda, C. Genetic differentiation of Pinus sylvestris L. and Pinus mugo aggr. populations in Switzerland. 1994, 43, 207–214. [Google Scholar]
- Christensen, K.I.; Dar, G.H. A morphometric analysis of spontaneous and artificial hybrids of Pinus mugo ×sylvestris (Pinaceae). Nord. J. Bot. 1997, 17, 77–86. [Google Scholar] [CrossRef]
- Boratyńska, K.; Bobowicz, M.A. Pinus uncinata Ramond taxonomy based on needle characters. Plant. Syst. Evol. 2001, 227, 183–194. [Google Scholar] [CrossRef]
- Boratyńska, K.; Boratyński, A.; Lewandowski, A. Morphology of Pinus uliginosa (Pinaceae) needles from populations exposed to and isolated from the direct influence of Pinus sylvestris. Bot. J. Linn. Soc. 2003, 142, 83–91. [Google Scholar] [CrossRef]
- Boratyńska, K.; Boratyński, A. Taxonomic differences among closely related pines Pinus sylvestris, P. mugo, P. uncinata, P. rotundata and P. uliginosa as revealed in needle sclerenchyma cells. Flora 2007, 202, 555–569. [Google Scholar] [CrossRef]
- Boratyńska, K.; Jasińska, A.; Ciepłuch, E. Effect of tree age on needle morphology and anatomy of Pinus uliginosa and Pinus silvestris—Species-specific character separation during ontogenesis. Flora Morphol. Distrib. Funct. Ecol. Plants 2008, 203, 617–626. [Google Scholar] [CrossRef]
- Venturas, M.; García Álvarez, S.; Alcantra, M.F.; Collada, C.; Gil, L. Species selection for reforestations: What happens with historic local extinctions and habitat protection zones? A case study in the Cantabrian Range. Eur. J. Res. 2013, 132, 107–120. [Google Scholar] [CrossRef]
- Lewandowski, A.; Smoćko, J.; Boratynska, K.; Boratynski, A. Genetic differences between two Polish populations of Pinus uliginosa, compared to P. sylvestris and P. mugo. Dendrobiology 2002, 48, 51–57. [Google Scholar]
- Wachowiak, W.; Celinski, K.; Prus-Glowacki, W. Evidence of natural reciprocal hybridisation between Pinus uliginosa and P. sylvestris in the sympatric population of the species. Flora 2005, 200, 563–568. [Google Scholar] [CrossRef]
- Wachowiak, W.; Lewandowski, A.; Prus-Głowacki, W. Reciprocal controlled crosses between Pinus sylvestris and P. mugo verified by a species-specific cpDNA marker. J. Appl. Genet. 2005, 46, 41–43. [Google Scholar]
- Kormutak, A.; Vookova, B.; Manka, P.; Salaj, J.; Camek, V.; Gömöry, D. Abortive embryogenesis in hybrid swarm populations of Pinus sylvestris L. and Pinus mugo Turra. Trees 2008, 22, 657–662. [Google Scholar] [CrossRef]
- Wachowiak, W.; Prus-Glowacki, W. Hybridisation processes in sympatric populations of pines Pinus sylvestris L., P.mugo Turra and P.uliginosa Neumann. Plant Syst. Evol. 2008, 271, 29–40. [Google Scholar] [CrossRef]
