Antimicrobial Resistance in Food Animals and the Environment in Nigeria: A Review
Abstract
1. Introduction
2. Materials and Methods
2.1. Research Question(s)
2.2. Search Design
2.3. Analysis
3. Results
3.1. Antimicrobial Resistance Studies (AMRS)
3.1.1. Antimicrobial Resistance According to Generation of Antibiotics
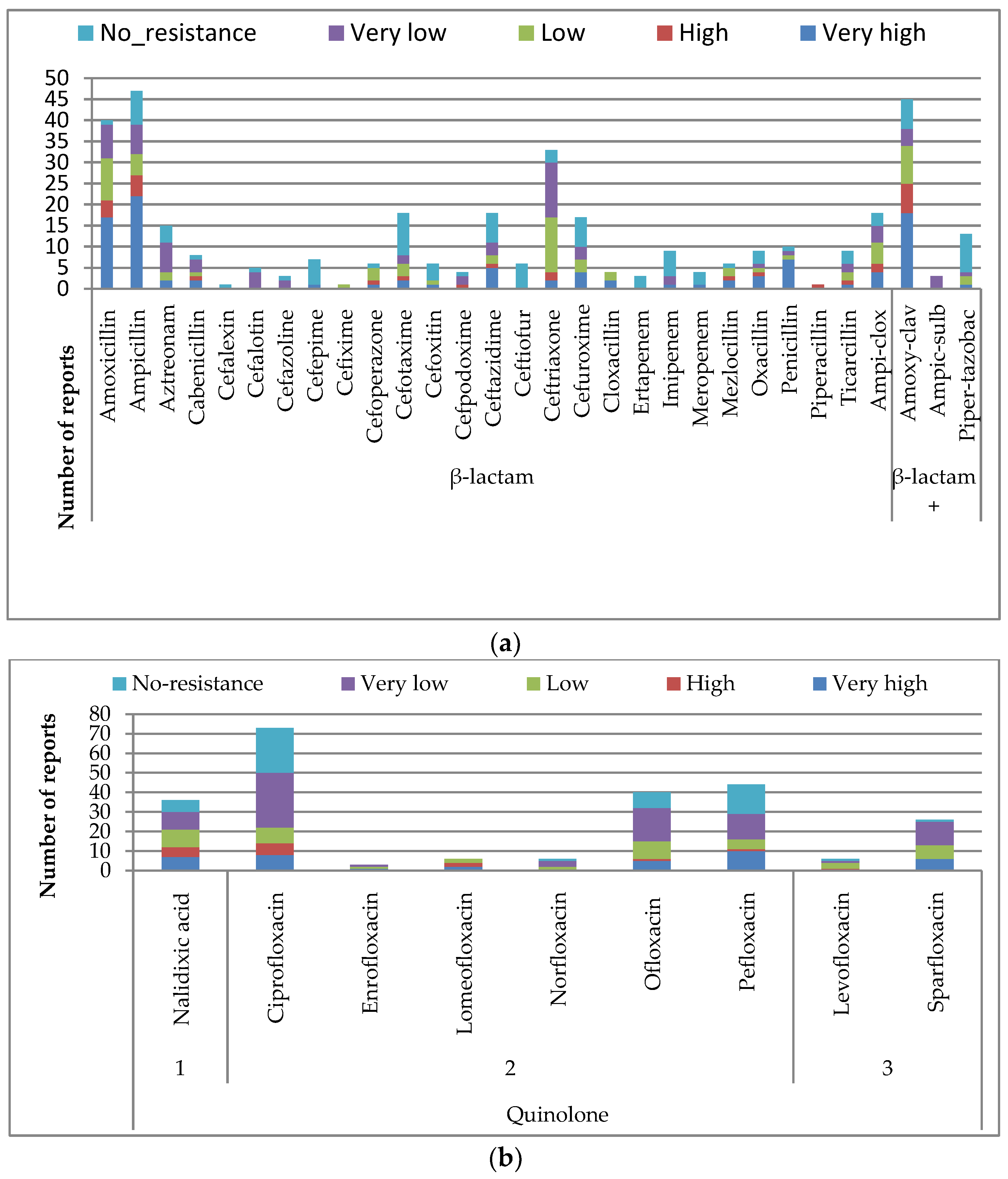
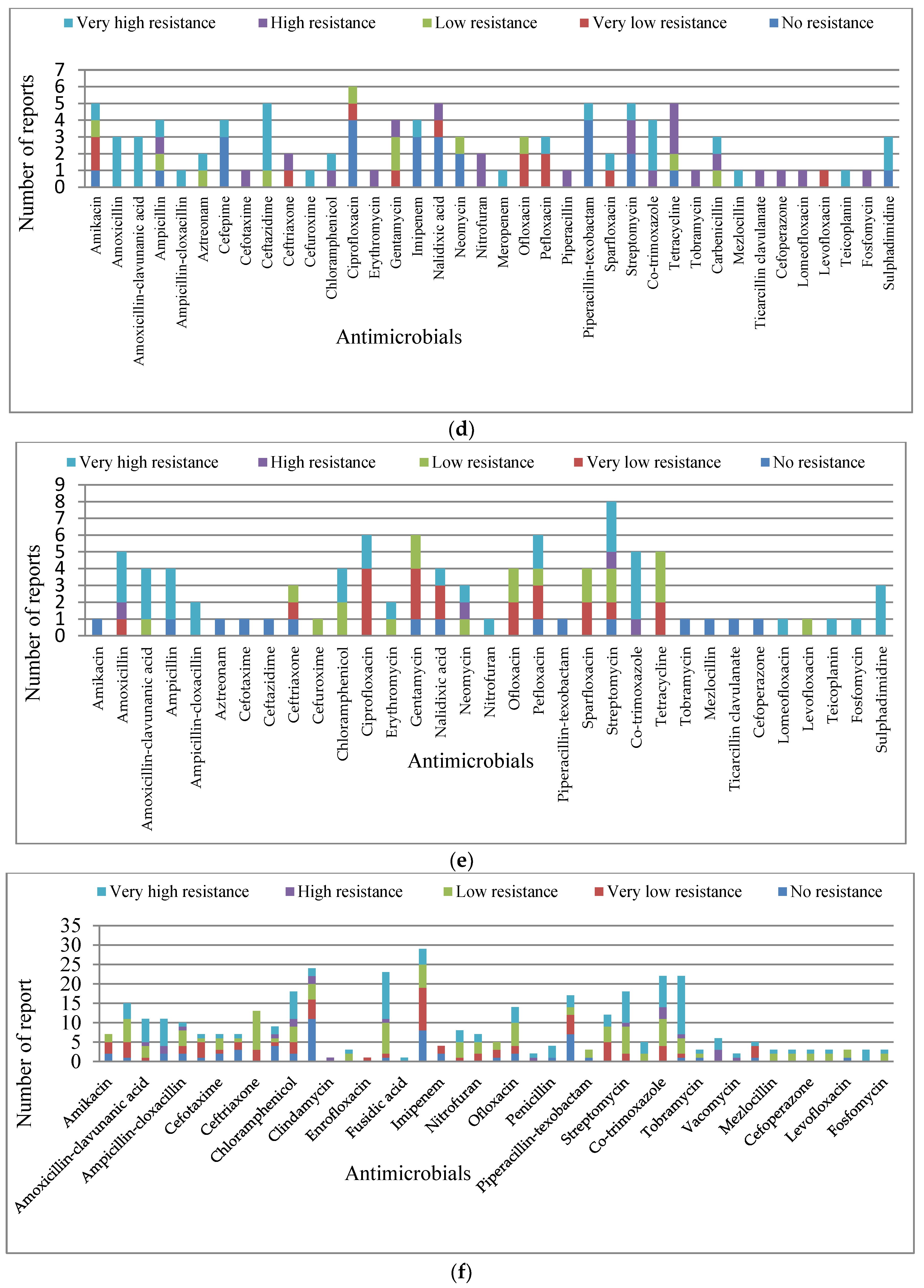
3.1.2. Resistance Level within the Classes of Antibiotics
β-lactam Derivatives
Quinolones
Aminoglycosides
Macrolide, Phenicol, and Tetracycline
Sulfonamides Derivatives
Other Classes of Antibiotics
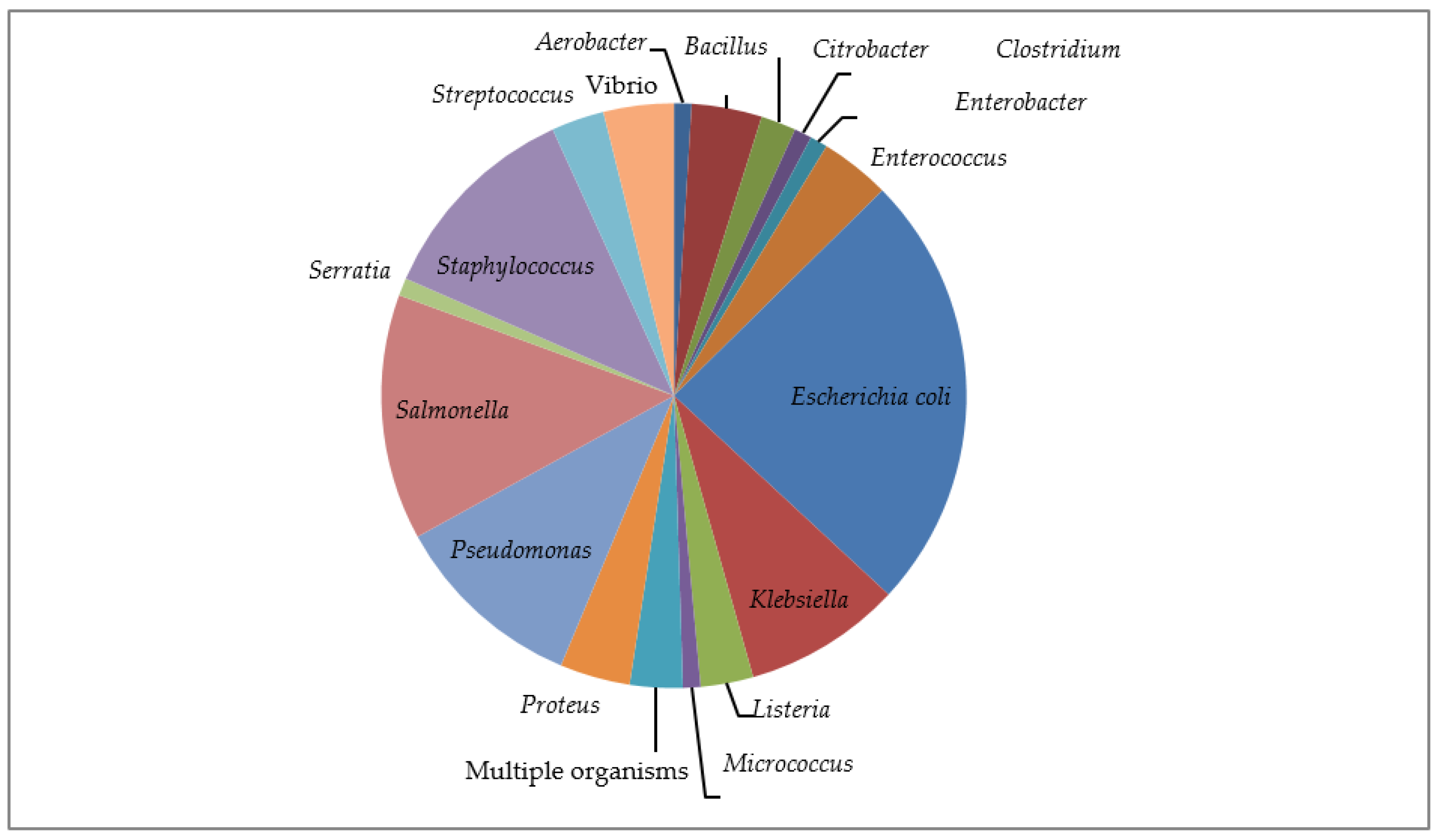
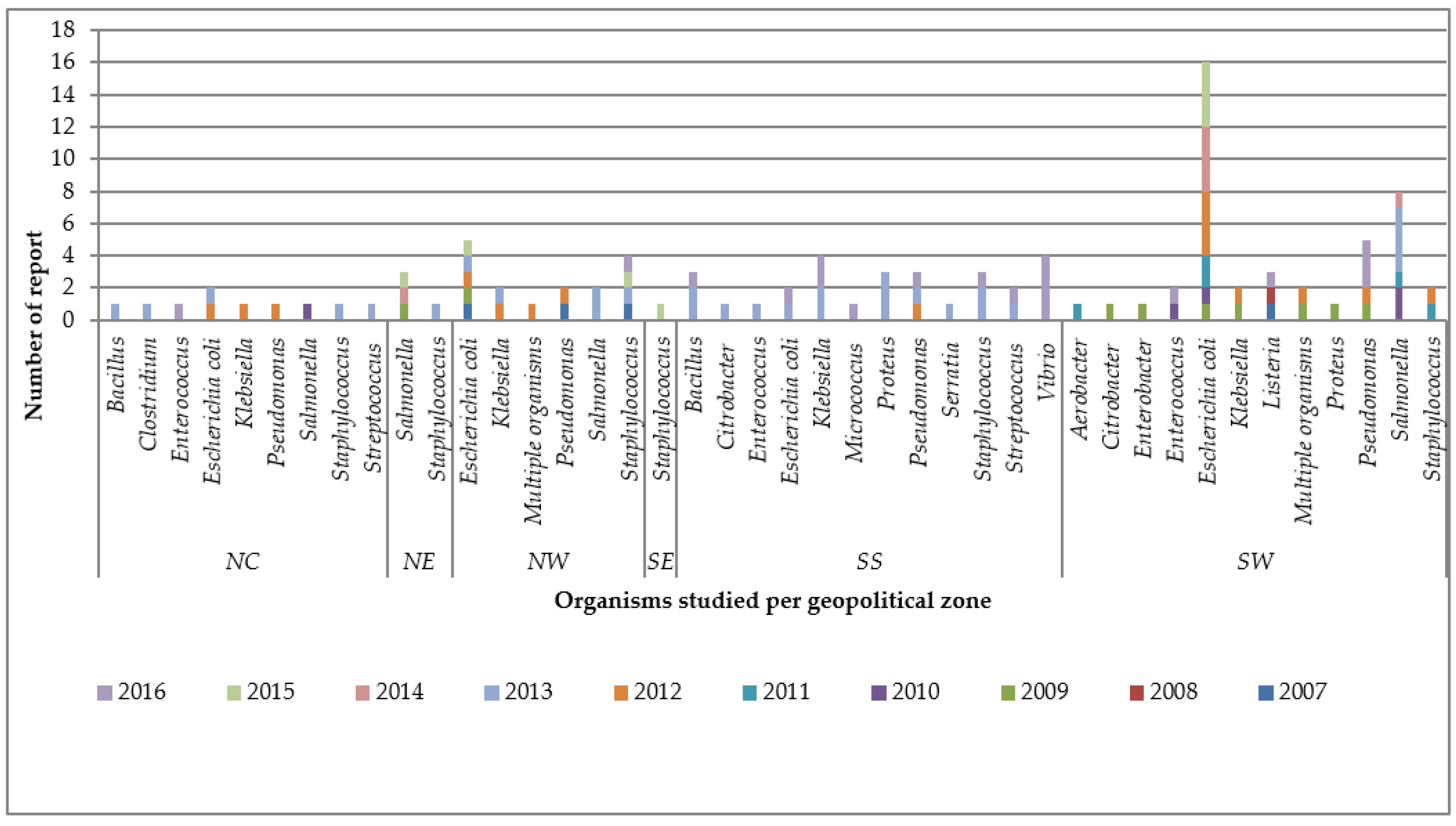
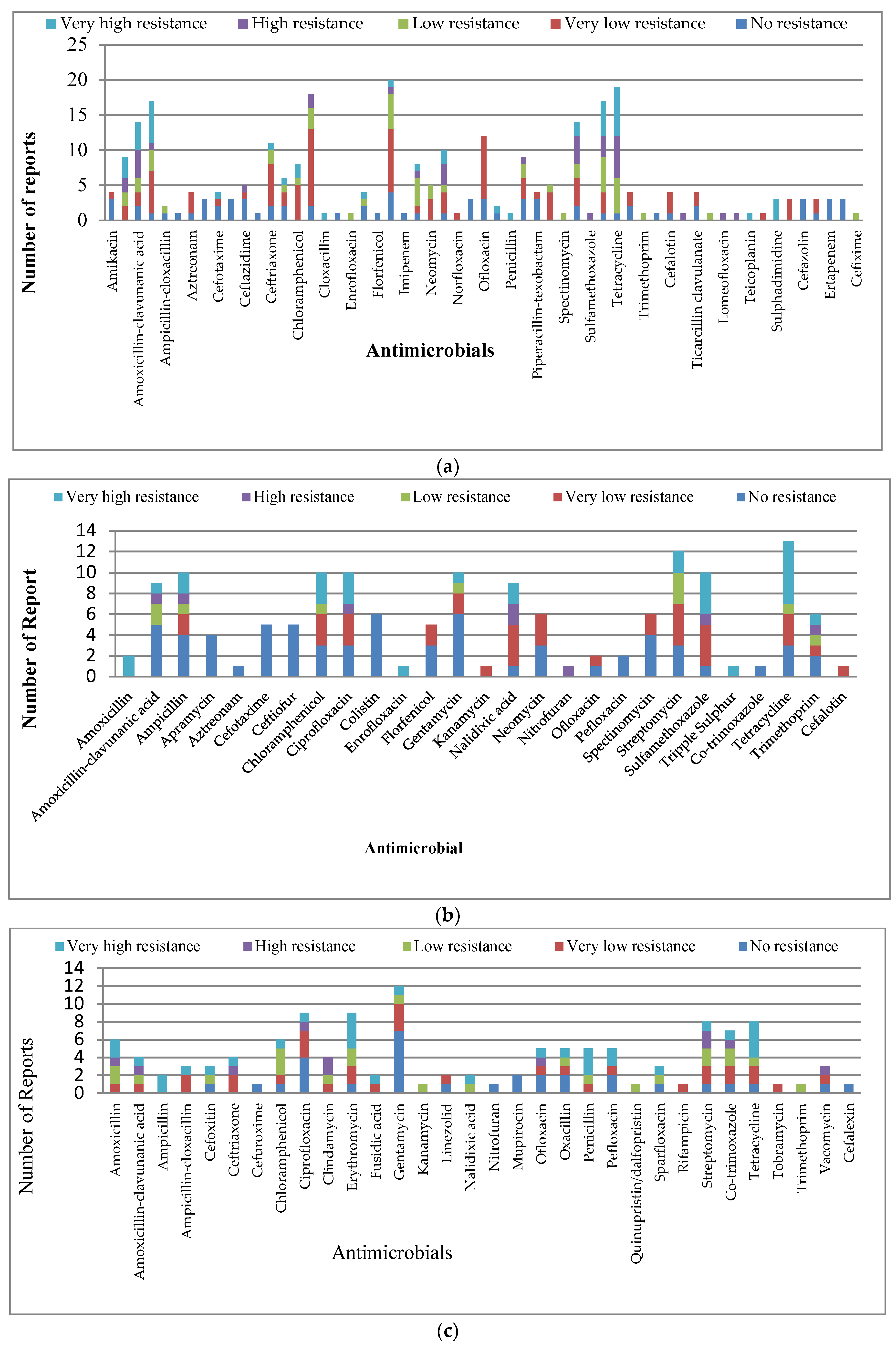
3.1.3. Resistance along the Organisms Studied
Escherichia coli (E. coli)
