A Review of Atrial Fibrillation Detection Methods as a Service
Abstract
1. Introduction
2. Background
2.1. Photoplethysmogram
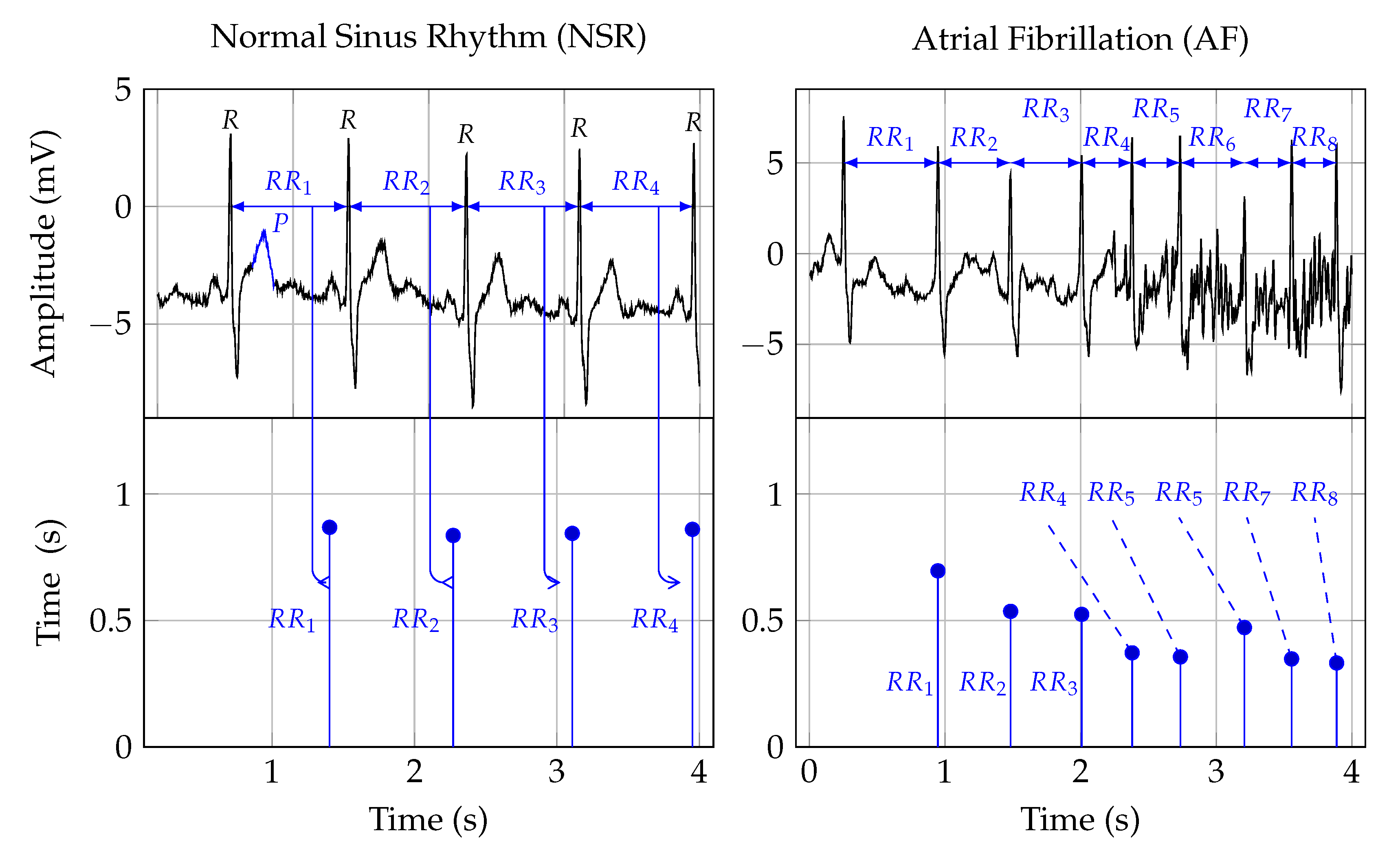
2.2. Electrocardiogram
2.3. Heart Rate
3. Materials
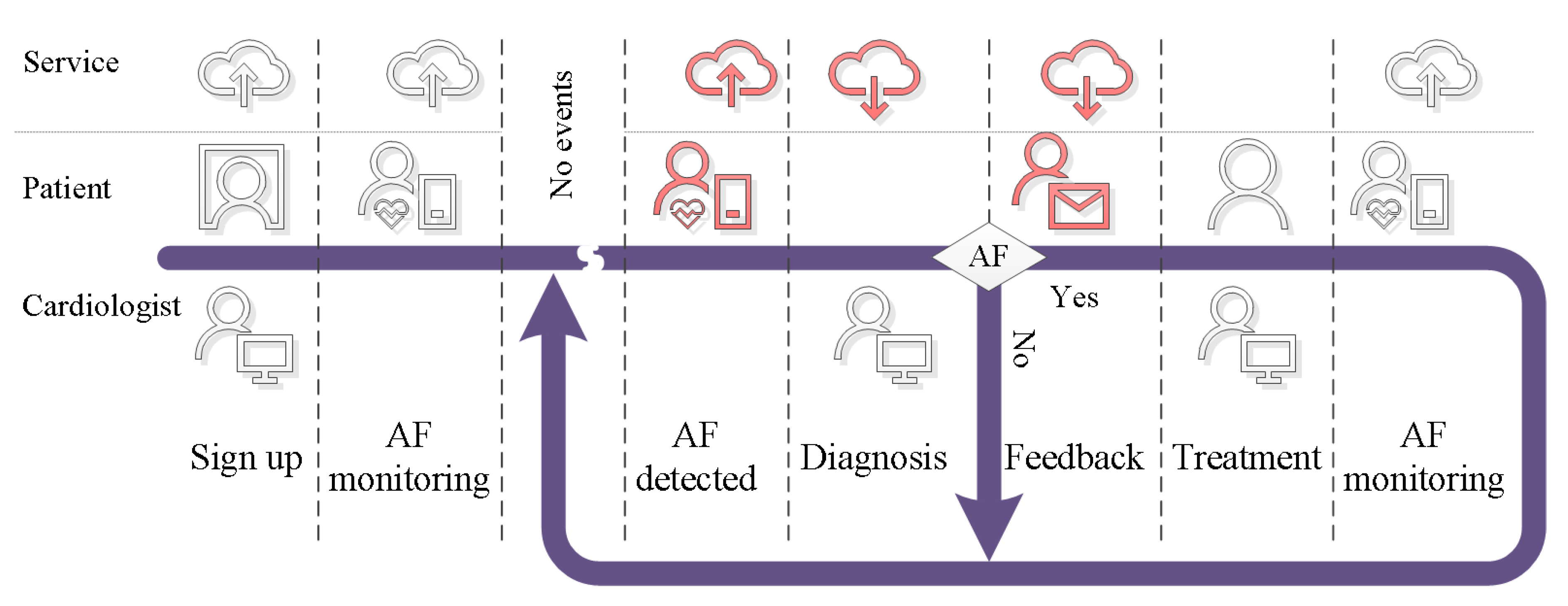
3.1. Service
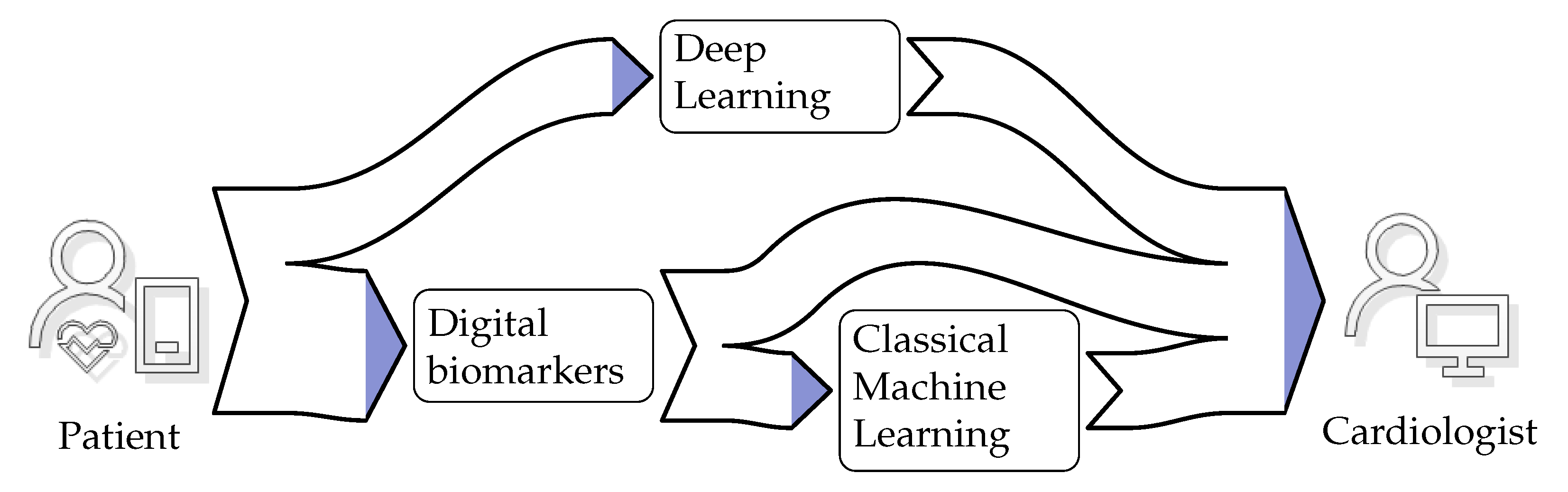
3.2. System Architecture
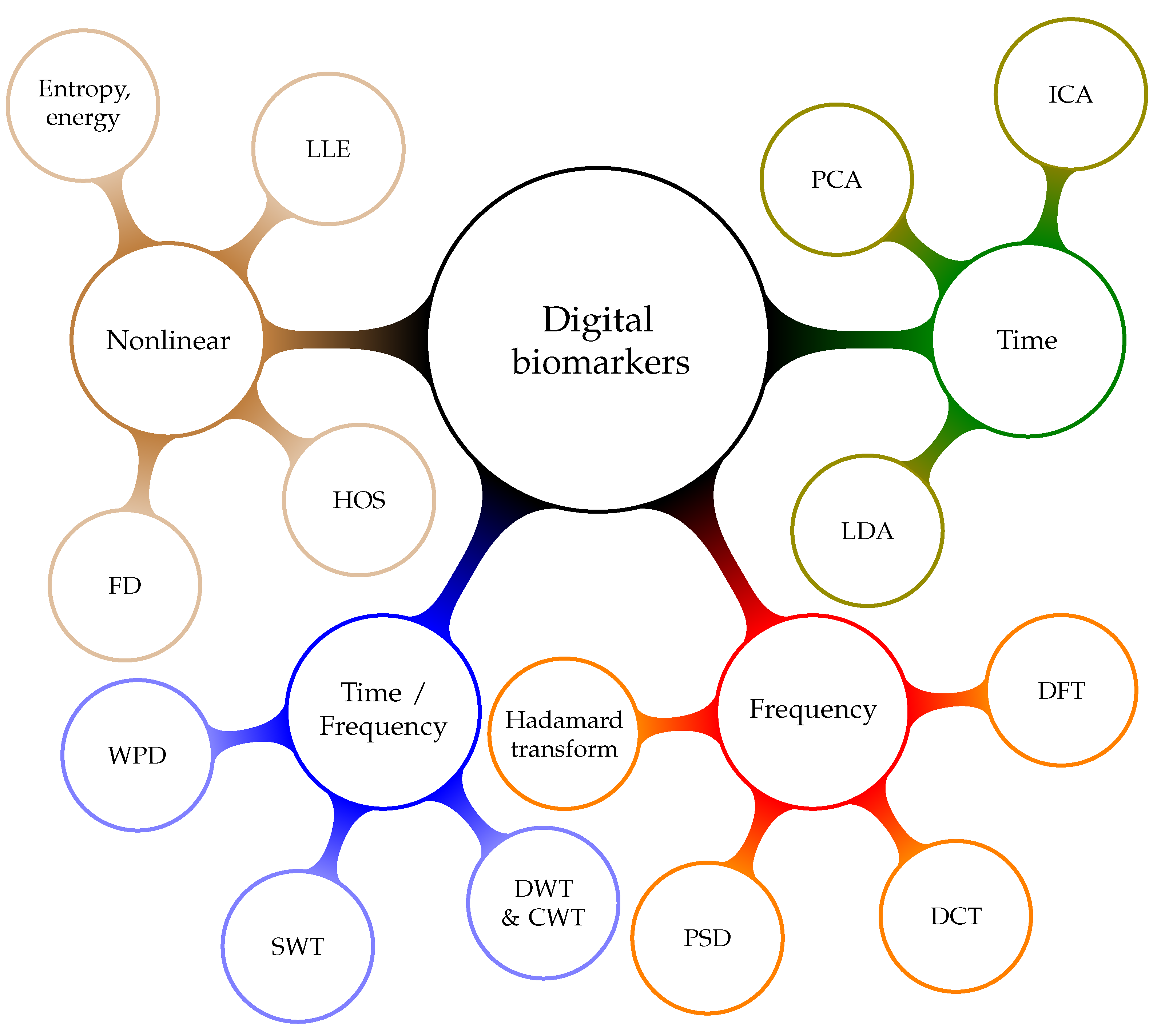
3.2.1. Digital Biomarkers
- Time-domain biomarkers result from algorithms that analyze signals over time. They can be used to model the dynamic behavior of the human heart. These are the most explainable digital biomarkers, because they quantify some of the observations that could be gained through visual inspection of the signal. However, these biomarkers fail to explain the nonlinear characteristics of heart’s oscillation process. Hence, they do not provide all the available information. Examples of time-domain biomarker algorithms are differential equations [102], Principal Component Analysis (PCA) [103,104], the Hadamard transform [105,106], Linear Discriminant Analysis (LDA) [107], and Independent Component Analysis (ICA) [108].
