Abstract
The herbicide atrazine is widely used across the globe, which is a great concern. To investigate its potential toxicity in the human body, human serum albumin (HSA) was selected as a model protein. The interaction between atrazine and HSA was investigated using steady-state fluorescence spectroscopy, synchronous fluorescence spectroscopy, UV-Vis spectroscopy, three-dimensional (3D) fluorescence spectroscopy and circular dichroism (CD) spectroscopy. The intrinsic fluorescence of HSA was quenched by the atrazine through a static quenching mechanism. Fluorescence spectra at two excitation wavelengths (280 and 295 nm) showed that the fluorescence quenched in HSA was mainly contributed to by tryptophan residues. In addition, the atrazine bound to HSA, which induced changes in the conformation and secondary structure of HSA and caused an energy transfer. Thermodynamic parameters revealed that this binding is spontaneous. Moreover, electrostatic interactions play a major role in the combination of atrazine and HSA. One atrazine molecule can only bind to one HSA molecule to form a complex, and the atrazine molecule is bound at site II (subdomain IIIA) of HSA. This study furthers the understanding of the potential effects posed by atrazine on humans at the molecular level.
1. Introduction
Atrazine (ATR) is a triazine herbicide with an excellent herbicidal activity. ATR can kill a broad spectrum of plants and has been widely used to control broadleaf and grass weeds in past decades [1,2,3]. ATR plays a key role in inhibiting the photosynthesis of plants by targeting the D1 protein of photosystem II (PSII) and causes the death of weeds to achieve the function of weed prevention and treatment [4,5,6,7,8]. Such herbicides have played a significant role in agriculture but can unfortunately enter surface waters through various paths and can further enter groundwater [9,10,11]. These compounds may remain in crops and can directly or indirectly enter the food chain and then the human body, potentially harming human health [12,13,14]. The use of ATR, simazine, prometryn, and terbutryn, as four classic types of triazine herbicides, has been limited by the European Parliament Agency. For example, ATR plays a significant role in inhibiting the specific binding of various types of proteins such as oestrogens and progesterone receptors; the extent of its harm varies and can even lead to cancer [15,16,17]. ATR may harm the central nervous systems, endocrine systems, immune systems, and reproductive systems of amphibians, rats, and pigs, and other animals, and ATR can produce genotoxic damage in fish [18,19,20]. However, up to now, there have been no studies regarding the interactions between ATR and physiologically important proteins using fluorescence spectroscopy; these studies are needed to provide important insight into the interaction of plasma proteins with ATR.
Human serum albumin (HSA) is one of the most abundant proteins in plasma, accounting for approximately 60% of the total serum protein content, and is an important carrier of transport proteins [21,22]. Approximately 80% of the blood osmotic pressure is provided by HSA. Many of the endogenous and exogenous substances that enter the human body bind to HSA through reversible noncovalent binding and are transported to various parts of the body after binding [23]. Therefore, HSA has important physiological functions, such as bodily protection and information storage, and is the main carrier through which various endogenous and exogenous substances are transported and eliminated [24]. The binding of small herbicide molecules with HSA will affect its structure and function, so it is important to study the interaction of herbicides with HSA to understand the metabolism and biological effects in vivo.
Many physical and chemical methods exist to determine the structure and function of bioactive molecules as well as their mechanism of action and other information. Some of these molecules emit optical signals and can therefore be determined spectroscopically [25,26]. Spectroscopy is often used to study the material interactions in a method and has many advantages, including its simple operation, ease of use, good detection performance, high sensitivity and wide selectivity. Spectroscopic techniques mainly include fluorescence spectroscopy, UV-Vis absorption spectroscopy, infrared spectroscopy, and circular dichroism (CD) spectroscopy, which are the commonly used methods for studying the interactions between small molecules and proteins [27,28].
In this study, we investigated ATR, and the interaction between small herbicidal molecule and proteins was mainly explored using fluorescence methods. The aim of this work was to study the potential influence of pesticide residues to the human body, and HSA was selected as the research object. The mechanism was elucidated at the molecular level to understand the effect on human health by studying the interaction of ATR and proteins.
2. Experimental Section
2.1. Materials
HSA without fatty acids (≥99.9%, A1887) and digitoxin (Dig, D5878) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Flufenamic acid (FA, 186419) and phenylbutazone (PB, 451567) were obtained from J&K (Beijing, China) and were used without further purification. Three drugs, digitoxin, flufenamic acid and phenylbutazone, were used as labelled drugs for binding at sites I, II and III of HSA, respectively. The specific binding site of the ligand on the HSA can be determined by the labelling competition experiment. Metsulfuron-methyl, prometryn and ATR were purchased from ANPEL Laboratory Technologies Inc. (Shanghai, China), and stock solutions of these compounds were prepared using anhydrous ethanol. All other commercial reagents were obtained from J&K Scientific, Alfa Aesar (Ward Hill, MA, USA) and Sigma-Aldrich. A Tris-HCl buffer solution (pH 7.4) was prepared with 0.20 mol/L trihydroxymethylaminomethane and 0.10 mol/L hydrochloric acid. An HSA solution (3 × 10−5 M) was prepared in Tris-HCl buffer and was stored in the refrigerator at 4 °C. Other chemicals and reagents used in this study were all analytical grade. All water used in these experiments was deionized water.
