Comparison of Word Embeddings for Extraction from Medical Records
Abstract
:1. Introduction
2. Materials and Methods
2.1. Data
2.1.1. Labelled Dataset Gathering
2.1.2. Embedding Models
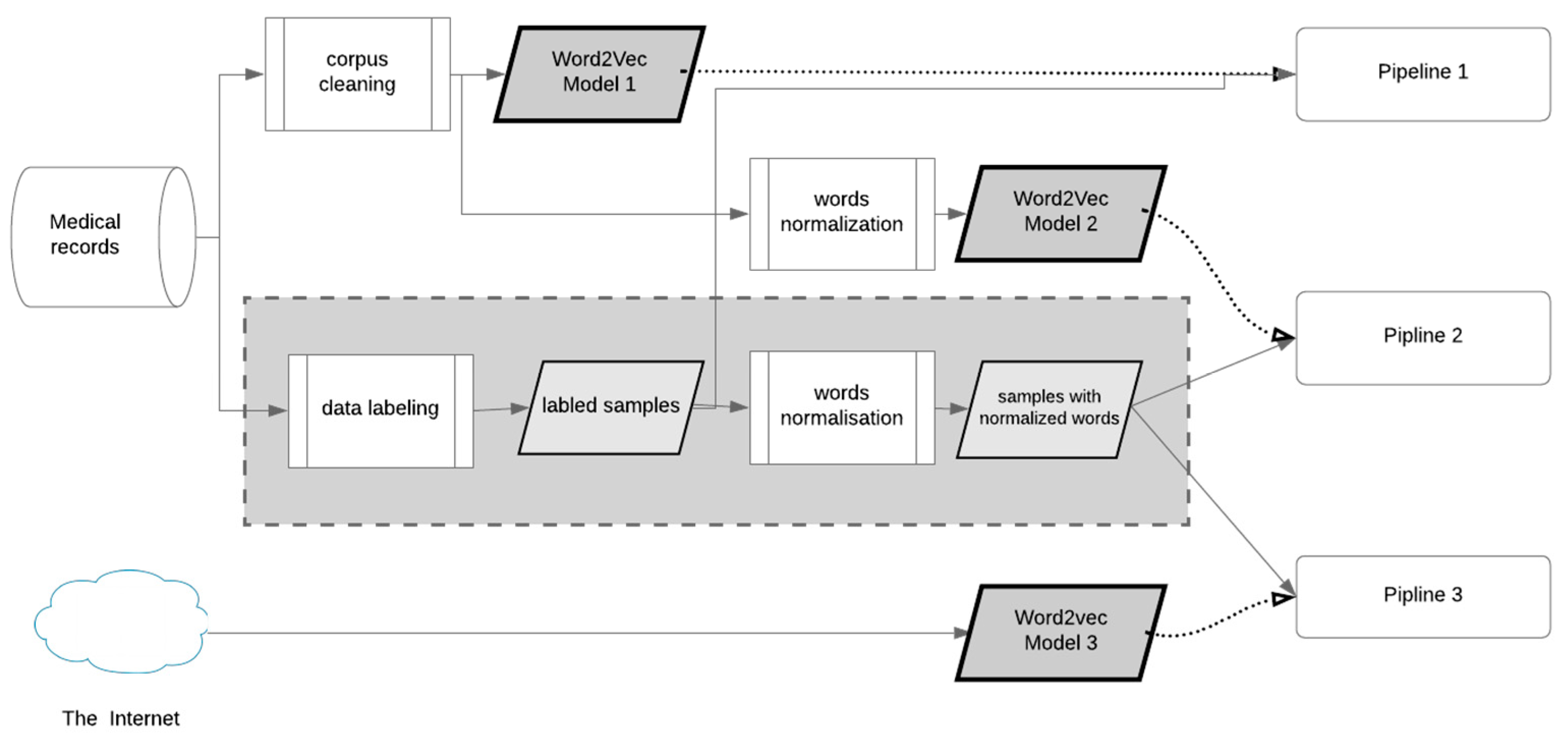
2.2. Preprocessing Pipelines
2.3. Algorithm Evaluation
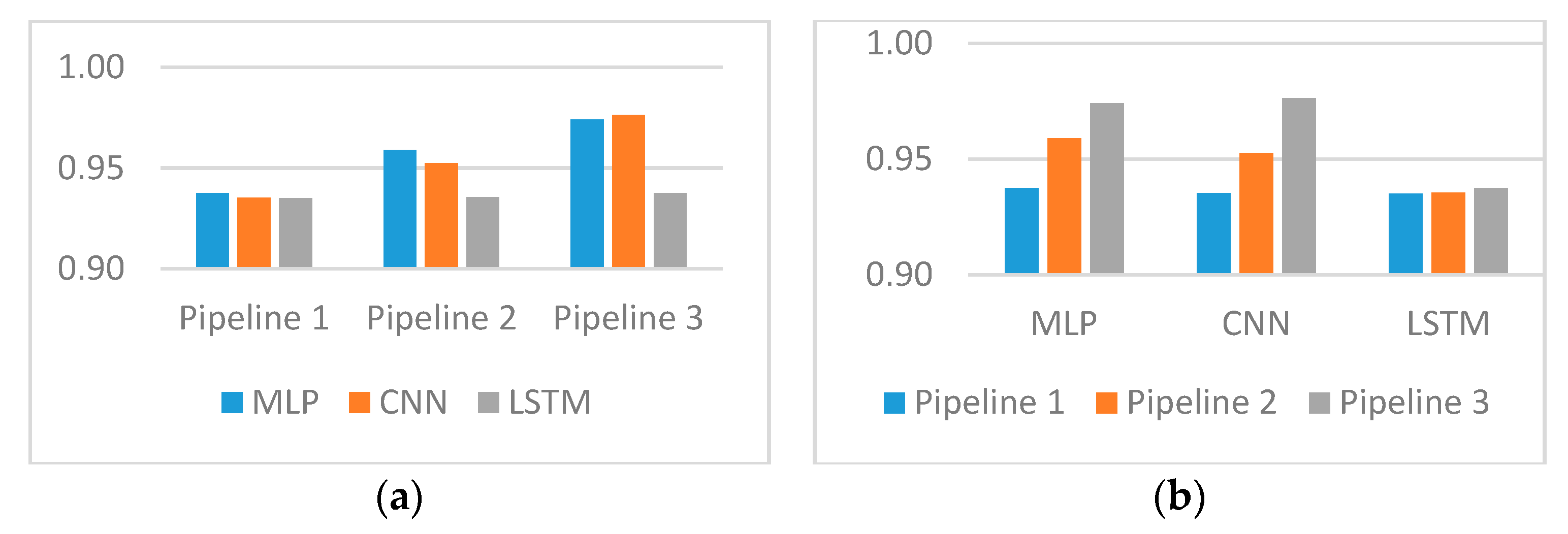
3. Results
4. Discussion and Further Work
Author Contributions
Funding
Conflicts of Interest
References
- Dudchenko, A.; Dudchenko, P.; Ganzinger, M.; Kopanitsa, G. Extraction from Medical Records. Stud. Health Technol. Inform. 2019, 261, 62–67. [Google Scholar] [PubMed]
- Dhamdhere, S.P.; Harmsen, J.; Hebbar, R.; Mandalapu, S.; Mehra, A.; Rajan, S. ELPP 2016: Big Data for Healthcare; University of California Berkeley: Berkeley, CA, USA, 2016. [Google Scholar]
- Hirschberg, J.; Manning, C.D. Advances in natural language processing. Science 2015, 349, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, G.G. Natural Language Processing: Deep Neural Networks with Multitask Learning; Association for Information Science and Technology: Silver Spring, MD, USA, 2003; Volume 37. [Google Scholar]
- Collobert, R.; Weston, J. A Unified Architecture for Natural Language Processing Deep Neural Networks with Multitask Learning; Association for Computing Machinery (ACM): New York, NY, USA, 2008; pp. 160–167. [Google Scholar]
- Sebastiani, F. Machine Learning in Automated Text Categorization. ACM Comput. Surv. 2002, 34, 1–47. [Google Scholar] [CrossRef]
