Molecular Epidemiology for Vector Research on Leishmaniasis
Abstract
:1. Leishmaniasis
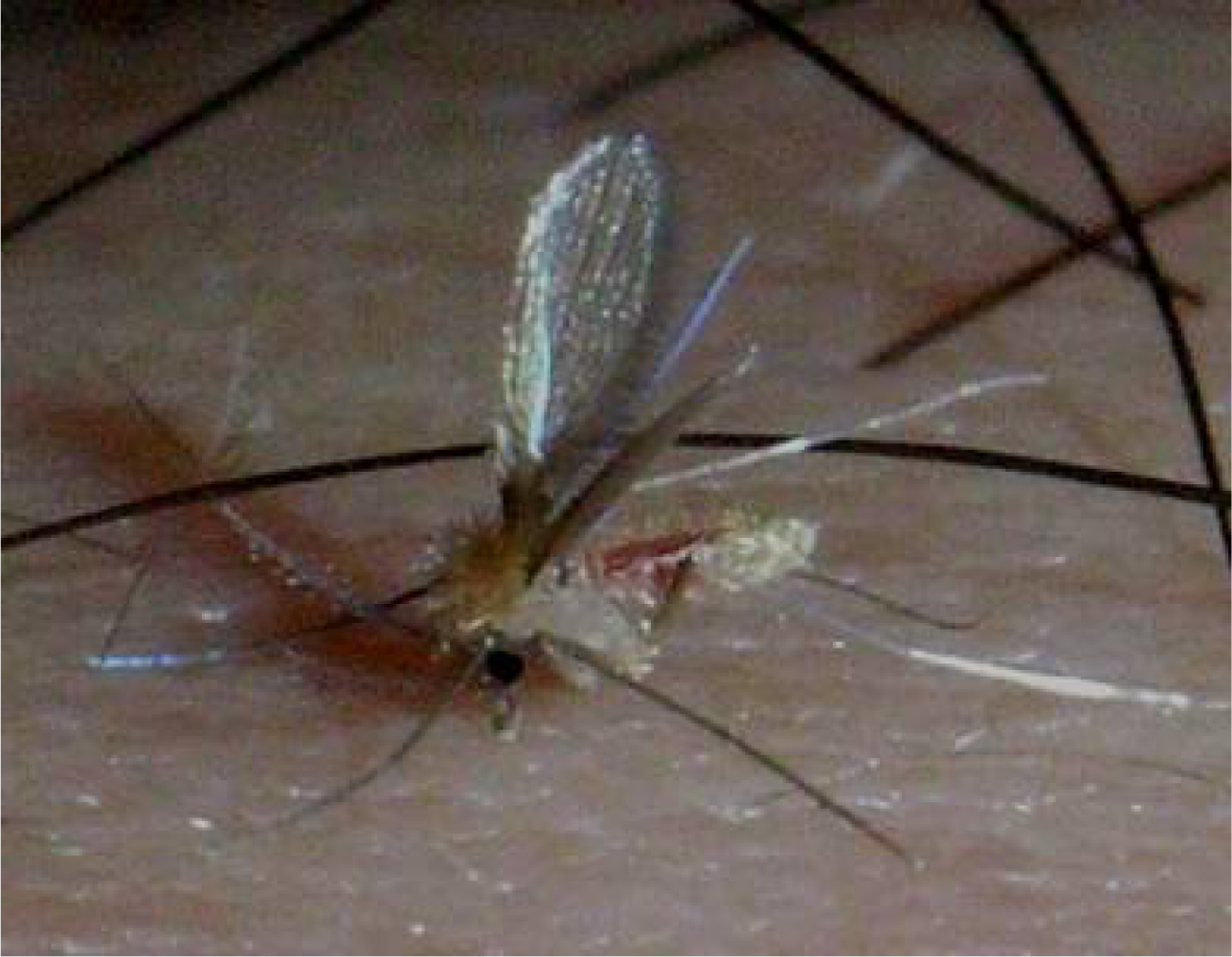
2. Phlebotomine Sand Flies as a Vector of Leishmaniasis
3. Advances in Sand Fly Taxonomy
4. Advances in the Detection and Identification of Leishmania Species within Naturally Infected Sand Flies
5. Concluding Remarks
References
- Desjeux, P. The increase of risk factors for leishmaniasis worldwide. Trans. R. Soc. Trop. Med. Hyg 2001, 95, 239–243. [Google Scholar]
- Desjeux, P. Leishmaniasis: current situation and new perspectives. Comp. Immunol. Microbiol. Infect. Dis 2004, 27, 305–318. [Google Scholar]
- Choi, CM; Lerner, EA. Leishmaniasis as an emerging infection. J. Invest. Dermatol. Symp. Proc 2001, 6, 175–182. [Google Scholar]
- Grimaldi, G, Jr; Tesh, RB; McMahon-Pratt, D. A review of the geographic distribution and epidemiology of leishmaniasis in the New World. Am. J. Trop. Med. Hyg 1989, 41, 687–725. [Google Scholar]
- Marsden, PD. Mucosal leishmaniasis (“espundia” Escomel, 1911). Trans. R. Soc. Trop. Med. Hyg 1986, 80, 859–876. [Google Scholar]
- Naiff, RD; Talhari, S; Barrett, TV. Isolation of Leishmania guyanensis from lesions of the nasal mucosa. Mem. Inst. Oswaldo Cruz 1988, 83, 529–530. [Google Scholar]
- Barral, A; Pedral-Sampaio, D; Grimaldi, G, Jr; Momen, H; McMahon-Pratt, D; Ribeiro de Jesus, A; Almeida, R; Badaro, R; Barral-Netto, M; Carvalho, EM; Johnson, WD, Jr. Leishmaniasis in Bahia, Brazil: evidence that Leishmania amazonensis produces a wide spectrum of clinical disease. Am. J. Trop. Med. Hyg 1991, 44, 536–546. [Google Scholar]
- Osorio, LE; Castillo, CM; Ochoa, MT. Mucosal leishmaniasis due to Leishmania (Viannia) panamensis in Colombia: clinical characteristics. Am. J. Trop. Med. Hyg 1998, 59, 49–52. [Google Scholar]
- Bryceson, ADM. Leishmaniasis. In Manson’s Tropical Disease, 20th ed; Cook, CC, Ed.; W.B. Saunders Comp. Ltd: London, Philadelphia, Tronto, Sydney and Tokyo, 1996; pp. 1213–1245. [Google Scholar]
