Prevalence of Multiple Antibiotics Resistant (MAR) Pseudomonas Species in the Final Effluents of Three Municipal Wastewater Treatment Facilities in South Africa
Abstract
:1. Introduction
2. Materials and Method
2.1. Study Site and Sampling
2.2. Physicochemical Analyses
2.3. Isolation, Enumeration and Identification of Pseudomonas Species
2.4. Antibiogram Assay
2.4.1. Antimicrobial Agents
2.4.2. Antibiotic Susceptibility Test
2.5. Statistical Analysis
3. Results
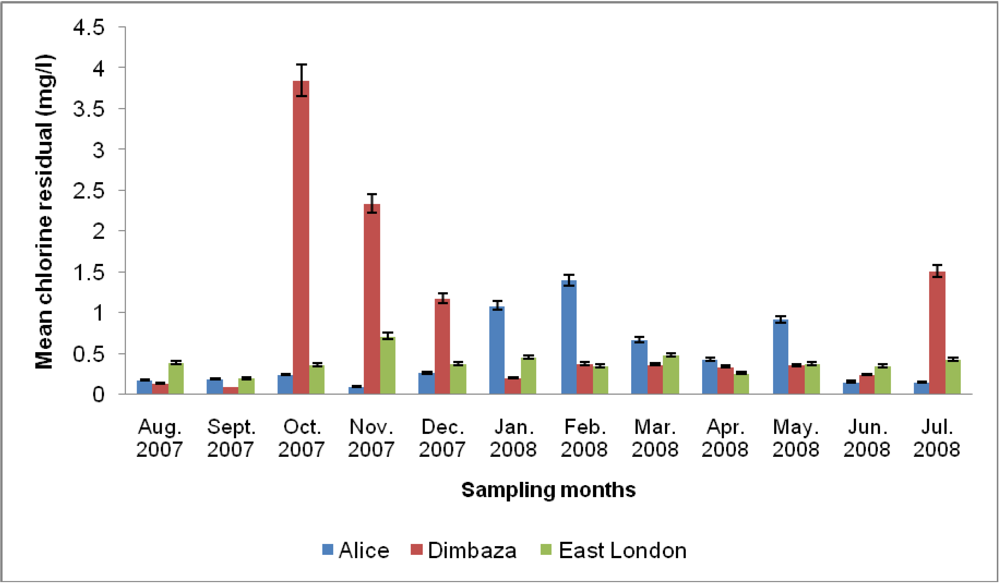
3.1. Physicochemical Analyses
| Seasons | Sampling Site | pH | Temperature (°C) | Turbidity (NTU) | a TDS (mg/L) | b DO (mg/L) | c COD (mg/L) | NO3− (mg/L) | NO2− (mg/L) | PO43− (mg/L) |
|---|---|---|---|---|---|---|---|---|---|---|
| Spring | d AL | 6.8 ± 0.32 | 20 ± 2.22 | 3.6 ± 2.10 | 162 ± 13 | 4.9 ± 1.52 | 57 ± 21 | 11.3 ± 1.0 | 0.29 ± 0.14 | 3.50 ± 0.88 |
| e DMB | 7.3 ± 0.44 | 19 ± 0.74 | 17.6 ± 15.7 | 148 ± 9 | 5.1 ± 0.06 | 25 ± 8 | 1.4 ± 0.78 | 0.1 ± 0.015 | 0.91 ± 0.15 | |
| f EL | 7.1 ± 0.20 | 20 ± 1.17 | 12.5 ± 2.53 | 372 ± 26 | 5.4 ± 0.13 | 68 ± 0 | 3.8 ± 3.21 | 2.4 ± 3.82 | 0.38 ± 0.08 | |
| Summer | d AL | 7.3 ± 1.9 | 24 ± 3.36 | 12.2 ± 9 | 132 ± 10 | 5.7 ± 4.07 | 323 ± 457 | 8.1 ± 2.15 | 0.21 ± 0.12 | 1.88 ± 0.53 |
| e DMB | 7.0 ± 0.12 | 21 ± 1.28 | 8.2 ± 1.81 | 128 ± 14 | 4.9 ± 0.30 | 330 ± 523 | 5.9 ± 0.61 | 0.37 ± 0.09 | 2.83 ± 0.85 | |
| f EL | 7.2 ± 0.19 | 24 ± 0.95 | 4.10 ± 1.75 | 367 ± 80 | 4.2 ± 0.19 | 462 ± 599 | 6.5 ± 0.28 | 0.21 ± 0.15 | 0.32 ± 0.13 | |
| Autumn | d AL | 6.4 ± 0.28 | 24 ± 1.66 | 7.39 ± 3 | 140 ± 8 | 4.9 ± 0.65 | 78 ± 59 | 12.7 ± 5.45 | 0.13 ± 0.06 | 1.31 ± 0.91 |
| e DMB | 7.1 ± 0.34 | 21 ± 2.27 | 8.7 ± 3.21 | 115 ± 0.51 | 4.8 ± 0.34 | 37 ± 11 | 5.9 ± 1.17 | 0.33 ± 0.12 | 2.97 ± 1.6 | |
| f EL | 7.5 ± 0.17 | 25 ± 1.77 | 3.8 ± 1.10 | 470 ± 232 | 3.9 ± 0.98 | 48 ± 29 | 3.4 ± 3.0 | 0.23 ± 0.05 | 0.37 ± 0.31 | |
| Winter | d AL | 6.0 ± 0.55 | 15 ± 2.02 | 3.51 ± 1.4 | 142 ± 9 | 4.6 ± 1.68 | 50 ± 31 | 9.3 ± 6.51 | 0.22 ± 0.17 | 1.39 ± 2.15 |
| e DMB | 6.9 ± 0.21 | 17 ± 2.21 | 10.5 ± 2.49 | 117 ± 6 | 5.3 ± 0.75 | 78 ± 57 | 2.2 ± 2.05 | 0.34 ± 0.28 | 1.25 ± 2.08 | |