- Jasińska, A.K.; Wachowiak, W.; Muchewicz, E.; Boratyńska, K.; Montserrat, J.M.; Boratyński, A. Cryptic hybrids between Pinus uncinata and P. sylvestris. Bot. J. Linn. Soc. 2010, 163, 473–485. [Google Scholar] [CrossRef]
- Wachowiak, W.; Cavers, S.; Żukowska, W.B. Interspecific gene flow and ecological selection in a pine (Pinus sp.) contact zone. Plant Syst. Evol. 2015, 301, 1643–1652. [Google Scholar] [CrossRef]
- Wachowiak, W.; Zukowska, W.B.; Wojkiewicz, B.; Cavers, S.; Litkowiec, M. Hybridization in contact zone between temperate European pine species. Tree Genet. Genomes 2016, 12. [Google Scholar] [CrossRef]
- Lewandowski, A.; Dering, M. Crossability between Pinus uliginosa and its putative parental species Pinus sylvestris and Pinus mugo. Silvae Genet. 2006, 55, 52–54. [Google Scholar] [CrossRef][Green Version]
- Kormutak, A.; Galgoci, M.; Manka, P.; Koubova, M.; Jopcik, M.; Sukenikova, D.; Bolecek, P.; Gőmőry, D. Field-based artificial crossings indicate partial compatibility of reciprocal crosses between Pinus sylvestris and Pinus mugo and unexpected chloroplast DNA inheritance. Tree Genet. Genomes 2017, 13, 68. [Google Scholar] [CrossRef]
- Liston, A.; Parker-Defeniks, M.; Syring, J.; Willyard, A.; Cronn, R. Interspecific phylogenetic analysis enhances intraspecific phylogeographical inference: A case study in Pinus lambertiana. Mol. Ecol. 2007, 16, 3926–3937. [Google Scholar] [CrossRef]
- Senjo, M.; Kimura, K.; Watano, Y.; Ueda, K.; Shimizu, T. Extensive mitochondrial introgression from Pinus pumila to P. parviflora var. pentaphylla (Pinaceae). J. Plant Res. 1999, 112, 97–105. [Google Scholar] [CrossRef]
- Ma, X.-F.; Szmidt, A.E.; Wang, X.-R. Genetic structure and evolutionary history of a diploid hybrid pine pinus densata inferred from the nucleotide variation at seven gene loci. Mol. Biol. Evol. 2006, 23, 807–816. [Google Scholar] [CrossRef]
- Staffa, M. Słownik Geografii Turystycznej Sudetów: Góry Stołowe [Lexicon of the Turistic Geography of the Sudetes. 13. Stołowe Mountains]; PTTK Kraj: Kraków, Poland, 1992. [Google Scholar]
- Latocha, A.; Migoń, P. Geneza i przemiany krajobrazu kulturowego Gór Stołowych [Genezis and transformations of the cultural lanscape of the Stołowe Mountains]. In Góry Stołowe—Przyroda i Ludzie [The Stołowe Mountains—Nature and People]; Kabała, C., Ed.; Park Narodowy Gór Stołowych: Kudowa Zdrój, Poland, 2018; pp. 47–61. [Google Scholar]
- Latałowa, M.; Tobolski, K.; Nalepka, D. Pinus L. subgenus Pinus (subgen. Diploxylon (Koehne) Pilger). In Late Glacial and Holocene History of Vegetation in Poland Based on Isopollen Maps; Ralska-Jasiewiczowa, M., Ed.; W. Szafer Institute of Botany: Kraków; Poland, 2004; pp. 165–177. [Google Scholar]
- Treml, V.; Jankovská, V.; Petr, L. Holocene dynamics of the alpine timberline in the High Sudetes. Biologia 2008, 63, 73–80. [Google Scholar] [CrossRef]