2.2. Instrumentation
The fluorescence spectra were measured on a 970 CRT fluorescence spectrophotometer (Shanghai Electronic Science Instrument Factory, Shanghai, China). An HH-6 digital constant temperature water bath (Ronghua Instrument Factory, Changzhou, China) was used for temperature control. The three-dimensional (3D) fluorescence spectra and the synchronous fluorescence spectra were measured on a Cary Eclipse fluorescence spectrophotometer (Agilent Technologies, Palo Alto, CA, USA). The UV spectra were recorded using a UV-1800 UV spectrophotometer (Shimadzu, Kyoto, Japan). All the above measurements were made using quartz cuvettes with a 1 cm path length and 3 mL volume. The pH of the buffer solution was measured using a PHS-25 digital pH metre (Shanghai REX Instrument Factory, Shanghai, China). The CD spectra were measured using a Jasco-810 spectrometer (Jasco, Tsukuba, Japan).
2.3. Steady-State Fluorescence Spectra
Steady-state fluorescence was determined using a 970CRT fluorescence spectrophotometer (Shanghai Electronic Science Instrument Factory, Shanghai, China) equipped with a 1.0 cm quartz cuvette. The temperature was controlled with an HH-6 digital constant temperature water bath. The excitation wavelength (λex) are set to 280 nm and 295 nm, respectively. And the maximum emission wavelength (λem) of HSA were 340 nm and 350 nm, respectively. The emission spectrum was scanned in a wavelength range of 300–500 nm, and the excitation and emission slit widths were fixed at 5.0 nm. The scanning speed was 240 nm/min. The corresponding concentration of the Tris-HCl buffer was subtracted from all fluorescence spectra.
2.4. Fluorescence Titration Experiments
The 3 × 10−5 mol/L HSA stock solution was diluted to a 3 × 10−6 mol/L HSA solution. The ATR stock solution (6 × 10−4 mol/L), which was prepared in ethanol, was added continuously at three temperatures (298, 307 and 316 K), and the change in fluorescence intensity was measured by manual titration using a syringe (λex = 280 nm and 295 nm, λem = 340 nm and 350 nm). The experimental temperature was controlled with a water bath that was had a digital constant temperature.
In solution, due to the inner filter effect, ATR may absorb some radiation generated by the fluorophore at the excitation or emission wavelength of HSA, or may influence excited radiation that reaches the fluorophore. To decrease the inner filter effects, the absorbance of the ligand at each concentration and the absorbance of the protein without ligand at the excitation and emission wavelength were recorded, respectively. The observed fluorescence intensity was corrected using the following equation:
where Fcor and Fobs refer to the corrected and observed fluorescence intensities, respectively. Aex and Aem represent the absorption of the ligand-protein solution at the excitation and the emission wavelengths, respectively. According to Makarska’s study [29], the intensity was obtained by deducting the ATR curve from the HSA-ATR spectra, at each step of titration with ATR.
2.5. Site Marker Competition Experiments
The binding sites of the ATR on the HSA were determined by adding three classical binding sites to create binding competition amongst competitive drugs (PB, FA and Dig) during the fluorescence titration. First, the three site markers were bound to HSA. Then, the ATR solution was gradually added to the mixture of the three markers and HSA, and the change in fluorescence intensity was measured.
2.6. UV-Vis Spectra
The 10−6 mol/L sample was diluted from a 10−5 mol/L HSA stock solution to which the ATR was added (from 0.3 × 10−6 mol/L to 2.4 × 10−6 mol/L at intervals of 0.3 × 10−6 mol/L). The mixed sample was placed in a 1.0 cm quartz cuvette, and the absorption spectra of the system at wavelengths of 200–500 nm were recorded using a UV-1800 UV spectrophotometer (Shimadzu, Kyoto, Japan) at room temperature (298 K) with a scanning speed of 300 nm/min. The background absorbance of the buffer solution was subtracted from the sample absorbance.
2.7. CD Spectra
The change in the secondary structure of HSA in the presence and absence of the ATR was measured using a Jasco-810 CD spectrometer under a constant nitrogen atmosphere in a quartz cuvette with a 0.1 cm path length over a scanning range of 200–260 nm. The step resolution was 0.1 nm, and the scanning speed was 50 nm/min. The value of the buffer solution was subtracted from all CD spectral data. The change in the secondary structure of HSA was calculated using the Jasco secondary structure estimation software that accompanied the spectrometer.
