Detection of COVID-19 Patients Using Machine Learning Techniques: A Nationwide Chilean Study
Abstract
:1. Introduction
2. Related Work
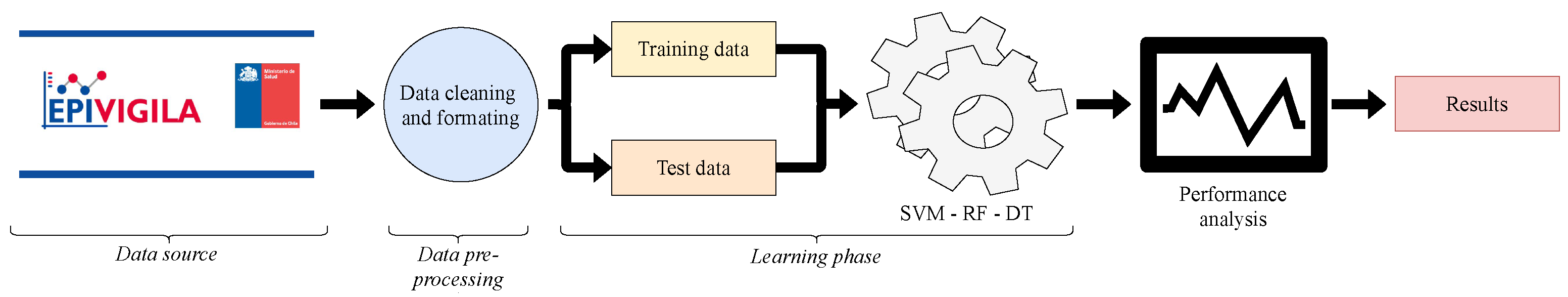
3. Materials and Methods
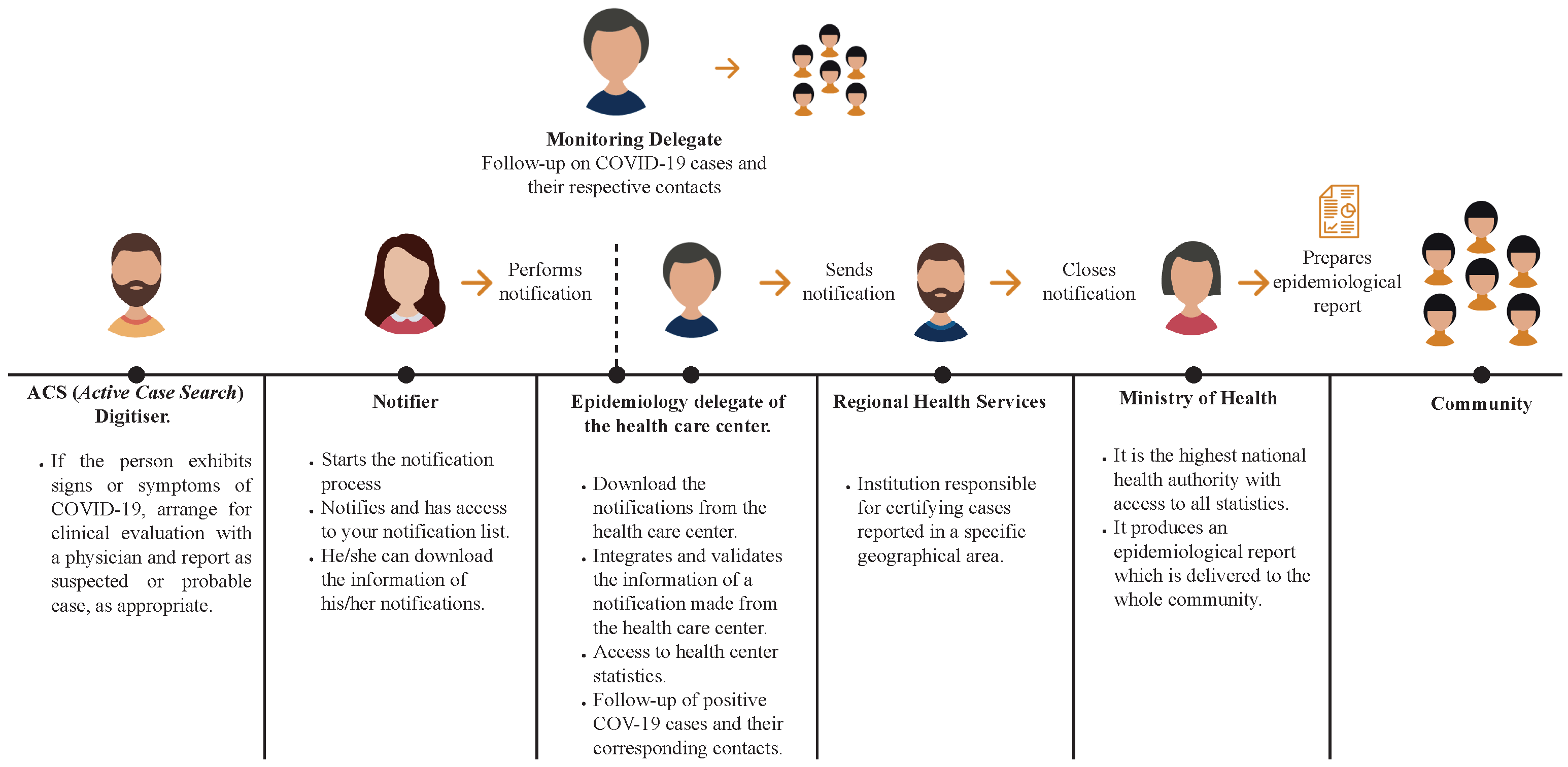
3.1. Data Source
3.2. Data Pre-Processing
3.3. Learning Phase
3.3.1. Support-Vector Machine (SVM)
3.3.2. Decision Tree (DT)
3.3.3. Random Forest (RF)
3.4. Performance Analysis
4. Results
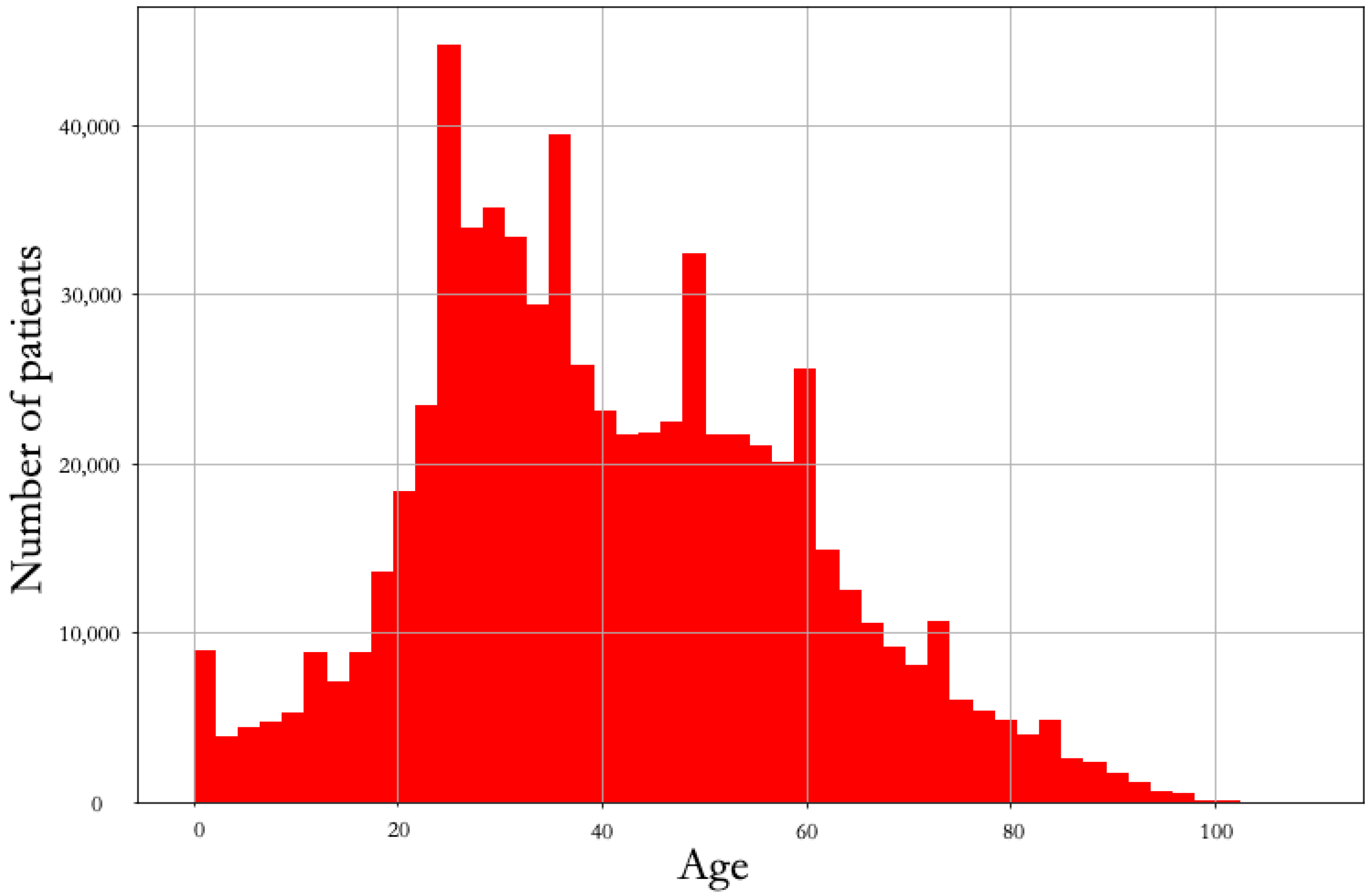
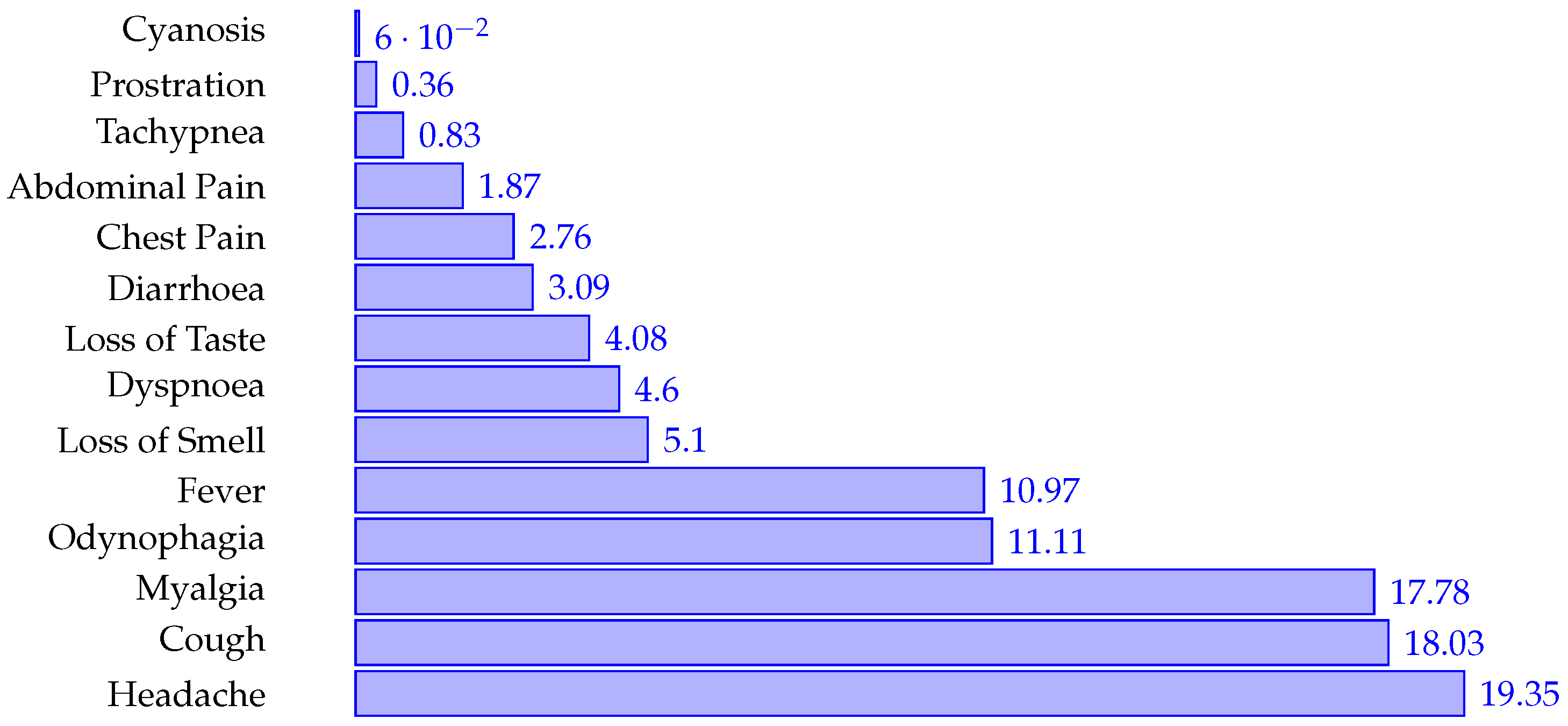
4.1. Demographics
4.2. Machine Learning Analysis