| f EL | 6.8 ± 0.10 | 20 ± 2.03 | 5.6 ± 0.42 | 387 ± 17 | 4.3 ± 0.50 | 53 ± 31 | 5.6 ± 1.85 | 0.73 ± 0.40 | 0.29 ± 0.13 | |
| RecommendedTarget Limits | g 6–9 | g ≤25 | g 0–1;h ≤5 | g 0–450 | i ≥5 | j 30 | g 6; j 1–5 | g 0–6; k <0.5 | k 0.005 | |
3.2. Total Pseudomonas Counts (TPC)
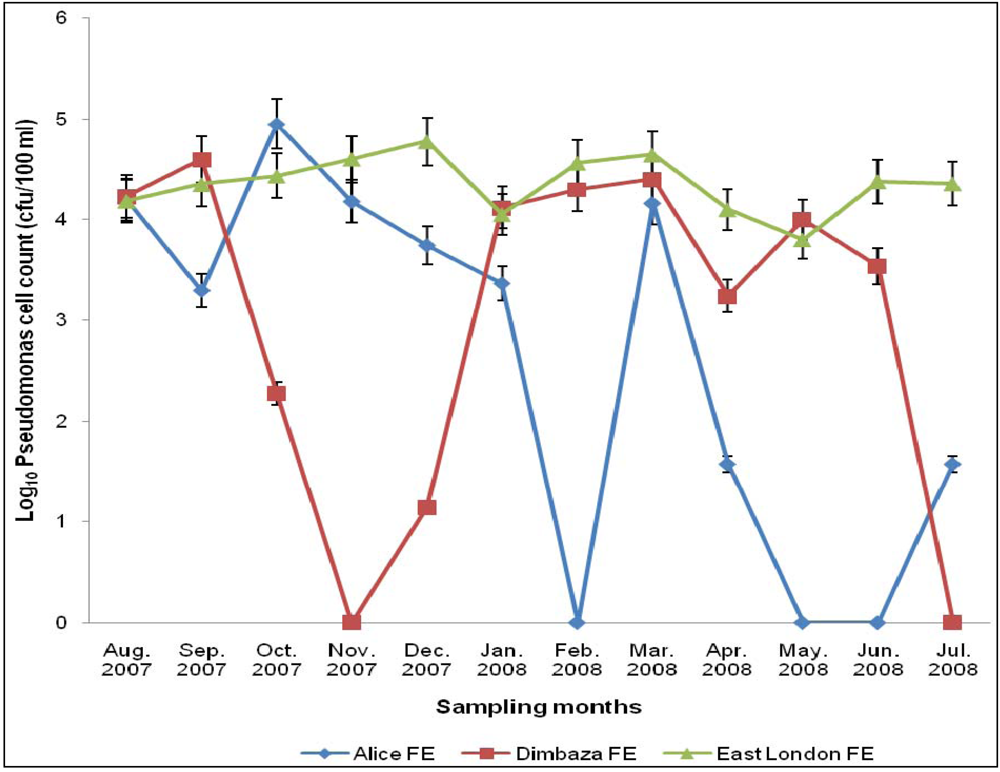
3.3. Pseudomonas Isolates and Antibiogram
| Antibiotics Class | Antibiotics | Number of isolates (%) | ||
|---|---|---|---|---|
| Sensitivity | Intermediate | Resistant | ||
| Penicillins | Penicillin G | 0(0) | 0(0) | 10(100) |
| Ampicillin | 0(0) | 1(10) | 9(90) | |
| Oxacillin | 0(0) | 0(0) | 10(100) | |
| Cephems | Cefotaxime | 1(10) | 2(20) | 7(70) |
| Cefepime | 3(30) | 0(0) | 7(70) | |
| Cephalothin | 0(0) | 3(30) | 7(70) | |
| Folate Pathway Inhibitors | Sulphamethoxazole | 0(0) | 1(10) | 9(90) |
| Ansamycins | Rifampin | 0(0) | 1(10) | 9(90) |
| Quinolones | Nalidixic acid | 1(10) | 7(70) | 2(20) |
| β-Lactam/β-Lactamase Inhibitor Combinations | a Ampicillin/sulbactam | 1(10) | 5(50) | 3(30) |
| Phenicols | Chloramphenicol | 4(40) | 5(50) | 1(10) |
| Tetracyclines | Tetracycline | 5(50) | 4(40) | 1(10) |
| Minocycline | 2(20) | 6(60) | 2(20) | |
| Aminoglycosides | Gentamicin | 10(100) | 0(0) | 0(0) |
| Fluoroquinolones | Ofloxacin | 10(100) | 0(0) | 0(0) |
| Macrolides | Erythromycin | 9(90) | 1(10) | 0(0) |
| Glycopeptides | Vancomycin | 1(10) | 6(60) | 3(30) |
| Nitrofurantoins | Nitrofurantoin | 8(80) | 1(10) | 1(10) |
| Lincosamides | Clindamycin | 9(90) | 0(0) | 1(10) |
| Isolates Code | Organism Identity | Antibiotics | MAR Index |
|---|---|---|---|
| Control | P. aeruginosa (ATCC 27853) | AMP, CEP, CHL, CLI, MINO, NAL, NIT, OXA, PEN, RIF,SMX, TET, VAN, SAM | 0.74 |
| AL 1 | P. aeruginosa | CEP, CHL, CLI, NIT, OXA, PEN, SMX, VAN | 0.42 |
| AL 2 | P. fluorescens | AMP, CTX, CPM, OXA, PEN, RIF, SMX, SAM | 0.42 |
| DB 1 | P. aeruginosa | AMP, CTX, CEP, CPM, NAL, OXA, PEN, RIF, SMX, TET, VAN | 0.58 |
| DB 2 | P. fluorescens | AMP, CTX, CEP, CPM, OXA, PEN, RIF, SMX | 0.42 |
| EL 1 | P. fluorescens | AMP, CTX, OXA, PEN, RIF | 0.26 |