- Malkiewicz, M.; Waroszewski, J.; Bojko, O.; Egli, M.; Kabala, C. Holocene vegetation history and soil development reflected in the lake sediments of the Karkonosze Mountains (Poland). Holocene 2016, 26, 890–905. [Google Scholar] [CrossRef]
- Marek, S. Rozwój Wielkiego Torfowiska Batorowskiego w świetle badań biostratygraficznych [Development of Wielkie Torfowisko Barowskie mire in the light of biostratygraphic investigations]. Szczeliniec 1988, 2, 49–88. [Google Scholar]
- Madeyska, E. The history of the Zieleniec Mire and the surrounding areas based on the palynological research. Monogr. Bot. 2005, 94, 145–157. [Google Scholar]
- Baranowska-Kącka, A. Historia roślinności Gór Stołowych w świetle badań palinologicznych Sudetów [The history of vegetation of the Stołowe Mts in the light of palynological studies]. In Przyroda Parku Narodowego Gór Stołowych [Nature of National Park of the Stołowe Mts]; Witkowski, A., Pokryszko, B.M., Ciężkowski, W., Eds.; National Park of the Stołowe Mountains: Kudowa Zdrój, Poland, 2013; pp. 130–136. [Google Scholar]
- Boratyński, A. Pinus uliginosa Neumann in the Błedne Skały reserve in Stolowe Mountains. Dendrobiology 1978, 23, 261–267. [Google Scholar]
- Glina, B.; Malkiewicz, M.; Mendyk, Ł.; Bogacz, A.; Woźniczka, P. Human-affected disturbances in vegetation cover and peatland development in the late Holocene recorded in shallow mountain peatlands (Central Sudetes, SW Poland). Boreas 2017, 46, 294–307. [Google Scholar] [CrossRef]
- Sarvas, R. Investigations on the flowering and seed crop of Pinus silvestris. Commun. Inst. Fenn. 1962, 53, 198. [Google Scholar]
- Sarvas, R. Investigations on the annual cycle of development of forest trees. Active period. Commun. Inst. Fenn. 1972, 76, 110. [Google Scholar]
- Poska, A.; Pidek, I.A. Pollen dispersal and deposition characteristics of Abies alba, Fagus sylvatica and Pinus sylvestris, Roztocze region (SE Poland). Veg. Hist. Archaeobot. 2010, 19, 91–101. [Google Scholar] [CrossRef]
- Robledo-Arnuncio, J.J.; Gil, L. Patterns of pollen dispersal in a small population of Pinus sylvestris L. revealed by total-exclusion paternity analysis. Heredity 2005, 94, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Burczyk, J.; Chalupka, W. Flowering and cone production variability and its effect on parental balance in a Scots pine clonal seed orchard. Ann. Sci. 1997, 54, 129–144. [Google Scholar] [CrossRef]
- Marcysiak, K. Interpopulational variability of Pinus uncinata Ramond ex DC. in Lam. & DC (Pinaceae) cone characters. Dendrobiology 2004, 51, 43–51. [Google Scholar]
- Marcysiak, K.; Boratyńska, K.; Mazur, M. Variability of Pinus uliginosa cones from the peat-bog in Węgliniec. Dendrobiology 2003, 49, 43–47. [Google Scholar]
