Convolution- and Attention-Based Neural Network for Automated Sleep Stage Classification
Abstract
1. Introduction
- A neural network based on convolution and attention mechanism is built. The network uses a CNN to extract local signal features and multilayer attention networks to learn intra- and inter-epoch features. The recursive architecture is completely deprecated in our model.
- For the unbalanced dataset, the proposed method uses a weighted loss function during training to improve model performance on minority classes.
- The model outperforms other methods on sleep-edf and sleep-edfx datasets utilizing various training and testing set partitioning methods without changing the model’s structure or any of its parameters.
2. Materials and Methods
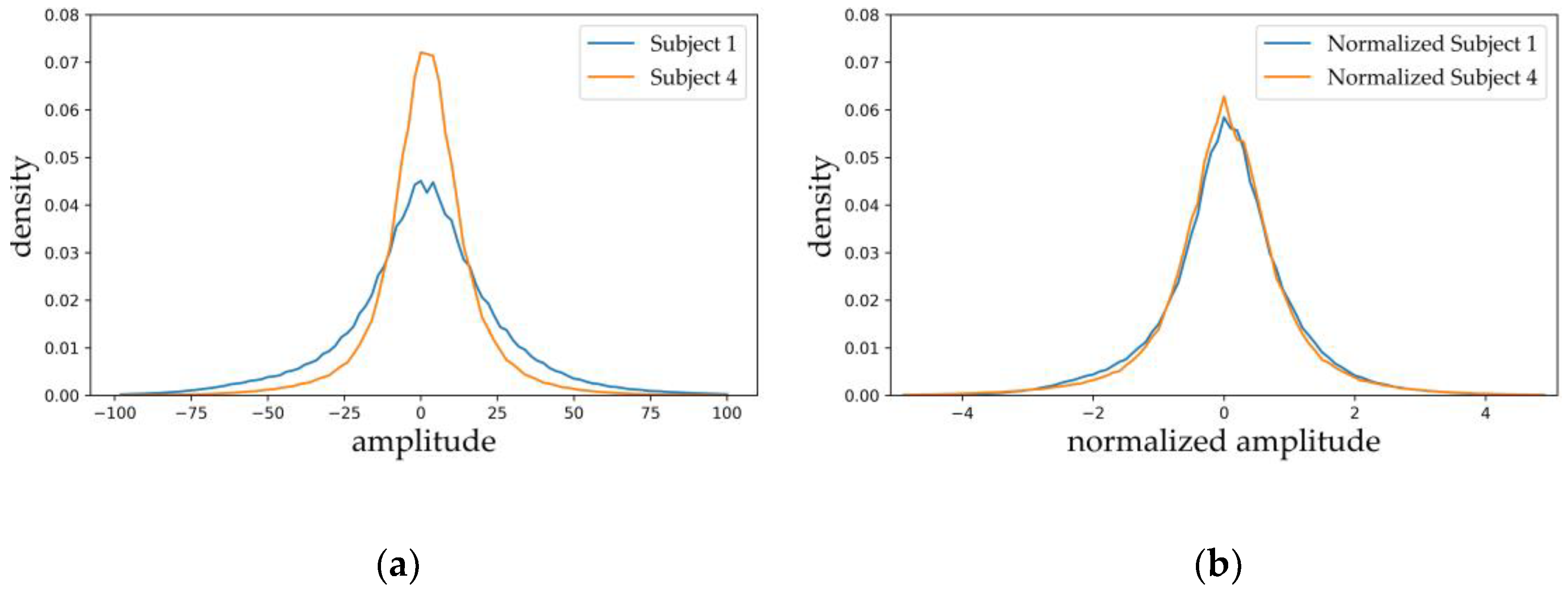
2.1. Dataset and Preprocessing
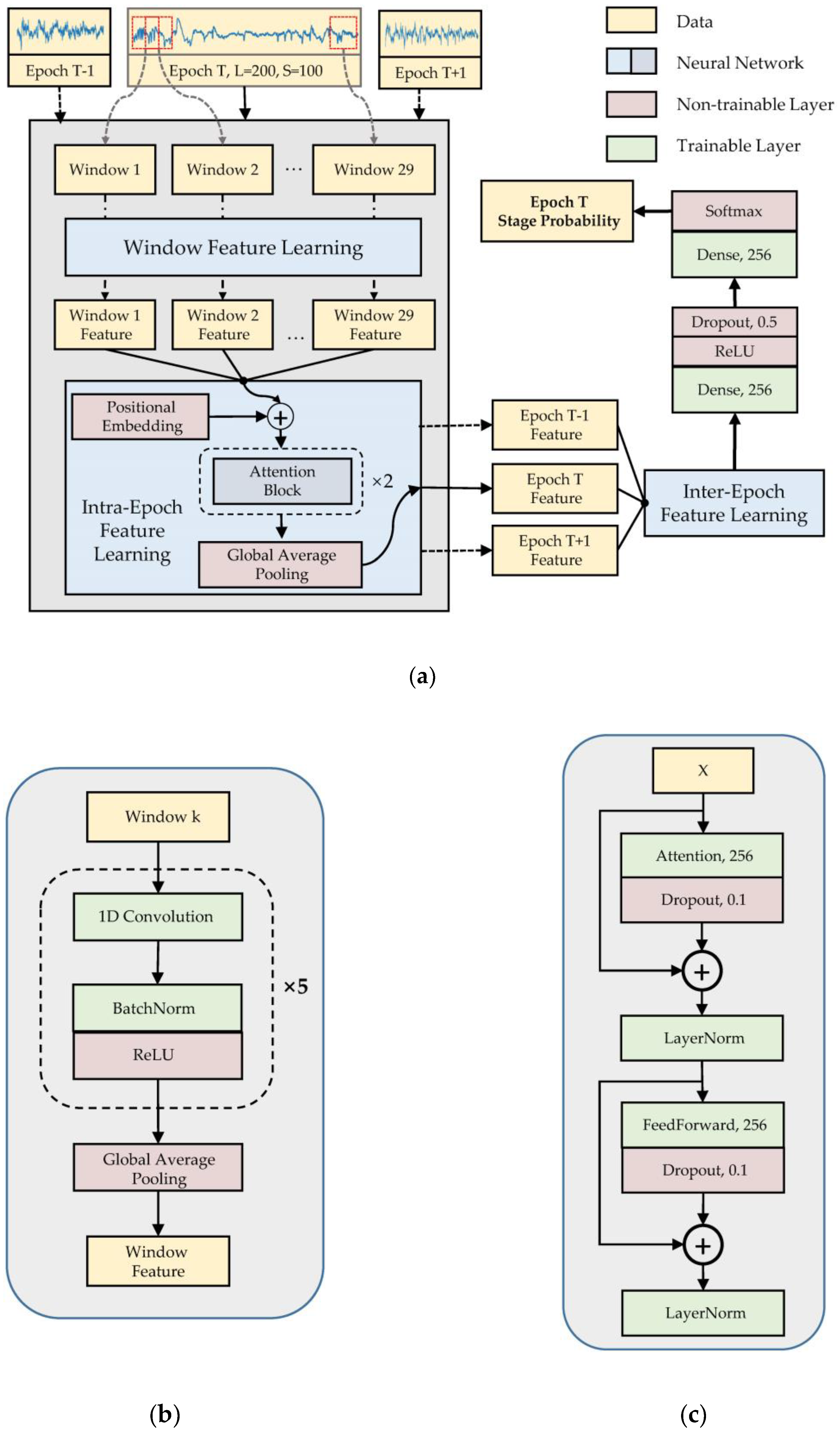
2.2. Model Architecture
2.3. Training and Testing
2.3.1. Training Parameters
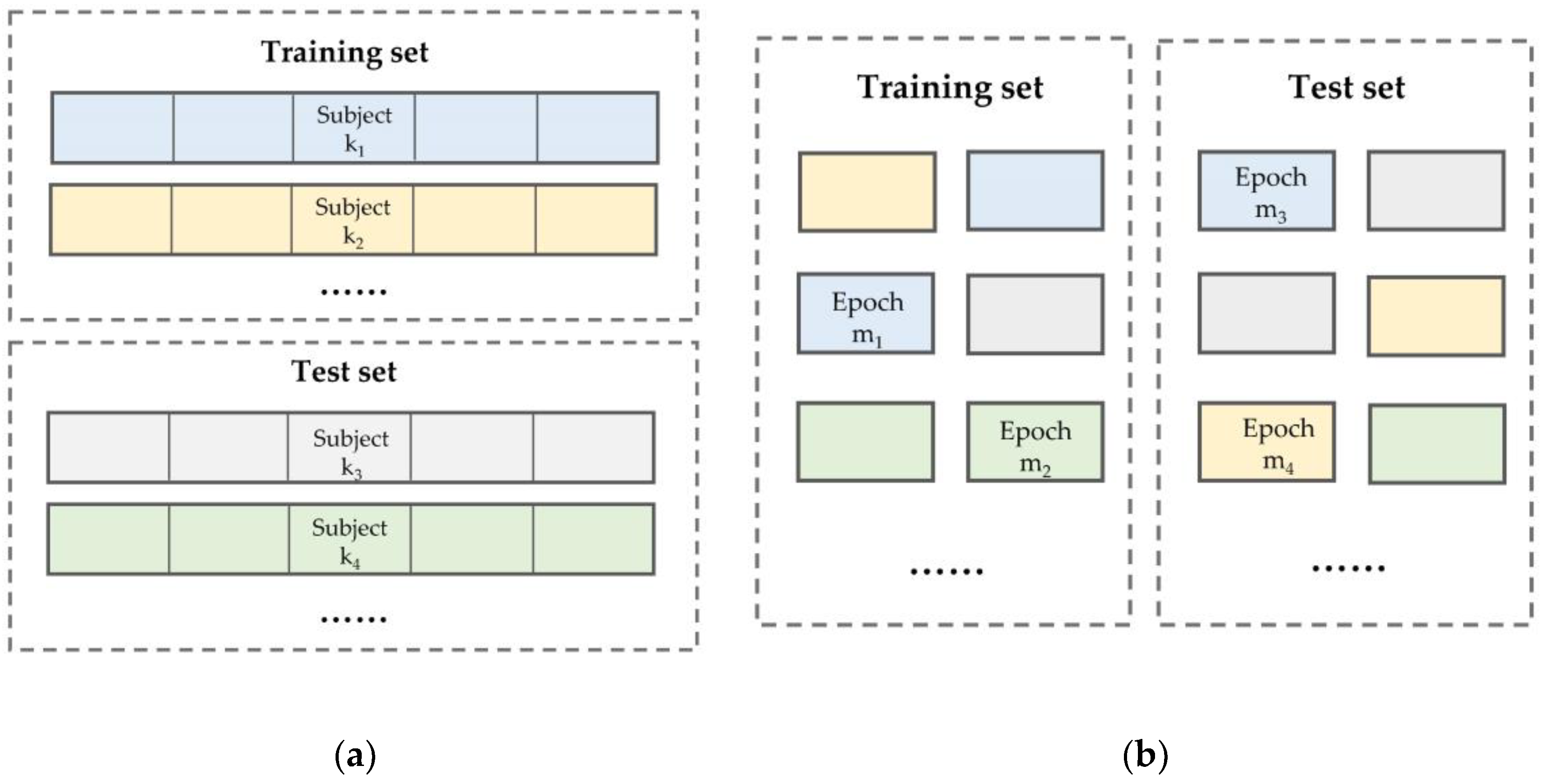
2.3.2. Testing Method
2.3.3. Performance Metrics
3. Results
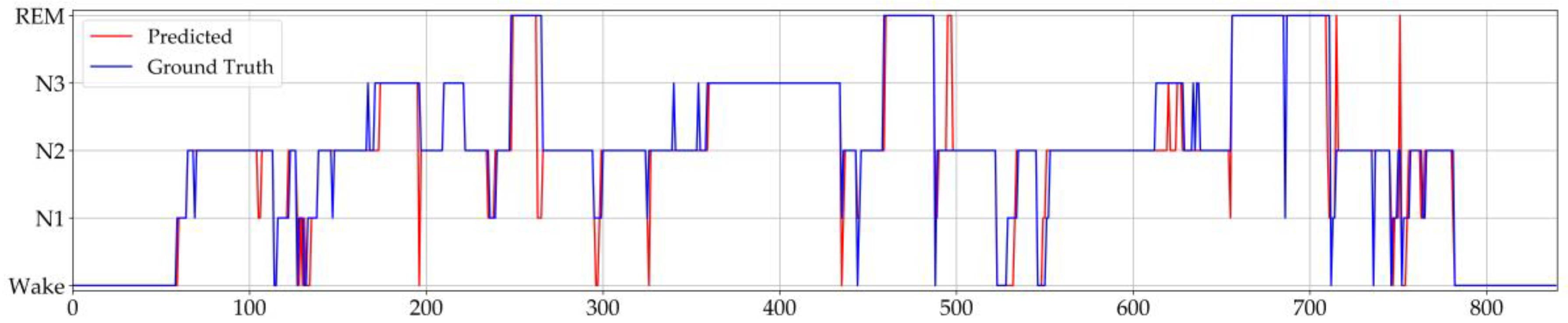
3.1. Model Performance
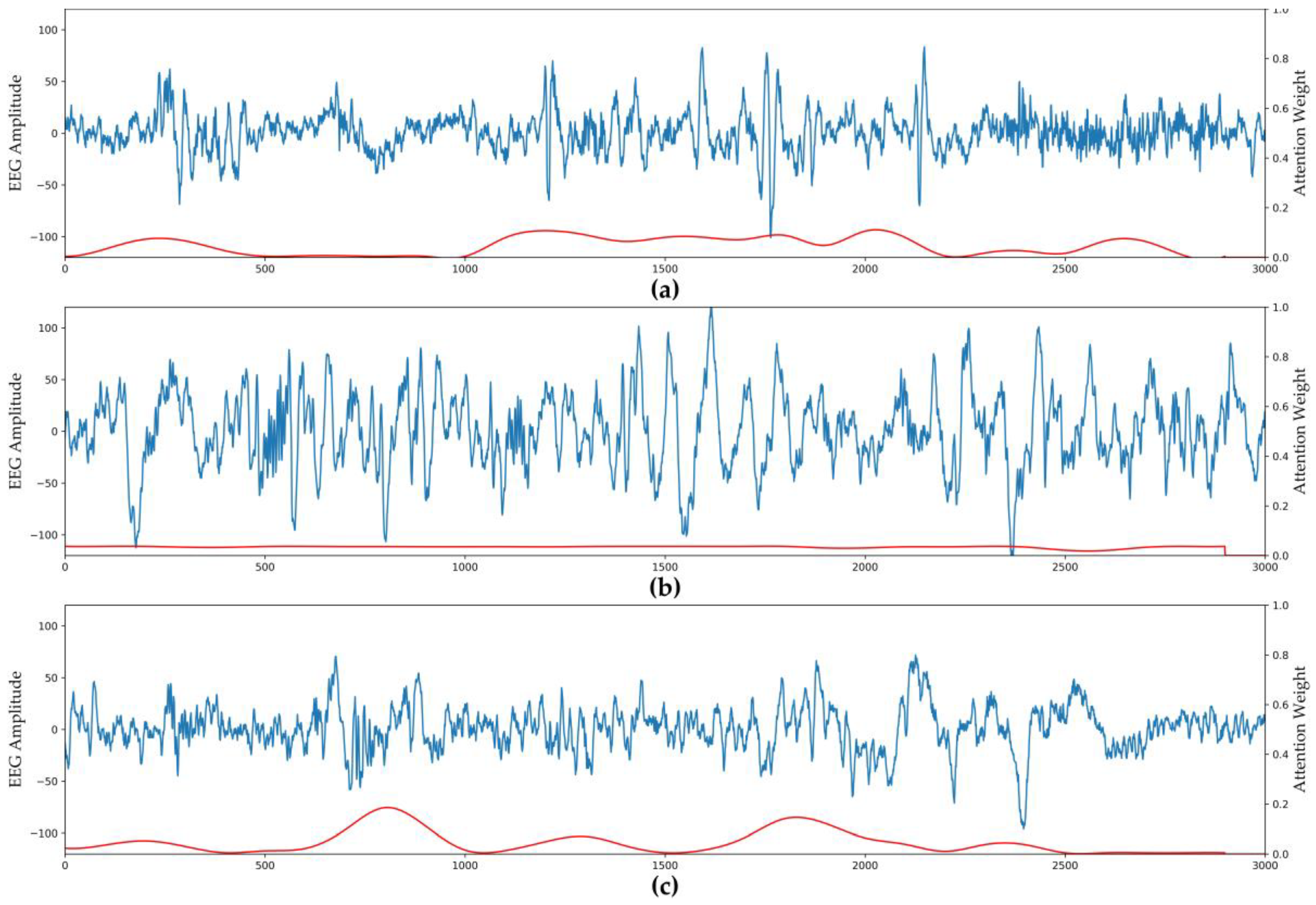
3.2. Visualization of Attention Weights
3.3. Ablation Analysis of Model Components
3.4. Comparison with Other Methods
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Bjørnarå, K.A.; Dietrichs, E.; Toft, M. Longitudinal Assessment of Probable Rapid Eye Movement Sleep Behaviour Disorder in Parkinson’s Disease. Eur. J. Neurol. 2015, 22, 1242–1244. [Google Scholar] [CrossRef] [PubMed]