- Singhal, A.; Simmons, M.; Lu, Z. Text mining for precision medicine: Automating disease-mutation relationship extraction from biomedical literature. J. Am. Med. Inform. Assoc. 2016, 23, 766–772. [Google Scholar] [CrossRef] [PubMed]
- Lample, G.; Ballesteros, M.; Subramanian, S.; Kawakami, K.; Dyer, C. Neural Architectures for Named Entity Recognition. In Proceedings of the NAACL-HLT, San Diego, CA, USA, 12 July 2016; Association for Computational Linguistics: San Diego, CA, USA, 2016. [Google Scholar]
- Hasan, S.A.; Liu, J.; Datla, V.; Shamsuzzaman, M.; Abdullah Al Hafiz Khan, M.; Sorower, M.S.; Mankovich, G.; van Ommering, R.; Pilato, S.; Dimitrova, N. Improving the State-of-the-Art for Disease Named Entity Recognition. In Proceedings of the NAACL-HLT, San Diego, CA, USA, 12 July 2016; Association for Computational Linguistics: San Diego, CA, USA, 2016. [Google Scholar]
- Chiu, J.P.C.; Nichols, E. Named Entity Recognition with Bidirectional LSTM-CNNs. Trans. Assoc. Comput. Linguist. 2016, 4, 357–370. [Google Scholar] [CrossRef]
- Kim, Y. Convolutional Neural Networks for Sentence Classification. In Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP), Doha, Qatar, 25–29 October 2014. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems; Curran Associates: Lake Tahoe, NV, USA, 2012; pp. 1097–1105. [Google Scholar]
- Nogueira, C.; Santos, D.; Gatti, M. Deep Convolutional Neural Networks for Sentiment Analysis of Short Texts; Dublin City University and Association for Computational Linguistics: Dublin, Ireland, 2014; pp. 69–78. [Google Scholar]
- Hochreiter, S.; Schmidhuber, J. Long Short-Term Memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef] [PubMed]
