A Double-Activity (Green Algae Toxicity and Bacterial Genotoxicity) 3D-QSAR Model Based on the Comprehensive Index Method and Its Application in Fluoroquinolones’ Modification
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Sources of Quinolones’ Toxicity
2.2. Calculation of Double-Activity Characterization Values of Green Algae Toxicity and Bacterial Genotoxicity on the Basis of the Comprehensive Index Method
2.3. Construction Method of the CoMSIA Model of the Double-Activity Quinolones’ Green Algae Toxicity and Bacterial Genotoxicity
2.4. Environmental Stability Evaluation Method of FQ Derivatives on the Basis of Gaussian Calculations
2.5. Environmentally Friendly Evaluation Method for FQ Derivatives on the Basis of EPI
2.6. Mechanism Analysis of Green Algae Toxicity and Bacterial Genotoxicity Based on Molecular Docking
3. Results and Discussion
3.1. Construction and Evaluation of the CoMSIA Model of Double-Activity Quinolones’ Green Algae Toxicity and Bacterial Genotoxicity
3.1.1. Calculation of Double-Activity Characterization Values of Quinolones’ Green Algae Toxicity and Bacterial Genotoxicity
3.1.2. Construction of the CoMSIA Model of the Double-Activity Quinolones’ Green Algae Toxicity and Bacterial Genotoxicity
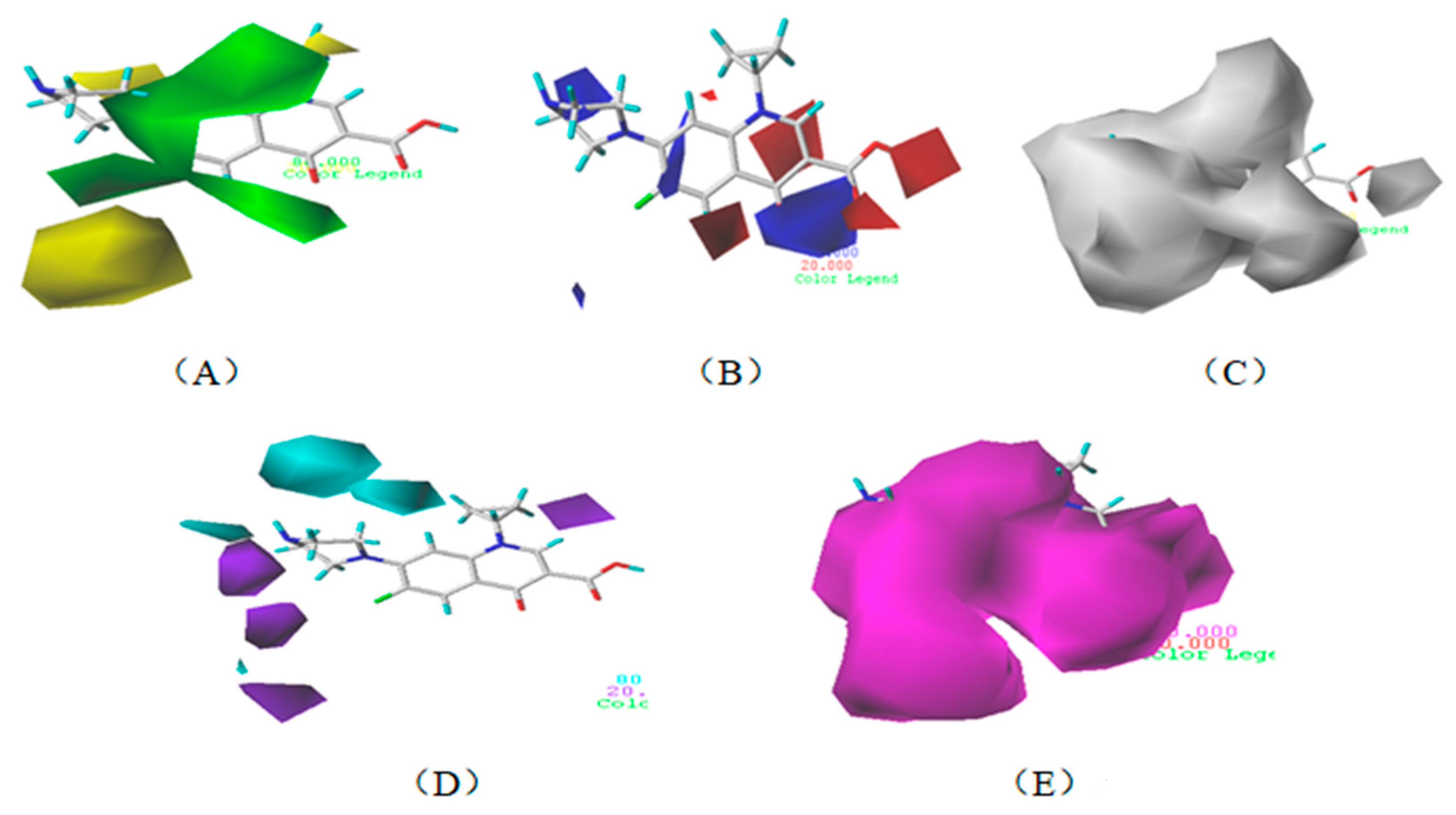
3.1.3. Contour Map Analysis of the CoMSIA Model for Double-Activity Quinolones’ Green Algae Toxicity and Bacterial Genotoxicity
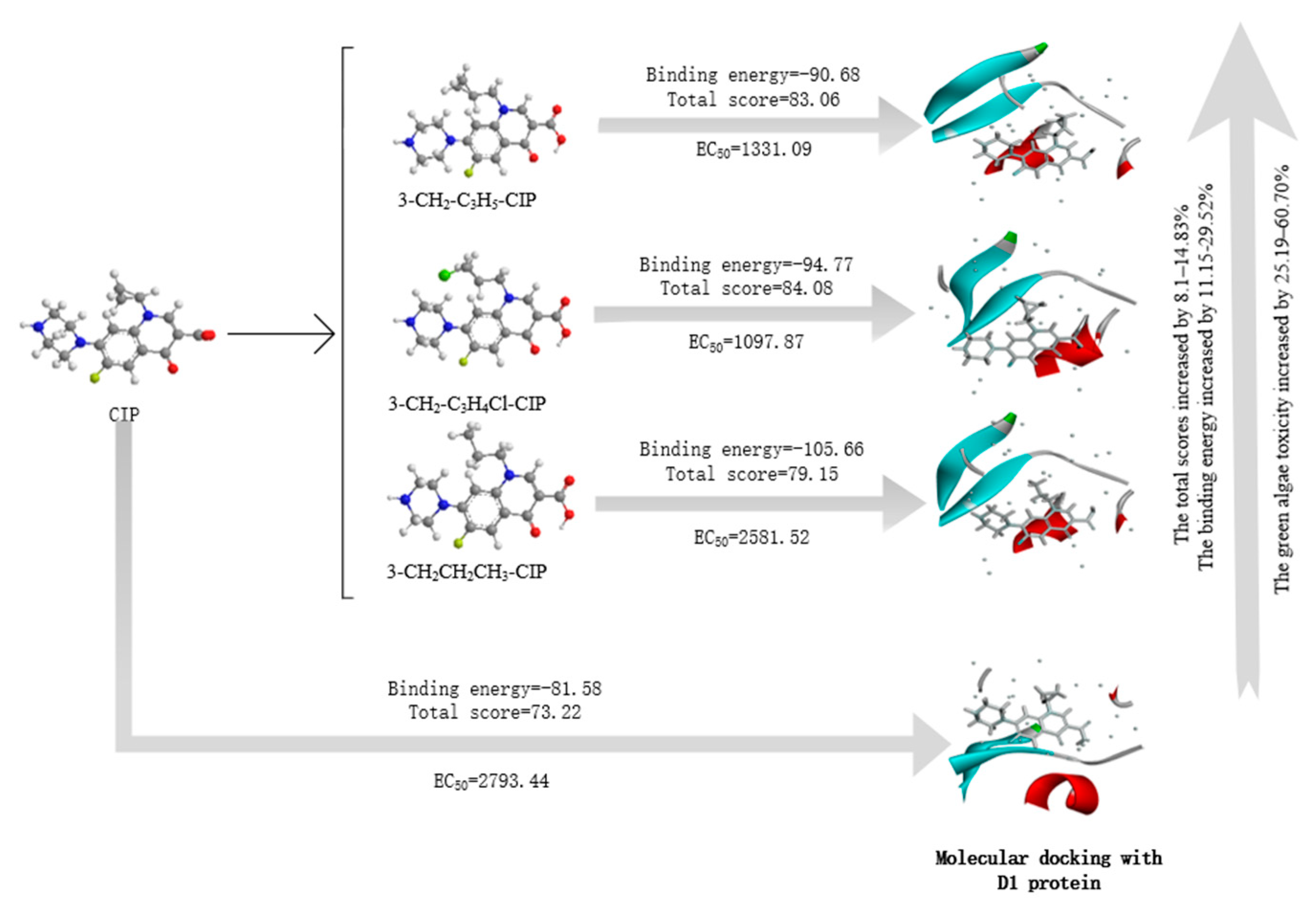
3.2. Molecular Design of CIP Derivatives with Enhanced Double-Activity FQs’ Green Algae Toxicity and Bacterial Genotoxicity
3.3. Environmental Stability of the CIP Derivatives on the Basis of Gaussian Calculations
3.4. Environmentally Friendly Evaluation of CIP Derivatives on the Basis of EPI
3.5. Validation of Double-Activity Qunolones’ Toxicities in the CoMSIA Model
3.5.1. Validation of the CoMSIA Model on the Basis of the Single Activities of Green Algae Toxicity and Bacterial Genotoxicity
3.5.2. Reliability Analysis of the CoMSIA Model on the Basis of the Contour Maps of a Single Activity of Green Algae Toxicity and Bacterial Genotoxicity