Salmonella
Staphylococcus
Pseudomonas
Klebsiella
Other Organisms
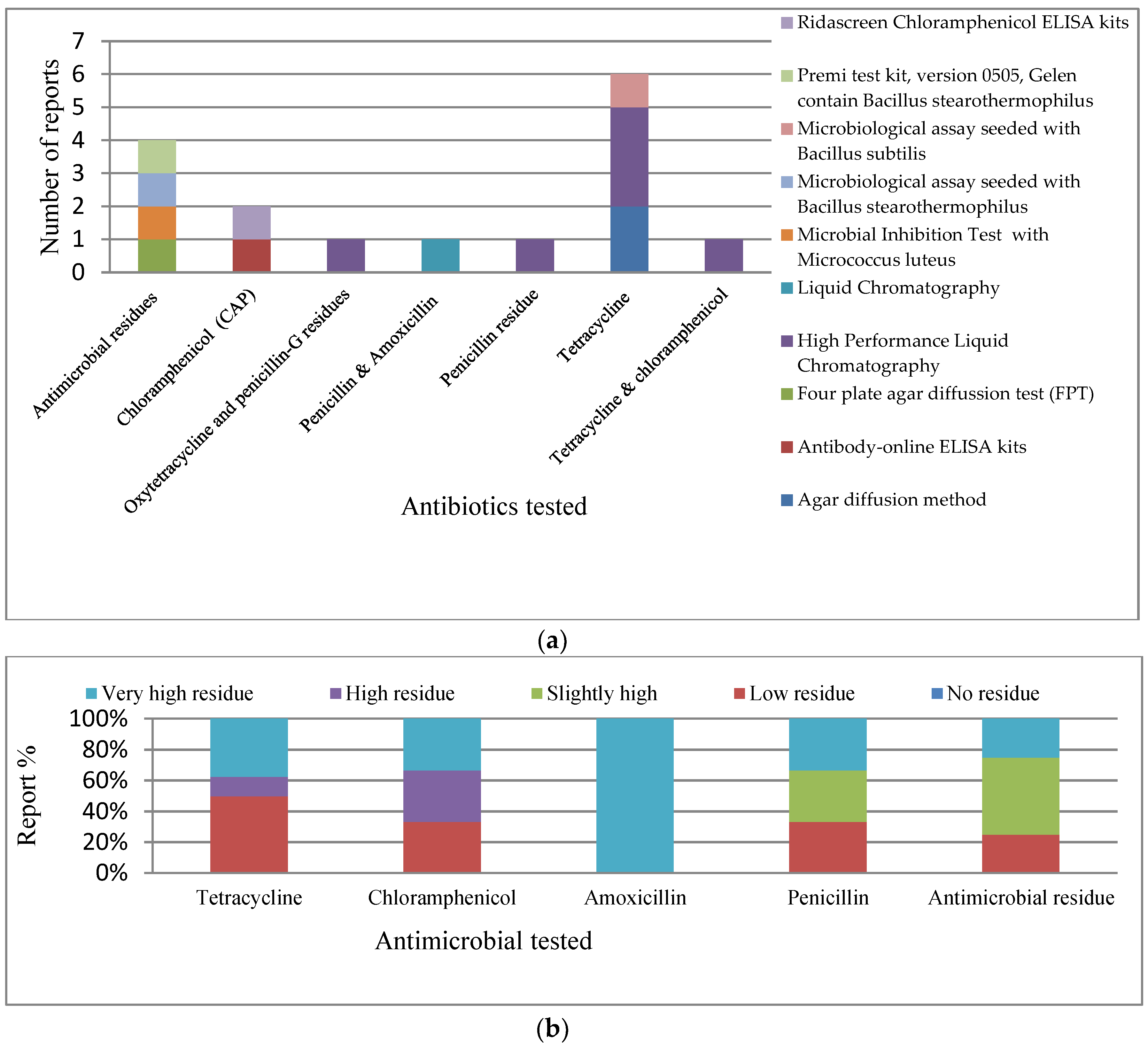
3.2. Antimicrobial Residue Studies (ARS)
3.3. Antiseptics and Disinfectant Chemicals
4. Discussion
5. Limitations
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AMR | antimicrobial resistance |
| AMRS | antimicrobial resistance studies |
| AMX | Amoxicillin |
| AR | Antimicrobial residue |
| ARS | antimicrobial residue studies |
| B.S 2 | Bacillus stearothermophilus |
| BS 1 | Bacillus subtilis |
| CHL (CAP) | Chloramphenicol |
| ELISA | Enzyme-linked immune sorbent assay |
| FAO | Food and Agriculture Organization of the United Nations |
| FPT | Four plate agar diffusion test |
| HPLC | High Performance Liquid Chromatography |
| MDR | multidrug resistance |
| NC | North Central |
| NCDC | Nigerian Centre for Disease Control |
| NE | North East |
| NGC | no generational classification |
| NW | North West |
| PEN | Penicillin |
| SDA | Surface disinfectants and antiseptics |
| SE | South East |
| SS | South South |
| SW | South West |
| TET | Tetracycline |
| WHO | World Health Organization |
References
- Kingston, W. Antibiotics, invention and innovation. Res. Policy 2000, 29, 679–710. [Google Scholar] [CrossRef]
- O’Neill, J. Antimicrobials in Agriculture and the Environment: Reducing Unnecessary Use and Waste; Wellcome Trust: London, UK, 2015; Available online: https://amr-review.org/sites/default/files/Antimicrobials%20in%20agriculture%20and%20the%20environment%20-%20Reducing%20unnecessary%20use%20and%20waste.pdf (accessed on 29 November 2017).