- Schnabel, T.; Labutov, I.; Mimno, D.; Joachims, T. Evaluation Methods for Unsupervised Word Embeddings; Association for Computational Linguistics: Stroudsburg, PA, USA, 2015. [Google Scholar]
- Mikolov, T.; Chen, K.; Corrado, G.; Dean, J. Efficient Estimation of Word Representations in Vector Space. In Proceedings of the International Conference on Learning Representations (ICLR), Scottsdale, AZ, USA, 2–4 May 2013. arXiv preprint arXiv:1301.3781. [Google Scholar]
- Danilov, G.; Kotik, K.; Shifrin, M.; Strunina, U.; Pronkina, T.; Potapov, A. Prediction of Postoperative Hospital Stay with Deep Learning Based on 101 654 Operative Reports in Neurosurgery. Stud. Health Technol. Inform. 2019, 258, 125–129. [Google Scholar] [PubMed]
- Zhou, X.; Xiong, H.; Zeng, S.; Fu, X.; Wu, J. An approach for medical event detection in Chinese clinical notes of electronic health records. BMC Med. Inform. Decis. Mak. 2019, 19, 54. [Google Scholar] [CrossRef] [PubMed]
- Musto, C.; Semeraro, G.; de Gemmis, M.; Lops, P. Learning word embeddings from wikipedia for content-based recommender systems. In Proceedings of the Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Berlin, Germany, 2016; Volume 9626, pp. 729–734. [Google Scholar]
- Wang, Y.; Liu, S.; Afzal, N.; Rastegar-Mojarad, M.; Wang, L.; Shen, F.; Kingsbury, P.; Liu, H. A comparison of word embeddings for the biomedical natural language processing. J. Biomed. Inform. 2018, 87, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Russian National Corpus. Available online: http://www.ruscorpora.ru/old/en/index.html (accessed on 27 August 2019).