2.8. 3D Fluorescence Spectra
3D fluorescence spectra were scanned on a Cary Eclipse fluorescence spectrophotometer (Agilent Technologies, Palo Alto, CA, USA) using a quartz cuvette with a 1-cm path length and a 3-mL volume. The initial excitation wavelength was set to 200 nm, the emission wavelength scanning range was 200–500 nm, the Ex and Em slit widths were both 10 nm, and the scanning frequency was 16.
2.9. Synchronous Fluorescence Spectra
Synchronous fluorescence spectra were scanned on a Cary Eclipse fluorescence spectrophotometer (Agilent Technologies) using a quartz cuvette with a 1-cm path length and a 3-mL volume. The HSA concentration was 3 × 10−6 mol/L, and the ATR solution was added to the HSA solution at concentrations ranging from 0 to 2.4 × 10−6 mol/L; λex = λem. When Δλ = 15 nm, the scan range was set to 260–320 nm, and when Δλ = 60 nm, the scanning range was 240–320 nm.
3. Results and Discussion
3.1. Study of the Quenching Mechanism of HSA by ATR
HSA produces UV fluorescence because it contains three amino acids with aromatic ring structures, including tryptophan (Trp), tyrosine (Tyr) and phenylalanine (Phe) [30,31]. Among them, Phe fluorescence is not often observed, so Trp and Tyr are commonly used as endogenous HSA fluorescent probes. When λex = 280 nm, the fluorescence is mainly contributed to by Trp and Tyr residues, and when the excitation wavelength is 295 nm, only the fluorescence of the Trp residue is excited [32,33,34]. In this experiment, 280 nm and 295 nm were used as excitation wavelengths. Under the experimental conditions, the fluorescence emission spectrum of the HSA solution at a certain concentration was scanned. Under the same conditions, the fluorescence spectra of mixtures of HSA with different concentrations of ATR were scanned. The results are shown in Figure 1. For the ATR-HSA system, with an increase in the ATR concentration, the fluorescence intensity of the HSA emission peak decreased, indicating an interaction between the ATR and HSA and quenching of the endogenous fluorescence of HSA. Compared to the results of excitation at 280 nm and excitation at 295 nm, the fluorescence intensity of the Trp groups is more pronounced than the Tyr groups that are present in HSA, indicating that Trp is mainly involved in the quenching of HSA.

Figure 1.
Steady-state fluorescence spectra of HSA in the presence of different concentrations of ATR at three different temperature (298 K, 307 K and 316 K). (a–j) 0, 0.3, 0.6, 0.9, 1.2, 1.5, 1.8, 2.1, and 2.4 × 10−6 mol/L respectively. (x) only ATR.
The mechanism by which small molecule ligands quench the fluorescence of plasma albumin is mainly divided into dynamic quenching and static quenching. In dynamic quenching, the excited molecules of the fluorescent substance collide with the quencher molecules, and through energy transfer or charge transfer, the excited molecules lose their excitation energy and return to the ground state without emitting photons. In static quenching, the quenching agent and fluorescent substance in the ground state form a complex, leading to fluorescence quenching. The static and dynamic quenching mechanisms follow the Stern–Volmer equation [35,36,37]:
where F0 and F are the fluorescence intensity of the system without and with the quencher, respectively; Kq is the quenching rate constant of the biomolecule; τ0 is the average life of the biomolecule in the absence of the quencher (for most biomolecules, τ0 is approximately 10−8 s−1) [38]; [Q] is the concentration of the quencher; and KSV is the quenching constant as the ratio of the bimolecular quenching rate constant to the single molecule decay rate constant. The Stern–Volmer diagram of the ATR-HSA system at 298 K can be calculated from the Stern–Volmer equation and the Stern–Volmer quenching constant for the interaction between ATR and HSA at three different temperatures (Table 1).

Table 1.
Stern-Volmer quenching constants for the interactions of ATR with HSA at different temperatures.
As shown in Figure 2, the Stern–Volmer diagrams of the ATR-HSA system have very high linear correlation coefficients, and it is speculated that the ATR-HSA combination has only one quenching mechanism. Table 1 lists the value of Kq for different excitation wavelengths. The Kq ranges from 6.66 × 1012 to 8.90 × 1012 mol/L at an excitation wavelength of 280 nm, and when the excitation wavelength set to 295 nm, the value of Kq ranges from 6.07 × 1012 to 8.12 × 1012 mol/L. It is apparent from the data that the quenching constants at different excitation wavelengths are very close, indicating that ATR mainly quenches the fluorescence of Trp residues and is more than 100 times the maximum dispersion collision quenching constant (2.0 × 1010 mol/L) [39,40] of the various quenchers and biological macromolecules. Therefore, this reaction produces a complex of ATR and HSA, and the endogenous fluorescence of HSA is quenched via the static quenching mechanism.