3.6. Mechanism Analysis of Green Algae Toxicity and Bacterial Genotoxicity on the Basis of Molecular Docking
3.6.1. Molecular Docking of the CIP Derivatives with the D1 Protein and DNA Gyrase
3.6.2. Molecular Docking of the CIP Derivatives with Ferredoxin-NADP (H) Reductases (FNRs)
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Vincent, T.A. The Quinolones: Past, Present, and Future. Clin. Infect. Dis. 2005, 40, 425–431. [Google Scholar]
- Hamad, B. The antibiotics market. Nat. Rev. Drug Discov. 2010, 9, 675–676. [Google Scholar] [CrossRef] [PubMed]
- Pico, Y.; Andreu, V. Fluoroquinolones in soil-risks and challenges. Analyt. Bioanalyt. Chem. 2007, 387, 1287–1299. [Google Scholar] [CrossRef] [PubMed]
- Rowen, R.C.; Michel, D.; Thompson, J.C. Norfloxacin: Clinical pharmacology and clinical use. Pharmacotherapy 1987, 7, 92–110. [Google Scholar] [CrossRef] [PubMed]
- Wise, R. Norfloxacin-a review of pharmacology and tissue penetration. J. Antimicrob. Chemother. 1984, 13, 59–64. [Google Scholar] [CrossRef]
- Kolpin, D.W.; Furlong, E.T.; Meyer, M.T.; Thurman, E.M.; Zaugg, S.D.; Barber, L.B.; Buxton, H.T. Pharmaceuticals, hormones, and other organic wastewater contaminants in U.S. streams, 1999–2000: A national reconnaissance. Environ. Sci. Technol. 2002, 36, 1202–1211. [Google Scholar] [CrossRef]
- Watkinson, A.; Murby, E.J.; Kolpin, D.W.; Costanzo, S.D. The occurrence of antibiotics in an urban watershed: From wastewater to drinking water. Sci. Total Environ. 2009, 407, 2711–2723. [Google Scholar] [CrossRef]
- Huang, Y.H.; Liu, Y.; Du, P.P.; Zeng, L.J.; Mo, C.H.; Li, Y.W.; Lü, H.X.; Cai, Q.Y. Occurrence and distribution of antibiotics and antibiotic resistant genes in water and sediments of urban rivers with black-odor water in Guangzhou, South China. Sci. Total Environ. 2019, 670, 170–180. [Google Scholar] [CrossRef]
- Liu, X.H.; Zhang, G.D.; Liu, Y.; Lu, S.Y.; Qin, P.; Guo, X.C.; Bi, B.; Wang, L.; Xi, B.D.; Wu, F.C.; et al. Occurrence and fate of antibiotics and antibiotic resistance genes in typical urban water of Beijing, China. Environ. Pollut. 2019, 246, 163–173. [Google Scholar] [CrossRef]