- Zhong, G.; Naismith, S.L.; Rogers, N.L.; Lewis, S.J.G. Sleep–Wake Disturbances in Common Neurodegenerative Diseases: A Closer Look at Selected Aspects of the Neural Circuitry. J. Neurol. Sci. 2011, 307, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Buysse, D.J. Sleep Health: Can We Define It? Does It Matter? Sleep 2014, 37, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Wolpert, E.A. A Manual of Standardized Terminology, Techniques and Scoring System for Sleep Stages of Human Subjects. Arch. Gen. Psychiatry 1969, 20, 246–247. [Google Scholar] [CrossRef]
- Iber, C.; Ancoli-Israel, S.; Chesson, A.L.; Quan, S.F. The AASM Manual for the Scoring of Sleep and Associated Events: Rules, Terminology and Technical Specification; American Academy of Sleep Medicine: Westchester, NY, USA, 2007. [Google Scholar]
- Fiorillo, L.; Puiatti, A.; Papandrea, M.; Ratti, P.-L.; Favaro, P.; Roth, C.; Bargiotas, P.; Bassetti, C.L.; Faraci, F.D. Automated Sleep Scoring: A Review of the Latest Approaches. Sleep Med. Rev. 2019, 48, 101204. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adam, M.; Gertych, A.; Tan, R.S. A Deep Convolutional Neural Network Model to Classify Heartbeats. Comput. Biol. Med. 2017, 89, 389–396. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.-Z.; Ni, D.; Chou, Y.-H.; Qin, J.; Tiu, C.-M.; Chang, Y.-C.; Huang, C.-S.; Shen, D.; Chen, C.-M. Computer-Aided Diagnosis with Deep Learning Architecture: Applications to Breast Lesions in US Images and Pulmonary Nodules in CT Scans. Sci. Rep. 2016, 6, 24454. [Google Scholar] [CrossRef] [PubMed]
- Talo, M.; Baloglu, U.B.; Yıldırım, Ö.; Rajendra Acharya, U. Application of Deep Transfer Learning for Automated Brain Abnormality Classification Using MR Images. Cogn. Syst. Res. 2019, 54, 176–188. [Google Scholar] [CrossRef]