| EL 2 | P. aeruginosa | AMP, CEP, CPM, NAL, OXA, PEN, SMX, VAN, SAM | 0.47 |
| EL 3 | P. fluorescens | AMP, CTX, CEP, CPM, OXA, PEN, RIF, SMX | 0.42 |
| EL 4 | P. aeruginosa | AMP, MINO, NAL, OXA, PEN, RIF, SMX | 0.37 |
| EL 5 | P. aeruginosa | AMP, CTX, CEP, CPM, OXA, PEN, RIF, SMX, SAM | 0.47 |
| EL 6 | P.. fluorescens | AMP, CTX, CEP, CPM, OXA, PEN, RIF, SMX | 0.42 |
4. Discussion
5. Conclusions
Conflict of Interest Statement
Acknowledgements
References
- Nwachcuku, N.; Gerba, C.P. Emerging waterborne pathogens: Can we kill them all? Curr. Opin. Biotechnol. 2004, 15, 175–180. [Google Scholar] [CrossRef]
- Sharma, S.; Sachdeva, P.; Virdi, J.S. Emerging water-borne pathogens. Appl. Microbiol. Biotechnol. 2003, 61, 424–428. [Google Scholar]
- Bert, F.; Maubec, E.; Bruneau, B.; Berry, P.; Lambert-Zechovsky, N. Multi-resistant Pseudomonas aeruginosa outbreak associated with contaminated tap water in a neurosurgery intensive care unit. J. Hosp. Infect. 1998, 39, 53–62. [Google Scholar] [CrossRef]
- Widmer, F.; Seidler, R.J.; Gillevet, P.M.; Watrud, L.S.; di Giovanni, D. A highly selective PCR protocol for detecting 16S rRNA genes of the genus Pseudomonas (Sensustricto) in environmental samples. App. Environ. Microbiol. 1998, 64, 2545–2553. [Google Scholar]
- Goldberg, J.B. Pseudomonas: Global bacteria. Trends Microbiol. 2000, 8, 55–57. [Google Scholar] [CrossRef]
- Pirnay, J.-P.; Matthijs, S.; Colak, H.; Chablain, P.; Bilocq, F.; van Eldere, J.; de Vos, D.; Zizi, M.; Triest, L.; Cornelis, P. Global Pseudomonas aeruginosa biodiversity as reflected in a Belgian river. Environ. Microbiol. 2005, 7, 969–980. [Google Scholar]
- Mena, K.D.; Gerba, C.P. Risk assessment of Pseudomonas aeruginosa in water. Rev. Environ. Contam. Toxicol. 2009, 201, 71–115. [Google Scholar]
- D’Agata, E.M. Rapidly rising prevalence of nosocomial multidrug-resistant Gram-negative bacilli: A 9-year surveillance study. Infect. Control Hosp. Epidemiol. 2004, 25, 842–846. [Google Scholar]
