What Role Can Process Mining Play in Recurrent Clinical Guidelines Issues? A Position Paper
Abstract
1. Introduction
2. The Data: The Starting Point
2.1. Data Sources
2.2. Giving Meaning to Data
- Data semantics are concerned about the content and structure of observations over a biological subject or organization. For example, the data structure for representing the items contained in a urine analysis (color, pH, specific gravity, presence of nitrites, glucose measurement, etc.).
- Contextual semantics are needed to correctly interpret data semantics. They concern contextual aspects such as times of events, display used for a specific measurement, performer of an activity, and place where an event occurred. They are often tightly linked to data semantics in most of the standards for representing clinical information.
- Workflow semantics concern the specification of the order of biomedical events. These semantics are needed to understand temporal, conditional, and causal relationships among various events. Examples include which event occurred before a nosocomial infection, which activity is executed after detecting stroke in the emergency ward, and what order should preoperative activities follow. These kinds of semantics are represented by workflow specification standards such as the openEHR Task Model, GLIF, or BPM.
3. Clinical Guidelines
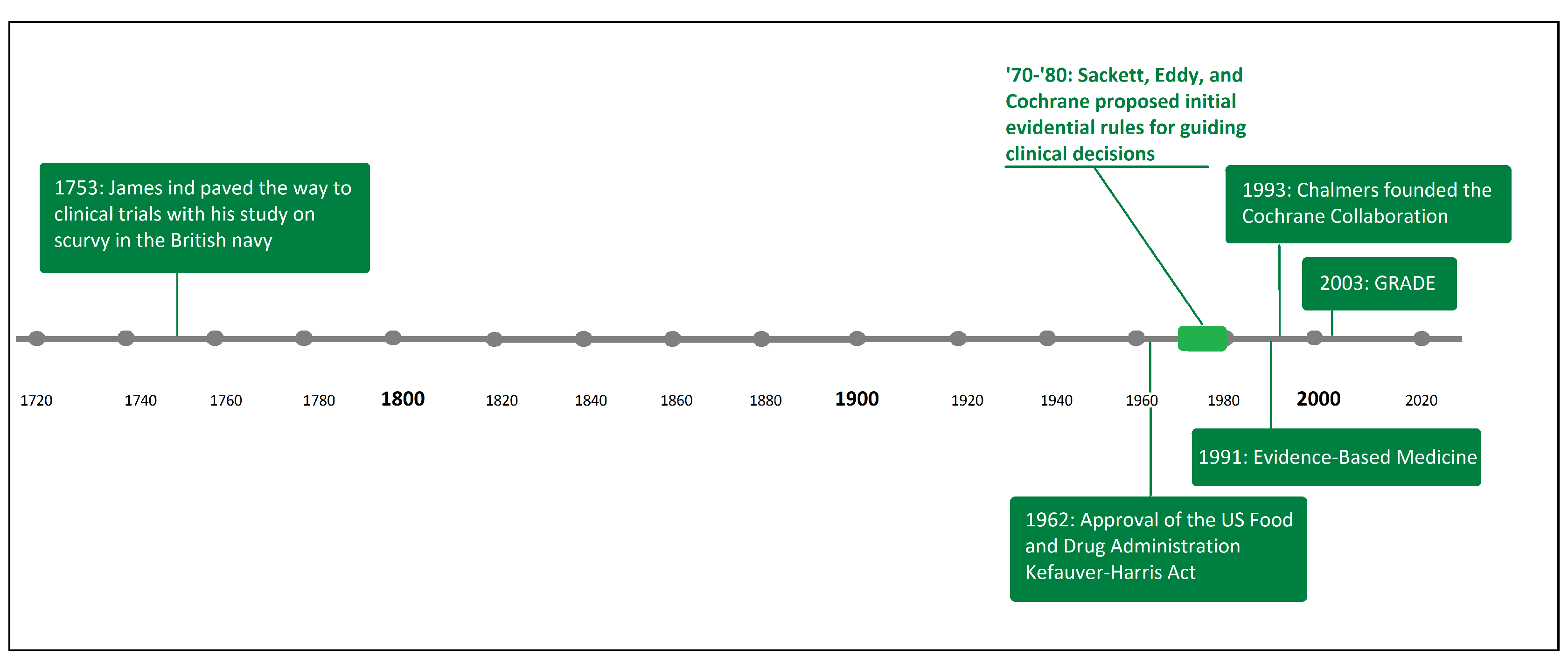
3.1. Definition and History of CGs
3.2. Between the Shades of CGs
- the full guidelines, that provide complete coverage of a health topic/disease, include recommendations in relation to all aspects of the topic (e.g., surveillance, diagnosis, public health, and clinical interventions), and need to be fully based on systematic reviews of the evidence for each aspect;
- the standard guidelines, that are produced in response to a request for guidance in relation to a change in practice or controversy in a single clinical or policy area, and that are supported by systematic reviews of the evidence, but not expected to cover the full scope of the condition or public health problem;
- the rapid advice guidelines, that are produced in response to a public health emergency and, for this reason, are mainly only evidence-informed and may not be supported by full reviews of the evidence; and
- the compilations of guidelines, that contain current recommendations from WHO and other sources, but does not include any new recommendations.
- The clinical consensus statements are a collection of opinions on a particular aspect of medical knowledge, generally agreed by a representative group of experts in that area upon an evidence-based, state-of-the-art knowledge. In contrast to CGs, which are based primarily on high-level evidence, clinical consensus statements are more applicable to situations where evidence is limited or lacking, yet there are still opportunities to reduce uncertainty and improve quality of care [41]. Moreover, the consensus statements synthesize new information that may have implications for revaluation of routine medical practices, and they do not provide specific algorithms or guidelines for practice because these depend on cost, available expertise and technology, and local practice circumstances [42].
- After their introduction in the practice in 2008 [45], clinical checklists are experiencing a wide spread [46]. Structured as a schematic list of actions or controls, checklists are inserted into various points of the clinical process, ensuring that providers do not forget crucial steps during either routine, mundane tasks or dynamic, emergent events [47].
3.3. Guidelines in Health Management
- CGs may avoid unnecessary diagnostic tests, which are routinely performed in daily practice in hospitals, helping doctors to select the most suitable ones based on predefined procedural indications. This may permit increased hospital efficiency, avoiding wasting resources on unnecessary testing, while maintaining the level of care provided to patients [49,54]. In addition, a reduction in the number of test requests also leads to an improvement in the service level and care effectiveness because it allows to cut the waiting lists (queues) for the tests and therefore to reduce the waiting times for patients who really need them [51,55]. This rationale, explained for diagnostic tests, can be extended also to medical treatments although probably to a lesser extent.