- Heart rhythm is associated with the frequency content of the signal. Frequency-domain digital biomarkers extract measures that indicate the characteristics of the rhythm change. They are especially sensitive to rhythm change caused by AF. Essentially, frequency analysis methods are linear. Hence, frequency-domain digital biomarkers fail to reflect nonlinear properties that might be a relevant source of information. Algorithms that extract these biomarkers are based on the theory of Fourier Transform (FT) and Power Spectral Density (PSD) [109].
- Mixed-domain digital biomarkers are based on the idea of combining the analytical strength of multiple domains [110]. Most practical realizations are based on the fact that the spectrum changes as time progresses [111]. It is possible to track spectral changes over time [112]. Based on the premise that a rhythm disorder will emerge in the spectral domain, time resolution allows us to estimate the time of an AF event [113]. These biomarkers are a great tool for explaining the arrhythmia locations. However, the analysis does not go beyond linear relationships. Examples for mixed domain transformation algorithms are: Discrete Wavelet Transform (DWT) [110], Discrete Cosine Transform (DCT) [114], Continuous Wavelet Transform (CWT) [115], Wavelet Packet Decomposition (WPD) [116], Stationary Wavelet Transform (SWT), Short-Time Fourier Transform (STFT) [117,118], Empirical Mode Decomposition (EMD) [119], and Time-Varying Coherence Function (TVCF) [120].
- Nonlinear digital biomarkers aim to reflect the nonlinear nature of the human heart [121,122]. Surrogate data tests [123] show that nonlinearity cannot be ruled out in ECG and RR interval signals. Performance testing of nonlinear digital biomarkers, with statistical and classification methods, indicates that they are an independent information source. Examples of algorithms that provide parameters for such biomarkers are: Recurrence Quantification Analysis (RQA) [124,125], Higher Order Spectra (HOS) [126], Fractal Dimension (FD) [127], entropy [128], energy [129], and Largest Lyapunov Exponent (LLE) [130].RQA defines a set of digital biomarkers that result from analyzing recurrence plots. Quantifying both the number and duration of recurrences in the physiological signal can help us to understand the phase space trajectory. Changes in the phase space trajectory might indicate the presence of AF [124]. The HOS is a spectral representation of third-order and higher moments and cumulants. It yields nonlinear digital biomarkers that quantify the nonlinear correlations between several frequency components [131,132,133,134]. These measures are sensitive to rhythm changes, which make them useful for arrhythmia detection. FD and LLE are chaos measures of signal complexity [12,132]. Entropy gauges the information content of a signal [135,136,137,138]. In general, signals with less structure, such as RR interval signals, have more information as compared with signals that have more structure, like the ECG. Energy biomarkers quantify the rhythmicity and regularity of signals.
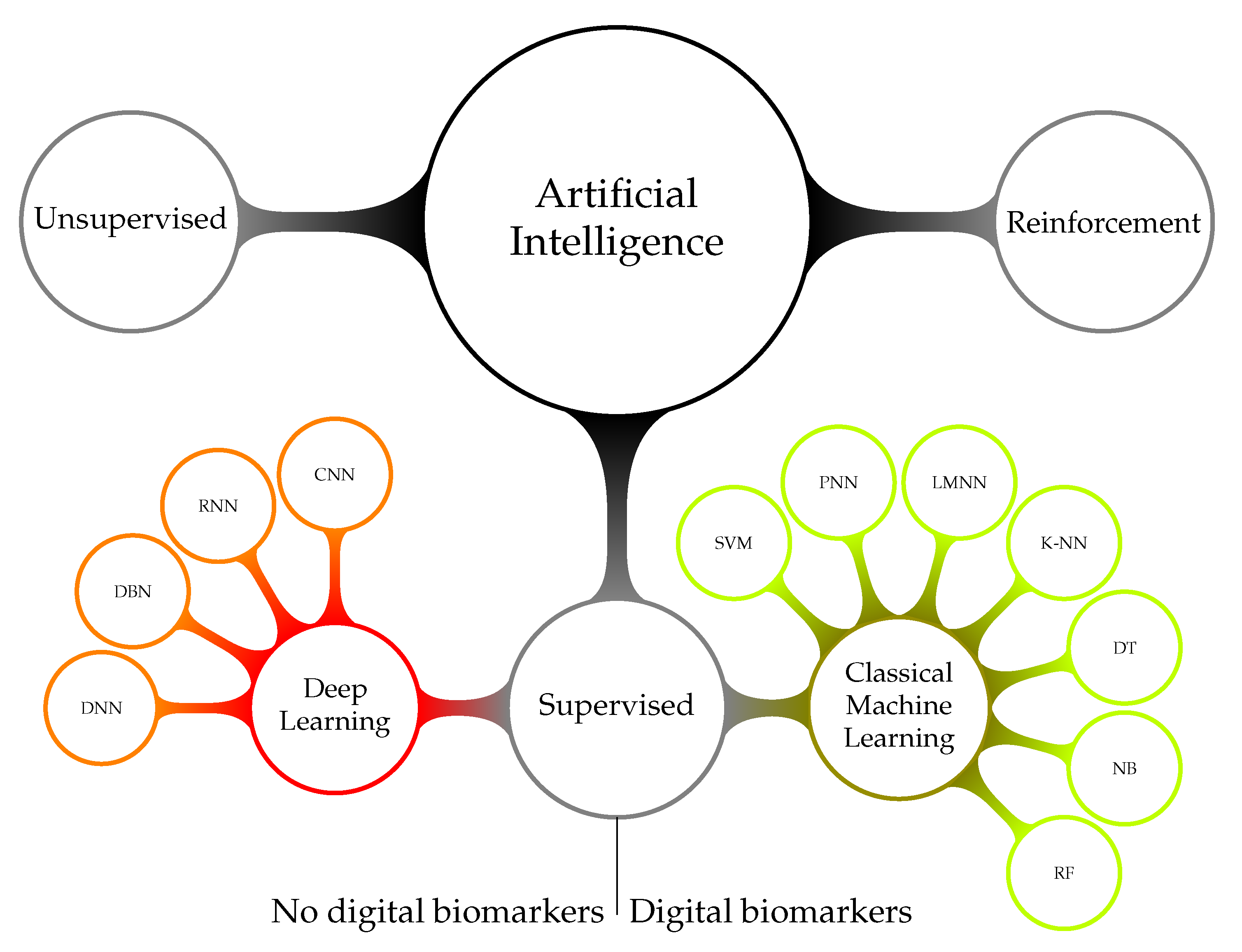
3.2.2. Artificial Intelligence
4. Discussion
- Holter monitors have been fitted with more memory and better energy measurement to extend the measurement time. State-of-the-art devices achieve 72 h ECG recordings in clinical practice. Data analysis and indeed a diagnosis happens after the measurement when the device is returned to the stroke clinic. This process usually incurs a delay of two to three weeks before the diagnosis is communicated to the patient.
- Patient-triggered event recorders are a way to circumvent the energy and storage limitations of Holter monitors. These devices depend on a patient suspecting an AF episode and acting on this suspicion by triggering a measurement. Therefore, such patient-triggered event recorders have a high intra- and inter-observer variability. During a state of reduced consciousness, e.g., sleep, the patient cannot trigger the recorder. The consistency with which the device is used varies from patient to patient, dependent on their mental capability [175]. For example, elderly patients might not be able to interact with the devices to initiate the required measurements [176]. The ideas of patient-triggered event recorders underpin a number of commercial products, such as CardiaMobile from AliveCor (web page (12.04.2020): https://www.alivecor.com/kardiamobile) and Apple Watch (web page (12.04.2020): https://support.apple.com/en-us/HT208955). The services these devices offer are aimed at the patient, and as such, they can only establish a suspicion that AF is present. A diagnosis requires further measurement, in most cases with Holter monitors.
- Cardiac event recorders have replaced patients with algorithm-based triggers, thereby addressing the same energy and storage limitations while reducing intra- and inter-observer variability. The idea is machine algorithms never sleep, and they arrive at the same conclusion for the same input. The diagnosis support quality of these devices is directly related to the sensitivity and specificity of the algorithms used to trigger the recording. These performance measures tend to go up when the computational complexity of the algorithm is increased. Unfortunately, computational complexity is also positively correlated with the energy consumption of the processing platform that executes the algorithm. Hence, these cannot overcome the energy limitation completely. The trade-off between energy consumption and computational complexity of the algorithm has to be resolved during design time. Another limitation of this recording device is the delay between the time when an event was recorded and the diagnosis. During clinical studies, patients wore event triggers, such as the ER910AF Cardiac Event Monitor from Braemar (web page (12.04.2020): http://braemarllc.com/event_products.html), for 30 days [177]. That prolonged observation duration increased the AF detection five-fold.