- Qin, Y.W.; Wen, Q.; Ma, Y.Q.; Yang, C.C.; Liu, Z.C. Antibiotics pollution in Gonghu Bay in the period of water diversion from Yangtze River to Taihu Lake. Environ. Earth Sci. 2018, 47, 419. [Google Scholar] [CrossRef]
- Donald, M.A.; Patricia, M.G.; Joann, M.B. Harmful algal blooms and eutrophication: Nutrient sources, composition, and consequences. Estuaries 2002, 25, 704–726. [Google Scholar]
- Huo, Y.Z.; Zhang, J.H.; Chen, L.P.; Hu, M.; Yu, K.F.; Chen, Q.F.; He, Q.; He, P.M. Green algae blooms caused by Ulva prolifera in the southern Yellow Sea: Identification of the original bloom location and evaluation of biological processes occurring during the early northward floating period. Limnol. Oceanogr. 2013, 58, 2206–2218. [Google Scholar] [CrossRef]
- Zhao, X.H.; Wang, X.L.; Li, Y. Combined HQSAR method and molecular docking study on genotoxicity mechanism of quinolones with higher genotoxicity. Environ. Sci. Pollut. Res. 2019, 26, 34830–34853. [Google Scholar] [CrossRef] [PubMed]
- Ebert, I.; Bachmann, J.; Kühnen, U.; Küster, A.; Kussatz, C.; Maletzki, D.; Schlüter, C. Toxicity of the fluoroquinolone antibiotics enrofloxacin and ciprofloxacin to photoautotrophic aquatic organisms. Environ. Toxic. Chem. 2011, 30, 2786–2792. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.H.; Zhao, Y.Y.; Ren, Z.X.; Li, Y. Combined QSAR/QSPR and molecular docking study on fluoroquinolones to reduce biological enrichment. Comput. Biol. Chem. 2019, 79, 177–184. [Google Scholar] [CrossRef]
- Zhao, X.H.; Chu, Z.H.; Li, Y. Molecular Design of Lower Photodegradation Fluoroquinolone Antibiotics and Their Photolysis Paths Inference. Chem. J. Chin. U. 2018, 39, 2707–2718. [Google Scholar]
- Ren, Z.X.; Wang, Y.W.; Xu, H.L.; Li, Y.F.; Han, S. Fuzzy Comprehensive Evaluation Assistant 3D-QSAR of Environmentally Friendly FQs to Reduce ADRs. Int. J. Environ. Res. Public Health 2019, 16, 3161. [Google Scholar] [CrossRef]
- Li, M.; Wei, D.B.; Zhao, H.M.; Du, Y.G. Genotoxicity of quinolones: Substituents contribution and transformation products QSAR evaluation using 2D and 3D models. Chemosphere 2014, 95, 220–226. [Google Scholar] [CrossRef]