- CGs establish a benchmark to periodically evaluate the care paths of patients affected by a specific disease [52,56]. This makes it possible to match the activities which were actually executed against CGs. Therefore, it permits providing detailed feedback on the decisions of medical staff and/or on the unit management according to the different specific diseases. Such indications may allow doctors to identify their evaluation errors and to improve their future attitude.
- CGs may help health managers in resource planning, especially when a new center/unit must be set up [57]. Indeed, CGs may be exploited to estimate the activities needed for an “average patient” admitted with a certain disease.
- CGs can be exploited by insurance companies or health authorities to analyze whether hospitals are, for a particular patient group, complying with CGs when executing their diagnostic and treatments activities [58,59]. Non-compliant behavior, if large and economically advantageous, may be linked to abuses or mistakes.
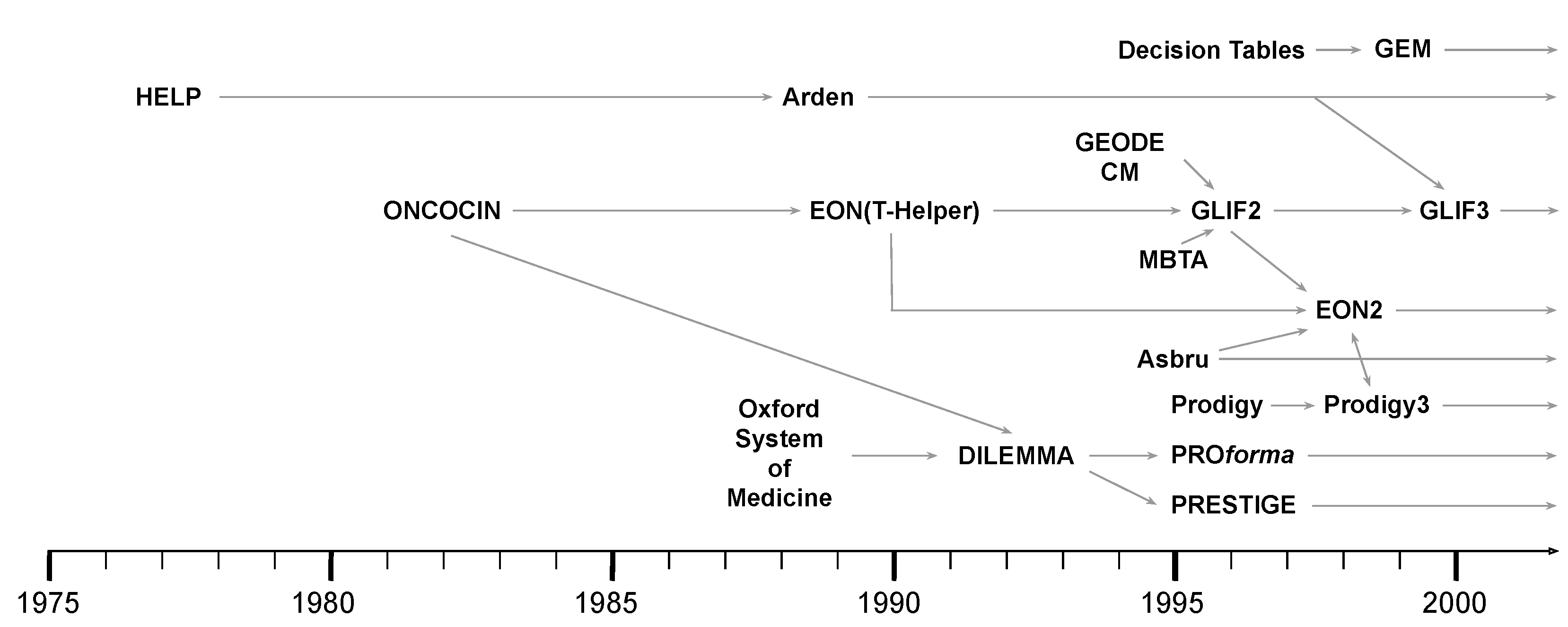
4. Computer Interpretable Clinical Guidelines
Linking CIGs to Data
5. CIGs and BPM
6. PM for Healthcare: The Perspectives on CGs
7. Conclusions
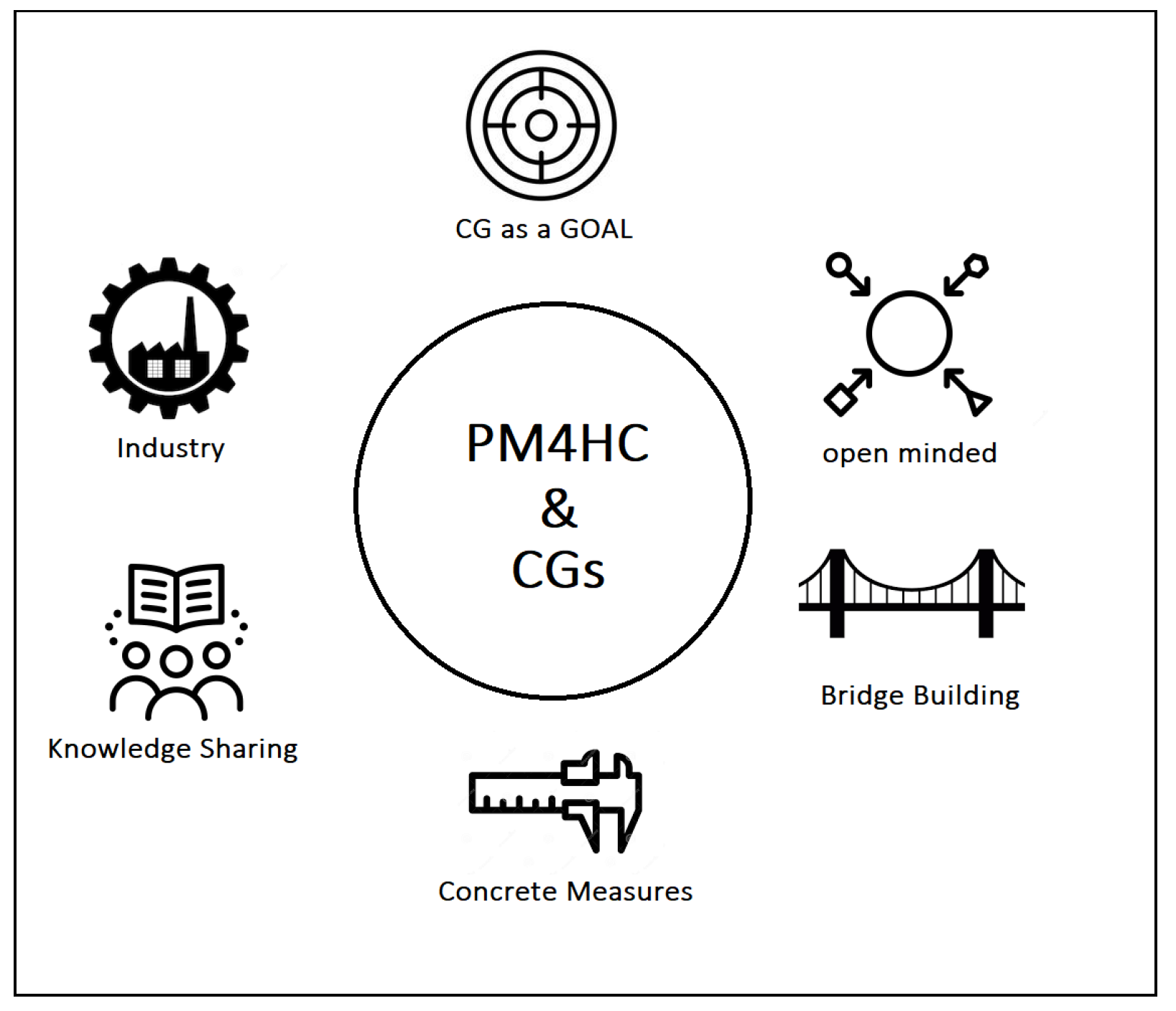
- CG as a goal. CGs represent an essential point in healthcare and PM4HC. Measuring process performance by considering real-world data is a primary aim of CC and CGs, and represents low-hanging fruits. The synergy with process discovery and process enhancement, in particular, open exciting scenarios about the potential of PM4HC, e.g., the former to potentially identify the most promising pattern of care given some successful clinical pathways, the latter to try to improve a given CGs with real-world data from a specific care unit.
- Open-minded. The fragmented approach in coping with CGs (CIGs, BPM, CBR, etc.) and the lack of communication among the members of these sub-communities caused the potential contribution of some technologies to be unexplored. The PM4HC community should be open-minded with regards to the results of other disciplines, and should be inclusive with respect to successful and promising methods which can be acquired and adapted. It is well positioned to play a unifying role for the broader field.
- Bridge Building. The cultural gap between computer scientists and healthcare providers is one of the main challenges, and distinguishing features, of PM4HC. This gap is mainly due to many years of specialization and can play a critical role in leading a joint project to success or failure. To make the communication more efficient, and increase the chances to come to successful results, physicians, nurses, caregivers, and administrative workers in healthcare should be invited to play an active role in all the steps of the projects and, more in general, also invited to contribute in terms of vision for the future of the discipline. Beside the clear communication-related issues of the cultural gap, there is also the remarkable aspect of sustainability. Bridging the gap between computer scientists and healthcare experts also encompasses the fact that proposed technological solutions must not become overwhelming with respect to the daily clinical workload. In other words, bridging the cultural gap also means to propose solutions able to fit with the actual working environment and that can provide clear benefits beside the additional effort that they may require (e.g., time spent for data entry, meetings, etc.).
- Concrete measures. In many cases, CGs presented in papers did not lead to concrete applications. Unfortunately, CGs still remain on paper in many hospitals. We think that being able to validate our future proposals with feedback from the domain experts, and providing concrete measures of the benefits generated by PM4HC solutions could convince physicians to adopt PM4HC tools. In this direction, shared methods to validate on the field future works and tools must be encouraged, when possible.
- Knowledge sharing. We strongly believe in teamwork and in the cooperation among research centers and healthcare institutions across different countries. To this end, initiatives aimed at sharing knowledge about community members working on CGs implementations should be promoted and supported.