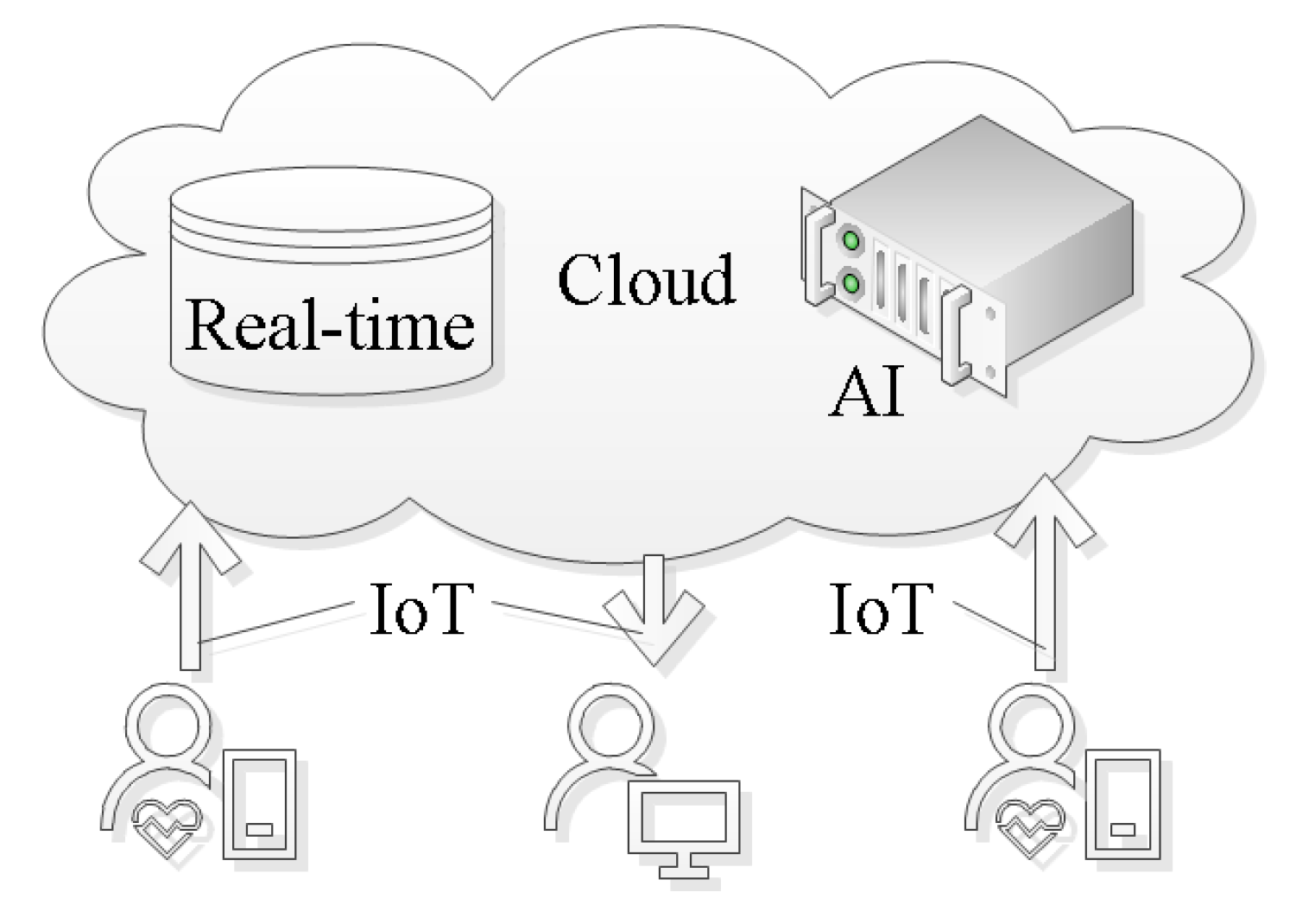
- Mobile cardiovascular telemetry employs state-of-the-art IoT technology to communicate the measurement signals to a central cloud server [178,179]. In that cloud server, the data are stored and processed with an AI system. The results and indeed the measurement data are made available for the reading cardiologist. Figure 6 provides an overview diagram that details the interplay of technologies used for AF detection. Storing the data on a central location is beneficial when it comes to retraining the AI model. One possible way to facilitate the retraining is to use the diagnosis from a cardiologist to label specific segments of the data on the cloud server. This will increase the available knowledge, which can be used to retrain AI models in order to make them more robust.
Open Questions and Future Work
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| Acc | Accuracy |
| AF | Atrial Fibrillation |
| ANN | Artificial Neural Network |
| AI | Artificial Intelligence |
| DL | Deep Learning |
| CNN | Convolutional Neural Network |
| CWT | Continuous Wavelet Transform |
| DBN | Deep Belief Network |
| DCT | Discrete Cosine Transform |
| DFT | Discrete Fourier Transform |
| DL | Deep Learning |
| DNN | Deep Neural Network |
| DT | Decision Tree |
| DWT | Discrete Wavelet Transform |
| ECG | Electrocardiogram |
| EMD | Empirical Mode Decomposition |
| FD | Fractal Dimension |
| FT | Fourier Transform |
| HOS | Higher Order Spectra |
| HR | Heart Rate |
| ICA | Independent Component Analysis |
| IoT | Internet of Things |
| K-NN | K-Nearest Neighbor |
| LDA | Linear Discriminant Analysis |
| LLE | Largest Lyapunov Exponent |
| LMNN | Levenberg–Marquardt Neural Network |
| LSTM | Long Short-Term Memory |
| ML | Machine Learning |
| NB | Naive Bayes |
| NSR | Normal Sinus Rhythm |
| PCA | Principal Component Analysis |
| PNN | Probabilistic Neural Network |
| PPG | Photoplethysmogram |
| PSD | Power Spectral Density |
| RF | Random Forest |
| RNN | Recurrent Neural Network |
| RQA | Recurrence Quantification Analysis |
| SWT | Stationary Wavelet Transform |
| Sen | Sensitivity |
| Spe | Specificity |
| STFT | Short-Time Fourier Transform |
| SWT | Stationary Wavelet Transform |
| SVM | Support Vector Machine |
| TVCF | Time-Varying Coherence Function |
| WHO | World Health Organization |
| WPD | Wavelet Packet Decomposition |
References
- Stergiou, G.; Karpettas, N.; Protogerou, A.; Nasothimiou, E.; Kyriakidis, M. Diagnostic accuracy of a home blood pressure monitor to detect atrial fibrillation. J. Hum. Hypertens. 2009, 23, 654. [Google Scholar] [CrossRef] [PubMed]
- Zoni-Berisso, M.; Lercari, F.; Carazza, T.; Domenicucci, S. Epidemiology of atrial fibrillation: European perspective. Clin. Epidemiol. 2014, 6, 213. [Google Scholar] [CrossRef] [PubMed]
- Fitzmaurice, D.A.; Hobbs, F.R.; Jowett, S.; Mant, J.; Murray, E.T.; Holder, R.; Raftery, J.; Bryan, S.; Davies, M.; Lip, G.Y.; et al. Screening versus routine practice in detection of atrial fibrillation in patients aged 65 or over: Cluster randomised controlled trial. BMJ 2007, 335, 383. [Google Scholar] [CrossRef] [PubMed]
- Cadilhac, D.A. The economics of atrial fibrillation: A time for review and prioritization. Int. J. Stroke 2012, 7, 477–479. [Google Scholar] [CrossRef]
- Hijazi, Z.; Oldgren, J.; Siegbahn, A.; Granger, C.B.; Wallentin, L. Biomarkers in atrial fibrillation: A clinical review. Eur. Heart J. 2013, 34, 1475–1480. [Google Scholar] [CrossRef]
- Heeringa, J.; van der Kuip, D.A.; Hofman, A.; Kors, J.A.; van Herpen, G.; Stricker, B.H.C.; Stijnen, T.; Lip, G.Y.; Witteman, J.C. Prevalence, incidence and lifetime risk of atrial fibrillation: The Rotterdam study. Eur. Heart J. 2006, 27, 949–953. [Google Scholar] [CrossRef]
- Schnabel, R.B.; Yin, X.; Gona, P.; Larson, M.G.; Beiser, A.S.; McManus, D.D.; Newton-Cheh, C.; Lubitz, S.A.; Magnani, J.W.; Ellinor, P.T.; et al. 50 year trends in atrial fibrillation prevalence, incidence, risk factors, and mortality in the Framingham Heart Study: A cohort study. Lancet 2015, 386, 154–162. [Google Scholar] [CrossRef]
- Go, A.S.; Hylek, E.M.; Phillips, K.A.; Chang, Y.; Henault, L.E.; Selby, J.V.; Singer, D.E. Prevalence of diagnosed atrial fibrillation in adults: National implications for rhythm management and stroke prevention: The AnTicoagulation and Risk Factors in Atrial Fibrillation (ATRIA) Study. JAMA 2001, 285, 2370–2375. [Google Scholar] [CrossRef]
- Furberg, C.D.; Psaty, B.M.; Manolio, T.A.; Gardin, J.M.; Smith, V.E.; Rautaharju, P.M.; CHS Collaborative Research Group. Prevalence of atrial fibrillation in elderly subjects (the Cardiovascular Health Study). Am. J. Cardiol. 1994, 74, 236–241. [Google Scholar] [CrossRef]
- Kernan, W.N.; Ovbiagele, B.; Black, H.R.; Bravata, D.M.; Chimowitz, M.I.; Ezekowitz, M.D.; Fang, M.C.; Fisher, M.; Furie, K.L.; Heck, D.V.; et al. Guidelines for the prevention of stroke in patients with stroke and transient ischemic attack: A guideline for healthcare professionals from the American Heart Association/American Stroke Association. Stroke 2014, 45, 2160–2236. [Google Scholar] [CrossRef]
- Wiesel, J.; Arbesfeld, B.; Schechter, D. Comparison of the Microlife blood pressure monitor with the Omron blood pressure monitor for detecting atrial fibrillation. Am. J. Cardiol. 2014, 114, 1046–1048. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Fujita, H.; Adam, M.; Lih, O.S.; Hong, T.J.; Sudarshan, V.K.; Koh, J.E. Automated characterization of arrhythmias using nonlinear features from tachycardia ECG beats. In Proceedings of the 2016 IEEE International Conference on Systems, Man, and Cybernetics (SMC), Budapest, Hungary, 9–12 October 2016; pp. 533–538. [Google Scholar]
- Benjamin, E.J.; Levy, D.; Vaziri, S.M.; D’Agostino, R.B.; Belanger, A.J.; Wolf, P.A. Independent risk factors for atrial fibrillation in a population-based cohort: The Framingham Heart Study. JAMA 1994, 271, 840–844. [Google Scholar] [CrossRef] [PubMed]
- Nieuwlaat, R.; Capucci, A.; Camm, A.J.; Olsson, S.B.; Andresen, D.; Davies, D.W.; Cobbe, S.; Breithardt, G.; Le Heuzey, J.Y.; Prins, M.H.; et al. Atrial fibrillation management: A prospective survey in ESC member countries: The Euro Heart Survey on Atrial Fibrillation. Eur. Heart J. 2005, 26, 2422–2434. [Google Scholar] [CrossRef] [PubMed]
- de Bruijn, R.F.; Heeringa, J.; Wolters, F.J.; Franco, O.H.; Stricker, B.H.; Hofman, A.; Koudstaal, P.J.; Ikram, M.A. Association between atrial fibrillation and dementia in the general population. JAMA Neurol. 2015, 72, 1288–1294. [Google Scholar] [CrossRef]
- Miyasaka, Y.; Barnes, M.E.; Petersen, R.C.; Cha, S.S.; Bailey, K.R.; Gersh, B.J.; Casaclang-Verzosa, G.; Abhayaratna, W.P.; Seward, J.B.; Iwasaka, T.; et al. Risk of dementia in stroke-free patients diagnosed with atrial fibrillation: Data from a community-based cohort. Eur. Heart J. 2007, 28, 1962–1967. [Google Scholar] [CrossRef]
- Soliman, E.Z.; Safford, M.M.; Muntner, P.; Khodneva, Y.; Dawood, F.Z.; Zakai, N.A.; Thacker, E.L.; Judd, S.; Howard, V.J.; Howard, G.; et al. Atrial fibrillation and the risk of myocardial infarction. JAMA Intern. Med. 2014, 174, 107–114. [Google Scholar] [CrossRef]
- Soliman, E.Z.; Lopez, F.; O’Neal, W.T.; Chen, L.Y.; Bengtson, L.; Zhang, Z.M.; Loehr, L.; Cushman, M.; Alonso, A. Atrial fibrillation and risk of ST-segment–elevation versus non–ST-segment–elevation myocardial infarction: The Atherosclerosis Risk in Communities (ARIC) Study. Circulation 2015, 131, 1843–1850. [Google Scholar] [CrossRef]
- Wolf, P.A.; Abbott, R.D.; Kannel, W.B. Atrial fibrillation as an independent risk factor for stroke: The Framingham Study. Stroke 1991, 22, 983–988. [Google Scholar] [CrossRef]
- Gustafsson, C.; Blombäck, M.; Britton, M.; Hamsten, A.; Svensson, J. Coagulation factors and the increased risk of stroke in nonvalvular atrial fibrillation. Stroke 1990, 21, 47–51. [Google Scholar] [CrossRef]
- Benjamin, E.J.; Wolf, P.A.; D’Agostino, R.B.; Silbershatz, H.; Kannel, W.B.; Levy, D. Impact of atrial fibrillation on the risk of death: The Framingham Heart Study. Circulation 1998, 98, 946–952. [Google Scholar] [CrossRef]
- January, C.T.; Wann, L.S.; Calkins, H.; Chen, L.Y.; Cigarroa, J.E.; Cleveland, J.C.; Ellinor, P.T.; Ezekowitz, M.D.; Field, M.E.; Furie, K.L.; et al. 2019 AHA/ACC/HRS focused update of the 2014 AHA/ACC/HRS guideline for the management of patients with atrial fibrillation: A report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines and the Heart Rhythm Society. J. Am. Coll. Cardiol. 2019, 74, 104–132. [Google Scholar] [PubMed]
- Potter, B.J.; Le Lorier, J. Taking the pulse of atrial fibrillation. Lancet 2015, 386, 113–115. [Google Scholar] [CrossRef]
- Dorian, P.; Jung, W.; Newman, D.; Paquette, M.; Wood, K.; Ayers, G.M.; Camm, J.; Akhtar, M.; Luderitz, B. The impairment of health-related quality of life in patients with intermittent atrial fibrillation: Implications for the assessment of investigational therapy. J. Am. Coll. Cardiol. 2000, 36, 1303–1309. [Google Scholar] [CrossRef]
- Steinberg, B.A.; Kim, S.; Fonarow, G.C.; Thomas, L.; Ansell, J.; Kowey, P.R.; Mahaffey, K.W.; Gersh, B.J.; Hylek, E.; Naccarelli, G.; et al. Drivers of hospitalization for patients with atrial fibrillation: Results from the Outcomes Registry for Better Informed Treatment of Atrial Fibrillation (ORBIT-AF). Am. Heart J. 2014, 167, 735–742. [Google Scholar] [CrossRef] [PubMed]
- Healey, J.S.; Oldgren, J.; Ezekowitz, M.; Zhu, J.; Pais, P.; Wang, J.; Commerford, P.; Jansky, P.; Avezum, A.; Sigamani, A.; et al. Occurrence of death and stroke in patients in 47 countries 1 year after presenting with atrial fibrillation: A cohort study. Lancet 2016, 388, 1161–1169. [Google Scholar] [CrossRef]