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Chilean Ministry of Health | MINSAL |
| Epidemiological surveillance system | Epivigila |
| Support-vector machine | SVM |
| Random forest | RF |
| Decision frees | DT |
| Harmonic mean between precision and recall | -score |
| The area under the curve | AUC |
References
- Kolbe, L.J. An Epidemiological Surveillance System to Monitor the Prevalence of Youth Behaviors That Most Affect Health. Health Educ. 1990, 21, 44–48. [Google Scholar] [CrossRef]
- Thacker, S.; Parrish, R.G.; Trowbridge, F. A method for evaluating systems of epidemiological surveillance. World Health Stat. Q. Rapp. Trimest. Stat. Sanit. Mond. 1988, 41, 11–18. [Google Scholar]
- Nogueira, R.; Davies, J.; Gupta, R.; Hassan, A.; Devlin, T.; Haussen, D.; Mohammaden, M.; Kellner, C.; Arthur, A.; Elijovich, L.; et al. Epidemiological surveillance of the impact of the COVID-19 pandemic on stroke care using artificial intelligence. Stroke 2021, 52, 1682–1690. [Google Scholar] [CrossRef] [PubMed]
- Chilean Ministry of Health. Epidemiological Surveillance System EPIVIGILA. 2022. Available online: http://epi.minsal.cl/sistema-de-vigilancia-epidemiologica-Epivigila-antecedentes/ (accessed on 20 June 2022).