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adeli, H. Deep Convolutional Neural Network for the Automated Detection and Diagnosis of Seizure Using EEG Signals. Comput. Biol. Med. 2018, 100, 270–278. [Google Scholar] [CrossRef] [PubMed]
- Voets, M.; Møllersen, K.; Bongo, L.A. Reproduction Study Using Public Data of: Development and Validation of a Deep Learning Algorithm for Detection of Diabetic Retinopathy in Retinal Fundus Photographs. PLoS ONE 2019, 14, e0217541. [Google Scholar] [CrossRef] [PubMed]
- Svetnik, V.; Ma, J.; Soper, K.A.; Doran, S.; Renger, J.J.; Deacon, S.; Koblan, K.S. Evaluation of Automated and Semi-Automated Scoring of Polysomnographic Recordings from a Clinical Trial Using Zolpidem in the Treatment of Insomnia. Sleep 2007, 30, 1562–1574. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Macaš, M.; Grimová, N.; Gerla, V.; Lhotská, L. Semi-Automated Sleep EEG Scoring with Active Learning and HMM-Based Deletion of Ambiguous Instances. Proceedings 2019, 31, 46. [Google Scholar] [CrossRef]
- Chambon, S.; Galtier, M.N.; Arnal, P.J.; Wainrib, G.; Gramfort, A. A Deep Learning Architecture for Temporal Sleep Stage Classification Using Multivariate and Multimodal Time Series. IEEE Trans. Neural Syst. Rehabil. Eng. 2018, 26, 758–769. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Goyal, D.; Achuth, P.V.; Acharya, U.R. An Accurate Sleep Stages Classification System Using a New Class of Optimally Time-Frequency Localized Three-Band Wavelet Filter Bank. Comput. Biol. Med. 2018, 98, 58–75. [Google Scholar] [CrossRef] [PubMed]
- Liang, S.-F.; Kuo, C.-E.; Hu, Y.-H.; Pan, Y.-H.; Wang, Y.-H. Automatic Stage Scoring of Single-Channel Sleep EEG by Using Multiscale Entropy and Autoregressive Models. IEEE Trans. Instrum. Meas. 2012, 61, 1649–1657. [Google Scholar] [CrossRef]