- Kim, M.H.; Johnston, S.S.; Chu, B.C.; Dalal, M.R.; Schulman, K.L. Estimation of total incremental health care costs in patients with atrial fibrillation in the United States. Circulation 2011, 4, 313–320. [Google Scholar] [CrossRef]
- Stewart, S.; Murphy, N.; Walker, A.; McGuire, A.; McMurray, J. Cost of an emerging epidemic: An economic analysis of atrial fibrillation in the UK. Heart 2004, 90, 286–292. [Google Scholar] [CrossRef]
- Wolowacz, S.; Samuel, M.; Brennan, V.; Jasso-Mosqueda, J.G.; Van Gelder, I. The cost of illness of atrial fibrillation: A systematic review of the recent literature. Europace 2011, 13, 1375–1385. [Google Scholar] [CrossRef]
- Developed with the special contribution of the European Heart Rhythm Association (EHRA); Endorsed by the European Association for Cardio-Thoracic Surgery (EACTS); Authors/Task Force Members; Camm, A.J.; Kirchhof, P.; Lip, G.Y.; Schotten, U.; Savelieva, I.; Ernst, S.; Van Gelder, I.C. Guidelines for the management of atrial fibrillation: The Task Force for the Management of Atrial Fibrillation of the European Society of Cardiology (ESC). Eur. Heart J. 2010, 31, 2369–2429. [Google Scholar] [CrossRef]
- Hendriks, J.M.; Heidbüchel, H. The management of atrial fibrillation: An integrated team approach–insights of the 2016 European Society of Cardiology guidelines for the management of atrial fibrillation for nurses and allied health professionals. Eur. J. Cardiovasc. Nurs. 2019, 18, 88–95. [Google Scholar] [CrossRef]
- Yu, H.C.; Tsai, Y.F.; Chen, M.C.; Yeh, C.H. Underuse of antithrombotic therapy caused high incidence of ischemic stroke in patients with atrial fibrillation. Int. J. Stroke 2012, 7, 112–117. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.J.; Cheng, M.H.; Lee, C.H.; Wung, D.C.; Cheng, C.L.; Yang, Y.H.K. Compliance with antithrombotic prescribing guidelines for patients with atrial fibrillation—A nationwide descriptive study in Taiwan. Clin. Ther. 2008, 30, 1726–1736. [Google Scholar] [CrossRef] [PubMed]
- Harris, K.; Edwards, D.; Mant, J. How can we best detect atrial fibrillation? J. R. Coll. Physicians Edinb. 2012, 42, 5–22. [Google Scholar] [CrossRef] [PubMed]
- Kearley, K.; Selwood, M.; Van den Bruel, A.; Thompson, M.; Mant, D.; Hobbs, F.R.; Fitzmaurice, D.; Heneghan, C. Triage tests for identifying atrial fibrillation in primary care: A diagnostic accuracy study comparing single-lead ECG and modified BP monitors. BMJ Open 2014, 4, e004565. [Google Scholar] [CrossRef] [PubMed]
- Camm, A.J.; Corbucci, G.; Padeletti, L. Usefulness of continuous electrocardiographic monitoring for atrial fibrillation. Am. J. Cardiol. 2012, 110, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Humphries, K.H.; Kerr, C.R.; Connolly, S.J.; Klein, G.; Boone, J.A.; Green, M.; Sheldon, R.; Talajic, M.; Dorian, P.; Newman, D. New-onset atrial fibrillation: Sex differences in presentation, treatment, and outcome. Circulation 2001, 103, 2365–2370. [Google Scholar] [CrossRef]
- Defaye, P.; Dournaux, F.; Mouton, E.; AIDA Multichnter Study Group. Prevalence of supraventricular arrhythmias from the automated analysis of data stored in the DDD pacemakers of 617 patients: The AIDA study. Pacing Clin. Electrophysiol. 1998, 21, 250–255. [Google Scholar] [CrossRef]
- Tsang, T.S.; Petty, G.W.; Barnes, M.E.; O’Fallon, W.M.; Bailey, K.R.; Wiebers, D.O.; Sicks, J.D.; Christianson, T.J.; Seward, J.B.; Gersh, B.J. The prevalence of atrial fibrillation in incident stroke cases and matched population controls in Rochester, Minnesota: Changes over three decades. J. Am. Coll. Cardiol. 2003, 42, 93–100. [Google Scholar] [CrossRef]
- Guo, F.; He, D.; Zhang, W.; Walton, R.G. Trends in prevalence, awareness, management, and control of hypertension among United States adults, 1999 to 2010. J. Am. Coll. Cardiol. 2012, 60, 599–606. [Google Scholar] [CrossRef]
- Lee, J.; Reyes, B.A.; McManus, D.D.; Maitas, O.; Chon, K.H. Atrial fibrillation detection using an iPhone 4S. IEEE Trans. Biomed. Eng. 2012, 60, 203–206. [Google Scholar] [CrossRef]
- Chugh, S.S.; Havmoeller, R.; Narayanan, K.; Singh, D.; Rienstra, M.; Benjamin, E.J.; Gillum, R.F.; Kim, Y.H.; McAnulty, J.H., Jr.; Zheng, Z.J.; et al. Worldwide epidemiology of atrial fibrillation: A Global Burden of Disease 2010 Study. Circulation 2014, 129, 837–847. [Google Scholar] [CrossRef] [PubMed]
- Meister, S.; Deiters, W.; Becker, S. Digital health and digital biomarkers–enabling value chains on health data. Curr. Directions Biomed. Eng. 2016, 2, 577–581. [Google Scholar] [CrossRef]
- Silva, B.M.; Rodrigues, J.J.; de la Torre Díez, I.; López-Coronado, M.; Saleem, K. Mobile-health: A review of current state in 2015. J. Biomed. Inform. 2015, 56, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Meister, S.; Deiters, W. Information logistics solutions to cope with big data challenges in AAL and Telemedicine. In Ambient Assisted Living; Springer: Cham, Switzerland, 2015; pp. 77–88. [Google Scholar]
- World Health Organization. New Horizons for Health through Mobile Technologies; World Health Organization: Geneva, Switzerland, 2011. [Google Scholar]
- Tomlinson, M.; Rotheram-Borus, M.J.; Swartz, L.; Tsai, A.C. Scaling up mHealth: Where is the evidence? PLoS Med. 2013, 10, e1001382. [Google Scholar] [CrossRef]
- Heerden, A.v.; Tomlinson, M.; Swartz, L. Point of care in your pocket: A research agenda for the field of m-health. Bull. World Health Organ. 2012, 90, 393–394. [Google Scholar] [CrossRef]
- Giebel, G.D.; Gissel, C. Accuracy of mHealth devices for atrial fibrillation screening: systematic review. JMIR mHealth uHealth 2019, 7, e13641. [Google Scholar] [CrossRef]
- Dorn, S.D. Digital health: Hope, hype, and Amara’s Law. Gastroenterology 2015, 149, 516–520. [Google Scholar] [CrossRef]
- Scully, C.G.; Lee, J.; Meyer, J.; Gorbach, A.M.; Granquist-Fraser, D.; Mendelson, Y.; Chon, K.H. Physiological parameter monitoring from optical recordings with a mobile phone. IEEE Trans. Biomed. Eng. 2011, 59, 303–306. [Google Scholar] [CrossRef]
- Jonathan, E.; Leahy, M. Investigating a smartphone imaging unit for photoplethysmography. Physiol. Meas. 2010, 31, N79. [Google Scholar] [CrossRef]
- Gregoski, M.J.; Mueller, M.; Vertegel, A.; Shaporev, A.; Jackson, B.B.; Frenzel, R.M.; Sprehn, S.M.; Treiber, F.A. Development and validation of a smartphone heart rate acquisition application for health promotion and wellness telehealth applications. Int. J. Telemed. Appl. 2012, 2012, 696324. [Google Scholar] [CrossRef]
- Grimaldi, D.; Kurylyak, Y.; Lamonaca, F.; Nastro, A. Photoplethysmography detection by smartphone’s videocamera. In Proceedings of the 6th IEEE International Conference on Intelligent Data Acquisition and Advanced Computing Systems, Prague, Czech Republic, 15–17 September 2011; Volume 1, pp. 488–491. [Google Scholar]
- Oh, S.L.; Ng, E.Y.; San Tan, R.; Acharya, U.R. Automated beat-wise arrhythmia diagnosis using modified U-net on extended electrocardiographic recordings with heterogeneous arrhythmia types. Comput. Biol. Med. 2019, 105, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Sanfilippo, A.J.; Abascal, V.M.; Sheehan, M.; Oertel, L.B.; Harrigan, P.; Hughes, R.A.; Weyman, A.E. Atrial enlargement as a consequence of atrial fibrillation. A prospective echocardiographic study. Circulation 1990, 82, 792–797. [Google Scholar] [CrossRef] [PubMed]
- Michałkiewicz, D.; Dziuk, M.; Kamiński, G.; Olszewski, R.; Cholewa, M.; Cwetsch, A.; Markuszewski, L. Detection of patients at risk for paroxysmal atrial fibrillation (PAF) by signal averaged P wave, standard ECG and echocardiography. Polski merkuriusz lekarski: Organ Polskiego Towarzystwa Lekarskiego 2006, 20, 69–72. [Google Scholar]
- Klein, A.L.; Grimm, R.A.; Murray, R.D.; Apperson-Hansen, C.; Asinger, R.W.; Black, I.W.; Davidoff, R.; Erbel, R.; Halperin, J.L.; Orsinelli, D.A.; et al. Use of transesophageal echocardiography to guide cardioversion in patients with atrial fibrillation. N. Engl. J. Med. 2001, 344, 1411–1420. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, C.; Mota, T.; Morais, H.; Matias, F.; Costa, C.; Oliveira, A.G.; Ceia, F.; Investigators, E. The value of the electrocardiogram and chest X-ray for confirming or refuting a suspected diagnosis of heart failure in the community. Eur. J. Heart Fail. 2004, 6, 807–812. [Google Scholar] [CrossRef]
- Thomas, J.T.; Kelly, R.F.; Thomas, S.J.; Stamos, T.D.; Albasha, K.; Parrillo, J.E.; Calvin, J.E. Utility of history, physical examination, electrocardiogram, and chest radiograph for differentiating normal from decreased systolic function in patients with heart failure. Am. J. Med. 2002, 112, 437–445. [Google Scholar] [CrossRef]
- Schnabel, R.B.; Larson, M.G.; Yamamoto, J.F.; Sullivan, L.M.; Pencina, M.J.; Meigs, J.B.; Tofler, G.H.; Selhub, J.; Jacques, P.F.; Wolf, P.A.; et al. Relations of biomarkers of distinct pathophysiological pathways and atrial fibrillation incidence in the community. Circulation 2010, 121, 200. [Google Scholar] [CrossRef]
- Dilaveris, P.E.; Gialafos, E.J.; Sideris, S.K.; Theopistou, A.M.; Andrikopoulos, G.K.; Kyriakidis, M.; Gialafos, J.E.; Toutouzas, P.K. Simple electrocardiographic markers for the prediction of paroxysmal idiopathic atrial fibrillation. Am. Heart J. 1998, 135, 733–738. [Google Scholar] [CrossRef]
- Dash, S.; Chon, K.; Lu, S.; Raeder, E. Automatic real time detection of atrial fibrillation. Ann. Biomed. Eng. 2009, 37, 1701–1709. [Google Scholar] [CrossRef]
- Babaeizadeh, S.; Gregg, R.E.; Helfenbein, E.D.; Lindauer, J.M.; Zhou, S.H. Improvements in atrial fibrillation detection for real-time monitoring. J. Electrocardiol. 2009, 42, 522–526. [Google Scholar] [CrossRef]
- Park, J.; Lee, S.; Jeon, M. Atrial fibrillation detection by heart rate variability in Poincare plot. Biomed. Eng. Online 2009, 8, 38. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.A. Heart rate turbulence: A review. Indian Pacing Electrophysiol. J. 2003, 3, 10. [Google Scholar] [PubMed]