- Industry. Proof of concepts or prototypes are the most common way for the scientific community to deliver their intellectual product. However, to make the jump from prototypes to commercial products which efficiently support daily clinical practice, the interaction with healthcare companies cannot be avoided and should be elicited as one of the aims of our community. The creation of a process-oriented culture with commercial vendors, in terms of data entry/presentation, ontologies, and data export should be pursued and joint project should be encouraged.
Author Contributions
Funding
Conflicts of Interest
References
- Guyatt, G.; Cairns, J.; Churchill, D.; Cook, D.; Haynes, B.; Hirsh, J.; Irvine, J.; Levine, M.; Levine, M.; Nishikawa, J.; et al. Evidence-Based Medicine: A New Approach to Teaching the Practice of Medicine. JAMA 1992, 268, 2420–2425. [Google Scholar] [CrossRef] [PubMed]
- Hripcsak, G.; Ludemann, P.; Pryor, T.; Wigertz, O.; Clayton, P. Rationale for the Arden Syntax. Comput. Biomed. Res. Int. J. 1994, 27, 291–324. [Google Scholar] [CrossRef] [PubMed]
- Peleg, M. Computer-interpretable Clinical Guidelines: A Methodological Review. J. Biomed. Inform. 2013, 46, 744–763. [Google Scholar] [CrossRef] [PubMed]
- Van de Velde, S.; Heselmans, A.; Delvaux, N.; Brandt, L.; Marco-Ruiz, L.; Spitaels, D.; Cloetens, H.; Kortteisto, T.; Roshanov, P.; Kunnamo, I.; et al. A systematic review of trials evaluating success factors of interventions with computerised clinical decision support. Implement. Sci. 2018, 13, 114. [Google Scholar] [CrossRef]
- Rawson, T.M.; Moore, L.S.P.; Hernandez, B.; Charani, E.; Castro-Sanchez, E.; Herrero, P.; Hayhoe, B.; Hope, W.; Georgiou, P.; Holmes, A.H. A systematic review of clinical decision support systems for antimicrobial management: Are we failing to investigate these interventions appropriately? Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2017, 23, 524–532. [Google Scholar] [CrossRef]
- Greenes, R.A.; Bates, D.W.; Kawamoto, K.; Middleton, B.; Osheroff, J.; Shahar, Y. Clinical decision support models and frameworks: Seeking to address research issues underlying implementation successes and failures. J. Biomed. Inform. 2018, 78, 134–143. [Google Scholar] [CrossRef]
- van der Aalst, W. Process Mining: Discovery, Conformance and Enhancement of Business Processes, 1st ed.; Springer Publishing Company, Incorporated: New York, NY, USA, 2011. [Google Scholar]
- van der Aalst, W.; Adriansyah, A.; de Medeiros, A.K.A.; Arcieri, F.; Baier, T.; Blickle, T.; Bose, J.C.; van den Brand, P.; Brandtjen, R.; Buijs, J.; et al. Process Mining Manifesto. In Business Process Management Workshops; Springer: Berlin/Heidelberg, Germany, 2011; pp. 169–194. [Google Scholar]
- dos Santos Garcia, C.; Meincheim, A.; Junior, E.R.F.; Dallagassa, M.R.; Sato, D.M.V.; Carvalho, D.R.; Santos, E.A.P.; Scalabrin, E.E. Process Mining Techniques and Applications—A Systematic Mapping Study. Expert Syst. Appl. 2019, 133, 260–295. [Google Scholar] [CrossRef]
- Bhargava, A.; Kim, T.; Quine, D.B.; Hauser, R.G. A 20-Year Evaluation of LOINC in the United States’ Largest Integrated Health System. Arch. Pathol. Lab. Med. 2019, 144. [Google Scholar] [CrossRef]
- Lee, D.; de Keizer, N.; Lau, F.; Cornet, R. Literature review of SNOMED CT use. J. Am. Med. Inform. Assoc. 2014, 21, e11–e19. [Google Scholar] [CrossRef]
- World Health Organization. ICD-10: International Statistical Classification of Diseases and Related Health Problems: Tenth Revision; World Health Organization: Geneva, Switzerland, 2004. [Google Scholar]
- Trotti, A.; Colevas, A.D.; Setser, A.; Rusch, V.; Jaques, D.; Budach, V.; Langer, C.; Murphy, B.; Cumberlin, R.; Coleman, C.N.; et al. CTCAE v3.0: Development of a comprehensive grading system for the adverse effects of cancer treatment. Semin. Radiat. Oncol. 2003, 13, 176–181. [Google Scholar] [CrossRef]
- Daniel, C.; Kalra, D. Section Editors for the IMIA Yearbook Section on Clinical Research Informatics. Clinical Research Informatics: Contributions from 2018. Yearb. Med. Inform. 2019, 28, 203–205. [Google Scholar] [PubMed]
- Marco-Ruiz, L.; Nicolaisen, K.; Pedersen, R.; Makhlysheva, A.; Bakkevoll, P.A. Ontology-Based Terminologies for Healthcare-Impact Assessment and Transitional Consequences for Implementation, 1st ed.; Norwegian Centre for E-health Research: Tromsø, Norway, 2017; Volume 1. [Google Scholar]
- Marco-Ruiz, L.; Pedrinaci, C.; Maldonado, J.A.; Panziera, L.; Chen, R.; Bellika, J.G. Publication, discovery and interoperability of Clinical Decision Support Systems: A Linked Data approach. J. Biomed. Inform. 2016, 62, 243–264. [Google Scholar] [CrossRef] [PubMed]