- Noble, W. What is a support vector machine? Nat. Biotechnol. 2006, 24, 1565–1567. [Google Scholar] [CrossRef] [PubMed]
- Cutler, A.; Cutler, D.R.; Stevens, J.R. Random forests. In Ensemble Machine Learning; Springer: Berlin/Heidelberg, Germany, 2012; pp. 157–175. [Google Scholar] [CrossRef]
- Safavian, S.R.; Landgrebe, D. A survey of decision tree classifier methodology. IEEE Trans. Syst. Man Cybern. 1991, 21, 660–674. [Google Scholar] [CrossRef] [Green Version]
- Ahamad, M.; Aktar, S.; Rashed-Al-Mahfuz, M.; Uddin, S.; Liò, P.; Xu, H.; Summers, M.A.; Quinn, J.M.W.; Moni, M.A. A machine learning model to identify early stage symptoms of SARS-Cov-2 infected patients. Expert Syst. Appl. 2020, 160, 113661. [Google Scholar] [CrossRef] [PubMed]
- Aydin, N.; Yurdakul, G. Assessing countries’ performances against COVID-19 via WSIDEA and machine learning algorithms. Expert Syst. Appl. 2020, 97, 106792. [Google Scholar] [CrossRef]
- Awal, M.; Masud, M.; Hossain, M.S.; Bulbul, A.A.M.; Mahmud, S.H.; Bairagi, A.K. A novel bayesian optimization-based machine learning framework for COVID-19 detection from inpatient facility data. IEEE Access 2020, 97, 106792. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, C.; Larkin, J.W.; Chaudhuri, S.; Han, H.; Jiao, Y.; Bermudez, K.M.; Weinhandl, E.D.; Dahne-Steuber, I.A.; Belmont, K.; Neri, L.; et al. Machine Learning for Prediction of Hemodialysis Patients with an Undetected SARS-CoV-2 Infection. Kidney360 2021, 2, 456–468. [Google Scholar] [CrossRef]
- Shaban, W.; Rabie, A.H.; Saleh, A.I.; Abo-Elsoud, M.A. Detecting COVID-19 patients based on fuzzy inference engine and Deep Neural Network. Appl. Soft Comput. 2021, 99, 106906. [Google Scholar] [CrossRef]
- Jain, V.; Yuan, J.M. Predictive symptoms and comorbidities for severe COVID-19 and intensive care unit admission: A systematic review and meta-analysis. Int. J. Public Health 2020, 65, 533–546. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Radenkovic, D.; Perez, K.; Nadeau, K.; Verdin, E.; Furman, D. Modeling Predictive Age-Dependent and Age-Independent Symptoms and Comorbidities of Patients Seeking Treatment for COVID-19: Model Development and Validation Study. J. Med. Internet Res. 2021, 23, e25696. [Google Scholar] [CrossRef] [PubMed]
- Hsia, J.Y.; Lin, C.J. Parameter selection for linear support vector regression. IEEE Trans. Neural Netw. Learn. Syst. 2020, 31, 5639–5644. [Google Scholar] [CrossRef]