- Yaghouby, F.; Ayatollahi, A.; Bahramali, R.; Yaghouby, M.; Alavi, A.H. Towards automatic detection of atrial fibrillation: A hybrid computational approach. Comput. Biol. Med. 2010, 40, 919–930. [Google Scholar] [CrossRef] [PubMed]
- Lian, J.; Wang, L.; Muessig, D. A simple method to detect atrial fibrillation using RR intervals. Am. J. Cardiol. 2011, 107, 1494–1497. [Google Scholar] [CrossRef] [PubMed]
- Lake, D.E.; Moorman, J.R. Accurate estimation of entropy in very short physiological time series: The problem of atrial fibrillation detection in implanted ventricular devices. Am. J. Physiol.-Heart Circulatory Physiol. 2010, 300, H319–H325. [Google Scholar] [CrossRef]
- Larburu, N.; Lopetegi, T.; Romero, I. Comparative study of algorithms for atrial fibrillation detection. In Proceedings of the 2011 Computing in Cardiology, Hangzhou, China, 18–21 September 2011; pp. 265–268. [Google Scholar]
- Hochstadt, A.; Chorin, E.; Viskin, S.; Schwartz, A.L.; Lubman, N.; Rosso, R. Continuous heart rate monitoring for automatic detection of atrial fibrillation with novel bio-sensing technology. J. Electrocardiol. 2019, 52, 23–27. [Google Scholar] [CrossRef]
- Allen, J. Photoplethysmography and its application in clinical physiological measurement. Physiol. Meas. 2007, 28, R1. [Google Scholar] [CrossRef]
- Bonomi, A.G.; Schipper, F.; Eerikäinen, L.M.; Margarito, J.; Aarts, R.M.; Babaeizadeh, S.; de Morree, H.M.; Dekker, L. Atrial fibrillation detection using photo-plethysmography and acceleration data at the wrist. In Proceedings of the 2016 Computing in Cardiology Conference (CinC), Vancouver, BC, Canada, 11–14 September 2016; pp. 277–280. [Google Scholar]
- Yang, C.; Veiga, C.; Rodríguez-Andina, J.J.; Farina, J.; Iniguez, A.; Yin, S. Using PPG signals and wearable devices for atrial fibrillation screening. IEEE Trans. Ind. Electron. 2019, 66, 8832–8842. [Google Scholar] [CrossRef]
- Nemati, S.; Ghassemi, M.M.; Ambai, V.; Isakadze, N.; Levantsevych, O.; Shah, A.; Clifford, G.D. Monitoring and detecting atrial fibrillation using wearable technology. In Proceedings of the 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Orlando, FL, USA, 16–20 August 2016; pp. 3394–3397. [Google Scholar]
- Sološenko, A.; Petrėnas, A.; Marozas, V.; Sörnmo, L. Modeling of the photoplethysmogram during atrial fibrillation. Comput. Biol. Med. 2017, 81, 130–138. [Google Scholar] [CrossRef]
- Aliamiri, A.; Shen, Y. Deep learning based atrial fibrillation detection using wearable photoplethysmography sensor. In Proceedings of the 2018 IEEE EMBS International Conference on Biomedical & Health Informatics (BHI), Las Vegas, NV, USA, 4–7 March 2018; pp. 442–445. [Google Scholar]
- Sanna, T.; Diener, H.C.; Passman, R.S.; Di Lazzaro, V.; Bernstein, R.A.; Morillo, C.A.; Rymer, M.M.; Thijs, V.; Rogers, T.; Beckers, F.; et al. Cryptogenic stroke and underlying atrial fibrillation. N. Engl. J. Med. 2014, 370, 2478–2486. [Google Scholar] [CrossRef]
- Moody, G.B.; G, M.R. A new method for detecting atrial fibrillation using R-R intervals. Comput. Cardiol. 1983, 10, 227–230. [Google Scholar]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef] [PubMed]
- Budeus, M.; Hennersdorf, M.; Perings, C.; Strauer, B.E. Detection of atrial late potentials with P wave signal averaged electrocardiogram among patients with paroxysmal atrial fibrillation. Zeitschrift fur Kardiologie 2003, 92, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Khorrami, H.; Moavenian, M. A comparative study of DWT, CWT and DCT transformations in ECG arrhythmias classification. Expert Syst. Appl. 2010, 37, 5751–5757. [Google Scholar] [CrossRef]
- Censi, F.; Calcagnini, G.; Mattei, E.; Gargaro, A.; Biancalana, G.; Capucci, A. Simulation of monitoring strategies for atrial arrhythmia detection. Annali dell’Istituto Superiore di Sanita 2013, 49, 176–182. [Google Scholar] [PubMed]
- Capucci, A.; Calcagnini, G.; Mattei, E.; Triventi, M.; Bartolini, P.; Biancalana, G.; Gargaro, A.; Puglisi, A.; Censi, F. Daily distribution of atrial arrhythmic episodes in sick sinus syndrome patients: Implications for atrial arrhythmia monitoring. Europace 2012, 14, 1117–1124. [Google Scholar] [CrossRef] [PubMed]
- Petrutiu, S.; Ng, J.; Nijm, G.M.; Al-Angari, H.; Swiryn, S.; Sahakian, A.V. Atrial fibrillation and waveform characterization. IEEE Eng. Med. Biol. Mag. 2006, 25, 24–30. [Google Scholar] [CrossRef]
- Huang, C.; Ye, S.; Chen, H.; Li, D.; He, F.; Tu, Y. A novel method for detection of the transition between atrial fibrillation and sinus rhythm. IEEE Trans. Biomed. Eng. 2010, 58, 1113–1119. [Google Scholar] [CrossRef]
- Tateno, K.; Glass, L. A method for detection of atrial fibrillation using RR intervals. In Proceedings of the 2000 Annual International IEEE Conference Computers in Cardiology, Cambridge, MA, USA, 24–27 September 2008; pp. 391–394. [Google Scholar]
- Hadhoud, M.M.; Eladawy, M.I.; Farag, A. Computer aided diagnosis of cardiac arrhythmias. In Proceedings of the 2006 International Conference on Computer Engineering and Systems, Cairo, Egypt, 5–7 November 2006; pp. 262–265. [Google Scholar]
- Hagiwara, Y.; Fujita, H.; Oh, S.L.; Tan, J.H.; San Tan, R.; Ciaccio, E.J.; Acharya, U.R. Computer-aided diagnosis of atrial fibrillation based on ECG signals: A review. Inf. Sci. 2018, 467, 99–114. [Google Scholar] [CrossRef]
- Israel, C.W.; Grönefeld, G.; Ehrlich, J.R.; Li, Y.G.; Hohnloser, S.H. Long-term risk of recurrent atrial fibrillation as documented by an implantable monitoring device: Implications for optimal patient care. J. Am. Coll. Cardiol. 2004, 43, 47–52. [Google Scholar] [CrossRef]
- Page, R.L.; Wilkinson, W.E.; Clair, W.K.; McCarthy, E.A.; Pritchett, E. Asymptomatic arrhythmias in patients with symptomatic paroxysmal atrial fibrillation and paroxysmal supraventricular tachycardia. Circulation 1994, 89, 224–227. [Google Scholar] [CrossRef] [PubMed]
- Andresen, D.; Brüggemann, T. Heart rate variability preceding onset of atrial fibrillation. J. Cardiovasc. Electrophysiol. 1998, 9, S26-9. [Google Scholar] [PubMed]
- Murgatroyd, F.D.; Xie, B.; Copie, X.; Blankoff, I.; Camm, A.J.; Malik, M. Identification of Atrial Fibrillation Episodes in Ambulatory Electrocardiographic Recordings: Validation of a Method for Obtaining Labeled R-R Interval Files. Pacing Clin. Electrophysiol. 1995, 18, 1315–1320. [Google Scholar] [CrossRef] [PubMed]
- Stickdorn, M.; Hormess, M.E.; Lawrence, A.; Schneider, J. This Is Service Design Doing: Applying Service Design Thinking in the Real World; O’Reilly Media, Inc.: Boston, MA, USA, 2018. [Google Scholar]
- Healey, J.S.; Connolly, S.J.; Gold, M.R.; Israel, C.W.; Van Gelder, I.C.; Capucci, A.; Lau, C.; Fain, E.; Yang, S.; Bailleul, C.; et al. Subclinical atrial fibrillation and the risk of stroke. N. Engl. J. Med. 2012, 366, 120–129. [Google Scholar] [CrossRef]
- Pahl, C.; Lee, B. Containers and clusters for edge cloud architectures—A technology review. In Proceedings of the 2015 3rd International Conference on Future Internet of Things and Cloud, Rome, Italy, 24–26 August 2015; pp. 379–386. [Google Scholar]
- Faust, O.; Ciaccio, E.J.; Majid, A.; Acharya, U.R. Improving the safety of atrial fibrillation monitoring systems through human verification. Saf. Sci. 2019, 118, 881–886. [Google Scholar] [CrossRef]
- Hlatky, M.A.; Greenland, P.; Arnett, D.K.; Ballantyne, C.M.; Criqui, M.H.; Elkind, M.S.; Go, A.S.; Harrell, F.E., Jr.; Hong, Y.; Howard, B.V.; et al. Criteria for evaluation of novel markers of cardiovascular risk: A scientific statement from the American Heart Association. Circulation 2009, 119, 2408–2416. [Google Scholar] [CrossRef]
- Ridker, P.; Rifai, N. C-reactive protein in the primary prevention of myocardial infarction and stroke. C-Reactive Protein and Cardiovascular Disease; MediEdition: St. Laurent, QC, USA, 2006; pp. 1–23. [Google Scholar]
- Working, G.B.D. Biomarkers and surrogate endpoints: Preferred definitions and conceptual framework. Clin. Pharmacol. Ther. 2001, 69, 89–95. [Google Scholar]
- Acharya, U.R.; Faust, O.; Sree, S.V.; Ghista, D.N.; Dua, S.; Joseph, P.; Ahamed, V.T.; Janarthanan, N.; Tamura, T. An integrated diabetic index using heart rate variability signal features for diagnosis of diabetes. Comput. Methods Biomech. Biomed. Eng. 2013, 16, 222–234. [Google Scholar] [CrossRef]
- Abdul-Kadir, N.A.; Safri, N.M.; Othman, M.A. Dynamic ECG features for atrial fibrillation recognition. Comput. Methods Progr. Biomed. 2016, 136, 143–150. [Google Scholar] [CrossRef]
- Pearson, K. LIII. On lines and planes of closest fit to systems of points in space. Lond. Edinb. Dublin Philos. Mag. J. Sci. 1901, 2, 559–572. [Google Scholar] [CrossRef]
- Faust, O.; Martis, R.J.; Min, L.; Zhong, G.L.Z.; Yu, W. Cardiac arrhythmia classification using electrocardiogram. J. Med. Imaging Health Inf. 2013, 3, 448. [Google Scholar] [CrossRef]
- Manz, J. A sequency-ordered fast Walsh transform. IEEE Trans. Audio Electroacoust. 1972, 20, 204–205. [Google Scholar] [CrossRef]
- Kunt, M. In place computation of the Hadamard transform in Cal-Sal order. Signal Process. 1979, 1, 227–231. [Google Scholar] [CrossRef]
- Fisher, R.A. The use of multiple measurements in taxonomic problems. Ann. Eugen. 1936, 7, 179–188. [Google Scholar] [CrossRef]
- Jutten, C.; Herault, J. Blind separation of sources, part I: An adaptive algorithm based on neuromimetic architecture. Signal Process. 1991, 24, 1–10. [Google Scholar] [CrossRef]
- Bracewell, R.N.; Bracewell, R.N. The Fourier Transform and Its Applications; McGraw-Hill: New York, NY, USA, 1986; Volume 31999. [Google Scholar]
- Addison, P.S. Wavelet transforms and the ECG: A review. Physiol. Meas. 2005, 26, R155. [Google Scholar] [CrossRef]
- Faust, O.; Acharya, R.; Krishnan, S.; Min, L.C. Analysis of cardiac signals using spatial filling index and time-frequency domain. Biomed. Eng. Online 2004, 3, 30. [Google Scholar] [CrossRef]