- Bates, PA. Transmission of Leishmania metacyclic promastigotes by phlebotomine sand flies. Int. J. Parasitol 2007, 37, 1097–1106. [Google Scholar]
- Bates, PA; Rogers, ME. New insights into the developmental biology and transmission mechanisms of Leishmania. Curr. Mol. Med 2004, 4, 601–609. [Google Scholar]
- Kamhawi, S. Phlebotomine sand flies and Leishmania parasites: friends or foes? Trends Parasitol 2006, 22, 439–445. [Google Scholar]
- Peters, NC; Sacks, DL. The impact of vector-mediated neutrophil recruitment on cutaneous leishmaniasis. Cell. Microbiol 2009, 11, 1290–1296. [Google Scholar]
- Laskay, T; van Zandbergen, G; Solbach, W. Neutrophil granulocytes—Trojan horses for Leishmania major and other intracellular microbes? Trends Microbiol 2003, 11, 210–214. [Google Scholar]
- Ribeiro-Gomes, FL; Otero, AC; Gomes, NA; Moniz-De-Souza, MC; Cysne-Finkelstein, L; Arnholdt, AC; Calich, VL; Coutinho, SG; Lopes, MF; DosReis, GA. Macrophage interactions with neutrophils regulate Leishmania major infection. J. Immunol 2004, 172, 4454–4462. [Google Scholar]
- van Zandbergen, G; Klinger, M; Mueller, A; Dannenberg, S; Gebert, A; Solbach, W; Laskay, T. Cutting edge: neutrophil granulocyte serves as a vector for Leishmania entry into macrophages. J. Immunol 2004, 173, 6521–6525. [Google Scholar]
- Peters, NC; Egen, JG; Secundino, N; Debrabant, A; Kimblin, N; Kamhawi, S; Lawyer, P; Fay, MP; Germain, RN; Sacks, D. In vivo imaging reveals an essential role for neutrophils in leishmaniasis transmitted by sand flies. Science 2008, 321, 970–974. [Google Scholar]
- Jochim, RC; Teixeira, C. Leishmania commandeers the host inflammatory response through neutrophils. Trends Parasitol 2009, 25, 145–147. [Google Scholar]
- Young, DG; Duncan, MA. Guide to the identification and geographic distribution of Lutzomyia sand flies in Mexico, the West Indies, Central and South America (Diptera: Psychodidae), Memoirs of the American Entomological Institute, 54, Associated Publishers—American Entomological Institute, Gainsville, FL, USA, 1994.