- Tang, T.; Zhai, Y.J.; Huang, K. Research on Evaluation of the Pipeline Direct Drinking Water Quality With Improved Comprehensive Index Method. Manag. Sci. Eng. 2011, 5, 95–100. [Google Scholar]
- Svante, W.; Michael, S.; Lennart, E. PLS-regression: A basic tool of chemometrics. Chemometr. Intell. Lab. 2001, 58, 109–130. [Google Scholar]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A.; et al. GAUSSIAN 09 (Revision, A.02); Gaussian, Inc.: Wallingford, CT, USA, 2009. [Google Scholar]
- Qu, R.J.; Liu, H.X.; Feng, M.B.; Yang, X.; Wang, Z.Y. Investigation on intramolecular hydrogenbon and some thermodynamic properties of polyhydroxylated pnthraquinones. Chem. Eng. Data 2012, 57, 2442–2455. [Google Scholar] [CrossRef]
- Yang, L.Z.; Liu, M. 3D-QSAR Model of Polybrominated Biphenyls Tri-effect Modified by Standard Deviation Standardization Method and Its Application in Environmental Friendly Molecular Modification. Chem. J. Chin. U. 2019, 40, 2471–2479. [Google Scholar]
- Gu, W.W.; Chen, Y.; Li, Y. Attenuation of the Atmospheric Migration Ability of Polychlorinated Naphthalenes (PCN-2) Based on Three-dimensional QSAR Models with Full Factor Experimental Design. Bull. Environ. Contam. Toxicol. 2017, 99, 276–280. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.L.; Gu, W.W.; Guo, E.M.; Cui, C.Y.; Li, Y. Assessment of long-range transport potential of polychlorinated Naphthalenes based on three-dimensional QSAR models. Environ. Sci. Pollut. Res. 2017, 24, 14802–14818. [Google Scholar] [CrossRef]
- Sheng, C.Q.; Zhang, W.N.; Ji, H.T.; Zhang, M.; Song, Y.L.; Xu, H.; Zhu, J.; Miao, Z.Y.; Jiang, Q.F.; Yao, J.Z.; et al. Structure-Based Optimization of Azole Antifungal Agents by CoMFA, CoMSIA, and Molecular Docking. J. Medic. Chem. 2006, 49, 2512–2525. [Google Scholar] [CrossRef]
- Aruoja, V.; Sihtmäe, M.; Dubourguier, H.C.; Kahru, A. Toxicity of 58 substituted anilines and phenols to algae Pseudokirchneriella subcapitata and bacteria Vibrio fischeri: Comparison with published data and QSARs. Chemosphere 2011, 84, 1310–1320. [Google Scholar] [CrossRef]
- Chen, C.Y.; Lin, J.H. Toxicity of chlorophenols to Pseudokirchneriella subcapitata under air-tight test environment. Chemosphere 2006, 62, 503–509. [Google Scholar] [CrossRef]