- Gomes, P.R.; Soares, F.O.; Correia, J.; Lima, C. ECG data-acquisition and classification system by using wavelet-domain hidden markov models. In Proceedings of the 2010 Annual International Conference of the IEEE Engineering in Medicine and Biology, Buenos Aires, Argentina, 31 August–4 September 2010; pp. 4670–4673. [Google Scholar]
- Weng, B.; Wang, J.J.; Michaud, F.; Blanco-Velasco, M. Atrial fibrillation detection using stationary wavelet transform analysis. In Proceedings of the 2008 30th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Vancouver, BC, Canada, 20–25 August 2008; pp. 1128–1131. [Google Scholar]
- Ahmed, N.; Natarajan, T.; Rao, K.R. Discrete cosine transform. IEEE Trans. Comput. 1974, 100, 90–93. [Google Scholar] [CrossRef]
- Unser, M.; Aldroubi, A. A review of wavelets in biomedical applications. Proc. IEEE 1996, 84, 626–638. [Google Scholar] [CrossRef]
- Bradie, B. Wavelet packet-based compression of single lead ECG. IEEE Trans. Biomed. Eng. 1996, 43, 493–501. [Google Scholar] [CrossRef]
- Cohen, L.; Loughlin, P. leid1. Time-frequency analysis: Theory and applications. J. Acoust. Soc. Am. 2013, 134, 4002. [Google Scholar] [CrossRef]
- Nason, G.P.; Silverman, B.W. The stationary wavelet transform and some statistical applications. In Wavelets and Statistics; Springer: New York, NY, USA, 1995; pp. 281–299. [Google Scholar]
- Pal, S.; Mitra, M. Empirical mode decomposition based ECG enhancement and QRS detection. Comput. Biol. Med. 2012, 42, 83–92. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Lu, S.; Zou, R.; Ju, K.; Chon, K.H. Estimation of time-varying coherence function using time-varying transfer functions. Ann. Biomed. Eng. 2005, 33, 1582–1594. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Faust, O.; Sree, V.; Swapna, G.; Martis, R.J.; Kadri, N.A.; Suri, J.S. Linear and nonlinear analysis of normal and CAD-affected heart rate signals. Comput. Methods Progr. Biomed. 2014, 113, 55–68. [Google Scholar] [CrossRef] [PubMed]
- Faust, O.; Bairy, M.G. Nonlinear analysis of physiological signals: A review. J. Mech. Med. Biol. 2012, 12, 1240015. [Google Scholar] [CrossRef]
- Schreiber, T.; Schmitz, A. Improved surrogate data for nonlinearity tests. Phys. Rev. Lett. 1996, 77, 635. [Google Scholar] [CrossRef]
- Eckmann, J.; Kamphorst, S.O.; Ruelle, D. Recurrence plots of dynamical systems. World Sci. Ser. Nonlinear Sci. Ser. A 1995, 16, 441–446. [Google Scholar]
- Acharya, U.R.; Faust, O.; Kadri, N.A.; Suri, J.S.; Yu, W. Automated identification of normal and diabetes heart rate signals using nonlinear measures. Comput. Biol. Med. 2013, 43, 1523–1529. [Google Scholar] [CrossRef]
- Nikias, C.L.; Mendel, J.M. Signal processing with higher-order spectra. IEEE Signal Process. Mag. 1993, 10, 10–37. [Google Scholar] [CrossRef]
- Mandelbrot, B.B. The Fractal Geometry of Nature; WH Freeman: New York, NY, USA, 1983; Volume 173. [Google Scholar]
- Czogała, E.; Łeski, J. Application of entropy and energy measures of fuzziness to processing of ECG signal. Fuzzy Sets Syst. 1998, 97, 9–18. [Google Scholar] [CrossRef]
- Oppenheim, A.V.; Verghese, G.C. Signals, Systems and Inference; Pearson: London, UK, 2015. [Google Scholar]
- Rosenstein, M.T.; Collins, J.J.; De Luca, C.J. A practical method for calculating largest Lyapunov exponents from small data sets. Physica D 1993, 65, 117–134. [Google Scholar] [CrossRef]
- Martis, R.J.; Acharya, U.R.; Adeli, H.; Prasad, H.; Tan, J.H.; Chua, K.C.; Too, C.L.; Yeo, S.W.J.; Tong, L. Computer aided diagnosis of atrial arrhythmia using dimensionality reduction methods on transform domain representation. Biomed. Signal Process. Control 2014, 13, 295–305. [Google Scholar] [CrossRef]
- Martis, R.J.; Acharya, U.R.; Prasad, H.; Chua, C.K.; Lim, C.M.; Suri, J.S. Application of higher order statistics for atrial arrhythmia classification. Biomed. Signal Process. Control 2013, 8, 888–900. [Google Scholar] [CrossRef]
- Prasad, H.; Martis, R.J.; Acharya, U.R.; Min, L.C.; Suri, J.S. Application of higher order spectra for accurate delineation of atrial arrhythmia. In Proceedings of the 2013 35th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Osaka, Japan, 3–7 July 2013; pp. 57–60. [Google Scholar]
- Martis, R.J.; Acharya, U.R.; Lim, C.M.; Mandana, K.; Ray, A.K.; Chakraborty, C. Application of higher order cumulant features for cardiac health diagnosis using ECG signals. Int. J. Neural Syst. 2013, 23, 1350014. [Google Scholar] [CrossRef]
- Asgari, S.; Mehrnia, A.; Moussavi, M. Automatic detection of atrial fibrillation using stationary wavelet transform and support vector machine. Comput. Biol. Med. 2015, 60, 132–142. [Google Scholar] [CrossRef]
- Lee, J.; Nam, Y.; McManus, D.D.; Chon, K.H. Time-varying coherence function for atrial fibrillation detection. IEEE Trans. Biomed. Eng. 2013, 60, 2783–2793. [Google Scholar]
- Martis, R.J.; Acharya, U.R.; Adeli, H. Current methods in electrocardiogram characterization. Comput. Biol. Med. 2014, 48, 133–149. [Google Scholar] [CrossRef]
- Tripathy, R.; Paternina, M.R.A.; Arrieta, J.G.; Pattanaik, P. Automated detection of atrial fibrillation ECG signals using two stage VMD and atrial fibrillation diagnosis index. J. Mech. Med. Biol. 2017, 17, 1740044. [Google Scholar] [CrossRef]
- Bishop, C.M. Pattern Recognition and Machine Learning; Springer Science+Business Media: Berlin, Germany, 2006. [Google Scholar]
- MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proceedings of the Fifth Berkeley Symposium on Mathematical Statistics and Probability; University of California Press: Berkeley, CA, USA, 1967; Volume 1, pp. 281–297. [Google Scholar]
- Wang, Y.; Zhu, Y.S.; Thakor, N.V.; Xu, Y.H. A short-time multifractal approach for arrhythmia detection based on fuzzy neural network. IEEE Trans. Biomed. Eng. 2001, 48, 989–995. [Google Scholar] [CrossRef]
- Bellman, R. A Markovian decision process. J. Math. Mech. 1957, 679–684. [Google Scholar] [CrossRef]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. In Advances in Neural Information Processing Systems; Curran Associates, Inc.: Montreal, QC, Canada, 2014; pp. 2672–2680. [Google Scholar]
- Duda, R.O.; Hart, P.E.; Stork, D.G. Pattern Classification; John Wiley & Sons: Hoboken, NJ, USA, 2012. [Google Scholar]
- Faust, O.; Acharya, U.R.; Tamura, T. Formal design methods for reliable computer-aided diagnosis: A review. IEEE Rev. Biomed. Eng. 2012, 5, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Devroye, L.; Wagner, T. Distribution-free performance bounds for potential function rules. IEEE Trans. Inf. Theory 1979, 25, 601–604. [Google Scholar] [CrossRef]
- Specht, D.F. Probabilistic neural networks. Neural Netw. 1990, 3, 109–118. [Google Scholar] [CrossRef]
- Karatsiolis, S.; Schizas, C.N. Region based Support Vector Machine algorithm for medical diagnosis on Pima Indian Diabetes dataset. In Proceedings of the 2012 IEEE 12th International Conference on Bioinformatics & Bioengineering (BIBE), Larnaca, Cyprus, 11–13 November 2012; pp. 139–144. [Google Scholar]
- El-Naqa, I.; Yang, Y.; Wernick, M.N.; Galatsanos, N.P.; Nishikawa, R.M. A support vector machine approach for detection of microcalcifications. IEEE Trans. Med. Imaging 2002, 21, 1552–1563. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C. Kernel methods: A survey of current techniques. Neurocomputing 2002, 48, 63–84. [Google Scholar] [CrossRef]
- Guyon, I.; Weston, J.; Barnhill, S.; Vapnik, V. Gene selection for cancer classification using support vector machines. Mach. Learn. 2002, 46, 389–422. [Google Scholar] [CrossRef]
- Breiman, L.; Cutler, A. State of the art of data mining using Random forest. In Proceedings of the Salford Data Mining Conference, San Diego, CA, USA, 24–25 May 2012. [Google Scholar]
- Levenberg, K. A method for the solution of certain non-linear problems in least squares. Q. Appl. Math. 1944, 2, 164–168. [Google Scholar] [CrossRef]
- Marquardt, D.W. An algorithm for least-squares estimation of nonlinear parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Cover, T.; Hart, P. Nearest neighbor pattern classification. IEEE Trans. Inf. Theory 1967, 13, 21–27. [Google Scholar] [CrossRef]
- Breiman, L. Classification and Regression Trees; Routledge: Abingdon, UK, 2017. [Google Scholar]
- Sufi, F.; Khalil, I. Diagnosis of cardiovascular abnormalities from compressed ECG: A data mining-based approach. IEEE Trans. Inf. Technol. Biomed. 2010, 15, 33–39. [Google Scholar] [CrossRef]
- Rodriguez, J.J.; Kuncheva, L.I.; Alonso, C.J. Rotation forest: A new classifier ensemble method. IEEE Trans. Pattern Anal. Mach. Intell. 2006, 28, 1619–1630. [Google Scholar] [CrossRef] [PubMed]
- Bengio, Y. Learning deep architectures for AI. Found. Trends® Mach. Learn. 2009, 2, 1–127. [Google Scholar] [CrossRef]
- Poznanski, A.; Wolf, L. Cnn-n-gram for handwriting word recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 2305–2314. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolutional neural networks. In Proceedings of the Advances in Neural Information Processing Systems, Lake Tahoe, NV, USA, 3–6 December 2012; pp. 1097–1105. [Google Scholar]
- Lee, J.G.; Jun, S.; Cho, Y.W.; Lee, H.; Kim, G.B.; Seo, J.B.; Kim, N. Deep learning in medical imaging: General overview. Korean J. Radiol. 2017, 18, 570–584. [Google Scholar] [CrossRef] [PubMed]
- Graves, A.; Mohamed, A.r.; Hinton, G. Speech recognition with deep recurrent neural networks. In Proceedings of the 2013 IEEE International Conference on Acoustics, Speech and Signal Processing, Vancouver, BC, Canada, 26–31 May 2013; pp. 6645–6649. [Google Scholar]
- Faust, O.; Hagiwara, Y.; Hong, T.J.; Lih, O.S.; Acharya, U.R. Deep learning for healthcare applications based on physiological signals: A review. Comput. Methods Progr. Biomed. 2018, 161, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Tolias, G.; Sicre, R.; Jégou, H. Particular object retrieval with integral max-pooling of CNN activations. arXiv 2015, arXiv:1511.05879. [Google Scholar]