- Munstermann, LE. Phlebotomine sand flies, the Psychodidae. In Biology of Disease Vectors, 2nd ed; Marquardt, WC, Black, WC, Freier, JE, Hagedorn, HH, Hemingway, J, Higgs, S, James, AA, Kondratieff, B, Moore, CG, Eds.; Elsevier: San Diego, CA, USA, 2004; pp. 141–151. [Google Scholar]
- Killick-Kendrick, R. The biology and control of phlebotomine sand flies. Clin. Dermatol 1999, 17, 279–289. [Google Scholar]
- Sacks, DL. Leishmania-sand fly interactions controlling species-specific vector competence. Cell. Microbiol 2001, 3, 189–196. [Google Scholar]
- Sacks, DL; Modi, G; Rowton, E; Späth, G; Epstein, L; Turco, SJ; Beverley, SM. The role of phosphoglycans in Leishmania-sand fly interactions. Proc. Natl. Acad. Sci. USA 2000, 97, 406–411. [Google Scholar]
- Kamhawi, S; Ramalho-Ortigao, M; Pham, VM; Kumar, S; Lawyer, PG; Turco, SJ; Barillas-Mury, C; Sacks, DL; Valenzuela, JG. A role for insect galectins in parasite survival. Cell 2004, 119, 329–341. [Google Scholar]
- Bauzer, LG; Souza, NA; Maingon, RD; Peixoto, AA. Lutzomyia longipalpis in Brazil: a complex or a single species? A mini-review. Mem. Inst. Oswaldo Cruz 2007, 102, 1–12. [Google Scholar]
- Ready, PD; Smith, DF; Killick-Kendrick, R. DNA hybridizations on squash-blotted sandflies to identify both Phlebotomus papatasi and infecting Leishmania major. Med. Vet. Entomol 1988, 2, 109–116. [Google Scholar]
- Ready, PD; Lainson, R; Shaw, JJ; Souza, AA. DNA probes for distinguishing Psychodopygus wellcomei from Psychodopygus complexus (Diptera:Psychodidae). Mem. Inst. Oswaldo Cruz 1991, 86, 41–49. [Google Scholar]
- Ready, PD; de Souza, AA; Rebelo, JM; Day, JC; Silveira, FT; Campbell-Lendrum, D; Davies, CR; Costa, JM. Phylogenetic species and domesticity of Lutzomyia whitmani at the southeast boundary of Amazonian Brazil. Trans. R. Soc. Trop. Med. Hyg 1998, 92, 159–160. [Google Scholar]
- Booth, DR; Ready, PD; Smith, DF. Retrotransposons and evolution in phlebotomines. Parassitologia 1991, 33, 105–112. [Google Scholar]
- Booth, DR; Ready, PD; Smith, DF. Isolation of non-LTR retrotransposon reverse transcriptase-like sequences from phlebotomine sandflies. Insect Mol. Biol 1994, 3, 89–96. [Google Scholar]
- Booth, DR; Ready, PD; Smith, DF. Evolution of multiple families of non-LTR retrotransposons in phlebotomine sandflies. Genet. Res 1996, 67, 227–237. [Google Scholar]
- Esseghir, S; Ready, PD; Killick-Kendrick, R; Ben-Ismail, R. Mitochondrial haplotypes and phylogeography of Phlebotomus vectors of Leishmania major. Insect Mol. Biol 1997, 6, 211–225. [Google Scholar]
- Aransay, AM; Scoulica, E; Chaniotis, B; Tselentis, Y. Typing of sand flies from Greece and Cyprus by DNA polymorphism of 18S rRNA gene. Insect Mol. Biol 1999, 8, 179–184. [Google Scholar]
- Aransay, AM; Scoulica, E; Tselentis, Y; Ready, PD. Phylogenetic relationships of phlebotomine sandflies inferred from small subunit nuclear ribosomal DNA. Insect Mol. Biol 2000, 9, 157–168. [Google Scholar]
- Aransay, AM; Ready, PD; Morillas-Marquez, F. Population differentiation of Phlebotomus perniciosus in Spain following postglacial dispersal. Heredity 2003, 90, 316–325. [Google Scholar]