- Downing, G.J. Chapter 28 - Knowledge Management to Support Personalized Healthcare. In Genomic and Personalized Medicine, 2nd ed.; Willard, G.S.G.F., Ed.; Academic Press: New York, NY, USA, 2013; pp. 332–339. [Google Scholar]
- Karlsson, D.; Berzell, M.; Schulz, S. Information Models and Ontologies for Representing the Electronic Health Record. In International Conference on Biomedical Ontology; ceur-ws.org: Buffalo, NY, USA, 2011; pp. 153–157. [Google Scholar]
- Marcos, C.; González-Ferrer, A.; Peleg, M.; Cavero, C. Solving the interoperability challenge of a distributed complex patient guidance system: A data integrator based on HL7’s Virtual Medical Record standard. J. Am. Med. Inform. Assoc. JAMIA 2015, 22, 587–599. [Google Scholar] [CrossRef] [PubMed]
- Wulff, A.; Haarbrandt, B.; Tute, E.; Marschollek, M.; Beerbaum, P.; Jack, T. An interoperable clinical decision-support system for early detection of SIRS in pediatric intensive care using openEHR. Artif. Intell. Med. 2018, 89, 10–23. [Google Scholar] [CrossRef] [PubMed]
- Beale, T.; Kejžar, M.; Polajnar, M.; Naess, B.; Santos, D.; Fabjan, B. Task Planning Specification. 2020. [Google Scholar]
- Chen, C.; Chen, K.; Hsu, C.Y.; Chiu, W.T.; Li, Y.C.J. A guideline-based decision support for pharmacological treatment can improve the quality of hyperlipidemia management. Comput. Methods Programs Biomed. 2010, 97, 280–285. [Google Scholar] [CrossRef]
- Anani, N.; Mazya, M.V.; Chen, R.; Prazeres Moreira, T.; Bill, O.; Ahmed, N.; Wahlgren, N.; Koch, S. Applying openEHR’s Guideline Definition Language to the SITS international stroke treatment registry: A European retrospective observational study. Bmc Med. Inform. Decis. Mak. 2017, 17, 7. [Google Scholar] [CrossRef]
- Marco-Ruiz, L.; Budrionis, A.; Yigzaw, K.Y.Y.; Bellika, J.G. Interoperability Mechanisms of Clinical Decision Support Systems: A Systematic Review. In Proceedings of the 14th Scandinavian Conference on Health Informatics 2016, Gothenburg, Sweden, 6–7 April 2016; Linköping University Electronic Press: Linköping, Sweden, 2016; pp. 13–21. [Google Scholar]
- Lind, J. A treatise on the scurvy: In Three Parts, Containing an Inquiry into the Nature, Causes, and Cure, of That Disease; A. Millar: London, UK, 1757. [Google Scholar]
- Mathews, J.R. Quantification and the Quest for Medical Certainty; Princeton University Press: Princeton, NJ, USA, 1995. [Google Scholar]
- Sackett, D.L.; Haynes, R.B.; Tugwell, P.; Guyatt, G.H. Clinical Epidemiology: A Basic Science for Clinical Medicine; Little, Brown and Company: New York, NY, USA, 1985. [Google Scholar]
- Eddy, D.M. Clinical policies and the quality of clinical practice. N. Engl. J. Med. 1982, 307, 343–347. [Google Scholar] [CrossRef]
- Cochrane, A.L. Effectiveness and Efficiency: Random Reflections on Health Services; Nuffield Provincial Hospitals Trust London: London, UK, 1972; Volume 900574178. [Google Scholar]
- Guyatt, G.H. The way of the past. ACP J. Club 1991, 114, A16. [Google Scholar]
- Guyatt, G.H.; Keller, J.L.; Jaeschke, R.; Rosenbloom, D.; Adachi, J.D.; Newhouse, M.T. The n-of-1 randomized controlled trial: Clinical usefulness: Our three-year experience. Ann. Intern. Med. 1990, 112, 293–299. [Google Scholar] [CrossRef]
- Chalmers, I. The Cochrane collaboration: Preparing, maintaining, and disseminating systematic reviews of the effects of health care. Ann. N. Y. Acad. Sci. 1993, 703, 156–165. [Google Scholar] [CrossRef]
- Field, M.J.; Lohr, K.N. Clinical Practice Guidelines. In Directions for a New Program; National Academies Press (US): Washington, DC, USA, 1990; p. 1990. [Google Scholar]
- Graham, R.; Mancher, M.; Wolman, D.; Greenfield, S.; Steinberg, E. Clinical Practice Guidelines We Can Trust; Institute of Medicine Committee on Standards for Developing Trustworthy Clinical Practice Guidelines: Washington, DC, USA, 2011. [Google Scholar]
- Woolf, S.H.; Grol, R.; Hutchinson, A.; Eccles, M.; Grimshaw, J. Potential benefits, limitations, and harms of clinical guidelines. BMJ 1999, 318, 527–530. [Google Scholar] [CrossRef] [PubMed]
- Atkins, D.; Best, D.; Briss, P.A.; Eccles, M.; Falck-Ytter, Y.; Flottorp, S.; Guyatt, G.H.; Harbour, R.T.; Haugh, M.C.; Henry, D.; et al. Grading quality of evidence and strength of recommendations. BMJ 2004, 328, 1490. [Google Scholar]