- Sainath, T.N.; Vinyals, O.; Senior, A.; Sak, H. Convolutional, long short-term memory, fully connected deep neural networks. In Proceedings of the 2015 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Brisbane, Australia, 19–24 April 2015; pp. 4580–4584. [Google Scholar]
- Mikolov, T.; Karafiát, M.; Burget, L.; Černockỳ, J.; Khudanpur, S. Recurrent neural network based language model. In Proceedings of the Eleventh Annual Conference of the International Speech Communication Association, Makuhari, Japan, 26–30 September 2010. [Google Scholar]
- Kim, Y. Convolutional neural networks for sentence classification. arXiv 2014, arXiv:1408.5882. [Google Scholar]
- Lee, H.; Grosse, R.; Ranganath, R.; Ng, A.Y. Convolutional deep belief networks for scalable unsupervised learning of hierarchical representations. In Proceedings of the 26th Annual International Conference on Machine Learning, Montreal, QC, Canada, 14–18 June 2009; pp. 609–616. [Google Scholar]
- Malhotra, P.; Vig, L.; Shroff, G.; Agarwal, P. Long Short Term Memory Networks for Anomaly Detection in Time Series; Presses Universitaires de Louvain: Leuven, Belgium, 2015; p. 89. [Google Scholar]
- Acharya, U.R.; Faust, O.; Ciaccio, E.J.; Koh, J.E.W.; Oh, S.L.; San Tan, R.; Garan, H. Application of nonlinear methods to discriminate fractionated electrograms in paroxysmal versus persistent atrial fibrillation. Comput. Methods Progr. Biomed. 2019, 175, 163–178. [Google Scholar] [CrossRef]
- Klein, M.; Evans, S.J.L.; Blumberg, S.; Cataldo, L.; Bodenheimer, M.M. Use of P-wave-triggered, P-wave signal-averaged electrocardiogram to predict atrial fibrillation after coronary artery bypass surgery. Am. Heart J. 1995, 129, 895–901. [Google Scholar] [CrossRef]
- Gladstone, D.J.; Dorian, P.; Spring, M.; Panzov, V.; Mamdani, M.; Healey, J.S.; Thorpe, K.E.; Aviv, R.; Boyle, K. Atrial premature beats predict atrial fibrillation in cryptogenic stroke: Results from the EMBRACE trial. Stroke 2015, 46, 936–941. [Google Scholar] [CrossRef]
- Kochhäuser, S.; Dechering, D.G.; Dittrich, R.; Reinke, F.; Ritter, M.A.; Ramtin, S.; Duning, T.; Frommeyer, G.; Eckardt, L. Supraventricular premature beats and short atrial runs predict atrial fibrillation in continuously monitored patients with cryptogenic stroke. Stroke 2014, 45, 884–886. [Google Scholar] [CrossRef]
- Roche, F.; Gaspoz, J.M.; Pichot, V.; Costes, F.; Isaaz, K.; Ferron, C.; Roche, C.; Geyssant, A.; Lacour, J.R.; Barthélémy, J.C. Accuracy of an automatic and patient-triggered long-term solid memory ambulatory cardiac event recorder. Am. J. Cardiol. 1997, 80, 1095–1098. [Google Scholar] [CrossRef]
- Linzer, M.; Pritchett, E.L.; Pontinen, M.; McCarthy, E.; Divine, G.W. Incremental diagnostic yield of loop electrocardiographic recorders in unexplained syncope. Am. J. Cardiol. 1990, 66, 214–219. [Google Scholar] [CrossRef]
- Gladstone, D.J.; Spring, M.; Dorian, P.; Panzov, V.; Thorpe, K.E.; Hall, J.; Vaid, H.; O’Donnell, M.; Laupacis, A.; Côté, R.; et al. Atrial fibrillation in patients with cryptogenic stroke. N. Engl. J. Med. 2014, 370, 2467–2477. [Google Scholar] [CrossRef] [PubMed]
- Faust, O.; Shetty, R.; Sree, S.V.; Acharya, S.; Acharya, R.; Ng, E.; Poo, C.K.; Suri, J. Towards the systematic development of medical networking technology. J. Med. Syst. 2011, 35, 1431–1445. [Google Scholar] [CrossRef] [PubMed]
- Faust, O.; Sputh, B.H.; Acharya, U.; Allen, A.R. A pervasive design strategy for distributed health care systems. Open Med. Imaging J. 2008, 2, 58–69. [Google Scholar] [CrossRef]
- Yuan, C.; Yan, Y.; Zhou, L.; Bai, J.; Wang, L. Automated atrial fibrillation detection based on deep learning network. In Proceedings of the 2016 IEEE International Conference on Information and Automation (ICIA), Ningbo, China, 1–3 August 2016; pp. 1159–1164. [Google Scholar]
- Ivanovic, M.D.; Atanasoski, V.; Shvilkin, A.; Hadzievski, L.; Maluckov, A. Deep Learning Approach for Highly Specific Atrial Fibrillation and Flutter Detection based on RR Intervals. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; pp. 1780–1783. [Google Scholar]
- Faust, O.; Shenfield, A.; Kareem, M.; San, T.R.; Fujita, H.; Acharya, U.R. Automated detection of atrial fibrillation using long short-term memory network with RR interval signals. Comput. Biol. Med. 2018, 102, 327–335. [Google Scholar] [CrossRef]
- Wang, J.; Wang, P.; Wang, S. Automated detection of atrial fibrillation in ECG signals based on wavelet packet transform and correlation function of random process. Biomed. Signal Process. Control 2020, 55, 101662. [Google Scholar] [CrossRef]
- Cao, P.; Li, X.; Mao, K.; Lu, F.; Ning, G.; Fang, L.; Pan, Q. A novel data augmentation method to enhance deep neural networks for detection of atrial fibrillation. Biomed. Signal Process. Control 2020, 56, 101675. [Google Scholar] [CrossRef]
- Ia, M.; Biasiolli, L.; Masè, M.; Adami, A.; Ao, A.; Ravelli, F.; Nollo, G. Implementation and validation of real-time algorithms for atrial fibrillation detection on a wearable ECG device. Comput. Biol. Med. 2019, 116, 103540. [Google Scholar]
- Cao, X.C.; Yao, B.; Chen, B.Q. Atrial fibrillation detection using an improved multi-Scale decomposition enhanced residual convolutional neural network. IEEE Access 2019, 7, 89152–89161. [Google Scholar] [CrossRef]
- Liu, C.; Oster, J.; Reinertsen, E.; Li, Q.; Zhao, L.; Nemati, S.; Clifford, G.D. A comparison of entropy approaches for AF discrimination. Physiol. Meas. 2018, 39, 074002. [Google Scholar] [CrossRef]
- Xia, Y.; Wulan, N.; Wang, K.; Zhang, H. Detecting atrial fibrillation by deep convolutional neural networks. Comput. Biol. Med. 2018, 93, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Kora, P.; Annavarapu, A.; Yadlapalli, P.; Krishna, K.S.R.; Somalaraju, V. ECG based atrial fibrillation detection using sequency ordered complex Hadamard transform and hybrid firefly algorithm. Eng. Sci. Technol. Int. J. 2017, 20, 1084–1091. [Google Scholar] [CrossRef]
- Annavarapu, A.; Kora, P. ECG-based atrial fibrillation detection using different orderings of Conjugate Symmetric–Complex Hadamard Transform. Int. J. Cardiovasc. Acad. 2016, 2, 151–154. [Google Scholar] [CrossRef]
- Daqrouq, K.; Alkhateeb, A.; Ajour, M.N.; Morfeq, A. Neural network and wavelet average framing percentage energy for atrial fibrillation classification. Comput. Methods Progr. Biomed. 2014, 113, 919–926. [Google Scholar] [CrossRef]
- Martis, R.J.; Acharya, U.R.; Prasad, H.; Chua, C.K.; Lim, C.M. Automated detection of atrial fibrillation using Bayesian paradigm. Knowl. Based Syst. 2013, 54, 269–275. [Google Scholar] [CrossRef]
- Maji, U.; Mitra, M.; Pal, S. Automatic detection of atrial fibrillation using empirical mode decomposition and statistical approach. Procedia Technol. 2013, 10, 45–52. [Google Scholar] [CrossRef]
- Rincón, F.; Grassi, P.R.; Khaled, N.; Atienza, D.; Sciuto, D. Automated real-time atrial fibrillation detection on a wearable wireless sensor platform. In Proceedings of the 2012 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, San Diego, CA, USA, 28 August–1 September 2012; pp. 2472–2475. [Google Scholar]
- Fukunami, M.; Yamada, T.; Ohmori, M.; Kumagai, K.; Umemoto, K.; Sakai, A.; Kondoh, N.; Minamino, T.; Hoki, N. Detection of patients at risk for paroxysmal atrial fibrillation during sinus rhythm by P wave-triggered signal-averaged electrocardiogram. Circulation 1991, 83, 162–169. [Google Scholar] [CrossRef]
- Parvaresh, S.; Ayatollahi, A. Automatic atrial fibrillation detection using autoregressive modeling. In Proceedings of the 2011 International Conference on Biomedical Engineering and Technology, APCBEES, Kuala Lumpur, Malaysia, 4–5 June 2011; Volume 4. [Google Scholar]
- Andersen, R.S.; Peimankar, A.; Puthusserypady, S. A deep learning approach for real-time detection of atrial fibrillation. Expert Syst. Appl. 2019, 115, 465–473. [Google Scholar] [CrossRef]
- Henzel, N.; Wróbel, J.; Horoba, K. Atrial fibrillation episodes detection based on classification of heart rate derived features. In Proceedings of the 2017 MIXDES-24th International Conference Mixed Design of Integrated Circuits and Systems, Bydgoszcz, Poland, 22–24 June 2017; pp. 571–576. [Google Scholar]
- Cui, X.; Chang, E.; Yang, W.H.; Jiang, B.; Yang, A.; Peng, C.K. Automated detection of paroxysmal atrial fibrillation using an information-based similarity approach. Entropy 2017, 19, 677. [Google Scholar] [CrossRef]
- Islam, M.S.; Ammour, N.; Alajlan, N.; Aboalsamh, H. Rhythm-based heartbeat duration normalization for atrial fibrillation detection. Comput. Biol. Med. 2016, 72, 160–169. [Google Scholar] [CrossRef] [PubMed]
- García, M.; Ródenas, J.; Alcaraz, R.; Rieta, J.J. Application of the relative wavelet energy to heart rate independent detection of atrial fibrillation. Comput. Methods Progr. Biomed. 2016, 131, 157–168. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, A.; Finlay, D.D.; Guldenring, D.; Bond, R.R.; Moran, K.; McLaughlin, J. Automated detection of atrial fibrillation using RR intervals and multivariate-based classification. J. Electrocardiol. 2016, 49, 871–876. [Google Scholar] [CrossRef] [PubMed]
- Petrėnas, A.; Marozas, V.; Sörnmo, L. Low-complexity detection of atrial fibrillation in continuous long-term monitoring. Comput. Biol. Med. 2015, 65, 184–191. [Google Scholar] [CrossRef] [PubMed]
- Andersson, O.; Chon, K.H.; Sörnmo, L.; Rodrigues, J.N. A 290 mV Sub-VT ASIC for Real-Time Atrial Fibrillation Detection. IEEE Trans. Biomed. Circuits Syst. 2014, 9, 377–386. [Google Scholar] [CrossRef]
- Zhou, X.; Ding, H.; Ung, B.; Pickwell-MacPherson, E.; Zhang, Y. Automatic online detection of atrial fibrillation based on symbolic dynamics and Shannon entropy. Biomed. Eng. Online 2014, 13, 18. [Google Scholar] [CrossRef]