- Alaoui, H.L.; Oufdou, K.; Mezrioui, N. Occurrence and antibiotic resistance of Pseudomonas aeruginosa and faecal indicator bacteria: Risk assessment for groundwater supplies (Marrakesh, Morocco). Hung. Med. J. 2007, 1, 315–330. [Google Scholar] [CrossRef]
- Jung, R.; Fish, D.N.; Obritsch, M.D.; MacLaren, R. Surveillance of multi-drug resistant Pseudomonas aeruginosa in an urban tertiary-care teaching hospital. J. Hosp. Infect. 2004, 57, 105–111. [Google Scholar] [CrossRef]
- Navon-Venezia, S.; Ben-Ami, R.; Carmeli, Y. Update on Pseudomonas aeruginosa and Acinetobacter baumannii infections in the healthcare setting. Curr. Opin. Infect. Dis. 2005, 18, 306–313. [Google Scholar] [CrossRef]
- Szewzyk, U.; Szewzyk, R.; Manz, W.; Schleifer, K.H. Microbiological safety of drinking water. Annu. Rev. Microbiol. 2000, 54, 81–127. [Google Scholar] [CrossRef]
- Boles, B.R.; Thoendel, M.; Singh, P.K. Self-generated diversity produces “insurance effects” in biofilm communities. Proc. Natl. Acad. Sci. USA 2004, 101, 16630–16635. [Google Scholar] [CrossRef]
- Samie, A.; Obi, C.L.; Igumbor, J.O.; Momba, M.N.B. Focus on 14 sewage treatment plants in the Mpumalanga Province, South Africa in order to gauge the efficiency of wastewater treatment. Afr. J. Biotechnol. 2009, 8, 3276–3285. [Google Scholar]
- Xi, C.; Zhang, Y.; Marrs, C.F.; Ye, W.; Simon, C.; Foxman, B.; Nriagu, J. Prevalence of antibiotic resistance in drinking water treatment and distribution systems. Appl. Environ. Microbiol. 2009, 75, 5714–5718. [Google Scholar]
- Huang, J.-J.; Hu, H.-Y.; Tang, F.; Li, Y.; Lu, S.-Q.; Lu, Y. Inactivation and reactivation of antibiotic-resistant bacteria by chlorination in secondary effluents of a municipal wastewater treatment plant. Water Res. 2011, 45, 2775–2781. [Google Scholar]
- Shrivastava, R.; Upreti, R.K.; Jain, S.R.; Prasad, K.N.; Seth, P.K.; Chaturvedi, U.C. Suboptimal chlorine treatment of drinking water leads to selection of multidrug-resistant Pseudomonas aeruginosa. Ecotoxicol. Environ. Saf. 2004, 58, 277–283. [Google Scholar] [CrossRef]
- Momba, M.N.B.; Osode, A.N.; Sibewu, M. The impact of inadequate wastewater treatment on the receiving water bodies case study: Buffalo City and Nkonkonbe Municipalities of the Eastern Cape Province. Water South Afr. 2006, 32, 687–692. [Google Scholar]