- Baeza-Yates, R.; Ribeiro-Neto, B. Modern Information Retrieval; ACM Press: New York, NY, USA, 1999. [Google Scholar]
- Pepe, M. Receiver operating characteristic methodology. J. Am. Stat. Assoc. 2000, 95, 308–311. [Google Scholar] [CrossRef]
- Manríquez, R.; Guerrero-Nancuante, C.; Martínez, F.; Taramasco, C. Spread of Epidemic Disease on Edge-Weighted Graphs from a Database: A Case Study of COVID-19. Int. J. Environ. Res. Public Health 2021, 18, 4432. [Google Scholar] [CrossRef]
- Manríquez, R.; Guerrero-Nancuante, C.; Taramasco, C. Protection Strategy against an Epidemic Disease on Edge-Weighted Graphs Applied to a COVID-19 Case. Biology 2021, 10, 667. [Google Scholar] [CrossRef]
- Mahdavi, M.; Choubdar, H.; Zabeh, E.; Rieder, M.; Safavi-Naeini, S.; Jobbagy, Z.; Ghorbani, A.; Abedini, A.; Kiani, A.; Khanlarzadeh, V.L.R.; et al. A machine learning based exploration of COVID-19 mortality risk. PLoS ONE 2021, 16, e0252384. [Google Scholar] [CrossRef]
- Moulaei, K.; Shanbehzadeh, M.; Mohammadi-Taghiabad, Z.; Kazemi-Arpanahi, H. Comparing machine learning algorithms for predicting COVID-19 mortality. BMC Med. Inform. Decis. Mak. 2022, 22, 2. [Google Scholar] [CrossRef]
- Mondal, M.R.H.; Bharati, S.; Podder, P. Diagnosis of COVID-19 Using Machine Learning and Deep Learning: A Review. Curr. Med. Imaging 2021, 17, 1403–1418. [Google Scholar] [CrossRef]
- de Souza, W.M.; Buss, L.F.; Candido, D.S.; Carrera, J.P.; Li, S.; Zarebski, A.E.; Pereira, R.H.M.; Prete, C.A.; de Souza-Santos, A.A.; Parag, K.V.; et al. Epidemiological and clinical characteristics of the COVID-19 epidemic in Brazil. Nat. Hum. Behav. 2020, 4, 856–865. [Google Scholar] [CrossRef]
- Montazeri, M.; ZahediNasab, R.; Farahani, A.; Mohseni, H.; Ghasemian, F. Machine Learning Models for Image-Based Diagnosis and Prognosis of COVID-19: Systematic Review. JMIR Med. Inform. 2021, 9, e25181. [Google Scholar] [CrossRef] [PubMed]
- Sujath, R.; Chatterjee, J.M.; Hassanien, A.E. A machine learning forecasting model for COVID-19 pandemic in India. Stoch. Environ. Res. Risk Assess. 2020, 34, 959–972. [Google Scholar] [CrossRef] [PubMed]
- Syeda, H.B.; Syed, M.; Sexton, K.W.; Syed, S.; Begum, S.; Syed, F.; Prior, F.; Yu, F., Jr. Role of machine learning techniques to tackle the COVID-19 crisis: Systematic review. JMIR Med. Inform. 2021, 9, e23811. [Google Scholar] [CrossRef] [PubMed]
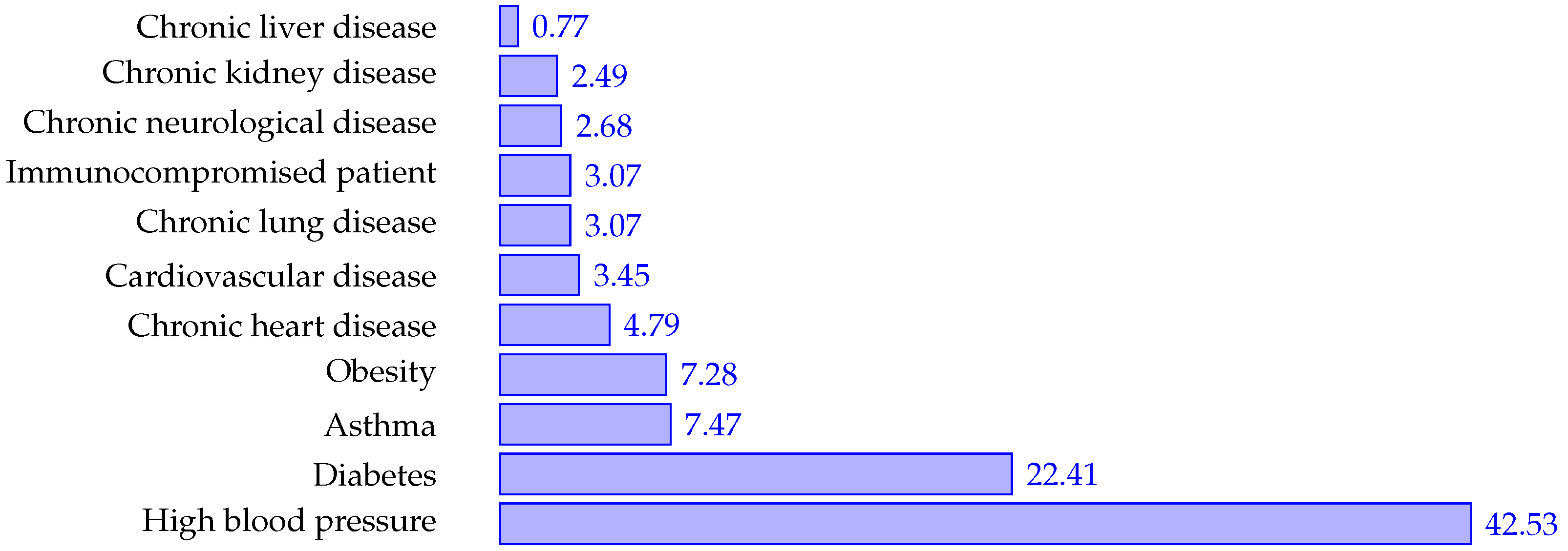