- Mohebbi, M.; Ghassemian, H. Prediction of paroxysmal atrial fibrillation using recurrence plot-based features of the RR-interval signal. Physiol. Meas. 2011, 32, 1147. [Google Scholar] [CrossRef]
- Ghodrati, A.; Marinello, S. Statistical analysis of RR interval irregularities for detection of atrial fibrillation. In Proceedings of the 2008 Annual International IEEE Conference Computers in Cardiology, Bologna, Italy, 14–17 September 2008; pp. 1057–1060. [Google Scholar]
- Ghodrati, A.; Murray, B.; Marinello, S. RR interval analysis for detection of atrial fibrillation in ECG monitors. In Proceedings of the 2008 30th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Vancouver, BC, Canada, 20–25 August 2008; pp. 601–604. [Google Scholar]
- Kikillus, N.; Hammer, G.; Lentz, N.; Stockwald, F.; Bolz, A. Three different algorithms for identifying patients suffering from atrial fibrillation during atrial fibrillation free phases of the ECG. In Proceedings of the 2007 Annual International IEEE Conference Computers in Cardiology, Durham, NC, USA, 30 September–3 October 2007; pp. 801–804. [Google Scholar]
- Logan, B.; Healey, J. Robust detection of atrial fibrillation for a long term telemonitoring system. In Proceedings of the 2005 Annual International IEEE Conference Computers in Cardiology, Lyon, France, 25–28 September 2005; pp. 619–622. [Google Scholar]
- Tateno, K.; Glass, L. Automatic detection of atrial fibrillation using the coefficient of variation and density histograms of RR and ΔRR intervals. Med. Biol. Eng. Comput. 2001, 39, 664–671. [Google Scholar] [CrossRef]
- Artis, S.G.; Mark, R.G.; Moody, G.B. Detection of atrial fibrillation using artificial neural networks. In Proceedings of the 1991 Annual International IEEE Conference Computers in Cardiology, Venice, Italy, 23–26 September 1991; pp. 173–176. [Google Scholar] [CrossRef]
- Balagopal, P.; de Ferranti, S.D.; Cook, S.; Daniels, S.R.; Gidding, S.S.; Hayman, L.L.; McCrindle, B.W.; Mietus-Snyder, M.L.; Steinberger, J. Nontraditional risk factors and biomarkers for cardiovascular disease: Mechanistic, research, and clinical considerations for youth: A scientific statement from the American Heart Association. Circulation 2011, 123, 2749–2769. [Google Scholar] [CrossRef]
- Beier, K.; Schweda, M.; Schicktanz, S. Taking patient involvement seriously: A critical ethical analysis of participatory approaches in data-intensive medical research. BMC Med. Inform. Decis. Mak. 2019, 19, 90. [Google Scholar] [CrossRef]
- Becker, S.; Miron-Shatz, T.; Schumacher, N.; Krocza, J.; Diamantidis, C.; Albrecht, U.V. mHealth 2.0: Experiences, possibilities, and perspectives. JMIR mHealth uHealth 2014, 2, e24. [Google Scholar] [CrossRef] [PubMed]
- Sriram, J.; Shin, M.; Kotz, D.; Rajan, A.; Sastry, M.; Yarvis, M. Challenges in data quality assurance in pervasive health monitoring systems. In Future of Trust in Computing; Springer: New York, NY, USA, 2009; pp. 129–142. [Google Scholar]
- Yıldırım, Ö.; Pławiak, P.; Tan, R.S.; Acharya, U.R. Arrhythmia detection using deep convolutional neural network with long duration ECG signals. Comput. Biol. Med. 2018, 102, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Yildirim, O.; Baloglu, U.B.; Tan, R.S.; Ciaccio, E.J.; Acharya, U.R. A new approach for arrhythmia classification using deep coded features and LSTM networks. Comput. Methods Progr. Biomed. 2019, 176, 121–133. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.; Das, M.K.; Ari, S. Automatic ECG arrhythmia classification using dual tree complex wavelet based features. AEU-Int. J. Electron. Commun. 2015, 69, 715–721. [Google Scholar] [CrossRef]
| Authors | Data | Digital Biomarkers | AI | Performance in % | Salient Features | ||
|---|---|---|---|---|---|---|---|
| Acc | Sen | Spe | |||||
| Wang et al., 2020 [183] | MIT-BIHAFDB | WPD followed by multivariate statistical features | ANN | 98.8 | 98.7 | 98.9 |
|
| Cao et al., 2020 [184] | CinCchallenge 2017 | Data augmentation | DNN | 78.35 | - | - |
|
| Marsili et al., 2019 [185] | MIT-BIH AFDB and measurements | Shannon entropy | Threshold | 98.1 | 99.2 | 97.3 |
|
| Yao et al., 2019 [186] | CinC challenge 2017 | DWT | Multi-scale CNN | 98.18 | 98.22 | 98.11 |
|
| Lui et al., 2018 [187] | MIT-BIH AFDB/NSR/Arrhythmia Database | Normalized fuzzy entropy | Threshold | - | - | - |
|
| Xia et al., 2018 [188] | MIT-BIH AFIB | STFT, SWT | CNN | 98.63 | 98.79 | 97.87 |
|
| Kora et al., 2017 [189] | MIT-BIH AFIB | CS-SCHT | LMNN | 99.30 | 96.97 | 99.43 |
|
| Tripathy et al., 2017 [138] | MIT-BIH AFIB | Sample entropy, VMD | DBN | 98.27 | 98.80 | 97.77 |
|
| Annavarapu and Padmavathi, 2016 [190] | MIT-BIH Arrhythmia Database | CS-SCHT | LMNN | 99.50 | 99.97 | 98.70 |
|
| Abdul-Kadir et al., 2016 [102] | MIT-BIH NSR/AFDB | Dynamic system | ANN, SVM | 95.00 |
| ||
| Yuan et al., 2016 [180] | MIT-BIH NSR/AFDB/LTAFDB | - | Autoencoder DL | 98.31 | 96.56 | 99.04 |
|
| Asgari et al., 2015 [135] | MIT-BIH AFDB | SWT, Log-energy entropy, peak-to-average power ratio | SVM | 97.10 | 97.00 | 97.10 |
|
| Daqrouq et al., 2014 [191] | MIT-BIH AFIB | WPD | PNN | 97.92 | - | - |
|
| Martis et al., 2013 [192] | MIT-BIH AFDB/Arrhythmia Database | DWT | NB | 99.33 | 99.32 | 99.33 |
|
| Majia et al., 2013 [193] | MIT-BIH Arrhythmia Database | HOS and EMD | Thresholded | - | 96 | - |
|
| Rincón et al., 2012 [194] | MIT-BIH AFIB | Statistical measures | Fuzzy classifier | - | 98.09 | 91.66 |
|
| Lee et al., 2012 [41] | MIT-BIH NSR/AFDB | RMSSD, sample entropy, Shannon entropy | Threshold | 98.44 | 97.63 | 99.61 |
|
| Fukunami et al., 1991 [195] | Measurement data | Frequency-domain | Statistical analysis | - | 91 | 76 |
|
| Parvaresh and Ayatollahi, 2011 [196] | MIT-BIH AFIB | Autoregressive model | Statistical classifier | - | 96.14 | 93.20 |
|
| Authors | Data | Digital Biomarkers | AI | Performance in % | Salient Features | ||
|---|---|---|---|---|---|---|---|
| Acc | Sen | Spe | |||||
| Ivanovic et al., 2019 [181] | Measurement data | - | DL | 89.67 | 94.20 | - |
|
| Anderson et al., 2019 [197] | MIT-BIH NSR/AFDB/Arrhythmia Database | - | CNN and RNN | - | 98.98% | 96.95% |
|
| Faust et al., 2018 [182] | MIT-BIH AFDB | - | LSTM | 98.51 | 98.32 | 98.67 |
|
| Henzel et al., 2017 [198] | MIT-BIH AFDB | Linear measures | Threshold | 93 | 90 | 95 |
|
| Cui et al., 2017 [199] | MIT-BIH NSR/AFDB | Ensemble model | Threshold | 97.78 | 97.04 | 97.96 |
|
| Islam et al., 2016 [200] | MIT-BIH AFDB | Entropy | Threshold | 96.38 | 96.39 | 96.38 |
|
| García et al., 2016 [201] | MIT-BIH AFDB/Arrhythmia Database | RWE, SWT | Threshold | 93.32 | 91.21 | 94.53 |
|
| Kennedy et al., 2016 [202] | SPIT1, THEW2, MIT-BIH Arrhythmia Database/AFDB/ LTAFDB/NSR/SVA | Sample entropy, CV, RMSSD, MAD | RF | - | 92.80 | 98.30 |
|
| Petrėnas et al., 2015 [203] | MIT-BIH NSR/AFDB | Linear measures | Threshold | - | 97.1 | 98.3 |
|
| Andersson et al., 2014 [204] | MIT-BIH Arrhythmia Database/AFDB | Statistical measures | Threshold | - | 94.6 | 95.8 |
|
| Zhou et al., 2014 [205] | MIT-BIH Arrhythmia Database/AFDB/NSR | Shannon entropy | Threshold | 96.05 | 96.72 | 95.07 |
|
| Lee et al., 2013 [136] | MIT-BIH Arrhythmia Database/NSR | TVCF, Shannon entropy | Threshold | - | 98.20 | 97.70 |
|
| Lake and Moorman 2011 [69] | MIT-BIH AFDB | Sample entropy | Threshold | - | 91 | 94 |
|
| Lian et al., 2011 [68] | MIT-BIH Arrhythmia Database/AFDB/ NSR/NSR2 | Statistical measures | Threshold | - | 95.9 | - |
|
| Lake and Moorman 2011 [69] | MIT-BIH AFDB | Sample entropy | Threshold | - | 91 | 94 |
|
| Mohebbi et al., 2011 [206] | MIT-BIH AFDB | RQA, recurrence plot | SVM | - | 97.00 | 100.00 |
|
| Yaghouby et al., 2010 [67] | MIT-BIH Arrhythmia Database | Statistical and geometrical | Genetic algorithm | 99.11 | - | - |
|
| Huang et al., 2010 [86] | MIT-BIH AFDB/NSR | Linear | Threshold | - | 96.10 | 98.10 |
|
| Dash et al., 2009 [63] | MIT-BIH AFDB | TPR, RMSSD, Shannon entropy | Threshold | - | 90.20 - 94.40 | 91.20 - 95.10 |
|
| Babaeizadeh et al., 2009 [64] | MIT-BIH AFDB | Statistical measures | DT | - | 98 | - |
|
| Ghodrati and Marinello, 2008 [207] | MIT-BIH Arrhythmia Database | Probability density function | Threshold | - | 92 | - |
|
| Ghodrati et al., 2008 [208] | Measurement data, MIT-BIH Arrhythmia Database/AFDB | Normalized absolute deviation and normalized absolute difference | Threshold | - | 89 | - |
|
| Kikillus et al., 2007 [209] | MIT-BIH NSR/AFDB | Linear, Frequency and RQA | Threshold | - | 94.1 | 93.4 |
|
| Logan and Healey, 2005 [210] | MIT-BIH AFDB | Statistical measures | Threshold | - | 94.4 | 97.2 |
|
| Tateno and Glass, 2001 [211] | MIT-BIH AFDB | CV & Kolmogorov–Smirnov test | Threshold | - | 91.20 | 96.08 |
|
| Artis et al., 1991 [212] | MIT-BIH AFDB | Statistical measures | ANN | - | 92.86 | - |
|
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Faust, O.; Ciaccio, E.J.; Acharya, U.R. A Review of Atrial Fibrillation Detection Methods as a Service. Int. J. Environ. Res. Public Health 2020, 17, 3093. https://doi.org/10.3390/ijerph17093093
Faust O, Ciaccio EJ, Acharya UR. A Review of Atrial Fibrillation Detection Methods as a Service. International Journal of Environmental Research and Public Health. 2020; 17(9):3093. https://doi.org/10.3390/ijerph17093093
Chicago/Turabian StyleFaust, Oliver, Edward J. Ciaccio, and U. Rajendra Acharya. 2020. "A Review of Atrial Fibrillation Detection Methods as a Service" International Journal of Environmental Research and Public Health 17, no. 9: 3093. https://doi.org/10.3390/ijerph17093093
APA StyleFaust, O., Ciaccio, E. J., & Acharya, U. R. (2020). A Review of Atrial Fibrillation Detection Methods as a Service. International Journal of Environmental Research and Public Health, 17(9), 3093. https://doi.org/10.3390/ijerph17093093