- Venter, S.N. Microbial water quality in the 21st century. South Afr. Water Bull. 2001, 27, 16–17. [Google Scholar]
- Mackintosh, G.; Colvin, C. Failure of rural schemes in South Africa to provide potable water. Environ. Geol. 2003, 44, 101–105. [Google Scholar]
- Obi, C.L.; Onabolu, B.; Momba, M.N.B.; Igumbor, J.O.; Ramalivahna, J.; Bossong, P.O.; van Rensburg, E.J.; Lukoto, M.; Green, E.; Mulaudzi, T.B. The interesting cross-paths of HIV/AIDS and water in Southern Africa with special reference to South Africa. Water South Afr. 2006, 32, 323–343. [Google Scholar]
- DWAF, Analytical Methods Manual, TR 151; Department of Water Affairs and Forestry: Pretoria, South Africa, 1992.
- CLSI, Performance Standards for Antimicrobial Susceptibility Testing; Sixteenth Informational Supplement, M100-S16; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2006; Volume 26, No. 1, p. 183.
- Blasco, M.D.; Esteve, C.; Alcaide, E. Multiresistant waterborne pathogens isolated from water reservoirs and cooling systems. J. Appl. Microbiol. 2008, 105, 469–475. [Google Scholar] [CrossRef]
- DWAF, South African Water Quality Guidelines: Domestic Use, 2nd edDepartment of Water Affairs and Forestry: Pretoria, South Africa, 1996; Volume 2.
- WHO, Rolling Revision of the WHO Guidelines for Drinking-Water Quality, Draft for Review and Comments. Nitrates and Nitrites in Drinking-Water; WHO/SDE/WSH/04.08/56; World Health Organization: Geneva, Switzerland, 2004.
- Fatoki, S.O.; Gogwana, P.; Ogunfowokan, A.O. Pollution assessment in the Keiskamma River and in the impoundment downstream. Water South Afr. 2003, 29, 183–187. [Google Scholar]
- SA Government Gazette, Requirements for the Purification of Wastewater or Effluent; Gazette No. 9225, Regulation 991; Minister of Environment Affairs and Fisheries: Pretoria, South Africa, 1984.
- DWAF, South African Water Quality Guidelines: Aquatic Ecosystems, 1st edDepartment of Water Affairs and Forestry: Pretoria, South Africa, 1996; Volume 7.