- Boratyńska, K.; Marcysiak, K.; Boratyński, A. Pinus mugo (Pinaceae) in the Abruzzi Mountains: High morphological variation in isolated populations. Bot. J. Linn. Soc. 2005, 147, 309–316. [Google Scholar] [CrossRef]
- Marcysiak, K.; Boratyński, A. Contribution to the taxonomy of Pinus uncinata (Pinaceae) based on cone characters. Plant Syst. Evol. 2007, 264, 57–73. [Google Scholar] [CrossRef]
- Sobierajska, K.; Boratynska, K. Variability of needle characters of Pinus mugo Turra populations in the Karkonosze Mountains in Poland. Dendrobiology 2008, 59, 41–49. [Google Scholar]
- Boratynska, K.; Lewandowska, D. Differences among three populations of Pinus uliginosa and their relation to P. sylvestris as expressed by the needle characters. Dendrobiology 2009, 61, 37–46. [Google Scholar]
- Sobierajska, K.; Boratyńska, K.; Marcysiak, K. Variation of cone characters in Pinus mugo (Pinaceae) populations in the Giant Mountains (Karkonosze, Sudetes). Dendrobiology 2010, 63, 33–41. [Google Scholar]
- Jasińska, A.; Boratyńska, K.; Dering, M.; Sobierajska, K.; Ok, T.; Romo Diez, A.M.; Boratyński, A. Distance between south-European and south-west Asiatic refugial areas involved morphological differentiation: Pinus sylvestris case study. Plant Syst. Evol. 2014, 300, 1487–1502. [Google Scholar] [CrossRef]
- Dzialuk, A.; Boratyńska, K.; Romo, A.; Boratyński, A. Taxonomic and geographic variation of the Pinus mugo complex on chloroplast microsatellite markers. Syst. Biodivers. 2016, 15, 464–479. [Google Scholar] [CrossRef]
- Wachowiak, W.; Boratynska, K.; Cavers, S. Geographical patterns of nucleotide diversity and population differentiation in three closely related European pine species in the Pinus mugo complex. Bot. J. Linn. Soc. 2013, 172, 225–238. [Google Scholar] [CrossRef][Green Version]
- Łabiszak, B.; Zaborowska, J.; Wachowiak, W. Patterns of mtDNA variation reveal complex evolutionary history of relict and endangered peat bog pine (Pinus uliginosa). AoB Plants 2019, 11. [Google Scholar] [CrossRef]
- Boratyńska, K.; Dzialuk, A.; Lewandowski, A.; Marcysiak, K.; Jasińska, A.; Sobierajska, K.; Tomaszewski, D.; Burczyk, J.; Boratyński, A. Geographic distribution of quantitative traits variation and genetic variability in natural populations of Pinus mugo in Central Europe. Dendrobiology 2014, 72, 65–84. [Google Scholar] [CrossRef]
- Donnelly, K.; Cottrell, J.; Ennos, R.A.; Vendramin, G.G.; A’Hara, S.; King, S.; Perry, A.; Wachowiak, W.; Cavers, S. Reconstructing the plant mitochondrial genome for marker discovery: A case study using Pinus. Mol. Ecol. Resour. 2017, 17, 943–954. [Google Scholar] [CrossRef]