- Guyatt, G.H.; Oxman, A.D.; Vist, G.E.; Kunz, R.; Falck-Ytter, Y.; Alonso-Coello, P.; Schünemann, H.J. GRADE: An emerging consensus on rating quality of evidence and strength of recommendations. BMJ 2008, 336, 924–926. [Google Scholar] [CrossRef]
- WHO. WHO Handbook for Guideline Development, 2nd ed.; World Health Organization: Geneva, Switzerland, 2014; p. 167. [Google Scholar]
- Hill, J.; Bullock, I.; Alderson, P. A summary of the methods that the National Clinical Guideline Centre uses to produce clinical guidelines for the National Institute for Health and Clinical Excellence. Ann. Intern. Med. 2011, 154, 752–757. [Google Scholar] [CrossRef]
- Qaseem, A.; Forland, F.; Macbeth, F.; Ollenschläger, G.; Phillips, S.; van der Wees, P. Guidelines International Network: Toward international standards for clinical practice guidelines. Ann. Intern. Med. 2012, 156, 525–531. [Google Scholar] [CrossRef]
- Rosenfeld, R.M.; Nnacheta, L.C.; Corrigan, M.D. Clinical consensus statement development manual. Otolaryngol. Head Neck Surg. 2015, 153, S1–S14. [Google Scholar] [CrossRef] [PubMed]
- De Boeck, K.; Castellani, C.; Elborn, J. Medical consensus, guidelines, and position papers: A policy for the ECFS. J. Cyst. Fibros. 2014, 13, 495–498. [Google Scholar] [CrossRef] [PubMed]
- Merriman, L.M.; Dale, J. Clinical Protocols. In Clinical Skills in Treating the Foot; Elsevier: Amsterdam, The Netherlands, 2005; pp. 37–52. [Google Scholar]
- Knowledge Management for Medical Care. Clinical Practical Guidelines. Available online: http://www.openclinical.org/guidelines.html (accessed on 6 April 2020).
- Haynes, A.B.; Weiser, T.G.; Berry, W.R.; Lipsitz, S.R.; Breizat, A.H.S.; Dellinger, E.P.; Herbosa, T.; Joseph, S.; Kibatala, P.L.; Lapitan, M.C.M.; et al. A surgical safety checklist to reduce morbidity and mortality in a global population. N. Engl. J. Med. 2009, 360, 491–499. [Google Scholar] [CrossRef]
- Grigg, E. Smarter clinical checklists: how to minimize checklist fatigue and maximize clinician performance. Anesth. Analg. 2015, 121, 570–573. [Google Scholar] [CrossRef]
- Hales, B.; Terblanche, M.; Fowler, R.; Sibbald, W. Development of medical checklists for improved quality of patient care. Int. J. Qual. Health Care 2008, 20, 22–30. [Google Scholar] [CrossRef]
- Greenfield, S. Clinical practice guidelines: Expanded use and misuse. JAMA 2017, 317, 594–595. [Google Scholar] [CrossRef] [PubMed]
- Vegting, I.L.; van Beneden, M.; Kramer, M.H.; Thijs, A.; Kostense, P.J.; Nanayakkara, P.W. How to save costs by reducing unnecessary testing: Lean thinking in clinical practice. Eur. J. Intern. Med. 2012, 23, 70–75. [Google Scholar] [CrossRef] [PubMed]
- Lenz, R.; Peleg, M.; Reichert, M. Healthcare Process Support: Achievements, Challenges, Current Research. Int. J. Knowl. Based Organ. 2012, 2, 784. [Google Scholar]
- Drummond, M. Clinical Guidelines: A NICE Way to Introduce Cost-Effectiveness Considerations? Value Health 2016, 19, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Burgers, J.; van der Weijden, T.; Grol, R. Clinical Practice Guidelines as a Tool for Improving Patient Care. In Improving Patient Care; Chapter 6; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2020; pp. 103–129. [Google Scholar]
- Prior, M.; Guerin, M.; Grimmer-Somers, K. The effectiveness of clinical guideline implementation strategies—A synthesis of systematic review findings. J. Eval. Clin. Pract. 2008, 14, 888–897. [Google Scholar] [CrossRef]
- Watts, C.G.; Dieng, M.; Morton, R.L.; Mann, G.J.; Menzies, S.W.; Cust, A.E. Clinical practice guidelines for identification, screening and follow-up of individuals at high risk of primary cutaneous melanoma: A systematic review. Br. J. Dermatol. 2015, 172, 33–47. [Google Scholar] [CrossRef]
- Woolf, S.; Schunemann, H.J.; Eccles, M.P.; Grimshaw, J.M.; Shekelle, P. Developing clinical practice guidelines: Types of evidence and outcomes; values and economics, synthesis, grading, and presentation and deriving recommendations. Implement. Sci. 2012, 7, 61. [Google Scholar] [CrossRef]
- Legido-Quigley, H.; Panteli, D.; Brusamento, S.; Knai, C.; Saliba, V.; Turk, E.; Sol, M.; Augustin, U.; Car, J.; McKee, M.; et al. Clinical guidelines in the European Union: Mapping the regulatory basis, development, quality control, implementation and evaluation across member states. Health Policy 2012, 107, 146–156. [Google Scholar] [CrossRef]