- Tsinalis, O.; Matthews, P.M.; Guo, Y. Automatic Sleep Stage Scoring Using Time-Frequency Analysis and Stacked Sparse Autoencoders. Ann. Biomed. Eng. 2016, 44, 1587–1597. [Google Scholar] [CrossRef] [PubMed]
- Kemp, B.; Zwinderman, A.H.; Tuk, B.; Kamphuisen, H.A.C.; Oberye, J.J.L. Analysis of a Sleep-Dependent Neuronal Feedback Loop: The Slow-Wave Microcontinuity of the EEG. IEEE Trans. Biomed. Eng. 2000, 47, 1185–1194. [Google Scholar] [CrossRef] [PubMed]
- Hassan, A.R.; Subasi, A. A Decision Support System for Automated Identification of Sleep Stages from Single-Channel EEG Signals. Knowl.-Based Syst. 2017, 128, 115–124. [Google Scholar] [CrossRef]
- Jiang, D.; Lu, Y.; Ma, Y.; Wang, Y. Robust Sleep Stage Classification with Single-Channel EEG Signals Using Multimodal Decomposition and HMM-Based Refinement. Expert Syst. Appl. 2019, 121, 188–203. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep Learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Schmidhuber, J. Deep Learning in Neural Networks: An Overview. Neural Netw. 2015, 61, 85–117. [Google Scholar] [CrossRef] [PubMed]
- LeCun, Y.; Boser, B.; Denker, J.S.; Henderson, D.; Howard, R.E.; Hubbard, W.; Jackel, L.D. Backpropagation Applied to Handwritten Zip Code Recognition. Neural Comput. 1989, 1, 541–551. [Google Scholar] [CrossRef]
- Andreotti, F.; Phan, H.; Cooray, N.; Lo, C.; Hu, M.T.M.; De Vos, M. Multichannel Sleep Stage Classification and Transfer Learning using Convolutional Neural Networks. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 18–21 July 2018; pp. 171–174. [Google Scholar] [CrossRef]