- Soranzo, N.; Alia, R.; Provan, J.; Powell, W. Patterns of variation at a mitochondrial sequence-tagged-site locus provides new insights into the postglacial history of European Pinus sylvestris populations. Mol. Ecol. 2000, 9. [Google Scholar] [CrossRef] [PubMed]
- Zaborowska, J.; Łabiszak, B.; Wachowiak, W. Population history of European mountain pines Pinus mugo and Pinus uncinata revealed by mitochondrial DNA markers. J. Syst. Evol. 2019. [Google Scholar] [CrossRef]
- Żukowska, W.B.; Boratyńska, K.; Wachowiak, W. Comparison of range-wide chloroplast microsatellite and needle trait variation patterns in Pinus mugo Turra (dwarf mountain pine). iForest 2017, 10, e1–e9. [Google Scholar] [CrossRef]
- Wachowiak, W.; Lesniewicz, K.; Odrzykoski, I.; Augustyniak, H.; Prus-Głowacki, W. Species specific cpDNA markers useful for studies on the hybridisation between Pinus mugo and P. sylvestris. Acta Soc. Bot. Pol. 2000, 69, 273–276. [Google Scholar] [CrossRef]
- Żukowska, W.B.; Wachowiak, W. Nuclear microsatellite markers reveal the low genetic structure of Pinus mugo Turra (dwarf mountain pine) populations in Europe. Plant Syst. Evol. 2017, 303, 641–651. [Google Scholar] [CrossRef]
- Bandelt, H.-J.; Forster, P.; Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D. Popart: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Excoffier, L.; Lischer, H.E.L. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 2010, 10, 564–567. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol. Ecol. Notes 2007, 7, 574–578. [Google Scholar] [CrossRef] [PubMed]
- Hubisz, M.J.; Falush, D.; Stephens, M.; Pritchard, J.K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 2009, 9, 1322–1332. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Earl, D.A.; Vonholdt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Boratyńska, K.; Tomaszewski, D.; Montserrat Martí, J.; Marek, S.; Boratyński, A. Taxonomic position of Pinus ceciliae (Pinaceae) endemic for Balearic Islands as revealed on needle characteristics. Dendrobiology 2019, 82, 8–16. [Google Scholar] [CrossRef]
- Staszkiewicz, J. Morphological variation of leaves, cones and seeds. In Scots Pine Biology; Białobok, S., Boratyński, A., Bugała, W., Eds.; Instytut Dendrologii Polskiej Akademii Nauk: Poznań—Kórnik, Poland, 1993; pp. 33–44. [Google Scholar]
- Zar, J.H. Biostatistical Analysis, 5th ed.; Prentice-Hall/Pearson: Upper Saddle River, NJ, USA, 2007. [Google Scholar]
- Sokal, R.R.; Rohlf, F.J. Biometry, 4th ed.; W.H. FREEMAN & CO LTD: New York, NY, USA, 2012. [Google Scholar]
- Watala, C. Biostatystyka: Wykorzystanie Metod Statystycznych w Pracy Badawczej w Naukach Biomedycznych [Biostatistics: Using Statistic Methods in Biomedical Scientific Investigations]; Alfamedica Press: Bielsko Biała, Poland, 2012. [Google Scholar]