- Odjadjare, E.E.O.; Okoh, A.I. Prevalence and distribution of Listeria pathogens in the final effluents of a rural wastewater treatment facility in the Eastern Cape Province of South Africa. World J. Microbiol. Biotechnol. 2010, 26, 297–307. [Google Scholar] [CrossRef]
- Odjadjare, E.E.O.; Okoh, A.I. Physicochemical quality of an urban municipal wastewater effluent and its impact on the receiving environment. Environ. Monit. Assess. 2010, 170, 383–394. [Google Scholar] [CrossRef]
- Odjadjare, E.O.; Obi, C.L.; Okoh, A.I. Municipal wastewater effluent as a source of Listerial pathogens in the aquatic milieu of the Eastern Cape Province of South Africa: A concern of public health. Int. J. Environ. Res. Public Health 2010, 7, 2376–2394. [Google Scholar] [CrossRef]
- Obi, C.L.; Igunmbo, J.O.; Momba, M.N.B.; Samie, A. Interplay factors involving chlorine dose, turbidity flow capacity and pH on microbial quality of drinking water in small treatment plants. Water South Afr. 2008, 34, 565–572. [Google Scholar]
- Obi, C.L.; Momba, M.N.B.; Samie, A.; Igumbor, J.O.; Green, E.; Musie, E. Microbiological, physicochemical and management parameters impinging on the efficiency of small water treatment plants in Limpopo and Mpumalanga Provinces of South Africa. Water South Afr. 2007, 33, 229–237. [Google Scholar]
- Environment Canada. The State of Municipal Wastewater Effluent in Canada; K1A 0H3; Minister of Public Works and Government Services Canada: Ottawa, ON, Canada, 2001. Available online: http://www.ec.gc.ca/soer-ree (accessed on 27 January 2009).
- Rowan, N.J. Defining established and emerging microbial risks in the aquatic environment: Current knowledge, implications, and outlooks. Int. J. Microbiol. 2011. [Google Scholar] [CrossRef]
- Havelaar, A.H.; During, M. C-390 as sole selective agent for isolation of pseudomonas aeruginosa from hospital waste water. Can. J. Microbiol. 1986, 32, 513–515. [Google Scholar] [CrossRef]
- LeChevallier, M.W.; Cawthan, C.D.; Lee, R.G. Factors promoting survival of bacteria in chlorinated water supplies. Appl. Environ. Microbiol. 1988, 54, 649–654. [Google Scholar]
- Price, D.; Ahearn, D.G. Incidence and persistence of Pseudomonas aeruginosa in whirlpools. J. Clin. Microbial. 1988, 26, 1650–1654. [Google Scholar]
- Hijnen, W.A.M.; Beerendonk, E.F.; Medema, G.J. Inactivation of UV radiation for viruses, bacteria, and protozoan (oo)cysts in water: A review. Water Res. 2006, 40, 3–22. [Google Scholar] [CrossRef]
- Daily Dispatch. Report Highlights Cholera Risk Profile. 2003. Available online: http://www.dispatch.co.za/2003/01/30/easterncape/BCHOLERA.HTM (accessed on 12 September 2008).
- Alonso, A.; Rojo, F.; Martinez, J.L. Environmental and clinical isolates of Pseudomonas aeruginosa show pathogenic and biodegradative properties irrespective of their origin. Environ. Microbiol. 1999, 1, 421–430. [Google Scholar] [CrossRef]
- Chang, J.-S.; Chou, C.; Chen, S.-Y. Decolorization of azo dyes with immobilized Pseudomonas luteola. Process Biochem. 2001, 36, 757–763. [Google Scholar] [CrossRef]
- Kao, C.M.; Liu, J.K.; Chen, Y.L.; Chai, C.T.; Chen, S.C. Factors affecting the biodegradation of PCP by Pseudomonas mendocina NSYSU. J. Hazard. Mater. 2005, 124, 68–73. [Google Scholar] [CrossRef]