- Yildirim, O.; Baloglu, U.; Acharya, U. A Deep Learning Model for Automated Sleep Stages Classification Using PSG Signals. Int. J. Environ. Res. Public Health 2019, 16, 599. [Google Scholar] [CrossRef] [PubMed]
- Phan, H.; Andreotti, F.; Cooray, N.; Chen, O.Y.; De Vos, M. DNN Filter Bank Improves 1-Max Pooling CNN for Single-Channel EEG Automatic Sleep Stage Classification. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 18–21 July 2018; pp. 453–456. [Google Scholar] [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long Short-Term Memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef] [PubMed]
- Michielli, N.; Acharya, U.R.; Molinari, F. Cascaded LSTM Recurrent Neural Network for Automated Sleep Stage Classification Using Single-Channel EEG Signals. Comput. Biol. Med. 2019, 106, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Supratak, A.; Dong, H.; Wu, C.; Guo, Y. DeepSleepNet: A Model for Automatic Sleep Stage Scoring Based on Raw Single-Channel EEG. IEEE Trans. Neural Syst. Rehabil. Eng. 2017, 25, 1998–2008. [Google Scholar] [CrossRef] [PubMed]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention Is All You Need. In Advances in Neural Information Processing Systems 30; Guyon, I., Luxburg, U.V., Bengio, S., Wallach, H., Fergus, R., Vishwanathan, S., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2017; pp. 5998–6008. [Google Scholar]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a New Research Resource for Complex Physiologic Signals. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef] [PubMed]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going Deeper with Convolutions. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. arXiv 2015, arXiv:1502.03167. [Google Scholar]