- Morrison, D.F. Multivariate Statistical Methods; Thomson/Brooks/Cole: Belmont, CA, USA, 2005. [Google Scholar]
- Bobowicz, M.; Danielewicz, W. Isoenzymatic variability in progeny of Pinus mugo Turra x Pinus sylvestris L. hybrids from Bór na Czerwonem, in experimental culture. Acta Soc. Bot. Pol. 2000, 69, 137–144. [Google Scholar] [CrossRef]
- Kremer, A.; Dupouey, J.-L.; Deans, J.; Cottrell, J.; Csaikl, U.; Finkeldey, R.; Espinel, S.; Jensen, J.; Kleinschmit, J.; Dam, B.C.; et al. Leaf morphological differentiation between Quercus robur and Quercus petraea is stable across western European mixed oak stands. Ann. For. Sci. 2002, 59, 777–787. [Google Scholar] [CrossRef]
- Petit, R.J.; Bodénès, C.; Ducousso, A.; Roussel, G.; Kremer, A. Hybridization as a mechanism of invasion in oaks. New Phytol. 2004, 161, 151–164. [Google Scholar] [CrossRef]
- Boratyński, A.; Marcysiak, K.; Jasińska, A.K.; Iszkuło, G. Interrelations among con-generic and co-occurring tree species: Asymmetric hybridization and the high success of Quercus petraea (Matt.) Liebl. regeneration in mixed Q. petraea Q. robur L. stands. Pol. J. Ecol. 2010, 58, 273–283. [Google Scholar]
- Petit, R.J. Biological invasions at the gene level. Divers. Distrib. 2004, 10, 159–165. [Google Scholar] [CrossRef]
- Petit, R.; Bialozyt, R.; Garnier-Géré, P.; Hampe, A. Ecology and genetics of tree invasions: From recent introductions to Quaternary migrations. Ecol. Manag. 2004, 197, 117–137. [Google Scholar] [CrossRef]
- Martinsen, G.D.; Whitham, T.G.; Turek, R.J.; Keim, P. Hybrid populations selectively filter gene introgression between species. Evolution 2001, 55, 1325–1335. [Google Scholar] [CrossRef]
- Fay, M.F.; Cowan, R.S.; Simpson, D.A. Hybridisation between Schoenoplectus tabernaemontani and S. triqueter (Cyperaceae) in the British Isles. Watsonia 2003, 24, 433–442. [Google Scholar]
- Álvarez, I.; Wendel, J.F. Cryptic Interspecific Introgression and Genetic Differentiation within Gossypium Aridum (Malvaceae) and Its Relatives. Evolution 2006, 60, 505–517. [Google Scholar] [CrossRef][Green Version]
- Wachowiak, W.; Trivedi, U.; Perry, A.; Cavers, S. Comparative transcriptomics of a complex of four European pine species. BMC Genom. 2015, 16, 234. [Google Scholar] [CrossRef]
- Dzialuk, A.; Muchewicz, E.; Boratyński, A.; Montserrat, J.M.; Boratyńska, K.; Burczyk, J. Genetic variation of Pinus uncinata (Pinaceae) in the Pyrenees determined with cpSSR markers. Plant Syst. Evol. 2009, 277, 197–205. [Google Scholar] [CrossRef]
- Dzialuk, A.; Boratyński, A.; Boratyńska, K.; Burczyk, J. Geographic patterns of genetic diversity of Pinus mugo (Pinaceae) in Central European mountains. Dendrobiology 2012, 68, 31–41. [Google Scholar]