- Campbell-Lendrum, DH; Brandão-Filho, SP; Pinto, MC; Vexenat, A; Ready, PD; Davies, CR. Domesticity of Lutzomyia whitmani (Diptera: psychodidae) populations: field experiments indicate behavioural differences. Bull. Entomol. Res 2000, 90, 41–48. [Google Scholar]
- Di Muccio, T; Marinucci, M; Frusteri, L; Maroli, M; Pesson, B; Gramiccia, M. Phylogenetic analysis of Phlebotomus species belonging to the subgenus Larroussius (Diptera, psychodidae) by ITS2 rDNA sequences. Insect Biochem. Mol. Biol 2000, 30, 387–393. [Google Scholar]
- Depaquit, J; Ferté, H; Léger, N; Lefranc, F; Alves-Pires, C; Hanafi, H; Maroli, M; Morillas-Marquez, F; Rioux, JA; Svobodova, M. Volf, P. ITS 2 sequences heterogeneity in Phlebotomus sergenti and Phlebotomus similis (Diptera, Psychodidae): possible consequences in their ability to transmit Leishmania tropica. Int. J. Parasitol 2002, 32, 1123–1131. [Google Scholar]
- Depaquit, J; Lienard, E; Verzeaux-Griffon, A; Ferté, H; Bounamous, A; Gantier, JC; Hanafi, HA; Jacobson, RL; Maroli, M; Moin-Vaziri, V; Müller, F; Ozbel, Y; Svobodova, M; Volf, P; Léger, N. Molecular homogeneity in diverse geographical populations of Phlebotomus papatasi (Diptera, Psychodidae) inferred from ND4 mtDNA and ITS2 rDNA epidemiological consequences. Infect. Genet. Evol 2008, 8, 159–170. [Google Scholar]
- Testa, JM; Montoya-Lerma, J; Cadena, H; Oviedo, M; Ready, PD. Molecular identification of vectors of Leishmania in Colombia: mitochondrial introgression in the Lutzomyia townsendi series. Acta Trop 2002, 84, 205–218. [Google Scholar]
- Torgerson, DG; Lampo, M; Velazquez, Y; Woo, PT. Genetic relationships among some species groups within the genus Lutzomyia (Diptera: Psychodidae). Am. J. Trop. Med. Hyg 2003, 69, 484–493. [Google Scholar]
- Beati, L; Caceres, AG; Lee, JA; Munstermann, LE. Systematic relationships among Lutzomyia sand flies (Diptera: Psychodidae) of Peru and Colombia based on the analysis of 12S and 28S ribosomal DNA sequences. Int. J. Parasitol 2004, 34, 225–234. [Google Scholar]
- Yahia, H; Ready, PD; Hamdani, A; Testa, JM; Guessous-Idrissi, N. Regional genetic differentiation of Phlebotomus sergenti in three Moroccan foci of cutaneous leishmaniasis caused by Leishmania tropica. Parasite 2004, 11, 189–199. [Google Scholar]
- Pesson, B; Ready, JS; Benabdennbi, I; Martín-Sánchez, J; Esseghir, S; Cadi-Soussi, M; Morillas-Marquez, F; Ready, PD. Sandflies of the Phlebotomus perniciosus complex: mitochondrial introgression and a new sibling species of P. longicuspis in the Moroccan Rif. Med. Vet. Entomol 2004, 18, 25–37. [Google Scholar]
- Kato, H; Uezato, H; Katakura, K; Calvopiña, M; Marco, JD; Barroso, PA; Gomez, EA; Mimori, T; Korenaga, M; Iwata, H; Nonaka, S; Hashiguchi, Y. Detection and identification of Leishmania species within naturally infected sand flies in the Andean areas of Ecuador by a polymerase chain reaction. Am. J. Trop. Med. Hyg 2005, 72, 87–93. [Google Scholar]