- Vinod, N.; Geoffrey, E.H. Rectified Linear Units Improve Restricted Boltzmann Machines. In Proceedings of the 27th International Conference on Machine Learning (ICML-10), Haifa, Israel, 21–24 June 2010; pp. 807–814. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems 25; Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2012; pp. 1097–1105. [Google Scholar]
- Ba, J.L.; Kiros, J.R.; Hinton, G.E. Layer Normalization. arXiv 2016, arXiv:1607.06450. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. arXiv 2017, arXiv:1412.6980. [Google Scholar]
- Zhang, M.R.; Lucas, J.; Hinton, G.; Ba, J. Lookahead Optimizer: K Steps Forward, 1 Step Back. arXiv 2019, arXiv:1907.08610. [Google Scholar]
- Danker-Hopfe, H.; Anderer, P.; Zeitlhofer, J.; Boeck, M.; Dorn, H.; Gruber, G.; Heller, E.; Loretz, E.; Moser, D.; Parapatics, S.; et al. Interrater Reliability for Sleep Scoring According to the Rechtschaffen & Kales and the New AASM Standard. J. Sleep Res. 2009, 18, 74–84. [Google Scholar] [CrossRef] [PubMed]
- Tsinalis, O.; Matthews, P.M.; Guo, Y.; Zafeiriou, S. Automatic Sleep Stage Scoring with Single-Channel EEG Using Convolutional Neural Networks. arXiv 2016, arXiv:1610.01683. [Google Scholar]
- Sharma, R.; Pachori, R.B.; Upadhyay, A. Automatic Sleep Stages Classification Based on Iterative Filtering of Electroencephalogram Signals. Neural Comput. Appl. 2017, 28, 2959–2978. [Google Scholar] [CrossRef]

| Dataset | W | N1 | N2 | N3 | REM | Total |
|---|---|---|---|---|---|---|
| Sleep-edfx | 8246 | 2804 | 17,799 | 5703 | 7717 | 42,269 |
| Sleep-edf | 8055 | 604 | 3621 | 1299 | 1609 | 15,188 |
| Module | Number of Filters | Kernel Size | Stride | Output Shape |
|---|---|---|---|---|