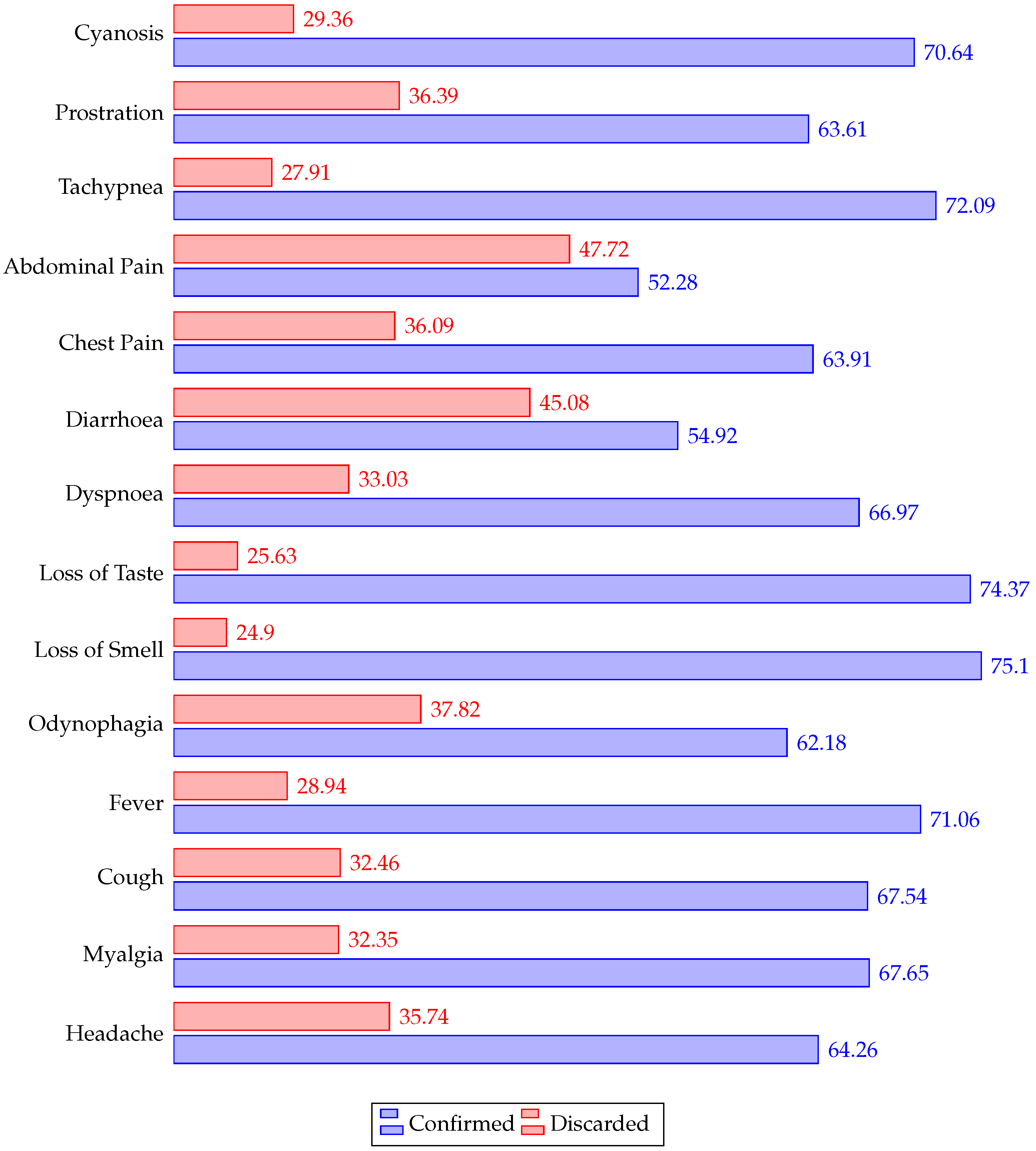
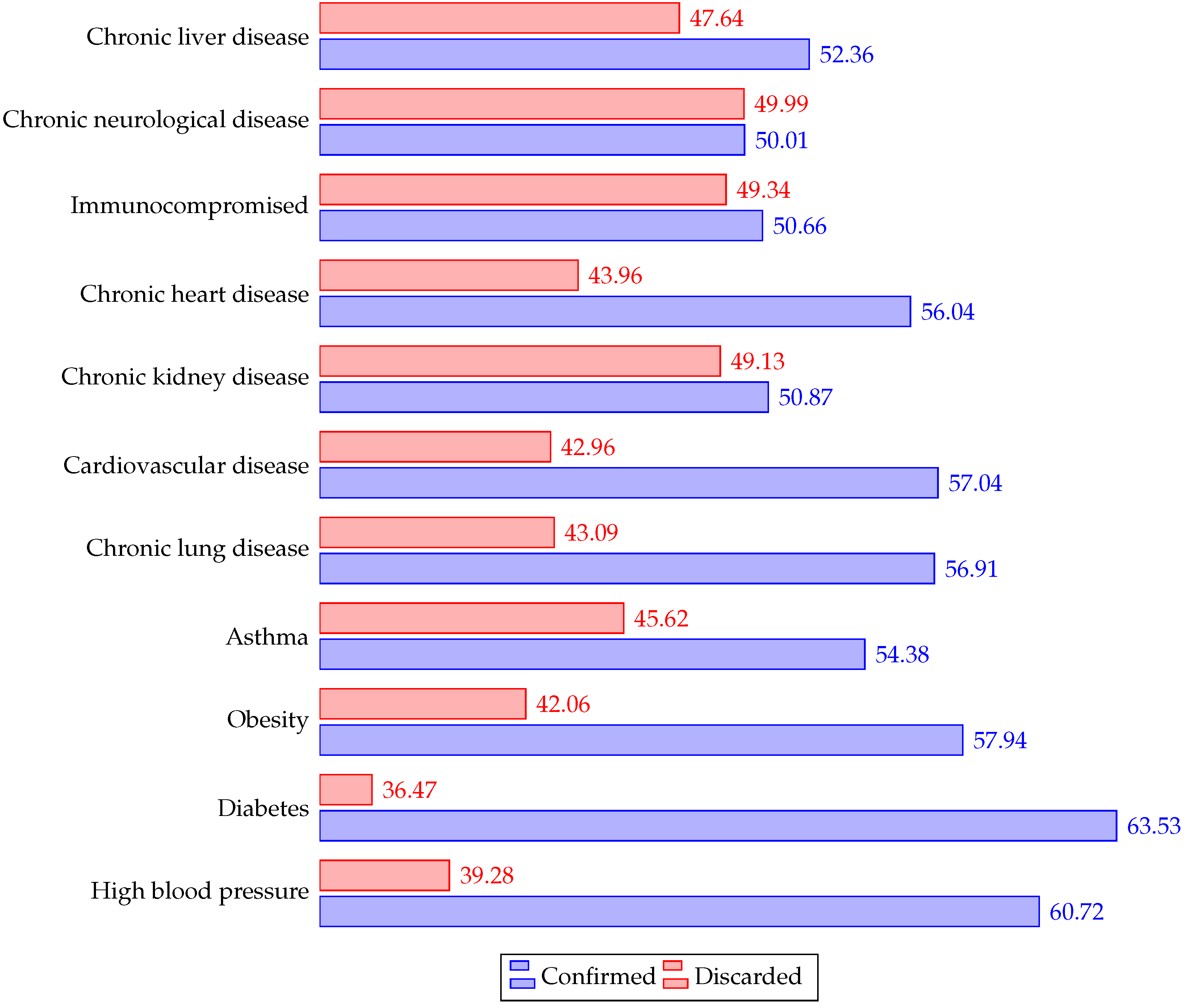
| Symptoms | Comorbidities |
|---|---|
| Tachypnoea | Asthma |
| Odynophagia | Chronic kidney disease |
| Cyanosis | Chronic lung disease |
| Abdominal pain | High blood pressure |
| Headache | Obesity |
| Fever | Immunocompromised patient |
| Diarrhoea | Chronic heart disease |
| Loss of taste | Diabetes |
| Myalgia | Chronic neurological disease |
| Chest pain | Chronic liver disease |
| Prostration | Cardiovascular disease |
| Dyspnoea | |
| Cough | |
| Loss of smell |
| All Patients | Suspected | Confirmed | Total |
|---|---|---|---|
| Mean age (interquartile range) | 37 (27–52) | 36 (26–51) | 39 (28–54) |
| Male gender (%) | 52.1% | 52.7% | 51.2% |
| Female gender (%) | 47.9% | 47.3% | 48.8% |
| Have symptoms (%) | 52.6% | 33.3% | 79.3% |
| Dataset | Technique | Top 5 Features | ||||
|---|---|---|---|---|---|---|
| 1st | 2nd | 3rd | 4th | 5th | ||
| Age (0–20) | SVM | Abdominal pain | Loss of taste | Chronic kidney disease | Tachypnea | Chronic liver disease |
| RF | Chronic heart disease | Odynophagia | Diarrhoea | Cough | Fever | |
| DT | Abdominal pain | Odynophagia | Diarrhoea | Loss of smell | Cough | |
| Age (21–60) | SVM | Abdominal pain | High blood pressure | Chronic kidney disease | Asthma | Diabetes |
| RF | Abdominal pain | Diarrhoea | Chronic heart disease | Cough | Fever | |
| DT | Abdominal pain | Cough | Odynophagia | Dyspnoea | Chronic heart disease | |
| Age (61–96) | SVM | Abdominal pain | Diabetes | Loss of taste | Odynophagia | Fever |
| RF | Abdominal pain | Cough | Diarrhoea | Chronic heart disease | Fever | |
| DT | Abdominal pain | Cough | Diarrhoea | Chronic heart disease | Fever | |
| Age (0–96) | SVM | Abdominal pain | Chronic lung disease | Immunocompromised patient | Diabetes | Loss of taste |
| RF | Abdominal pain | Cough | Diarrhoea | Chronic heart disease | Fever | |
| DT | Abdominal pain | Chronic heart disease | Cough | Cardiovascular disease | Odynophagia | |
| Dataset | Technique | Precision | Recall | -Score | Specificity | AUC |
|---|---|---|---|---|---|---|
| SVM | 0.613 | 0.680 | 0.645 | 0.574 | 0.640 | |
| Age (0–20) | RF | 0.628 | 0.604 | 0.616 | 0.712 | 0.636 |
| DT | 0.628 | 0.558 | 0.591 | 0.737 | 0.626 | |
| Age (21–60) | SVM | 0.717 | 0.785 | 0.749 | 0.705 | 0.739 |
| RF | 0.735 | 0.721 | 0.728 | 0.758 | 0.732 | |
| DT | 0.731 | 0.667 | 0.697 | 0.792 | 0.712 | |