- Kato, H; Uezato, H; Gomez, EA; Terayama, Y; Calvopiña, M; Iwata, H; Hashiguchi, Y. Establishment of a mass screening method of sand fly vectors for Leishmania infection by molecular biological methods. Am. J. Trop. Med. Hyg 2007, 77, 324–329. [Google Scholar]
- Kato, H; Cáceres, AG; Gomez, EA; Mimori, T; Uezato, H; Marco, JD; Barroso, PA; Iwata, H; Hashiguchi, Y. Molecular mass screening to incriminate sand fly vectors of Andean-type cutaneous leishmaniasis in Ecuador and Peru. Am. J. Trop. Med. Hyg 2008, 79, 719–721. [Google Scholar]
- Kato, H; Gomez, EA; Yamamoto, Y; Calvopiña, M; Guevara, AG; Marco, JD; Barroso, PA; Iwata, H; Hashiguchi, Y. Natural infection of Lutzomyia tortura with Leishmania (Viannia) naiffi in an Amazonian area of Ecuador. Am. J. Trop. Med. Hyg 2008, 79, 438–440. [Google Scholar]
- Barón, S; Martín-Sánchez, J; Gállego, M; Morales-Yuste, M; Boussaa, S; Morillas-Márquez, F. Intraspecific variability (rDNA ITS and mtDNA Cyt b) of Phlebotomus sergenti in Spain and Morocco. Acta Trop 2008, 107, 259–267. [Google Scholar]
- Barroso, PA; Marco, JD; Kato, H; Tarama, R; Rueda, P; Cajal, SP; Basombrío, MA; Korenaga, M; Taranto, NJ; Hashiguchi, Y. The identification of sand fly species, from an area of Argentina with endemic leishmaniasis, by the PCR-based analysis of the gene coding for 18S ribosomal RNA. Ann. Trop. Med. Parasitol 2007, 101, 247–253. [Google Scholar]
- Terayama, Y; Kato, H; Gomez, EAL; Uezato, H; Calvopiña, M; Iwata, H; Hashiguchi, Y. Molecular typing of sand fly species (Diptera, Psychodidae, Phlebotominae) from areas endemic for leishmaniasis in Ecuador by PCR-RFLP of 18S ribosomal RNA gene. J. Vet. Med. Sci 2008, 70, 907–913. [Google Scholar]
- Kuwahara, K; Kato, H; Gomez, EA; Uezato, H; Mimori, T; Yamamoto, YI; Calvopiña, M; Cáceres, AG; Iwata, H; Hashiguchi, Y. Genetic diversity of ribosomal RNA internal transcribed spacer sequences in Lutzomyia species from areas endemic for New World cutaneous leishmaniasis. Acta Trop 2009, 112, 131–136. [Google Scholar]
- Krzywinski, J; Besansky, NJ. Molecular systematics of Anopheles: from subgenera to subpopulations. Annu. Rev. Entomol 2003, 48, 111–139. [Google Scholar]
- Obsomer, V; Defourny, P; Coosemans, M. The Anopheles dirus complex: spatial distribution and environmental drivers. Malar. J 2007, 6, 26. [Google Scholar]
- Soto, SI; Lehmann, T; Rowton, ED; Vélez, BID; Porter, CH. Speciation and population structure in the morphospecies Lutzomyia longipalpis (Lutz & Neiva) as derived from the mitochondrial ND4 gene. Mol. Phylogenet. Evol 2001, 18, 84–93. [Google Scholar]
- Bottecchia, M; Oliveira, SG; Bauzer, LG; Souza, NA; Ward, RD; Garner, KJ; Kyriacou, CP; Peixoto, AA. Genetic divergence in the cacophony IVS6 intron among five Brazilian populations of Lutzomyia longipalpis. J. Mol. Evol 2004, 58, 754–761. [Google Scholar]
- Bauzer, LG; Souza, NA; Ward, RD; Kyriacou, CP; Peixoto, AA. The period gene and genetic differentiation between three Brazilian populations of Lutzomyia longipalpis. Insect. Mol. Biol 2002, 11, 315–323. [Google Scholar]