- Rashidian, A.; Eccles, M.P.; Russell, I. Falling on stony ground? A qualitative study of implementation of clinical guidelines’ prescribing recommendations in primary care. Health Policy 2008, 85, 148–161. [Google Scholar] [CrossRef]
- Yang, W.S.; Hwang, S.Y. A process-mining framework for the detection of healthcare fraud and abuse. Expert Syst. Appl. 2006, 31, 56–68. [Google Scholar] [CrossRef]
- Kose, I.; Gokturk, M.; Kilic, K. An interactive machine-learning-based electronic fraud and abuse detection system in healthcare insurance. Appl. Soft Comput. 2015, 36, 283–299. [Google Scholar] [CrossRef]
- Shortliffe, E.; Scott, A.; Bischoff, M.; Campbell, A.; Melle, W.; Jacobs, C. ONCOCIN: An Expert System for Oncology Protocol Management. In Proceedings of the 7th International Joint Conference on Artificial Intelligence (IJCAI ’81), Vancouver, BC, Canada, 24–28 August 1981; pp. 876–881. [Google Scholar]
- Pryor, T.A.; Gardner, R.M.; Clayton, P.D.; Warner, H.R. The HELP system. J. Med. Syst. 1983, 7, 87–102. [Google Scholar] [CrossRef] [PubMed]
- Shahar, Y.; Miksch, S.; Johnson, P. The Asgaard project: A task-specific framework for the application and critiquing of time-oriented clinical guidelines. Artif Intell. Med. 1998, 14, 29–51. [Google Scholar] [CrossRef]
- Boxwala, A.A.; Peleg, M.; Tu, S.; Ogunyemi, O.; Zeng, Q.T.; Wang, D.; Patel, V.L.; Greenes, R.A.; Shortliffe, E.H. GLIF3: A representation format for sharable computer-interpretable clinical practice guidelines. J. Biomed. Inform. 2004, 37, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Terenziani, P.; Molino, G.; Torchio, M. A modular approach for representing and executing clinical guidelines. Artif. Intell. Med. 2001, 23, 249–276. [Google Scholar] [CrossRef]
- Sutton, D.R.; Fox, J. The syntax and semantics of the PROforma guideline modeling language. J. Am. Med. Inform. Assoc. 2003, 10, 433–443. [Google Scholar] [CrossRef]
- Musen, M.A.; Tu, S.W.; Das, A.K.; Shahar, Y. EON: A component-based approach to automation of protocol-directed therapy. J. Am. Med. Inform. Assoc. 1996, 3, 367–388. [Google Scholar] [CrossRef]
- Ciccarese, P.; Caffi, E.; Quaglini, S.; Stefanelli, M. Architectures and tools for innovative Health Information Systems: The Guide Project. Int. J. Med. Inform. 2005, 74, 553–562. [Google Scholar] [CrossRef]
- Shiffman, R.N.; Greenes, R.A. Improving clinical guidelines with logic and decision-table techniques: Application to hepatitis immunization recommendations. Med. Decis. Mak. 1994, 14, 245–254. [Google Scholar] [CrossRef]
- Elkin, P.L.; Peleg, M.; Lacson, R.; Bernstam, E.; Tu, S.; Boxwala, A.; Greenes, R.; Shortliffe, E.H. Toward the standardization of electronic guidelines. MD Comput. 2000, 17, 39–44. [Google Scholar]
- Peleg, M.; Tu, S.; Bury, J.; Ciccarese, P.; Fox, J.; Greenes, R.; Hall, R.; Johnson, P.; Jones, N.; Kumar, A.; et al. Comparing computer-interpretable guideline models: A case-study approach. J. Am. Med. Inform. Assoc. 2003, 10, 52–68. [Google Scholar] [CrossRef] [PubMed]
- Sonnenberg, F.A.; Hagerty, C.G. Computer-interpretable clinical practice guidelines. Where are we and where are we going ? Yearb. Med. Inform. 2006, 45, 145–158. [Google Scholar]
- Kawamoto, K.; Greenes, R.A. Chapter 21—The Role of Standards: What We Can Expect and When. In Clinical Decision Support, 2nd ed.; Greenes, R.A., Ed.; Academic Press: Oxford, UK, 2014; pp. 599–615. [Google Scholar]
- Karadimas, H.; Ebrahiminia, V.; Lepage, E. User-defined functions in the Arden Syntax: An extension proposal. Artif. Intell. Med. 2018, 92, 103–110. [Google Scholar] [CrossRef]
- Kraus, S.; Toddenroth, D.; Unberath, P.; Prokosch, H.U.; Hueske-Kraus, D. An Extension of the Arden Syntax to Facilitate Clinical Document Generation. Stud. Health Technol. Inform. 2019, 259, 65–70. [Google Scholar] [PubMed]
- Jenders, R.A. Evaluation of the Fast Healthcare Interoperability Resources (FHIR) Standard for Representation of Knowledge Bases Encoded in the Arden Syntax. Stud. Health Technol. Inform. 2019, 264, 1692–1693. [Google Scholar] [PubMed]
- Sordo, M.; Ogunyemi, O.; Boxwala, A.A.; Greenes, R.A. GELLO: An object-oriented query and expression language for clinical decision support. AMIA Annu. Symp. Proc. 2003, 2003, 1012. [Google Scholar]