- Odjadjare, E.E.; Ajisebutu, S.O.; Igbinosa, E.O.; Aiyegoro, O.A.; Trejo-Hernandez, M.R.; Okoh, A.I. Escravos light crude oil degrading potentials of axenic and mixed bacteria cultures. J. Gen. Appl. Microbiol. 2008, 54, 277–284. [Google Scholar] [CrossRef]
- Hsueh, P.; Teng, L.-J.; Pan, H.-J.; Chen, Y.-C.; Sun, C.-C.; Ho, S.-W.; Luh, K.-T. Outbreak of Pseudomonas fluorescens bacteremia among oncology patients. J. Clin. Microbiol. 1998, 36, 2914–2917. [Google Scholar]
- Ergin, C.; Mutlu, G. Clinical distribution and antibiotic resistance of Pseudomonas species. Eastern J. Med. 1999, 4, 65–69. [Google Scholar]
- Casalta, J.-P.; Fournier, P.-E.; Habib, G.; Riberi, A.; Raoult, D. Prosthetic valve endocarditis caused by Pseudomonas luteola. BMC Infect. Dis. 2005, 5. [Google Scholar] [CrossRef]
- Nseir, W.; Taha, H.; Abid, A.; Khateeb, J. Pseudomonas mendocina sepsis in a healthy man. Israel Med. Assoc. J. 2011, 13, 375–376. [Google Scholar]
- Asthana, S.; Rusin, P.; Gerba, C.P. Influence of hydrocarbons on the virulence and antibiotic sensitivity associated with Pseudomonas aeruginosa. Int. J. Environ. Health Res. 1997, 7, 277–287. [Google Scholar] [CrossRef]
- Jalal, S.; Wretlind, B. Mechanisms of quinolone resistance in clinical strains of Pseudomonas aeruginosa. Microb. Drug Resist. 1998, 4, 257–261. [Google Scholar] [CrossRef]
- Lateef, A. The microbiology of a pharmaceutical effluent and its public health implications. World J. Microbiol. Biotechnol. 2004, 20, 167–171. [Google Scholar] [CrossRef]
- Iwane, T.; Urase, T.; Yamamoto, K. Possible impact of treated wastewater discharge on incidence of antibiotic resistant bacteria in river water. Water Sci. Technol. 2001, 43, 91–99. [Google Scholar]
- Schwartz, T.; Kohnen, W.; Jansen, B.; Obst, U. Detection of antibiotic-resistant bacteria and their resistance genes in wastewater, surface water, and drinking water biofilms. FEMS Microbiol. Ecol. 2003, 43, 325–335. [Google Scholar] [CrossRef]
- Volkmann, H.; Schwartz, T.; Bischoff, P.; Kirchen, S.; Obst, U. Detection of clinically relevant antibiotic-resistance genes in municipal wastewater using real-time PCR (TaqMan). J. Microbol. Methods 2004, 56, 277–286. [Google Scholar] [CrossRef]
- Nagata, T.; Mukae, H.; Kadota, J.; Hayashi, T.; Fujii, T.; Kuroki, M.; Shirai, R.; Yanagihara, K.; Tomono, K.; Kohno, S. Effect of erythromycin on chronic respiratory infection caused by Pseudomonas aeruginosa with biofilm formation in an experimental murine model. Antimicrob. Agents Chemother. 2004, 48, 2251–2259. [Google Scholar] [CrossRef]
- George, S.E.; Kohan, M.J.; Whitehouse, D.A.; Creason, J.P.; Claxton, L.D. Influence of antibiotics on intestinal tract survival and translocation of environmental Pseudomonas species. Appl. Environ. Microbiol. 1990, 56, 1559–1564. [Google Scholar]
- Shahid, M.; Malik, A. Plasmid mediated amikacin resistance in clinical isolates of Pseudomonas aeruginosa. Ind. J. Med. Microbiol. 2004, 22, 182–184. [Google Scholar]
- Jombo, G.T.A.; Jonah, P.; Ayeni, J.A. Multidrug resistant pseudomonas aeruginosa in contemporary medical practice: Findings from urinary isolates at a Nigerian University Teaching Hospital. Niger. J. Physiol. Sci. 2008, 23, 105–109. [Google Scholar]
- Ozumba, U.C. Antibiotic sensitivity of isolates of Pseudomonas aeruginosa in Enugu, Nigeria. Afr. J. Clin. Exp. Microbiol. 2003, 4, 48–51. [Google Scholar]