- O’Neill, J. Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations; Wellcome Trust: London, UK, 2014; Available online: http://www.amr-review.org (accessed on 29 November 2017).
- World Health Organization. Worldwide Country Situation Analysis: Response to Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2015; Available online: http://apps.who.int/iris/bitstream/10665/163468/1/9789241564946_eng.pdf (accessed on 29 November 2017).
- O’Neill, J. Tackling Drug-Resistant Infections Globally: Final Report and Recommendations: Final Report; Wellcome Trust: London, UK, 2016; Available online: https://amr-review.org/sites/default/files/160518_Final%20paper_with%20cover.pdf (accessed on 29 November 2017).
- United Nations. Press Release: High-Level Meeting on Antimicrobial Resistance. In Proceedings of the 71st General Assembly of the United Nations, New York, NY, USA, 21 September 2016; Available online: http://www.un.org/pga/71/2016/09/21/press-release-hl-meeting-on-antimicrobial-resistance/ (accessed on 29 November 2017).
- World Health Organization. Global Action Plan on Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2015; Available online: http://www.wpro.who.int/entity/drug_resistance/resources/global_action_plan_eng.pdf (accessed on 29 November 2017).
- Food and Agricultural Organization of the United Nation. The FAO Action Plan on Antimicrobial Resistance (2016–2020). 2016. Available online: http://www.fao.org/3/a-i5996e.pdf (accessed on 22 September 2017).
- Food and Agriculture Organization of the United Nations. Monitoring and Evaluation of the Global Action Plan on Antimicrobial Resistance. Produced Jointly by FAO, OIE and WHO. 2016. Available online: http://www.fao.org/3/a-i7711e.pdf (accessed on 29 November 2017).
- Marshall, B.M.; Levy, S.B. Food Animals and Antimicrobials: Impacts on Human Health. Clin. Microbiol. Rev. 2011, 4, 718–733. [Google Scholar] [CrossRef] [PubMed]
- Parmley, J.; Leung, Z.; Léger, D.; Finley, R.; Irwin, R.; Pintar, K.; Pollari, P.; Reid-Smith, R.; Waltner-Toews, D.; Karmali, M.; et al. A holistic approach toward enteric bacterial pathogens and antimicrobial resistance surveillance. In Improving Food Safety through a One Health Approach: Workshop Summary; National Academies Press: Washington, DC, USA, 2012. Available online: https://www.ncbi.nlm.nih.gov/books/NBK114511/ (accessed on 29 November 2017).
- Leonard, C.T.; Ward, D.; Longson, C. Antimicrobial resistance: A light at the end of the tunnel? Lancet 2017, 389, 803. [Google Scholar] [CrossRef]
- Nigeria Centre for Disease Control. Antimicrobial Use and Resistance in Nigeria. Situation Analysis and Recommendations. Produced by Federal Ministries of Agriculture, Environment and Health. Coordinated by Nigeria Centre for Disease Control. 2017. Available online: http://www.ncdc.gov.ng/themes/common/docs/protocols/56_1510840387.pdf (accessed on 29 November 2017).
- Mzungu, I. Isolation, Antibiotic and Heavy Metal Susceptibility Patterns of Some Pathogens from Domestic Dumpsites and Waste Water. Master’s Thesis, Ahmadu Bello University, Zaria, Nigeria, November 2007. [Google Scholar]
- David, O.M.; Odeyemi, A.T. Antibiotic resistant pattern of environmental isolates of Listeria monocytogenes from Ado-Ekiti, Nigeria. Afr. J. Biotechnol. 2007, 6, 2135–2139. [Google Scholar] [CrossRef]
- Adetunji, V.O.; Adegoke, G.O. Formation of biofilm by strains of Listeria monocytogenes isolated from soft cheese ‘wara’ and its processing environment. Afr. J. Biotechnol. 2008, 7, 2893–2897. [Google Scholar] [CrossRef]
- Adelowo, O.O.; Fagade, O.E. The tetracycline resistance gene tet39 is present in both Gram-negative and Gram-positive bacteria from a polluted river, Southwestern Nigeria. Lett. Appl. Microbiol. 2009, 48, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Raufu, I.; Hendriksen, R.S.; Ameh, J.A.; Aarestrup, F.M. Occurrence and characterization of Salmonella Hiduddify from chickens and poultry meat in Nigeria. Foodborne Pathog. Dis. 2009, 6, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Muhammad, M.; Muhammad, L.U.; Ambali, A.G.; Mani, A.U.; Azard, S.; Barco, L. Prevalence of Salmonella associated with chick mortality at hatching and their susceptibility to antimicrobial agents. Vet. Microbiol. 2010, 140, 131–135. [Google Scholar] [CrossRef] [PubMed]
- Ayeni, F.A.; Adeniyi, B.A.; Ogunbanwo, S.T.; Tabasco, R.; Paarup, T.; Peláez, C.; Requena, T. Inhibition of uropathogens by lactic acid bacteria isolated from dairy foods and cow’s intestine in western Nigeria. Arch. Microbiol. 2009, 191, 639–648. [Google Scholar] [CrossRef] [PubMed]
- Garba, I.; Tijjani, M.B.; Aliyu, M.S.; Yakubu, S.E.; Wada-Kura, A.; Olonitola, O.S. Prevalence of Escherichia coli in some public water sources in Gusau municipal, North-western Nigeria. Bayero J. Pure Appl. Sci. 2009, 2, 134–137. [Google Scholar] [CrossRef]
- Fashae, K.; Ogunsola, F.; Aarestrup, F.M.; Hendriksen, R.S. Antimicrobial susceptibility and serovars of Salmonella from chickens and humans in Ibadan, Nigeria. J. Infect. Dev. Ctries. 2010, 4, 484–494. [Google Scholar] [CrossRef] [PubMed]
- Odeyemi, A.T.; Dada, A.C.; Ogunbanjo, O.R.; Ojo, M.A. Bacteriological, physicochemical and mineral studies on Awedele spring water and soil samples in Ado Ekiti, Nigeria. Afr. J. Environ. Sci. Technol. 2010, 4, 319–327. [Google Scholar] [CrossRef]
- Ojo, O.E.; Ajuwape, A.T.; Otesile, E.B.; Owoade, A.A.; Oyekunle, M.A.; Adetosoye, A.I. Potentially zoonotic shiga toxin-producing Escherichia coli serogroups in the faeces and meat of food-producing animals in Ibadan, Nigeria. Int. J. Food Microbiol. 2010, 142, 214–221. [Google Scholar] [CrossRef] [PubMed]
- Adesiji, Y.O.; Alli, O.T.; Adekanle, M.A.; Jolayemi, J.B. Prevalence of Arcobacter, Escherichia coli, Staphylococcus aureus and Salmonella species in Retail Raw Chicken, Pork, Beef and Goat meat in Osogbo, Nigeria. Sierra Leone J. Biomed. Res. 2011, 3, 8–12. [Google Scholar] [CrossRef]
- Fortini, D.; Fashae, K.; García-Fernández, A.; Villa, L.; Carattoli, A. Plasmid-mediated quinolone resistance and β-lactamases in Escherichia coli from healthy animals from Nigeria. J. Antimicrob. Chemother. 2011, 66, 1269–1272. [Google Scholar] [CrossRef] [PubMed]
- Amosun, E.A.; Olatoye, I.O.; Adetosoye, A.I. Antimicrobial resistance in Escherichia coli, Klebsiella pneumonia and Pseudomonas aeruginosa isolated from the milk of dairy cows in three Nigerian cities. Niger. Vet. J. 2012, 33, 617–623. [Google Scholar]
- Damian. U.I. Antibiotics Susceptibility Studies of Some Bacterial Isolates from Packaged Milk Marketed in Zaria, Nigeria. Master’s Thesis, Ahmadu Bello University, Zaria, Nigeria, 2012.