| Age (61–96) | SVM | 0.730 | 0.811 | 0.768 | 0.687 | 0.753 |
| RF | 0.717 | 0.690 | 0.704 | 0.747 | 0.705 | |
| DT | 0.718 | 0.607 | 0.658 | 0.779 | 0.680 | |
| Age (0–96) | SVM | 0.727 | 0.798 | 0.761 | 0.681 | 0.748 |
| RF | 0.739 | 0.740 | 0.740 | 0.740 | 0.738 | |
| DT | 0.746 | 0.684 | 0.713 | 0.784 | 0.724 |
| 0–20 Years | 21–60 Years | 61–96 Years | 0–96 Years | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SVM | RF | DT | SVM | RF | DT | SVM | RF | DT | SVM | RF | DT | |
| Abdominal Pain | 0.9712 | 0.0711 | 0.1946 | 0.9962 | 0.1973 | 0.5565 | 0.9930 | 0.1723 | 0.4304 | 0.9919 | 0.1859 | 0.4294 |
| Asthma | 0.0016 | 0.0346 | 0.0527 | 0.0002 | 0.0211 | 0.0216 | 0.0000 | 0.0421 | 0.0359 | 0.0000 | 0.0053 | 0.0055 |
| Cardiovascular disease | 0.0000 | 0.0458 | 0.0379 | 0.0000 | 0.0289 | 0.0197 | 0.0000 | 0.0160 | 0.0231 | 0.0000 | 0.0308 | 0.0395 |
| Chest pain | 0.0001 | 0.0363 | 0.0484 | 0.0000 | 0.0042 | 0.0039 | 0.0000 | 0.0132 | 0.0169 | 0.0000 | 0.0312 | 0.0187 |
| Chronic heart disease | 0.0001 | 0.0946 | 0.0276 | 0.0000 | 0.1043 | 0.0292 | 0.0000 | 0.0676 | 0.0331 | 0.0000 | 0.0813 | 0.0657 |
| Chronic kidney disease | 0.1198 | 0.0052 | 0.0046 | 0.0632 | 0.0036 | 0.0043 | 0.0000 | 0.0137 | 0.0125 | 0.0000 | 0.0111 | 0.0141 |
| Chronic liver disease | 0.0849 | 0.0028 | 0.0049 | 0.0000 | 0.0152 | 0.0166 | 0.0000 | 0.0134 | 0.0104 | 0.0000 | 0.0039 | 0.0045 |
| Chronic lung disease | 0.0003 | 0.0116 | 0.0093 | 0.0000 | 0.0126 | 0.0170 | 0.0000 | 0.0224 | 0.0076 | 0.0001 | 0.0288 | 0.0247 |
| Chronic neurological disease | 0.0000 | 0.0029 | 0.0014 | 0.0000 | 0.0106 | 0.0094 | 0.0000 | 0.0088 | 0.0068 | 0.0000 | 0.0027 | 0.0003 |
| Cough | 0.0000 | 0.0793 | 0.0617 | 0.0000 | 0.0886 | 0.0334 | 0.0000 | 0.1032 | 0.0283 | 0.0000 | 0.0879 | 0.0566 |
| Cyanosis | 0.0002 | 0.0639 | 0.0516 | 0.0000 | 0.0308 | 0.0130 | 0.0000 | 0.0304 | 0.0302 | 0.0000 | 0.0326 | 0.0102 |
| Diabetes | 0.0007 | 0.0275 | 0.0367 | 0.0000 | 0.0193 | 0.0203 | 0.0000 | 0.0169 | 0.0170 | 0.0000 | 0.0148 | 0.0157 |
| Diarrhoea | 0.0000 | 0.0820 | 0.0680 | 0.0000 | 0.1221 | 0.0168 | 0.0000 | 0.0770 | 0.0432 | 0.0000 | 0.0863 | 0.0303 |
| Dyspnoea | 0.0001 | 0.0390 | 0.0394 | 0.0000 | 0.0268 | 0.0297 | 0.0000 | 0.0518 | 0.0465 | 0.0000 | 0.0403 | 0.0333 |
| Fever | 0.0000 | 0.0734 | 0.0464 | 0.0000 | 0.0744 | 0.0212 | 0.0000 | 0.0606 | 0.0093 | 0.0000 | 0.0652 | 0.0107 |