| Input | - | - | - | (200, 1) |
| Conv_1 | 64 | 5 | 3 | (66, 64) |
| Conv_2 | 64 | 5 | 3 | (21, 64) |
| Conv_3 | 128 | 3 | 2 | (10, 128) |
| Conv_4 | 128 | 3 | 1 | (8, 128) |
| Conv_5 | 256 | 3 | 1 | (6, 256) |
| GAP | - | - | - | (1, 256) |
| Stage | Predictions | Per-Class Metrics | ||||||
|---|---|---|---|---|---|---|---|---|
| Wake | N1 | N2 | N3 | REM | Precision | Recall | F1 | |
| W | 2388 | 33 | 6 | 1 | 5 | 99.1 | 98.2 | 98.6 |
| N1 | 15 | 83 | 25 | 1 | 35 | 52.9 | 52.2 | 52.5 |
| N2 | 2 | 28 | 1024 | 49 | 16 | 92.7 | 91.5 | 92.1 |
| N3 | 2 | 0 | 44 | 336 | 1 | 86.8 | 87.7 | 87.3 |
| REM | 2 | 13 | 6 | 0 | 437 | 88.5 | 95.4 | 91.8 |
| Stage | Predictions | Per-Class Metrics | ||||||
|---|---|---|---|---|---|---|---|---|
| W | N1 | N2 | N3 | REM | Precision | Recall | F1 | |
| W | 7287 | 586 | 89 | 57 | 149 | 91.5 | 89.2 | 90.3 |
| N1 | 279 | 1497 | 434 | 24 | 570 | 42.1 | 53.4 | 47.1 |
| N2 | 259 | 846 | 14,596 | 1388 | 710 | 90.5 | 82.1 | 86.0 |
| N3 | 39 | 31 | 586 | 5042 | 5 | 76.6 | 88.4 | 82.1 |
| REM | 103 | 598 | 422 | 69 | 6525 | 82.0 | 84.6 | 83.2 |
| Window Feature | Intra-Epoch Attention | Inter-Epoch Attention | Weighted Loss Function | Overall Performance | |||
|---|---|---|---|---|---|---|---|
| Subject-Wise | Epoch-Wise | ||||||
| Accuracy | MF1 | Accuracy | MF1 | ||||
| √ | √ | √ | √ | 82.8 | 77.8 | 93.7 | 84.5 |
| × | √ | √ | √ | 76.7 | 70.5 | 83.5 | 68.2 |
| √ | × | √ | √ | 81.3 | 76.3 | 92.3 | 82.2 |
| √ | √ | × | √ | 82.0 | 76.9 | 93.1 | 83.7 |
| √ | √ | √ | × | 82.8 | 75.8 | 93.8 | 84.1 |
| Methods | Samples | Per-Class F1-Score | Overall Performances | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Wake | N1 | N2 | N3 | REM | Accuracy | MF1 | |||