- Igbinosa, E.O.; Odjadjare, E.E.; Igbinosa, I.H.; Orhue, P.O.; Omoigberale, M.N.; Amhanre, N.I. Antibiotic synergy interaction against multidrug-resistant Pseudomonas aeruginosa isolated from an abattoir effluent environment. Sci. World J. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Akobi, B.; Aboderin, O.; Sasaki, T.; Shittu, A. Characterization of Staphylococcus aureus isolates from faecal samples of the straw-coloured fruit bat (Eidon helvum) in Obafemi Awolowo University (OAU), Nigeria. BMC Microbiol. 2012, 12, 279. [Google Scholar] [CrossRef] [PubMed]
- Ojo, O.E.; Awosile, B.; Agbaje, M.; Sonibare, A.O.; Oyekunle, M.A.; Kasali, O.B. Quinolone resistance in bacterial isolates from chicken carcasses in Abeokuta, Nigeria: A retrospective study from 2005–2011. Niger. Vet. J. 2012, 33, 483–491. [Google Scholar]
- Fashae, K.; Hendriksen, R.S. Diversity and antimicrobial susceptibility of Salmonella enterica serovars isolated from pig farms in Ibadan, Nigeria. Folia Microbiol. (Praha) 2013, 59, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Mailafia, S.; Michael, O.; Kwaja, E. Evaluation of microbial contaminants and antibiogram of Nigerian paper currency notes (Naira) circulation in Gwagwalada, Abuja, Nigeria. Niger. Vet. J. 2013, 34, 726–735. [Google Scholar]
- Oluduro, A.O. Antibiotic-resistant commensal Escherichia coli in faecal droplets from bats and poultry in Nigeria. Vet. Ital. 2012, 48, 297–308. [Google Scholar] [PubMed]
- Kawo, A.H.; Musa, A.M. Enumeration, isolation and antibiotic susceptibility profile of bacteria associated with mobile cellphones in a university environment. Niger. J. Basic Appl. Sci. 2013, 21, 39–44. [Google Scholar] [CrossRef]
- Ogunleye, A.O.; Ajuwape, A.T.P.; Adetosoye, A.I.; Carlson, S.A. Characterization of Salmonella enterica Ituri isolated from diseased poultry in Nigeria. Afr. J. Biotechnol. 2013, 12, 2125–2128. [Google Scholar] [CrossRef]
- Oviasogie, F.E.; Agbonlahor, D.E. The burden, antibiogram and pathogenicity of bacteria found in municipal waste dumpsites and on waste site workers in Benin City. J. Biomed. Sci. 2013, 12, 115–130. [Google Scholar]
- Raufu, I.; Bortolaia, V.; Svendsen, C.A.; Ameh, J.A.; Ambali, A.G.; Aarestrup, F.M.; Hendriksen, R.S. The first attempt of an active integrated laboratory-based Salmonella surveillance programme in the North-eastern region of Nigeria. J. Appl. Microbiol. 2013, 115, 1059–1067. [Google Scholar] [CrossRef] [PubMed]
- Suleiman, A.; Zaria, L.T.; Grema, H.A.; Ahmadu, P. Antimicrobial resistant coagulase positive Staphylococcus aureus from chickens in Maiduguri, Nigeria. Sokoto J. Vet. Sci. 2013, 11, 51–55. [Google Scholar] [CrossRef]
- Adelowo, O.O.; Fagade, O.E.; Agersø, Y. Antibiotic resistance and resistance genes in Escherichia coli from poultry farms, southwest Nigeria. J. Infect. Dev. Ctries. 2014, 8, 1103–1112. [Google Scholar] [CrossRef] [PubMed]
- Adefarakan, T.A.; Oluduro, A.O.; David, O.M.; Ajayi, A.O.; Ariyo, A.B.; Fashina, C.D. Prevalence of antibiotic resistance and molecular characterization of Escherichia coli from faeces of apparently healthy rams and goats in Ile-Ife, Southwest, Nigeria. Ife J. Sci. 2014, 16, 447–460. [Google Scholar]
- Adeyanju, G.T.; Ishola, O. Salmonella and Escherichia coli contamination of poultry meat from a processing plant and retail markets in Ibadan, Oyo State, Nigeria. Springerplus 2014, 3, 139. [Google Scholar] [CrossRef] [PubMed]
- Raufu, I.A.; Fashae, K.; Ameh, J.A.; Ambali, A.; Ogunsola, F.T.; Coker, A.O.; Hendriksen, R.S. Persistence of fluoroquinolone-resistant Salmonella enterica serovar Kentucky from poultry and poultry sources in Nigeria. J. Infect. Dev. Ctries. 2014, 8, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Raufu, I.A.; Odetokun, I.A.; Oladunni, F.S.; Adam, M.; Kolapo, U.T.; Akorede, G.J.; Ghali, I.M.; Ameh, J.A.; Ambali, A. Serotypes, antimicrobial profiles, and public health significance of Salmonella from camels slaughtered in Maiduguri central abattoir, Nigeria. Vet. World 2015, 8, 1068–1072. [Google Scholar] [CrossRef] [PubMed]
- Adenipekun, E.O.; Jackson, C.R.; Oluwadun, A.; Iwalokun, B.A.; Frye, J.G.; Barrett, J.B.; Hiott, L.M.; Woodley, T.A. Prevalence and antimicrobial resistance in Escherichia coli from food animals in Lagos, Nigeria. Microb. Drug Resist. 2015, 21, 358–365. [Google Scholar] [CrossRef] [PubMed]
- Olowe, O.A.; Adewumi, O.; Odewale, G.; Ojurongbe, O.; Adefioye, O.J. Phenotypic and molecular characterisation of extended-spectrum Beta-lactamase producing Escherichia coli obtained from animal fecals Samples in Ado Ekiti, Nigeria. J. Environ. Public Health 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Olonitola, O.S.; Fahrenfeld, N.; Pruden, A. Antibiotic resistance profiles among mesophilic aerobic bacteria in Nigerian chicken litter and associated antibiotic resistance genes1. Poult. Sci. 2015, 94, 867–874. [Google Scholar] [CrossRef] [PubMed]
- Ugwu, C.C.; Gomez-Sanz, E.; Agbo, I.C.; Torres, C.; Chah, K.F. Characterization of mannitol-fermenting methicillin-resistant staphylococci isolated from pigs in Nigeria. Braz. J. Microbiol. 2015, 46, 885–892. [Google Scholar] [CrossRef] [PubMed]
- Ayeni, F.A.; Odumosu, B.T.; Oluseyi, A.E.; Ruppitsch, W. Identification and prevalence of tetracycline resistance in enterococci isolated from poultry in Ilishan, Ogun State, Nigeria. J. Pharm. Bioallied Sci. 2016, 8, 69–73. [Google Scholar] [CrossRef] [PubMed]
- Eghomwanre, A.F.; Obayagbona, N.O.; Osarenotor, O.; Enagbonma, B.J. Evaluation of antibiotic resistance patterns and heavy metals tolerance of some bacteria isolated from contaminated soils and sediments from Warri, Delta State, Nigeria. J. Appl. Sci. Environ. Manag. 2016, 20, 287–291. [Google Scholar] [CrossRef]
- Igbinosa, E.O. Detection and antimicrobial resistance of Vibrio isolates in aquaculture environments: Implications for public health. Microb. Drug Resist. 2016, 22, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Ishola, O.O.; Mosugu, J.I.; Adesokan, H.K. Prevalence and antibiotic susceptibility profiles of Listeria monocytogenes contamination of chicken flocks and meat in Oyo State, south-western Nigeria: Public health implications. J. Prev. Med. Hyg. 2016, 57, E157–E163. [Google Scholar] [PubMed]