| Headache | 0.0283 | 0.0010 | 0.0022 | 0.0000 | 0.0001 | 0.0000 | 0.0000 | 0.0035 | 0.0026 | 0.0000 | 0.0014 | 0.0036 |
| High blood pressure | 0.0001 | 0.0035 | 0.0016 | 0.1019 | 0.0008 | 0.0012 | 0.0000 | 0.0152 | 0.0164 | 0.0000 | 0.0060 | 0.0054 |
| Immunocompromised patient | 0.0001 | 0.0010 | 0.0018 | 0.0000 | 0.0012 | 0.0020 | 0.0000 | 0.0173 | 0.0173 | 0.0000 | 0.0275 | 0.0299 |
| Loss of smell | 0.0001 | 0.0625 | 0.0645 | 0.0000 | 0.0329 | 0.0174 | 0.0000 | 0.0305 | 0.0268 | 0.0000 | 0.0375 | 0.0270 |
| Loss of taste | 0.4791 | 0.0521 | 0.0490 | 0.0000 | 0.0249 | 0.0268 | 0.0000 | 0.0228 | 0.0193 | 0.0000 | 0.0259 | 0.0281 |
| Myalgia | 0.0000 | 0.0487 | 0.0405 | 0.0000 | 0.0485 | 0.0257 | 0.0000 | 0.0158 | 0.0192 | 0.0000 | 0.0435 | 0.0253 |
| Obesity | 0.0000 | 0.0016 | 0.0013 | 0.0000 | 0.0030 | 0.0030 | 0.0000 | 0.0351 | 0.0168 | 0.0000 | 0.0193 | 0.0170 |
| Odynophagia | 0.0000 | 0.0820 | 0.0802 | 0.0000 | 0.0576 | 0.0316 | 0.0000 | 0.0277 | 0.0318 | 0.0000 | 0.0590 | 0.0380 |
| Prostration | 0.0000 | 0.0043 | 0.0056 | 0.0000 | 0.0013 | 0.0007 | 0.0000 | 0.0087 | 0.0129 | 0.0000 | 0.0078 | 0.0063 |
| Tachypnea | 0.1129 | 0.0061 | 0.0105 | 0.0000 | 0.0245 | 0.0233 | 0.0000 | 0.0376 | 0.0174 | 0.0000 | 0.0089 | 0.0047 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ormeño, P.; Márquez, G.; Guerrero-Nancuante, C.; Taramasco, C. Detection of COVID-19 Patients Using Machine Learning Techniques: A Nationwide Chilean Study. Int. J. Environ. Res. Public Health 2022, 19, 8058. https://doi.org/10.3390/ijerph19138058
Ormeño P, Márquez G, Guerrero-Nancuante C, Taramasco C. Detection of COVID-19 Patients Using Machine Learning Techniques: A Nationwide Chilean Study. International Journal of Environmental Research and Public Health. 2022; 19(13):8058. https://doi.org/10.3390/ijerph19138058
Chicago/Turabian StyleOrmeño, Pablo, Gastón Márquez, Camilo Guerrero-Nancuante, and Carla Taramasco. 2022. "Detection of COVID-19 Patients Using Machine Learning Techniques: A Nationwide Chilean Study" International Journal of Environmental Research and Public Health 19, no. 13: 8058. https://doi.org/10.3390/ijerph19138058
APA StyleOrmeño, P., Márquez, G., Guerrero-Nancuante, C., & Taramasco, C. (2022). Detection of COVID-19 Patients Using Machine Learning Techniques: A Nationwide Chilean Study. International Journal of Environmental Research and Public Health, 19(13), 8058. https://doi.org/10.3390/ijerph19138058