- Kawamoto, K.; Del Fiol, G.; Strasberg, H.R.; Hulse, N.; Curtis, C.; Cimino, J.J.; Rocha, B.H.; Maviglia, S.; Fry, E.; Scherpbier, H.J.; et al. Multi-National, Multi-Institutional Analysis of Clinical Decision Support Data Needs to Inform Development of the HL7 Virtual Medical Record Standard. AMIA Annu. Symp. Proc. 2010, 2010, 377–381. [Google Scholar]
- Peleg, M.; Keren, S.; Denekamp, Y. Mapping computerized clinical guidelines to electronic medical records: Knowledge-data ontological mapper (KDOM). J. Biomed. Inform. 2008, 41, 180–201. [Google Scholar] [CrossRef]
- German, E.; Leibowitz, A.; Shahar, Y. An architecture for linking medical decision-support applications to clinical databases and its evaluation. J. Biomed. Inform. 2009, 42, 203–218. [Google Scholar] [CrossRef][Green Version]
- Marcos, M.; Maldonado, J.A.; Martínez-Salvador, B.; Boscá, D.; Robles, M. Interoperability of clinical decision-support systems and electronic health records using archetypes: A case study in clinical trial eligibility. J. Biomed. Inform. 2013, 46, 676–689. [Google Scholar] [CrossRef]
- Marco-Ruiz, L.; Moner, D.; Maldonado, J.A.; Kolstrup, N.; Bellika, J.G. Archetype-based data warehouse environment to enable the reuse of electronic health record data. Int. J. Med Inform. 2015, 84, 702–714. [Google Scholar] [CrossRef] [PubMed]
- Gatta, R.; Vallati, M.; Cappelli, C.; De Bari, B.; Salvetti, M.; Finardi, S.; Muiesan, M.; Valentini, V.; Castellano, M. Bridging the Gap between Knowledge Representation and Electronic Health Records. In Proceedings of the 9th International Conference on Health Informatics, Rome, Italy, 21–23 February 2016; pp. 159–165. [Google Scholar]
- Gooch, P.; Roudsari, A. Computerization of workflows, guidelines, and care pathways: A review of implementation challenges for process-oriented health information systems. J. Am. Med. Inform. Assoc. 2011, 18, 738–748. [Google Scholar] [CrossRef] [PubMed]
- Quaglini, S.; Stefanelli, M.; Cavallini, A.; Micieli, G.; Fassino, C.; Mossa, C. Guideline-based careflow systems. Artif. Intell. Med. 2000, 20, 5–22. [Google Scholar] [CrossRef]
- Panzarasa, S.; Bellazzi, R.; Larizza, C.; Stefanelli, M. A Careflow Management System for Chronic Patients. In Studies in Health Technology and Informatics; MedInfo; Fieschi, M., Coiera, E.W., Li, J.Y.C., Eds.; IOS Press: Amsterdam, The Netherlands, 2004; Volume 107, pp. 673–677. [Google Scholar]
- Fox, J.; Black, E.; Chronakis, I.; Dunlop, R.; Patkar, V.; South, M.; Thomson, R. From guidelines to careflows: modelling and supporting complex clinical processes. In Studies in Health Technology and Informatics; IOS Press: Amsterdam, The Netherlands, 2008; Volume 139, pp. 44–62. [Google Scholar]
- Schadow, G.; Russler, D.C.; McDonald, C.J. Conceptual alignment of electronic health record data with guideline and workflow knowledge. Int. J. Med. Inform. 2001, 64, 259–274. [Google Scholar] [CrossRef]
- Gonzalez-Ferrer, A.; ten Teije, A.; Fdez-Olivares, J.; Milian, K. Automated generation of patient-tailored electronic care pathways by translating computer-interpretable guidelines into hierarchical task networks. Artif. Intell. Med. 2013, 57, 91–109. [Google Scholar] [CrossRef]
- Shabo, A.; Peleg, M.; Parimbelli, E.; Quaglini, S.; Napolitano, C. Interplay between Clinical Guidelines and Organizational Workflow Systems. Experience from the MobiGuide Project. Methods Inf. Med. 2016, 55, 488–494. [Google Scholar] [CrossRef]
- Mulyar, N.; van der Aalst, W.; Peleg, M. A pattern-based analysis of clinical computer-interpretable guideline modeling languages. J. Am. Med. Inform. Assoc. 2007, 14, 781–787. [Google Scholar] [CrossRef]
- Grando, M.A.; Glasspool, D.; Fox, J. A formal approach to the analysis of clinical computer-interpretable guideline modeling languages. Artif. Intell. Med. 2012, 54, 1–13. [Google Scholar] [CrossRef]
- Kaiser, K.; Marcos, M. Leveraging workflow control patterns in the domain of clinical practice guidelines. BMC Med. Inform. Decis. Mak. 2016, 16, 20. [Google Scholar] [CrossRef]
- Martínez-Salvador, B.; Marcos, M. Supporting the Refinement of Clinical Process Models to Computer-Interpretable Guideline Models. Bus. Inf. Syst. Eng. 2016, 58, 355–366. [Google Scholar] [CrossRef]
- Peleg, M.; González-Ferrer, A. Chapter 16—Guidelines and Workflow Models. In Clinical Decision Support (Second Edition), 2nd ed.; Academic Press: Oxford, UK, 2014; pp. 435–464. [Google Scholar]
- Object Management Group (OMG). Decision Model and Notation Version 1.0. 2015. Available online: https://www.omg.org/spec/DMN/1.0 (accessed on 15 July 2020).