- Huang, C.P.; Wang, Y.J.; Chen, C.Y. Toxicity and quantitative structure–activity relationships of nitriles based on Pseudokirchneriella subcapitata. Ecotoxicol. Environ. Saf. 2006, 67, 439–446. [Google Scholar] [CrossRef]
- Miguel, G.P.; Soledad, G.; Ismael, R.P.; Francisco, L.; Roberto, R.; Karina, B.; Eduardo, M.; Francisca, F.P. Toxicity of five antibiotics and their mixtures towards photosynthetic aquatic organisms: Implications for environmental risk assessment. Water Res. 2013, 47, 2050–2064. [Google Scholar]
- Trost, J.T.; Chisholm, D.A.; Jordan, D.B.; Diner, B.A. The D1 C-terminal processing protease of photosystem II from Scenedesmus obliquus. Protein purification and gene characterization in wild type and processing mutants. J. Biolog. Chemist. 1997, 272, 20348–20356. [Google Scholar] [CrossRef]
- Jansen, M.A.K.; Mattoo, A.K.; Edelman, M. D1-D2 protein degradation in the chloroplast. FEBS J. 1999, 260, 527–532. [Google Scholar] [CrossRef]
- Adir, N.; Zer, H.; Shochat, S.; Ohad, I. Photoinhibition-a historical perspective. Photosynth. Res. 2003, 76, 343. [Google Scholar] [CrossRef] [PubMed]
- Aro, E.M.; Suorsa, M.; Rokka, A.; Allahverdiyeva, Y.; Paakkarinen, V.; Saleem, A.; Battchikova, N.; Rintamaki, E. Dynamics of photosystem II: A proteomic approach to thylakoid protein complexes. J. Exp. Bot. 2005, 56, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Jacoby, G.A. Mechanisms of resistance to quinolones. Clin. Infect. Dis. 2005, 41, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Musumeci, M.A.; Botti, H.; Buschiazzo, A.; Ceccarelli, E.A. Swapping FAD Binding Motifs between Plastidic and Bacterial Ferredoxin-NADP(H.) Reductases. Biochemistry 2011, 50, 2111–2122. [Google Scholar] [CrossRef]
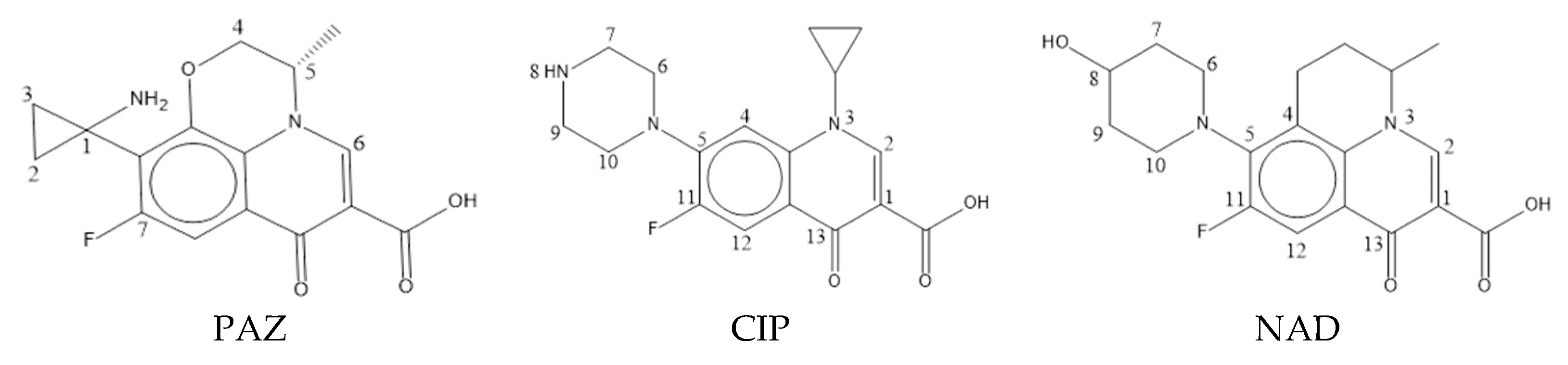
| No. | Name | pEC50 | pLOEC | Composite Value |
|---|---|---|---|---|
| 1 | Pipemidic acid (PIP) | −4.59 | 4.80 | −0.94 |
| 2 * | Cinoxacin (CIN) | −2.24 | 6.52 | −0.17 |
| 3 | Norfloxacin (NOR) | −3.64 | 7.67 | −0.48 |
| 4 * | Balofloxacin (BAL) | −2.83 | 7.31 | −0.28 |
| 5 | Ciprofloxacin (CIP) | −3.45 | 7.85 | −0.41 |