- Bauzer, LG; Gesto, JS; Souza, NA; Ward, RD; Hamilton, JG; Kyriacou, CP; Peixoto, AA. Molecular divergence in the period gene between two putative sympatric species of the Lutzomyia longipalpis complex. Mol. Biol. Evol 2002, 19, 1624–1627. [Google Scholar]
- Maingon, RD; Ward, RD; Hamilton, JG; Bauzer, LG; Peixoto, AA. The Lutzomyia longipalpis species complex: does population sub-structure matter to Leishmania transmission? Trends Parasitol 2008, 24, 2–7. [Google Scholar]
- Lins, RM; Souza, NA; Peixoto, AA. Genetic divergence between two sympatric species of the Lutzomyia longipalpis complex in the paralytic gene, a locus associated with insecticide resistance and lovesong production. Mem. Inst. Oswaldo Cruz 2008, 103, 736–740. [Google Scholar]
- Hamarsheh, O; Presber, W; Yaghoobi-Ershadi, MR; Amro, A; Al-Jawabreh, A; Sawalha, S; Al-Lahem, A; Das, ML; Guernaoui, S; Seridi, N; Dhiman, RC; Hashiguchi, Y; Ghrab, J; Hassan, M; Schönian, G. Population structure and geographical subdivision of the Leishmania major vector Phlebotomus papatasi as revealed by microsatellite variation. Med. Vet. Entomol 2009, 23, 69–77. [Google Scholar]
- Hashiguchi, Y; Gomez, EAL. A review of leishmaniasis in Ecuador. Bull. Pan Am. Hlth. Org 1991, 25, 64–76. [Google Scholar]
- Hashiguchi, Y. Leishmaniasis. In Progress of Medical Parasitology in Japan; Otsuru, M, Kamegai, S, Hayashi, S, Eds.; Megro Parasitological Museum: Tokyo, Japan, 2003; Volume 7, pp. 537–553. [Google Scholar]
- Aransay, AM; Scoulica, E; Tselentis, Y. Detection and identification of Leishmania DNA within naturally infected sand flies by seminested PCR on minicircle kinetoplast DNA. Appl. Environ. Microbiol 2000, 66, 1933–1938. [Google Scholar]
- Parvizi, P; Mauricio, I; Aransay, AM; Miles, MA; Ready, PD. First detection of Leishmania major in peridomestic Phlebotomus papatasi from Isfahan province, Iran: comparison of nested PCR of nuclear ITS ribosomal DNA and semi-nested PCR of minicircle kinetoplast DNA. Acta Trop 2005, 93, 75–83. [Google Scholar]
- Jorquera, A; González, R; Marchán-Marcano, E; Oviedo, M; Matos, M. Multiplex-PCR for detection of natural Leishmania infection in Lutzomyia spp. captured in an endemic region for cutaneous leishmaniasis in state of Sucre, Venezuela. Mem. Inst. Oswaldo Cruz 2005, 100, 45–48. [Google Scholar]
- Paiva, BR; Secundino, NF; Nascimento, JC; Pimenta, PF; Galati, EA; Junior, HF; Malafronte, RS. Detection and identification of Leishmania species in field-captured phlebotomine sandflies based on mini-exon gene PCR. Acta Trop 2006, 99, 252–259. [Google Scholar]
- Córdoba-Lanús, E; De Grosso, ML; Piñero, JE; Valladares, B; Salomón, OD. Natural infection of Lutzomyia neivai with Leishmania spp. in northwestern Argentina. Acta Trop 2006, 98, 1–5. [Google Scholar]
- Azizi, K; Rassi, Y; Javadian, E; Motazedian, MH; Asgari, Q; Yaghoobi-Ershadi, MR. First detection of Leishmania infantum in Phlebotomus (Larroussius) major (Diptera: Psychodidae) from Iran. J. Med. Entomol 2008, 45, 726–731. [Google Scholar]