- Ngbede, E.O.; Raji, M.A.; Kwanashie, C.N.; Kwaga, J.K. Antimicrobial resistance and virulence profile of enterococci isolated from poultry and cattle sources in Nigeria. Trop. Anim. Health Prod. 2017, 49, 451–458. [Google Scholar] [CrossRef] [PubMed]
- Odumosu, B.T.; Ajetumobi, O.; Adegbola, H.D.; Odutayo, I. Antibiotic susceptibility pattern and analysis of plasmid profiles of Pseudomonas aeruginosa from human, animal and plant sources. SpringerPlus 2016, 5, 1381. [Google Scholar] [CrossRef] [PubMed]
- Umaru, G.A.; Kwaga, J.K.P.; Bello, M.; Raji, M.A.; Maitala, Y.S. Antibiotic resistance of Staphylococcus aureus isolated from fresh cow milk in settled Fulani herds in Kaduna State, Nigeria. Bull. Anim. Health Prod. Afr. 2016, 64, 173–182. [Google Scholar]
- Dipeolu, M.A.; Alonge, D.O. Residues of tetracycline antibiotic in cattle meat marketed in Ogun and Lagos States of Nigeria. ASSET 2001, 1, 31–36. [Google Scholar]
- Kabir, J.; Umoh, J.U.; Umoh, V.J. Characterisation and screening for antimicrobial substances of slaughtered cattle in Zaria, Nigeria. Meat Sci. 2002, 61, 435–439. [Google Scholar] [CrossRef]
- Dipeolu, M.A. Residues of tetracycline antibiotic in marketed goats and pigs in Lagos and Ogun States Nigeria. Niger. J. Anim. Sci. 2002, 5. [Google Scholar] [CrossRef]
- Dipeolu, M.A.; Dada, K.O. Residues of tetracycline in imported frozen chickens in South West Nigeria. Trop. Vet. 2005, 23, 1–4. [Google Scholar] [CrossRef]
- Olatoye, I.O.; Ehinmowo, A.A. Oxytetracycline residues in edible tissues of cattle slaughtered in Akure, Nigeria. Niger. Vet. J. 2010, 31, 93–102. [Google Scholar] [CrossRef]
- Ezenduka, E.V.; Oboegbulem, S.I.; Nwanta, J.A.; Onunkwo, J.I. Prevalence of antimicrobial residues in raw table eggs from farms and retail outlets in Enugu State, Nigeria. Trop. Anim. Health Prod. 2011, 43, 557–559. [Google Scholar] [CrossRef] [PubMed]
- Adetunji, V.O.; Olaoye, O.O. Detection of β-Lactam antibiotics (Penicillin and Amoxicillin) residues in Goat milk. Nat. Sci. 2012, 10, 60–64. [Google Scholar]
- Olatoye, O.; Kayode, S.T. Oxytetracycline residues in retail chicken eggs in Ibadan, Nigeria. Food Addit. Contam. Part B Surveill. 2012, 5, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Ezenduka, E.V.; Ugwumba, C. Antimicrobial residues screening in pigs and goats slaughtered in Nsukka Municipal abattoir, Southeast Nigeria. Afr. J. Biotechnol. 2012, 11, 12138–12140. [Google Scholar] [CrossRef]
- Omeiza, K.G.; Otopa, E.A.; Okwoche, J.O. Assessment of drug residues in beef in Abuja, the Federal Capital Territory, Nigeria. Vet. Ital. 2012, 48, 283–289. [Google Scholar] [PubMed]
- Olatoye, I.O.; Oyelakin, E.F.; Adeyemi, I.G.; Call, D.R. Chloramphenicol use and prevalence of its residues in broiler chickens and eggs in Ibadan, Nigeria. Niger. Vet. J. 2012, 33, 643–650. [Google Scholar]
- Olusola, A.V.; Folashade, P.A.; Ayoade, O.I. Heavy metal (lead, Cadmium) and antibiotic (Tetracycline and Chloramphenicol) residues in fresh and frozen fish types (Clarias gariepinus, Oreochromis niloticus) in Ibadan, Oyo State, Nigeria. Pak. J. Biol. Sci. 2012, 15, 895–899. [Google Scholar] [CrossRef] [PubMed]
- Olusola, A.V.; Diana, B.E.; Ayoade, O.I. Assessment of tetracycline, lead and cadmium residues in frozen chicken vended in Lagos and Ibadan, Nigeria. Pak. J. Biol. Sci. 2012, 15, 839–844. [Google Scholar] [CrossRef] [PubMed]
- Adesokan, H.K.; Agada, C.A.; Adetunji, V.O.; Akanbi, I.M. Oxytetracycline and penicillin-G residues in cattle slaughtered in south-western Nigeria: Implications for livestock disease management and public health. J. S. Afr. Vet. Assoc. 2013, 84, E1–E5. [Google Scholar] [CrossRef] [PubMed]
- Mbodi, F.E.; Nguku, P.; Okolocha, E.; Kabir, J. Determination of chloramphenicol residues in commercial chicken eggs in the Federal Capital Territory, Abuja, Nigeria. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2014, 31, 1834–1839. [Google Scholar] [CrossRef] [PubMed]
- Olatoye, I.O.; Daniel, O.F.; Ishola, S.A. Screening of antibiotics and chemical analysis of penicillin residue in fresh milk and traditional dairy products in Oyo state, Nigeria. Vet. World 2015, 9, 948–954. [Google Scholar] [CrossRef] [PubMed]
- Adesida, S.A.; Coker, A.O.; Smith, S.I. Resistotyping of Campylobacter jejuni. Niger. Postgrad. Med. J. 2003, 10, 211–215. [Google Scholar] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; The PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef] [PubMed]
- Coates, A.R.; Halls, G.; Hu, Y. Novel classes of antibiotics or more of the same? Br. J. Pharmacol. 2011, 163, 184–194. [Google Scholar] [CrossRef] [PubMed]
- World Organization for Animal Health. List of Antimicrobial Agents of Veterinary Importance. 2015. Available online: http://www.oie.int/fileadmin/Home/eng/Our_scientific_expertise/docs/pdf/Eng_OIE_List_antimicrobials_May2015.pdf (accessed on 29 November 2017).
- World Health Organization. Critically Important Antimicrobials for Human Medicine. 5th Revision 2016. Ranking of Medically Important Antimicrobials for Risk Management of Antimicrobial Resistance Due to Non-Human Use. Updated June 2017. Available online: http://apps.who.int/iris/bitstream/10665/255027/1/9789241512220-eng.pdf (accessed on 29 November 2017).
- Israel, E.U.; Emmanuel, E.G.; Sylvester, E.G.; Chukwuma, E. Self-medication with antibiotics amongst Civil Servants in Uyo, Southern Nigeria. J. Adv. Med. Pharm. Sci. 2015, 2, 89–97. [Google Scholar] [CrossRef]
- Federal Ministries of Agriculture and Rural Development, Environment, and Health. Antimicrobial Use and Resistance in Nigeria: Situation Analysis and Recommendations. 2017. Available online: http://www.ncdc.gov.ng/themes/common/docs/protocols/56_1510840387.pdf (accessed on 14 May 2018).
- Adesokan, H.; Akanbi, I.; Akanbi, I.; Obaweda, R. Pattern of antimicrobial usage in livestock animals in south-western Nigeria: The need for alternative plans. Onderstepoort J. Vet. Res. 2015, 82, 1–6. [Google Scholar] [CrossRef] [PubMed]
- World Organization for Animal Health. OIE Terrestrial Animal Health Code. 2017. Available online: http://www.rr-africa.oie.int/docspdf/en/Codes/en_csat-vol1.pdf (accessed on 29 November 2017).