- Korobov, M. Morphological Analyzer and Generator for Russian and Ukrainian Languages; Springer: Berlin, Germany, 2015; pp. 320–332. [Google Scholar]
- Морфологический Анализатор Pymorphy2—Морфологический Анализатор Pymorphy2. Available online: https://pymorphy2.readthedocs.io/en/latest/ (accessed on 27 August 2019).
| SNOMED Code | SNOMED Names | Count |
|---|---|---|
| 443502000 | Atherosclerosis of coronary artery (disorder) | 235 |
| 57546000 | Asthma with status asthmaticus (disorder) | 11 |
| 72866009 | Varicose veins of lower extremity (disorder) | 22 |
| 4556007 | Gastritis (disorder) | 28 |
| 70153002 | Hemorrhoids (disorder) | 15 |
| 25064002 | Headache (finding) | 52 |
| 386705008 | Lightheadedness (finding) | 34 |
| 1201005 | Benign essential hypertension (disorder) | 125 |
| 84229001 | Fatigue (finding) | 71 |
| 235856003 | Disorder of liver (disorder) | 11 |
| 84089009 | Hiatal hernia (disorder) | 10 |
| 76581006 | Cholecystitis (disorder) | 18 |
| 45816000 | Pyelonephritis (disorder) | 29 |
| 44054006 | Diabetes mellitus type 2 (disorder) | 32 |
| 413838009 | Chronic ischemic heart disease (disorder) | 101 |
| 162864005 | Body mass index 30+—obesity (finding) | 25 |
| 266556005 | Calculus of kidney and ureter (disorder) | 20 |
| 298494008 | Scoliosis of thoracic spine (disorder) | 18 |
| 235494005 | Chronic pancreatitis (disorder) | 18 |
| 51868009 | Duodenal ulcer disease (disorder) | 24 |
| 191268006 | Chronic anemia (disorder) | 11 |
| 102572006 | Edema of lower extremity (finding) | 13 |
| 709044004 | Chronic kidney disease (disorder) | 34 |
| (other) | Snippets without any disorder or finding | 25 |
| Embedding Model | Corpus | Total Words | Normalization | Words in the Model | Vector Size |
|---|---|---|---|---|---|
| Word2vec model 1 | 220 medical records | 1,418,728 | no | 7505 | 50 |
| Word2vec model 2 | 220 medical records | 1,418,728 | yes | 3879 | 300 |
| Word2vec model 3 | Russian National Corpus (RNC) and Wikipedia | 788,000,000 | yes | 248,000 | 300 |
| Prediction Model | Pipeline 1 | Pipeline 2 | Pipeline 3 |
|---|---|---|---|
| Multi-layer perceptron (MLP) | 0.9374 | 0.9590 | 0.9741 |
| Convolutional neural networks (CNN) | 0.9353 | 0.9525 | 0.9763 |
| Long short-term memory networks (LSTMs) | 0.9351 | 0.9355 | 0.9375 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dudchenko, A.; Kopanitsa, G. Comparison of Word Embeddings for Extraction from Medical Records. Int. J. Environ. Res. Public Health 2019, 16, 4360. https://doi.org/10.3390/ijerph16224360
Dudchenko A, Kopanitsa G. Comparison of Word Embeddings for Extraction from Medical Records. International Journal of Environmental Research and Public Health. 2019; 16(22):4360. https://doi.org/10.3390/ijerph16224360
Chicago/Turabian StyleDudchenko, Aleksei, and Georgy Kopanitsa. 2019. "Comparison of Word Embeddings for Extraction from Medical Records" International Journal of Environmental Research and Public Health 16, no. 22: 4360. https://doi.org/10.3390/ijerph16224360
APA StyleDudchenko, A., & Kopanitsa, G. (2019). Comparison of Word Embeddings for Extraction from Medical Records. International Journal of Environmental Research and Public Health, 16(22), 4360. https://doi.org/10.3390/ijerph16224360