| 6 | Danofloxacin (DAN) | −3.17 | 7.66 | −0.35 |
| 7 * | Difloxacin (DIF) | −2.64 | 8.06 | −0.17 |
| 8 | Enrofloxacin (ENR) | −3.00 | 8.75 | −0.23 |
| 9 | Fleroxacin (FLE) | −3.46 | 6.77 | −0.49 |
| 10 | Gatifloxacin (GAT) | −3.16 | 8.28 | −0.30 |
| 11 | Lomefloxacin (LOM) | −3.26 | 7.54 | −0.38 |
| 12 | Moxifloxacin (MOX) | −0.32 | 8.89 | 0.53 |
| 13 | Nadifloxacin (NAD) | −1.62 | 8.08 | 0.11 |
| 14 | Ofloxacin (OFL) | −3.62 | 7.96 | −0.45 |
| 15 * | Pazufloxacin (PAZ) | −3.52 | 8.54 | −0.38 |
| 16 | Rufloxacin (RUF) | −3.55 | 6.89 | −0.51 |
| 17 | Sarafloxacin (SAR) | −2.77 | 7.92 | −0.22 |
| 18 * | Sparfloxacin (SPA) | −3.44 | 7.37 | −0.44 |
| 19 | Levofloxacin (LEV) | −3.62 | 7.75 | −0.47 |
| 20 * | Enoxacin (ENO) | −3.58 | 7.08 | −0.50 |
| 21 | Pefloxacin (PEF) | −3.51 | 7.96 | −0.42 |
| 22 * | Amifloxacin (AMI) | −4.39 | 8.00 | −0.66 |
| 23 * | Besifloxacin (BES) | −2.43 | 7.50 | −0.15 |
| 24 | Clinafloxacin (CLI) | −3.08 | 7.19 | −0.36 |
| 25 | Grepafloxacin (GRE) | −2.81 | 7.47 | −0.26 |
| 26 | Marbofloxacin (MAR) | −4.26 | 8.46 | −0.60 |
| 27 * | Orbifloxacin (ORB) | −2.67 | 7.51 | −0.22 |
| 28 * | Sitafloxacin (SIT) | −2.69 | 7.28 | −0.24 |
| 29 | Temafloxacin (TEM) | −2.38 | 8.47 | −0.07 |
| 30 | 1-CH3-PAZ | −0.98 | 8.05 | 0.29 |
| 31 | 1-H-PAZ | −1.27 | 7.67 | 0.18 |
| 32 | 5-OH-PAZ | −4.87 | 8.38 | −0.77 |
| 33 | 5-F-PAZ | −3.90 | 7.57 | −0.56 |
| 34 | 1-C2H3-5-F-PAZ | −1.14 | 7.13 | 0.18 |
| 35 | 1-CO-5-OH-PAZ | −3.39 | 7.72 | −0.41 |
| 36 | 1-CO-5-F-PAZ | −2.42 | 7.14 | −0.18 |
| 37 | 7-OH-CIP | −4.53 | 8.25 | −0.69 |
| 38 | 7-C2H5-CIP | −2.85 | 7.98 | −0.24 |
| 39 | 7-CN-CIP | −3.86 | 8.07 | −0.51 |
| 40 | 7-NO-CIP | −3.66 | 8.14 | −0.45 |
| 41 | 7-OCH3-CIP | −3.68 | 8.38 | −0.44 |
| 42 | 7-C2H3-CIP | −2.94 | 8.34 | −0.24 |
| 43 | 7-COOH-CIP | −4.70 | 8.31 | −0.73 |
| 44 | 9-F-NAD | −1.73 | 8.33 | 0.10 |
| 45 | 9-Cl-NAD | −1.53 | 8.32 | 0.15 |
| 46 | 9-Br-NAD | −1.52 | 8.28 | 0.15 |
| 47 | 2-C2H3-NAD | −1.02 | 8.32 | 0.29 |
| 48 | 2-C2H3-9-F-NAD | −1.13 | 8.18 | 0.25 |
| 49 | 2-C2H-9-F-NAD | −1.57 | 8.29 | 0.14 |
| 50 | 2-C2H3-9-Cl-NAD | −1.37 | 8.29 | −0.17 |