| a. Rate of Publication per Year | ||||
| Publication Year | AMRS | ARS | SDA | Total Reports |
| 2001 | 1 | 1 | ||
| 2002 | 2 | 2 | ||
| 2003 | 1 | 1 | ||
| 2005 | 1 | 1 | ||
| 2007 | 2 | 2 | ||
| 2008 | 1 | 1 | ||
| 2009 | 4 | 4 | ||
| 2010 | 4 | 1 | 5 | |
| 2011 | 2 | 1 | 3 | |
| 2012 | 6 | 7 | 13 | |
| 2013 | 7 | 1 | 8 | |
| 2014 | 4 | 1 | 5 | |
| 2015 | 5 | 5 | ||
| 2016 | 7 | 1 | 8 | |
| Total | 42 | 16 | 1 | 59 |
| b. Population Groups Identified in the Studies | ||||
| Sample Population | AMRS | ARS | SDA | Total Reports |
| Environment | 45 | - | 1 | 46 |
| Cattle | 28 | 6 | - | 34 |
| Poultry | 26 | 6 | - | 32 |
| Pig | 10 | 2 | - | 12 |
| Goat | 6 | 3 | - | 9 |
| Vegetables | 3 | - | - | 3 |
| Human | 3 | - | - | 3 |
| Bats | 2 | - | - | 2 |
| Camel | 2 | - | - | 2 |
| Sheep | 2 | - | - | 2 |
| Fish | 1 | 1 | - | 1 |
| Total | 128 | 18 | 1 | 146 |
| Group Scale | Categorization | Antimicrobial Resistance Studies | Antimicrobial Residue Studies |
|---|---|---|---|
| 1 | ≤1% | Sensitive or No resistance | No residue |
| 2 | >1 ≤ 24% | Moderately sensitive or very low resistance | Low residue |
| 3 | >24 ≤ 50% | Weakly sensitive or Low resistance | Slightly high residue |
| 4 | >50 ≤ 74% | Low sensitive or High resistance | High residue |
| 5 | >74% | Very low (no) sensitive or Very high resistance | Very high residue |
| Antibiotics in Peer-Reviewed Studies (n) | Class | Generation | Number of Reports & Category of Resistance Level | |||||
|---|---|---|---|---|---|---|---|---|
| Very High | High | Low | Very Low | No | Total | |||
| Amikacin (AMK) (5) | Aminoglycoside | NGC | 1 | 0 | 3 | 6 | 7 | 17 |
| Amoxicillin (AMX) (10) | β-lactam | 3 | 17 | 4 | 10 | 8 | 1 | 40 |
| Amoxycillin-clavunanic acid (AMC) (23) | β-lactam + | 4 | 18 | 7 | 9 | 4 | 7 | 45 |
| Ampicillin (AMP) (20) | β-lactam | 3 | 22 | 5 | 5 | 7 | 8 | 47 |
| Ampicillin-cloxacillin (APX) (3) | β-lactam | 4 | 4 | 2 | 5 | 4 | 3 | 18 |
| Ampicillin-sulbactam (AMS) (1) | β-lactam + | 4 | 0 | 0 | 0 | 3 | 0 | 3 |
| Apramycin (APR) (5) | Aminoglycoside | NGC | 0 | 0 | 0 | 0 | 5 | 5 |
| Aztreonam(AZT) (5) | β-lactam | 1 | 2 | 0 | 2 | 7 | 4 | 15 |
| Cabenicillin (CBN) (3) | β-lactam | 4 | 2 | 1 | 1 | 3 | 1 | 8 |
| Cefalexin (CLX) (1) | β-lactam | 2 | 0 | 0 | 0 | 0 | 1 | 1 |
| Cefalotin (CLT) (3) | β-lactam | 1 | 0 | 0 | 0 | 4 | 1 | 5 |
| Cefazoline (CFZ) (1) | β-lactam | 1 | 0 | 0 | 0 | 2 | 1 | 3 |
| Cefepime (CFP) (3) | β-lactam | 4 | 1 | 0 | 0 | 0 | 6 | 7 |
| Cefixime (CFX) (1) | β-lactam | 3 | 0 | 0 | 1 | 0 | 0 | 1 |
| Cefoperazone (CPZ) (1) | β-lactam | 3 | 1 | 1 | 3 | 0 | 1 | 6 |
| Cefotaxime (CTX) (10) | β-lactam | 3 | 2 | 1 | 3 | 2 | 10 | 18 |
| Cefoxitin (CXT) (4) | β-lactam | 2 | 1 | 0 | 1 | 0 | 4 | 6 |
| Cefpodoxime (CPM) (2) | β-lactam | 3 | 0 | 1 | 0 | 2 | 1 | 4 |
| Ceftazidime (CAZ) (6) | β-lactam | 3 | 5 | 1 | 2 | 3 | 7 | 18 |
| Ceftiofur (XNL) (6) | β-lactam | 3 | 0 | 0 | 0 | 0 | 6 | 6 |
| Ceftriaxone (CRO) (8) | β-lactam | 3 | 2 | 2 | 13 | 13 | 3 | 33 |
| Cefuroxime (CXM) (6) | β-lactam | 2 | 4 | 0 | 3 | 3 | 7 | 17 |
| Chloramphenicol (CHL) (21) | Phenicol | NGC | 16 | 3 | 11 | 12 | 6 | 48 |
| Ciprofloxacin (CIP) (30) | Quinolone | 2 | 8 | 6 | 8 | 28 | 23 | 73 |
| Clindamycin (CLI) (5) | Macrolide | NGC | 0 | 3 | 1 | 1 | 0 | 5 |
| Cloxacillin (CXL) (4) | β-lactam | 2 | 2 | 0 | 2 | 0 | 0 | 4 |
| Colistin (COL/CT) (7) | Polypeptide | 1 | 0 | 0 | 0 | 0 | 7 | 7 |
| Enrofloxacin (ENR) (3) | Quinolone | 2 | 1 | 0 | 1 | 1 | 0 | 3 |
| Ertapenem (ETP) (1) | β-lactam | NGC | 0 | 0 | 0 | 0 | 3 | 3 |
| Erythromycin (E) (17) | Macrolide | NGC | 18 | 2 | 12 | 3 | 4 | 39 |
| Florfenicol (FFC) (6) | Phenicol | NGC | 0 | 0 | 0 | 2 | 4 | 6 |
| Fosfomycin (FFM) (1) | Organophosphate | NGC | 2 | 1 | 2 | 1 | 0 | 6 |
| Fusidic acid (FUA) (3) | Steroid | NGC | 2 | 0 | 0 | 1 | 0 | 3 |
| Gentamycin (CN/GEN) (33) | Aminoglycoside | NGC | 7 | 2 | 17 | 29 | 26 | 81 |
| Imipenem (IMP) (4) | β-lactam | NGC | 1 | 0 | 0 | 2 | 6 | 9 |
| Kanamycin (K) (2) | Aminoglycoside | NGC | 0 | 0 | 1 | 1 | 0 | 2 |
| Levofloxacin (LVF) (1) | Quinolone | 3 | 0 | 1 | 3 | 1 | 1 | 6 |
| Linezolid (LIZ) (2) | Oxazolidinone | NGC | 0 | 0 | 0 | 1 | 1 | 2 |
| Lomeofloxacin (LMF) (1) | Quinolone | 2 | 2 | 2 | 2 | 0 | 0 | 6 |
| Nalidixic acid (NAL) (16) | Quinolone | 1 | 7 | 5 | 9 | 9 | 6 | 36 |
| Neomycin (N) (8) | Aminoglycoside | NGC | 1 | 1 | 4 | 6 | 5 | 17 |
| Nitrofuran (NIT) (8) | Furan | NGC | 5 | 6 | 4 | 5 | 2 | 22 |
| Norfloxacin (NOR) (3) | Quinolone | 2 | 0 | 0 | 2 | 3 | 1 | 6 |
| Meropenem (MPM) (2) | β-lactam | NGC | 1 | 0 | 0 | 0 | 3 | 4 |
| Mezlocillin (MZC) (1) | β-lactam | 4 | 2 | 1 | 2 | 0 | 1 | 6 |
| Mupirocin (MP) (2) | Carbolic acid | NGC | 0 | 0 | 0 | 0 | 2 | 2 |
| Ofloxacin (OFX) (11) | Quinolone | 2 | 5 | 1 | 9 | 17 | 8 | 40 |
| Oxacillin (OX) (7) | β-lactam | 2 | 3 | 1 | 1 | 1 | 3 | 9 |
| Penicillin (P) (6) | β-lactam | 1 | 7 | 0 | 1 | 1 | 1 | 10 |
| Pefloxacin (PEF) (9) | Quinolone | 2 | 10 | 1 | 5 | 13 | 15 | 44 |
| Piperacillin (PPC) (1) | β-lactam | 4 | 0 | 1 | 0 | 0 | 0 | 1 |
| Piperacillin-tazobactam (PTB) (4) | β-lactam + β-LI | 4 | 1 | 0 | 2 | 1 | 9 | 13 |
| Quinupristin (QUI) (1) | Streptogramins | 2 | 0 | 0 | 1 | 0 | 0 | 1 |
| Sparfloxacin (SPF) (4) | Quinolone | 3 | 6 | 0 | 7 | 12 | 1 | 26 |
| Rifampicin (RIF) (1) | Ansamycin | NGC | 0 | 0 | 0 | 1 | 0 | 1 |
| Spectinomycin (SPE) (6) | Aminoglycoside | NGC | 0 | 0 | 1 | 2 | 4 | 7 |
| Streptomycin (S) (22) | Aminoglycoside | NGC | 17 | 10 | 16 | 13 | 9 | 65 |
| Sulphadimidine (SDN) (1) | Sulfonamides | NGC | 8 | 0 | 0 | 0 | 1 | 9 |
| Sulfamethoxazole (SMX) (10) | Sulfonamides | NGC | 7 | 2 | 2 | 4 | 1 | 16 |