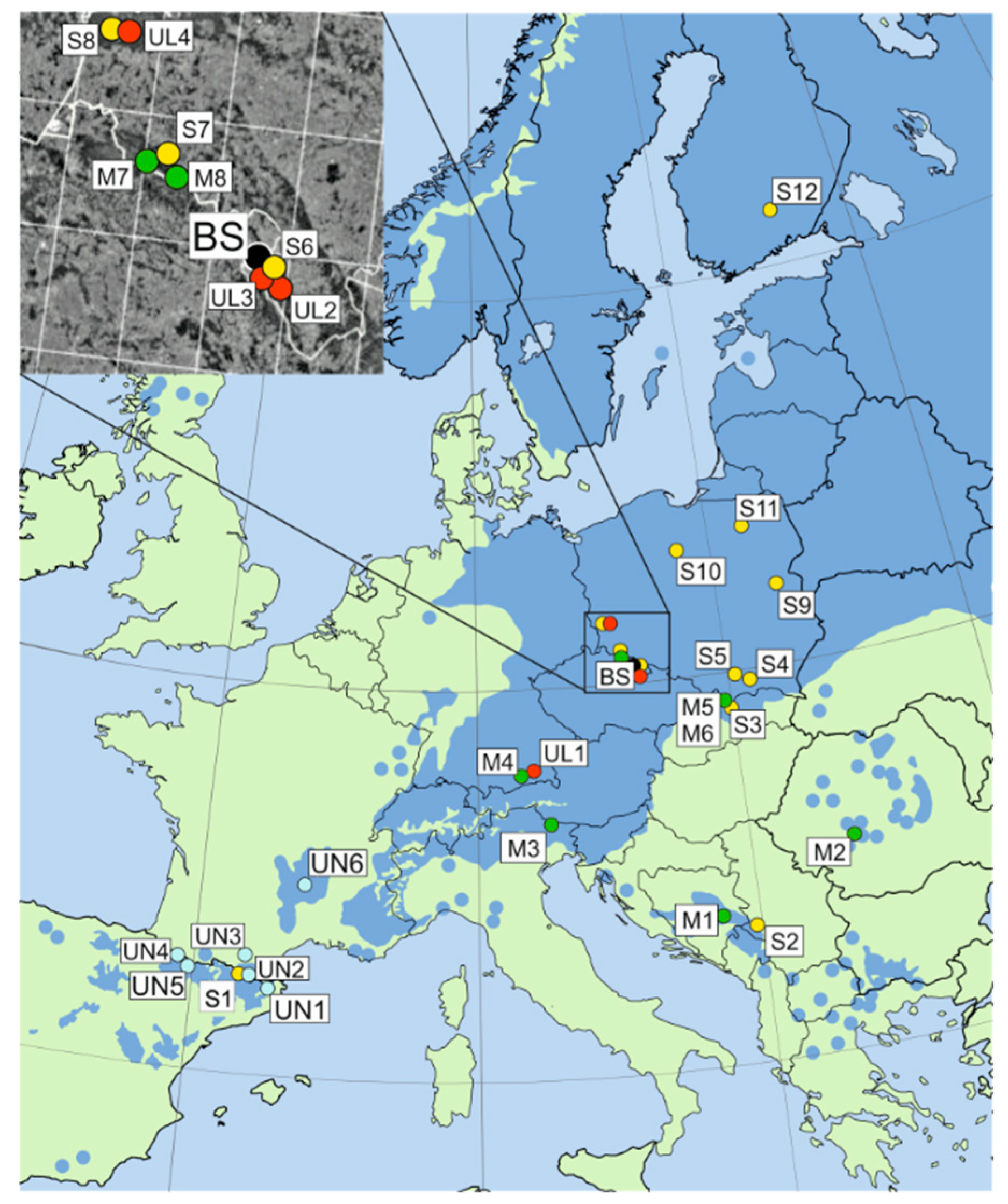
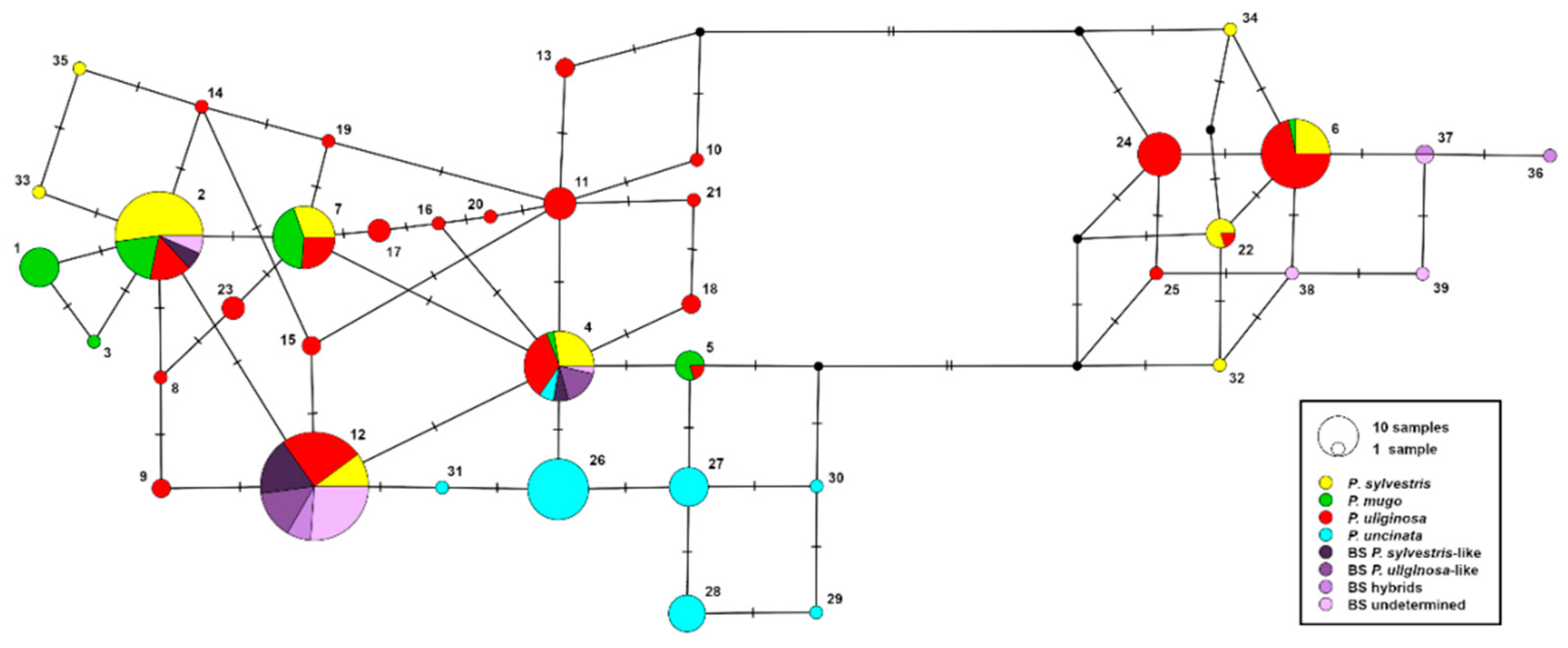

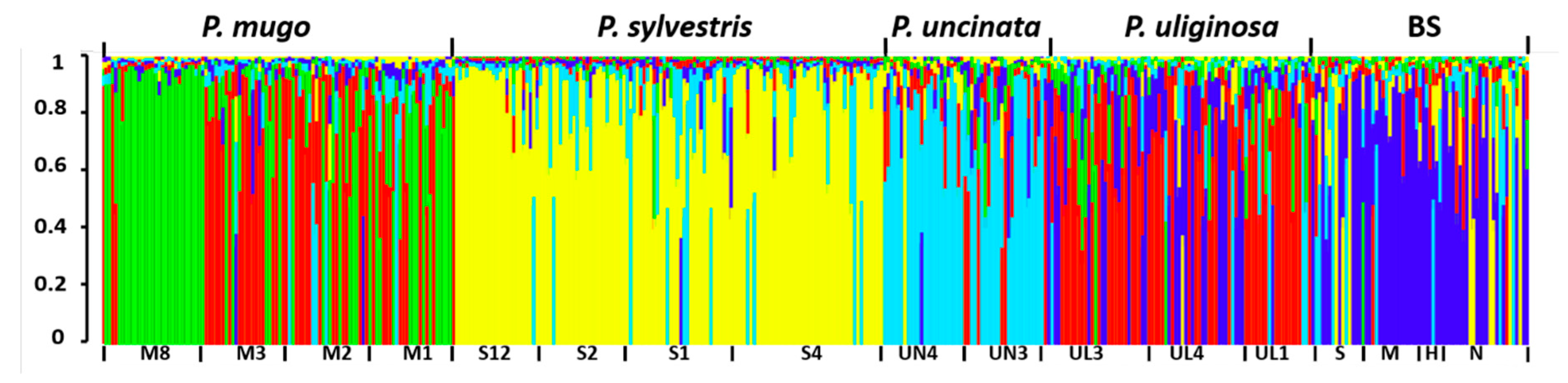
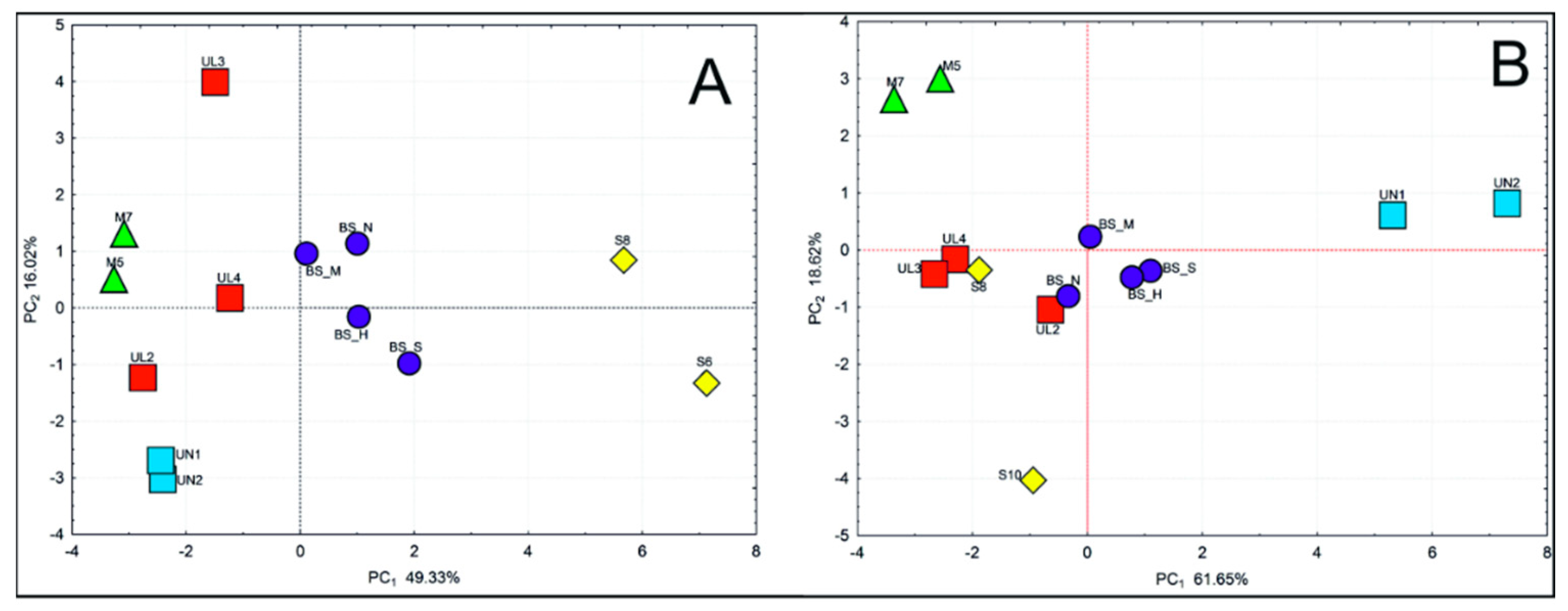
| Species | Code | Location | Latitude N | Longitude E | Altitude (m) | Nb (needles/cones) | Ng |
|---|---|---|---|---|---|---|---|
| Mixed BS | BS | Błedne Skały, Stołowe Mountains (Poland) | 50.48 | 16.29 | 850 | 64 (320/315) | 64 |
| Pinus sylvestris S | S1 | St. Miguel d’Engolasters (Andorra) | 42.51 | 1.56 | 1550 | - | 32 |
| S2 | Divčibare Mts. (Serbia) | 44.10 | 19.99 | 957 | - | 26 | |
| S3 | Tatra National Park (Poland) | 49.27 | 19.83 | 1000 | - | 12 | |
| S4 | Beskid (Poland) | 49.40 | 20.82 | 600 | - | 45 | |
| S5 | Pieniny National Park (Poland) | 49.42 | 20.36 | 800 | - | 21 | |
| S6 | Stołowe Mountains (Poland) | 50.43 | 16.23 | 900 | 30 (300/0) | ||
| S7 | Sudetes, Karkonosze (Poland) | 50.83 | 15.64 | 600 | - | 20 | |
| S8 | Silesian Lowland, Węgliniec (Poland) | 51.28 | 15.23 | 170 | 34 (340/46) | ||
| S9 | Liski Nature Reserve (Poland) | 51.93 | 22.82 | 150 | - | 11 | |
| S10 | Bory Tucholskie Forest, (Poland) | 53.32 | 17.94 | 90 | 50 (0/50) | ||
| S11 | Sosny Taborskie Reserve (Poland) | 53.78 | 20.02 | 115 | - | 13 | |
| S12 | Joutsa (Finland) | 61.76 | 26.10 | 125 | - | 25 | |
| Pinus mugo M | M1 | Dinaric Alps (Bosnia and Herzegovina) | 43.72 | 18.24 | 2020 | - | 25 |
| M2 | Carpathians, Fagaraş (Romania) | 45.61 | 24.60 | 1900 | - | 25 | |
| M3 | Alps, Carnic Alps, Pramollo (Italy) | 46.56 | 13.28 | 1600 | - | 25 | |
| M4 | Ammergau Alps (Germany) | 47.53 | 10.92 | 1870 | - | 25 | |
| M5 | Tatra National Park (Poland) | 49.12 | 20.05 | 1700 | 30 (300/330) | 12 | |
| M6 | Tatra National Park, Grześ (Poland) | 49.23 | 19.77 | 1620 | - | 25 | |
| M7 | Sudetes (Poland) | 50.78 | 15.58 | 1350 | 33 (330/310) | 30 | |
| M8 | Sudetes, Śnieżka (Poland) | 50.74 | 15.74 | 1435 | - | 30 | |
| Pinus uliginosa UL | UL1 | Alps, Mittelwalde (Germany) | 47.48 | 11.27 | 860 | - | 20 |
| UL2 | Zieleniec Nature Reserve (Poland) | 49.13 | 20.03 | 700 | 30 (300/50) | ||
| UL3 | Large Batorów Peatland (Poland) | 50.45 | 16.38 | 750 | 50 (500/50) | 31 | |
| UL4 | Węgliniec Nature Reserve (Poland) | 51.29 | 15.23 | 190 | 52 (520/230) | 30 | |
| Pinus uncinata UN | UN1 | Vall de Nuria (Spain) | 42.22 | 02.10 | 2200 | 28 (280/50) | |
| UN2 | Vall de Ransol (Andorra) | 42.38 | 01.37 | 2000 | 32 (320/49) | ||
| UN3 | Coll de Jau (France) | 42.69 | 2.25 | 1500 | - | 24 | |
| UN4 | Belagua (Spain) | 42.97 | -0.77 | 1760 | - | 24 | |
| UN5 | La Trapa near Jaca (Spain) | 42.69 | -0.54 | 1720 | - | 22 | |
| UN6 | Col de la Croix-Morand (France) | 45.60 | 2.85 | 1400 | - | 14 |
| Sample Name | Measures of Genetic Variation | ||||||