| Model | q2 | n | SEE | R2 | F | r2pred | SEP | Q2 | cSDEP | dq2/dr2yy |
|---|---|---|---|---|---|---|---|---|---|---|
| CoMSIA | 0.58 | 7 | 0.02 | 1 | 1265.33 | 0.73 | 0.17 | 0.54 | 0.25 | 0.8 |
| Model | S | E | H | D | A |
|---|---|---|---|---|---|
| CoMSIA | 10.9% | 19.8% | 32.7% | 13.8% | 22.8% |
| No. | Molecule | Comprehensive Value | Change Rate of Comprehensive Value (%) |
|---|---|---|---|
| 0 | CIP | −0.604 | - |
| 1 | 3-CH2-C3H5-CIP | −0.489 | 19.09 |
| 2 | 3-CH2-C3H4Cl-CIP | −0.529 | 12.47 |
| 3 | 3-CH2CH2CH3-CIP | −0.530 | 12.31 |
| No. | Molecule | Frequency | Total Energy | Change Rate of Total Energy |
|---|---|---|---|---|
| 0 | CIP | 26.78 | −1147.95 | - |
| 1 | 3-CH2-C3H5-CIP | 25.79 | −1187.27 | 3.43 |
| 2 | 3-CH2-C3H4Cl-CIP | 18.25 | −1646.78 | 43.45 |
| 3 | 3-CH2CH2CH3-CIP | 20.12 | −1149.18 | 0.11 |
| No. | Molecule | logKow | Change Rate of logKow | t1/2 (River) | Change Rate of t1/2 (River) |
|---|---|---|---|---|---|
| 0 | CIP | 0.28 | - | 8.72 × 1013 | - |
| 1 | 3-CH2-C3H5-CIP | 0.49 | 75 | 6.71 × 1013 | −23.12 |
| 2 | 3-CH2-C3H4Cl-CIP | 0.67 | 139.29 | 2.00 × 1014 | 129.02 |
| 3 | 3-CH2CH2CH3-CIP | 0.19 | −32.14 | 3.84 × 1013 | −55.98 |
| No. | Molecule | EC50 | LOEC | Change Rate of EC50 (%) | Change Rate of LOEC (%) | The Ratio of Two Toxicity Changes (Logarithmic Level) |
|---|---|---|---|---|---|---|
| 0 | CIP | 2793.44 | 1.40 × 10−8 | - | - | - |
| 1 | 3-CH2-C3H5-CIP | 1331.09 | 9.66 × 10−9 | 52.35 | 30.98 | 4.55 |
| 2 | 3-CH2-C3H4Cl-CIP | 1097.87 | 9.55 × 10−9 | 60.70 | 31.77 | 5.56 |
| 3 | 3-CH2CH2CH3-CIP | 2581.52 | 1.07 × 10−8 | 25.19 | 23.26 | 0.67 |
| Force Field | CoMSIA Model of Double-Activity | CoMSIA Model of Green Algae Toxicity | CoMSIA Model of Bacterial Genotoxicity |
|---|---|---|---|
| Steric | 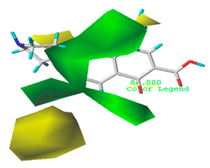 10.8% | 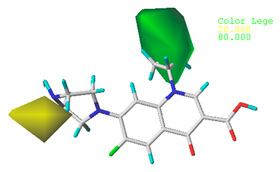 4.1% | 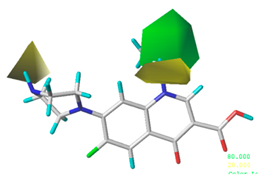 7.8% |
| Electrostatic | 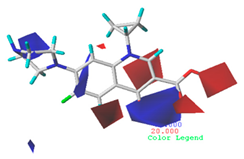 19.8% |  13.7% | 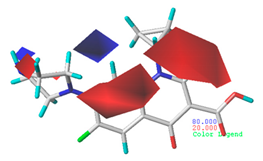 17.7% |
| Hydrophobic | 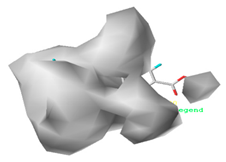 32.7% | 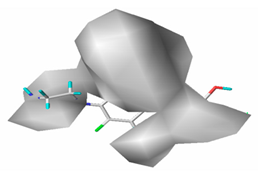 26.1% |  21.3% |