- Kaymak, U.; Mans, R.; van de Steeg, T.; Dierks, M. On Process Mining in Health Care. In Proceedings of the 2012 IEEE International Conference on Systems, Man, and Cybernetics (SMC), Seoul, Korea, 14–17 October 2012; pp. 1859–1864. [Google Scholar]
- Ghasemi, M.; Amyot, D. Process mining in healthcare: A systematised literature review. Int. J. Electron. Healthc. 2016, 9, 60. [Google Scholar] [CrossRef]
- Rebuge, Á.; Ferreira, D.R. Business process analysis in healthcare environments: A methodology based on process mining. Inf. Syst. 2012, 37, 99–116. [Google Scholar] [CrossRef]
- Mans, R.; van der Aalst, W.; Vanwersch, R.; Moleman, A. Process Mining in Healthcare: Data Challenges when Answering Frequently Posed Questions. In Process Support and Knowledge Representation in Health Care (BPM 2012 Joint Workshop, ProHealth 2012/KR4HC 2012, Tallinn, Estonia, 3 September 2012, Revised Selected Papers); Springer: Berlin/Heidelberg, Germany, 2013; pp. 140–153. [Google Scholar]
- Rovani, M.; Maggi, F.M.; de Leoni, M.; van der Aalst, W. Declarative Process Mining in Healthcare. Expert Syst. Appl. 2015, 42, 9236–9251. [Google Scholar] [CrossRef]
- Pesic, M.; van der Aalst, W. A Declarative Approach for Flexible Business Processes Management. In Business Process Management Workshops; Springer: Berlin/Heidelberg, Germany, 2006. [Google Scholar]
- Van Dongen, B.F.; de Medeiros, A.K.A.; Verbeek, H.; Weijters, A.; van der Aalst, W. The ProM Framework: A New Era in Process Mining Tool Support. In International Conference on Application and Theory of Petri Nets; Springer: Berlin/Heidelberg, Germany, 2005; pp. 444–454. [Google Scholar]
- Rojas, E.; Munoz-Gama, J.; Sepulveda, M.; Capurro, D. Process Mining in Healthcare: A literature review. J. Biomed. Inform. 2016, 61, 224–236. [Google Scholar] [CrossRef]
- Kurniati, A.P.; Johnson, O.; Hogg, D.; Hall, G. Process Mining in Oncology: A Literature Review. In Proceedings of the 2016 6th International Conference on Information Communication and Management (ICICM), Hatfield, UK, 29–31 October 2016; pp. 291–297. [Google Scholar]
- Johnson, O.A.; Dhafari, T.B.; Kurniati, A.; Fox, F.; Cordoba, E.R. The ClearPath Method for Care Pathway Process Mining and Simulation. In Business Process Management Workshops; Springer: Berlin/Heidelberg, Germany, 2018. [Google Scholar]
- Binder, M.; Dorda, W.; Duftschmid, G.; Dunkl, R.; Froeschl, K.; Gall, W.; Grossmann, W.; Harmankaya, K.; Hronsky, M.; Rinderle-Ma, S.; et al. On Analyzing Process Compliance in Skin Cancer Treatment: An Experience Report from the Evidence-Based Medical Compliance Cluster (EBMC2). In Advanced Information Systems Engineering; CAiSE 2012; Springer: Berlin/Heidelberg, Germany, 2012; pp. 398–413. [Google Scholar]
- Geleijnse, G.; Aklecha, H.; Vroling, M.; Verhoeven, R.; van Erning, F.N.; Vissers, P.A.; Buijs, J.C.A.M.; Verbeek, X.A. Using Process Mining to Evaluate Colon Cancer Guideline Adherence with Cancer Registry Data: A Case Study. In Proceedings of the AMIA 2018, American Medical Informatics Association Annual Symposium, San Francisco, CA, USA, 3–7 November 2018; AMIA: Bethesda, MD, USA, 2018. [Google Scholar]
- Lenkowicz, J.; Gatta, R.; Masciocchi, C.; Casà, C.; Cellini, F.; Damiani, A.; Dinapoli, N.; Valentini, V. Assessing the conformity to clinical guidelines in oncology: An example for the multidisciplinary management of locally advanced colorectal cancer treatment. Manag. Decis. 2018, 56, 2172–2186. [Google Scholar] [CrossRef]
- Fernández-Llatas, C.; Benedi, J.M.; García-Gómez, J.M.; Traver, V. Process mining for individualized behavior modeling using wireless tracking in nursing homes. Sensors 2013, 13, 15434–15451. [Google Scholar] [CrossRef]
- Valero-Ramon, Z.; Ibanez-Sanchez, G.; Traver, V.; Marco-Ruiz, L.; Fernandez-Lllatas, C. Towards Perceptual Spaces for Empowering Ergonomy in Workplaces by Using Interactive Process Mining. Transform. Ergon. Pers. Health Intell. Work. 2019, 25, 85. [Google Scholar]
- Fernandez-Llatas, C.; Valdivieso, B.; Traver, V.; Benedi, J.M. Using Process Mining for Automatic Support of Clinical Pathways Design. In Data Mining in Clinical Medicine; Springer: Berlin/Heidelberg, Germany, 2015; pp. 79–88. [Google Scholar]
- Porter, M.E.; Teisberg, E.O. Redefining Health Care: Creating Value-Based Competition on Results; Harvard Business Press: Boston, MA, USA, 2006. [Google Scholar]
- Valero-Ramon, Z.; Fernandez-Llatas, C.; Martinez-Millana, A.; Traver, V. Interactive Process Indicators for Obesity Modelling Using Process Mining. In Advanced Computational Intelligence in Healthcare-7; Springer: Berlin/Heidelberg, Germany, 2020; pp. 45–64. [Google Scholar]
- Qu, G.; Liu, Z.; Cui, S.; Tang, J. Study on Self-Adaptive Clinical Pathway Decision Support System Based on Case-Based Reasoning. Lect. Notes Electr. Eng. 2014, 269, 969–978. [Google Scholar]
- Van de Velde, S.; Roshanov, P.; Kortteisto, T.; Kunnamo, I.; Aertgeerts, B.; Vandvik, P.O.; Flottorp, S. Tailoring implementation strategies for evidence-based recommendations using computerised clinical decision support systems: Protocol for the development of the GUIDES tools. Implement. Sci. 2016, 11, 29. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gatta, R.; Vallati, M.; Fernandez-Llatas, C.; Martinez-Millana, A.; Orini, S.; Sacchi, L.; Lenkowicz, J.; Marcos, M.; Munoz-Gama, J.; Cuendet, M.A.; et al. What Role Can Process Mining Play in Recurrent Clinical Guidelines Issues? A Position Paper. Int. J. Environ. Res. Public Health 2020, 17, 6616. https://doi.org/10.3390/ijerph17186616
Gatta R, Vallati M, Fernandez-Llatas C, Martinez-Millana A, Orini S, Sacchi L, Lenkowicz J, Marcos M, Munoz-Gama J, Cuendet MA, et al. What Role Can Process Mining Play in Recurrent Clinical Guidelines Issues? A Position Paper. International Journal of Environmental Research and Public Health. 2020; 17(18):6616. https://doi.org/10.3390/ijerph17186616
Chicago/Turabian StyleGatta, Roberto, Mauro Vallati, Carlos Fernandez-Llatas, Antonio Martinez-Millana, Stefania Orini, Lucia Sacchi, Jacopo Lenkowicz, Mar Marcos, Jorge Munoz-Gama, Michel A. Cuendet, and et al. 2020. "What Role Can Process Mining Play in Recurrent Clinical Guidelines Issues? A Position Paper" International Journal of Environmental Research and Public Health 17, no. 18: 6616. https://doi.org/10.3390/ijerph17186616
APA StyleGatta, R., Vallati, M., Fernandez-Llatas, C., Martinez-Millana, A., Orini, S., Sacchi, L., Lenkowicz, J., Marcos, M., Munoz-Gama, J., Cuendet, M. A., de Bari, B., Marco-Ruiz, L., Stefanini, A., Valero-Ramon, Z., Michielin, O., Lapinskas, T., Montvila, A., Martin, N., Tavazzi, E., & Castellano, M. (2020). What Role Can Process Mining Play in Recurrent Clinical Guidelines Issues? A Position Paper. International Journal of Environmental Research and Public Health, 17(18), 6616. https://doi.org/10.3390/ijerph17186616