|---|---|---|---|---|---|---|---|
| N | Ah | Ae | AP | AR | Hd | D2sh | |
| Błędne Skały | 64 | 19 | 13 | 15 | 7.56 | 0.881 | 32.1 |
| P. mugo | 105 | 100 | 99 | 46 | 95.87 | 0.999 | 13.5 |
| P. sylvestris | 128 | 118 | 118 | 46 | 109.28 | 0.999 | 5.6 |
| P. uncinata | 48 | 33 | 32 | 32 | 20.57 | 0.972 | 7.2 |
| P. uliginosa | 81 | 40 | 32 | 28 | 22.70 | 0.968 | 8.2 |
| Sample Name | Measures of Genetic Variation | |||||||
|---|---|---|---|---|---|---|---|---|
| N | S | Sp | d | Hn | Hp | Hd (SD) | uH | |
| Błędne Skały | 64 | 9 | 0 | 1.83 | 7 | 4 | 0.487 (0.071) | 0.495 |
| P. mugo | 34 | 10 | 2 | 4.03 | 7 | 2 | 0.786 (0.034) | 0.810 |
| P. sylvestris | 61 | 9 | 0 | 3.31 | 10 | 4 | 0.796 (0.038) | 0.809 |
| P. uncinata | 44 | 5 | 1 | 1.06 | 7 | 6 | 0.687 (0.055) | 0.703 |
| P. uliginosa | 101 | 11 | 1 | 1.54 | 23 | 16 | 0.904 (0.014) | 0.913 |
| Sample Name | Measures of Genetic Variation | |||||
|---|---|---|---|---|---|---|
| N | Na | Ne | Ho | He | F | |
| Błędne Skały | 63 | 4.78 | 2.34 | 0.425 | 0.467 | 0.046 |
| P. mugo | 105 | 4.78 | 2.06 | 0.386 | 0.460 | 0.142 |
| P. sylvestris | 128 | 7.33 | 2.37 | 0.416 | 0.451 | 0.076 |
| P. uncinata | 48 | 3.89 | 2.17 | 0.372 | 0.410 | 0.051 |
| P. uliginosa | 81 | 4.78 | 2.09 | 0.393 | 0.438 | 0.101 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sobierajska, K.; Wachowiak, W.; Zaborowska, J.; Łabiszak, B.; Wójkiewicz, B.; Sękiewicz, M.; Jasińska, A.K.; Sękiewicz, K.; Boratyńska, K.; Marcysiak, K.; et al. Genetic Consequences of Hybridization in Relict Isolated Trees Pinus sylvestris and the Pinus mugo Complex. Forests 2020, 11, 1086. https://doi.org/10.3390/f11101086
Sobierajska K, Wachowiak W, Zaborowska J, Łabiszak B, Wójkiewicz B, Sękiewicz M, Jasińska AK, Sękiewicz K, Boratyńska K, Marcysiak K, et al. Genetic Consequences of Hybridization in Relict Isolated Trees Pinus sylvestris and the Pinus mugo Complex. Forests. 2020; 11(10):1086. https://doi.org/10.3390/f11101086
Chicago/Turabian StyleSobierajska, Karolina, Witold Wachowiak, Julia Zaborowska, Bartosz Łabiszak, Błażej Wójkiewicz, Maciej Sękiewicz, Anna K. Jasińska, Katarzyna Sękiewicz, Krystyna Boratyńska, Katarzyna Marcysiak, and et al. 2020. "Genetic Consequences of Hybridization in Relict Isolated Trees Pinus sylvestris and the Pinus mugo Complex" Forests 11, no. 10: 1086. https://doi.org/10.3390/f11101086
APA StyleSobierajska, K., Wachowiak, W., Zaborowska, J., Łabiszak, B., Wójkiewicz, B., Sękiewicz, M., Jasińska, A. K., Sękiewicz, K., Boratyńska, K., Marcysiak, K., & Boratyński, A. (2020). Genetic Consequences of Hybridization in Relict Isolated Trees Pinus sylvestris and the Pinus mugo Complex. Forests, 11(10), 1086. https://doi.org/10.3390/f11101086