| Sleep-edfx | Ref. [17] | 37,022 | 71.6 | 47.0 | 84.6 | 84.0 | 81.4 | 78.9 | 73.7 |
| Ref. [40] | 37,022 | 65.4 | 43.7 | 80.6 | 84.9 | 74.5 | 74.9 | 69.8 | |
| Ref. [29] | 41,950 | 84.7 | 46.6 | 85.9 | 84.8 | 82.4 | 82.0 | 76.9 | |
| Ref. [26] | 46,236 | 89.8 | 33.2 | 86.7 | 86.0 | 82.6 | 82.6 | 74.2 | |
| Proposed | 42,269 | 90.3 | 47.1 | 86.0 | 82.1 | 83.2 | 82.8 | 77.8 | |
| Sleep-edf | Ref. [19] | 15,188 | 96.9 | 49.1 | 88.9 | 84.2 | 81.2 | 90.8 | 80.1 |
| Ref. [41] | 15,136 | 97.8 | 30.4 | 89.0 | 85.5 | 82.5 | 91.3 | 77.0 | |
| Ref. [25] | 15,188 | 97.5 | 24.8 | 89.4 | 87.0 | 80.8 | 91.2 | 75.9 | |
| Proposed | 15,188 | 98.6 | 52.5 | 92.1 | 87.2 | 91.8 | 93.7 | 84.5 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, T.; Luo, W.; Yu, F. Convolution- and Attention-Based Neural Network for Automated Sleep Stage Classification. Int. J. Environ. Res. Public Health 2020, 17, 4152. https://doi.org/10.3390/ijerph17114152
Zhu T, Luo W, Yu F. Convolution- and Attention-Based Neural Network for Automated Sleep Stage Classification. International Journal of Environmental Research and Public Health. 2020; 17(11):4152. https://doi.org/10.3390/ijerph17114152
Chicago/Turabian StyleZhu, Tianqi, Wei Luo, and Feng Yu. 2020. "Convolution- and Attention-Based Neural Network for Automated Sleep Stage Classification" International Journal of Environmental Research and Public Health 17, no. 11: 4152. https://doi.org/10.3390/ijerph17114152
APA StyleZhu, T., Luo, W., & Yu, F. (2020). Convolution- and Attention-Based Neural Network for Automated Sleep Stage Classification. International Journal of Environmental Research and Public Health, 17(11), 4152. https://doi.org/10.3390/ijerph17114152