| Donor | 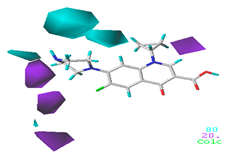 13.8% | 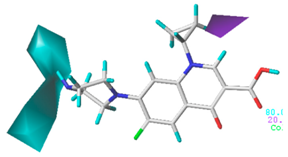 5.4% | 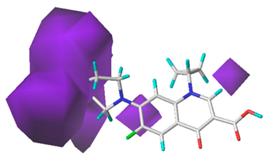 4.4% |
| Acceptor | 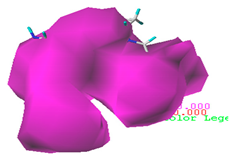 22.8% | 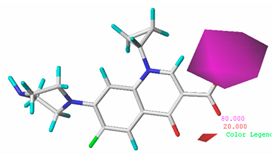 50.7% |  48.7% |
| No. | Molecule | Total Score (1FC6) | Change Rate | Binding Energy | Change Rate | Total Score (5Z9P) | Change Rate | Total Score (2XNC) | Change Rate |
|---|---|---|---|---|---|---|---|---|---|
| 0 | CIP | 73.22 | - | −81.58 | - | 59.22 | - | 71.08 | - |
| 1 | 3-CH2-C3H5-CIP | 83.06 | 13.44 | −90.68 | 11.15 | 70.89 | 19.72 | 77.82 | 9.48 |
| 2 | 3-CH2-C3H4Cl-CIP | 84.08 | 14.83 | −94.77 | 16.17 | 69.01 | 16.54 | 82.56 | 16.15 |
| 3 | 3-CH2CH2CH3-CIP | 79.15 | 8.1 | −105.66 | 29.52 | 67.18 | 13.44 | 77.17 | 8.58 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, L.-z.; Liu, M. A Double-Activity (Green Algae Toxicity and Bacterial Genotoxicity) 3D-QSAR Model Based on the Comprehensive Index Method and Its Application in Fluoroquinolones’ Modification. Int. J. Environ. Res. Public Health 2020, 17, 942. https://doi.org/10.3390/ijerph17030942
Yang L-z, Liu M. A Double-Activity (Green Algae Toxicity and Bacterial Genotoxicity) 3D-QSAR Model Based on the Comprehensive Index Method and Its Application in Fluoroquinolones’ Modification. International Journal of Environmental Research and Public Health. 2020; 17(3):942. https://doi.org/10.3390/ijerph17030942
Chicago/Turabian StyleYang, Lu-ze, and Miao Liu. 2020. "A Double-Activity (Green Algae Toxicity and Bacterial Genotoxicity) 3D-QSAR Model Based on the Comprehensive Index Method and Its Application in Fluoroquinolones’ Modification" International Journal of Environmental Research and Public Health 17, no. 3: 942. https://doi.org/10.3390/ijerph17030942
APA StyleYang, L.-z., & Liu, M. (2020). A Double-Activity (Green Algae Toxicity and Bacterial Genotoxicity) 3D-QSAR Model Based on the Comprehensive Index Method and Its Application in Fluoroquinolones’ Modification. International Journal of Environmental Research and Public Health, 17(3), 942. https://doi.org/10.3390/ijerph17030942