- Pita-Pereira, D; Souza, GD; Zwetsch, A; Alves, CR; Britto, C; Rangel, EF. First report of Lutzomyia (Nyssomyia) neivai (Diptera: Psychodidae: Phlebotominae) naturally infected by Leishmania (Viannia) braziliensis in a periurban area of south Brazil using a multiplex polymerase chain reaction assay. Am. J. Trop. Med. Hyg 2009, 80, 593–595. [Google Scholar]
- Silva, EA; Andreotti, R; Dias, ES; Barros, JC; Brazuna, JC. Detection of Leishmania DNA in phlebotomines captured in Campo Grande, Mato Grosso do Sul, Brazil. Exp. Parasitol 2008, 119, 343–348. [Google Scholar]
- Pandey, K; Pant, S; Kanbara, H; Shuaibu, MN; Mallik, AK; Pandey, BD; Kaneko, O; Yanagi, T. Molecular detection of Leishmania parasites from whole bodies of sandflies collected in Nepal. Parasitol. Res 2008, 103, 293–297. [Google Scholar]
- Savani, ES; Nunes, VL; Galati, EA; Castilho, TM; Zampieri, RA; Floeter-Winter, LM. The finding of Lutzomyia almerioi and Lutzomyia longipalpis naturally infected by Leishmania spp. in a cutaneous and canine visceral leishmaniases focus in Serra da Bodoquena, Brazil. Vet. Parasitol 2009, 160, 18–24. [Google Scholar]
- Luyo-Acero, G; Uezato, H; Oshiro, M; Kariya, K; Katakura, K; Gomez, EAL; Hashiguchi, Y; Nonaka, S. Sequence variation of the cytochrome b gene of various human infecting members of the genus Leishmania and their pathology. Parasitol 2004, 128, 483–491. [Google Scholar]
- Marco, JD; Bhutto, AM; Soomro, FR; Baloch, JH; Barroso, PA; Kato, H; Uezato, H; Katakura, K; Korenaga, M; Nonaka, S; Hashiguchi, Y. Multilocus enzyme electrophoresis and cytochrome b gene sequencing-based identification of Leishmania isolates from different foci of cutaneous leishmaniasis in Pakistan. Am. J. Trop. Med. Hyg 2006, 75, 261–266. [Google Scholar]
- Marco, JD; Uezato, H; Mimori, T; Barroso, PA; Korenaga, M; Nonaka, S; Basombrio, MA; Taranto, NJ; Hashiguchi, Y. Are cytochrome b gene sequencing and polymorphism-specific polymerase chain reaction as reliable as multilocus enzyme electrophoresis for identifying Leishmania spp. from Argentina? Am. J. Trop. Med. Hyg 2006, 75, 256–260. [Google Scholar]

© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kato, H.; Gomez, E.A.; Cáceres, A.G.; Uezato, H.; Mimori, T.; Hashiguchi, Y. Molecular Epidemiology for Vector Research on Leishmaniasis. Int. J. Environ. Res. Public Health 2010, 7, 814-826. https://doi.org/10.3390/ijerph7030814
Kato H, Gomez EA, Cáceres AG, Uezato H, Mimori T, Hashiguchi Y. Molecular Epidemiology for Vector Research on Leishmaniasis. International Journal of Environmental Research and Public Health. 2010; 7(3):814-826. https://doi.org/10.3390/ijerph7030814
Chicago/Turabian StyleKato, Hirotomo, Eduardo A. Gomez, Abraham G. Cáceres, Hiroshi Uezato, Tatsuyuki Mimori, and Yoshihisa Hashiguchi. 2010. "Molecular Epidemiology for Vector Research on Leishmaniasis" International Journal of Environmental Research and Public Health 7, no. 3: 814-826. https://doi.org/10.3390/ijerph7030814
APA StyleKato, H., Gomez, E. A., Cáceres, A. G., Uezato, H., Mimori, T., & Hashiguchi, Y. (2010). Molecular Epidemiology for Vector Research on Leishmaniasis. International Journal of Environmental Research and Public Health, 7(3), 814-826. https://doi.org/10.3390/ijerph7030814