| Triple sulphur (TS) (1) | Sulfonamides | NGC | 1 | 0 | 0 | 0 | 0 | 1 |
| Co-trimoxazole (COT) (17) | Sulfonamides + DI | NGC | 22 | 9 | 14 | 9 | 3 | 57 |
| Teicoplan (TCP) (1) | Glycopeptide | NGC | 6 | 0 | 0 | 0 | 0 | 6 |
| Tetracycline (T) (30) | Tetracycline | NGC | 32 | 10 | 15 | 8 | 7 | 72 |
| Ticarcillin (TCC) (2) | β-lactam | 4 | 1 | 1 | 2 | 2 | 3 | 9 |
| Tigecycline (TGC) (1) | Tetracycline | NGC | 0 | 0 | 0 | 0 | 3 | 3 |
| Tobramycin (TMN) (3) | Aminoglycoside | NGC | 1 | 0 | 2 | 3 | 4 | 10 |
| Trimethoprim (TMP) (10) | DI | NGC | 4 | 4 | 3 | 1 | 2 | 14 |
| Vancomycin (V) (4) | Glycopeptide | NGC | 1 | 2 | 0 | 1 | 2 | 6 |
| Total (42) | 289 | 100 | 223 | 266 | 261 | 1139 | ||
| Class of Antimicrobials | Number of Reports of Each Resistance Level Category | |||||
|---|---|---|---|---|---|---|
| Very High | High | Low | Very Low | No | Total n (%) | |
| Aminoglycoside | 27 | 13 | 44 | 60 | 60 | 204 (17.9%) |
| Ansamycin | 0 | 0 | 0 | 1 | 0 | 1 (0.09%) |
| Carbolic acid | 0 | 0 | 0 | 0 | 2 | 2 (0.18) |
| DPI | 4 | 4 | 3 | 1 | 2 | 14 (1.2) |
| Furan | 5 | 6 | 4 | 5 | 2 | 22 (1.9) |
| Glycopeptide | 7 | 2 | 0 | 1 | 2 | 12 (1.1%) |
| Macrolide | 18 | 5 | 13 | 4 | 4 | 44 (3.9) |
| Organophosphate | 2 | 1 | 2 | 1 | 0 | 6 (0.5%) |
| Oxazolidinone | 0 | 0 | 0 | 1 | 1 | 2 (0.16) |
| Phenicol | 16 | 3 | 11 | 14 | 10 | 54 (4.8%) |
| Polypeptide | 0 | 0 | 0 | 0 | 7 | 7 (0.6%) |
| Quinolone | 39 | 16 | 46 | 84 | 55 | 240 (21.1%) |
| Steroid | 2 | 0 | 0 | 1 | 0 | 3 (0.2%) |
| Streptogramins | 0 | 0 | 1 | 0 | 0 | 1 (0.08) |
| Sulfonamides | 16 | 2 | 2 | 4 | 2 | 26 (2.3%) |
| Sulfonamides + DI | 22 | 9 | 14 | 9 | 3 | 57 (5.0%) |
| Tetracycline | 32 | 10 | 15 | 8 | 10 | 75 (6.6%) |
| β-lactam | 80 | 22 | 57 | 64 | 85 | 308 (27.0%) |
| β-lactam + β-LI | 19 | 7 | 11 | 8 | 16 | 61 (5.4%) |
| Total | 289 | 100 | 223 | 266 | 261 | 1139 (100%) |
| Ref. | Sample | Zone | Test Procedure | Positive Tested Antimicrobial Residue Level | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Population | Type | Size | Site | TET | CHL | AMX | PEN | AR | |||
| 56 | Cattle | Liver, kidney & muscle | 180 | Ogun Lagos | SW | Agar diffusion method | Low (16.63%) | - | - | - | - |
| 57 | Cattle | Urine | 500 | Zaria | NW | Microbial Inhibition Test with Micrococcus luteus | - | - | - | Low (7.4%) | |
| 58 | Goat and pig | liver, kidney & muscle | 360 | Ogun Lagos | SW | Agar diffusion method | Low (15.6%) | - | - | - | - |
| 59 | Poultry | Imported layer birds meat | 100 | Ogun, Lagos, Oyo | SW | Microbiological assay seeded with B.S 1 | Low (14%) | - | - | - | - |
| 60 | Cattle | Beef | 180 | Akure | SW | High Performance Liquid Chromatography | High (54.4%) | - | - | - | - |
| 61 | Poultry | Eggs | 35 | Enugu | SE | Microbiological assay seeded with B.S 2 | - | - | - | - | Slightly high (30–36%) |
| 62 | Goat | Milk | 166 | Ibadan, | SW | Liquid Chromatography | - | - | Very high (100%) | Very high (100%) | - |
| 63 | Poultry | Chicken egg | 125 | Ibadan | SW | High Performance Liquid Chromatography | Very high >80% | - | - | - | - |
| 64 | Goat and pig | Muscle, liver & kidney | 240 | Nsukka | SE | Four plate agar diffusion test (FPT) | - | - | - | - | Slightly high 25–30% |
| 65 | Cattle | Kidney, Liver, Muscle, Urine | 448 | Abuja | NC | Premi test kit, version 0505, Gelen contain B.S 2 | - | - | - | - | Very high 89.3% |
| 66 | Poultry | Eggs, muscles, liver, & kidney | 168 | Ibadan | SW | Ridascreen CHL ELISA kits | - | High | - | - | - |
| 67 | Fish | Fresh & frozen fish | 60 | Ibadan | SW | High Performance Liquid Chromatography | Very high | Very high | - | - | - |
| 68 | Poultry | Frozen chicken | 100 | Lagos & Ibadan | SW | High Performance Liquid Chromatography | Very high | - | - | - | - |
| 69 | Cattle | Organs: kidney, liver, muscles | 90 | South west | SW | High Performance Liquid Chromatography | Low | - | - | Low | - |
| 70 | Poultry | Chicken eggs | 288 | Abuja | NC | Antibody-online ELISA kits | - | Low | - | - | - |
| 71 | Cattle | Dairy products | 598 | Oyo state | SW | High Performance Liquid Chromatography | - | - | - | Slightly high | - |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oloso, N.O.; Fagbo, S.; Garbati, M.; Olonitola, S.O.; Awosanya, E.J.; Aworh, M.K.; Adamu, H.; Odetokun, I.A.; Fasina, F.O. Antimicrobial Resistance in Food Animals and the Environment in Nigeria: A Review. Int. J. Environ. Res. Public Health 2018, 15, 1284. https://doi.org/10.3390/ijerph15061284
Oloso NO, Fagbo S, Garbati M, Olonitola SO, Awosanya EJ, Aworh MK, Adamu H, Odetokun IA, Fasina FO. Antimicrobial Resistance in Food Animals and the Environment in Nigeria: A Review. International Journal of Environmental Research and Public Health. 2018; 15(6):1284. https://doi.org/10.3390/ijerph15061284
Chicago/Turabian StyleOloso, Nurudeen Olalekan, Shamsudeen Fagbo, Musa Garbati, Steve O. Olonitola, Emmanuel Jolaoluwa Awosanya, Mabel Kamweli Aworh, Helen Adamu, Ismail Ayoade Odetokun, and Folorunso Oludayo Fasina. 2018. "Antimicrobial Resistance in Food Animals and the Environment in Nigeria: A Review" International Journal of Environmental Research and Public Health 15, no. 6: 1284. https://doi.org/10.3390/ijerph15061284
APA StyleOloso, N. O., Fagbo, S., Garbati, M., Olonitola, S. O., Awosanya, E. J., Aworh, M. K., Adamu, H., Odetokun, I. A., & Fasina, F. O. (2018). Antimicrobial Resistance in Food Animals and the Environment in Nigeria: A Review. International Journal of Environmental Research and Public Health, 15(6), 1284. https://doi.org/10.3390/ijerph15061284