- Gad, G.F.; El-Domany, R.A.; Zaki, S.; Ashour, H.M. Characterization of Pseudomonas aeruginosa isolated from clinical and environmental samples in Minia, Egypt: Prevalence, antibiogram and resistance mechanisms. J. Antimicrob. Chemother. 2007, 60, 1010–1017. [Google Scholar] [CrossRef]
- Emmanuel, I.; Joseph, N.; Kingsley, E.-I.; Egbebor, E.M.; Lawrence, E. Antibiotic susceptibility profiles of enteric bacterial isolates from dumpsite utisols and water sources in a rural community in Cross River State, Southern Nigeria. Nat. Sci. 2011, 9, 46–50. [Google Scholar]
- Li, D.; Yu, T.; Zhang, Y.; Yang, M.; Li, Z.; Liu, M.; Qi, R. Antibiotic resistance characteristics of environmental bacteria from an oxytetracycline production wastewater treatment plant and the receiving river. Appl. Environ. Microbiol. 2010, 76, 3444–3451. [Google Scholar] [CrossRef]
- Ashish, J.; Warghane, G.N.; Wagh, B.B.; Nag, S.P.; Jisnani, M.L.; Thaware, R.R.; Kitey, H.S. Isolation and characterization of Pseudomonas species from Godavari river sample. Asiat. J. Biotechol. Res. 2011, 2, 862–866. [Google Scholar]
- Cabrera, E.C.; Halos, S.C.; Melecia, A.; Velmonte, M.D. Antibiograms, O Serotypes and R Plasmids of nosocomial Pseudomonas aeruginosa isolates. Philipp. J. Microbiol. Infect. Dis. 1997, 26, 121–128. [Google Scholar]
- Murray, G.E.; Tobin, R.S.; Junkins, B.; Kushner, D.J. Effect of chlorination on antibiotic resistance profiles of sewage-related bacteria. Appl. Environ. Microbiol. 1984, 48, 73–77. [Google Scholar]
- Malekzadeh, F.; Abdi‐ali, A.; Levin, M.; Shahamat, M. Prevalence of Pseudomonas aeruginosa pyocin and antibiotic biotypes in four Tehran hospitals. Int. J. Environ. Health Res. 1995, 5, 229–238. [Google Scholar] [CrossRef]
- Paul, S.L.; Bezbaruah, R.L.; Roy, M.K.; Ghosh, A.C. Multiple antibiotic resistance index and its reversion in Pseudomonas aeruginosa. Lett. Appl. Microbiol. 1997, 24, 169–171. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Odjadjare, E.E.; Igbinosa, E.O.; Mordi, R.; Igere, B.; Igeleke, C.L.; Okoh, A.I. Prevalence of Multiple Antibiotics Resistant (MAR) Pseudomonas Species in the Final Effluents of Three Municipal Wastewater Treatment Facilities in South Africa. Int. J. Environ. Res. Public Health 2012, 9, 2092-2107. https://doi.org/10.3390/ijerph9062092
Odjadjare EE, Igbinosa EO, Mordi R, Igere B, Igeleke CL, Okoh AI. Prevalence of Multiple Antibiotics Resistant (MAR) Pseudomonas Species in the Final Effluents of Three Municipal Wastewater Treatment Facilities in South Africa. International Journal of Environmental Research and Public Health. 2012; 9(6):2092-2107. https://doi.org/10.3390/ijerph9062092
Chicago/Turabian StyleOdjadjare, Emmanuel E., Etinosa O. Igbinosa, Raphael Mordi, Bright Igere, Clara L. Igeleke, and Anthony I. Okoh. 2012. "Prevalence of Multiple Antibiotics Resistant (MAR) Pseudomonas Species in the Final Effluents of Three Municipal Wastewater Treatment Facilities in South Africa" International Journal of Environmental Research and Public Health 9, no. 6: 2092-2107. https://doi.org/10.3390/ijerph9062092
APA StyleOdjadjare, E. E., Igbinosa, E. O., Mordi, R., Igere, B., Igeleke, C. L., & Okoh, A. I. (2012). Prevalence of Multiple Antibiotics Resistant (MAR) Pseudomonas Species in the Final Effluents of Three Municipal Wastewater Treatment Facilities in South Africa. International Journal of Environmental Research and Public Health, 9(6), 2092-2107. https://doi.org/10.3390/ijerph9062092