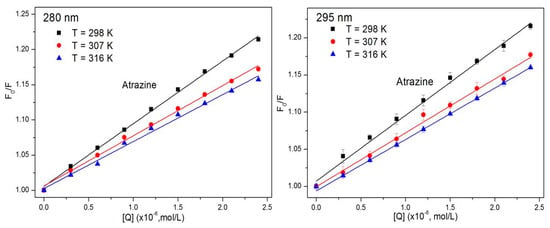
Figure 2.
Stern–Volmer plot describing HSA fluorescence quenching caused by ATR at different excitation wavelengths. Data are mean ± SE (bars) (n = 3) (T = 298 K, 307 K, 316 K).
3.2. UV-Vis Spectra of ATR Interacting with HSA
UV-Vis absorption spectroscopy is used to explore the structural changes and morphologies of complexes [41,42,43]. Dynamic quenching only affects the excited state of the fluorophore; therefore, dynamic quenching should not change the absorption spectrum. For static quenching, the changes in the UV-Vis absorption spectrum are due to the formation of the complex [44,45,46,47,48]. Figure 3 shows the absorption spectra of HSA after the addition of the ATR. The absorbance of HSA increased with the addition of different amounts of herbicides, accompanied by a 5-nm redshift. These data confirm that the spectral change is due to the formation of the complex and verifies the previous conclusion that the quenching mechanism of HSA and ATR is static quenching.
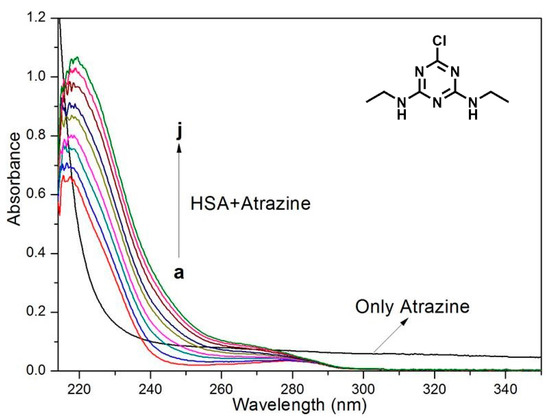
Figure 3.
Absorption spectra of HSA (3.0 × 10−6 mol/L) upon the addition of ATR. The arrows show the increase in the ATR concentration. The ATR concentrations were 0.3, 0.6, 0.9, 1.2, 1.5, 1.8, 2.1 and 2.4 × 10−6 mol/L, respectively.
3.3. Study of the Binding Sites and Binding Number
For static quenching, data from the fluorescence spectra can be utilized to calculate the binding constant (Ka) and number of binding sites (n) using the following formula [49]:
where F and F0 represent the fluorescence intensity with and without the quencher, respectively; n is the number of binding sites; PT is the total protein concentration; QT is the total quencher concentration; and Ka is a binding constant. nlogKa is the intercept of the linear equation represented by and . Figure 4 is a plot of log [(F0−F)/F] versus log Ka, and n can be calculated based on Equation (2). Ka decreases with increasing temperature; therefore, the decrease in the stability of the complex that formed from the ATR and HSA at a relatively high temperature is not conducive to the binding, which is also a characteristic of static quenching. Ka is large, which indicates a strong interaction of ATR with HSA. As shown in Table 2, the results at different excitation wavelengths are similar and the value of n is close to 1, which indicates that only one ATR molecule combines with one HSA molecule. This reaction proceeds at a ratio of 1:1.
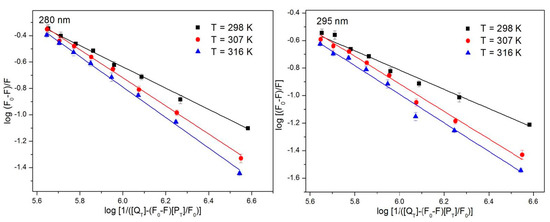
Figure 4.
The log(F0−F)/F against log[1/([QT]−(F0−F)[PT]/F0)] plots of ATR-HSA system at 298 K, 307 K and 316 K (n = 3).

Table 2.
Binding constants and thermodynamic parameters of the ATR to HSA.
Small molecule and protein binding sites can be confirmed by competitive marker experiments, providing a fast, simple and effective method [50,51,52]. To determine the binding sites of ATR on HSA, PB, FA and Dig were added to the ATR-HSA system as labelled drugs for binding at sites I, II and III of HSA, respectively. The binding constants of ATR with HSA at 298 K that were calculated using Equation (2) are listed in Table 3. It is apparent from the data in Table 3 that after the addition of PB and Dig to the ATR-HSA system, the binding constant Ka did not change much. However, the binding constant of the ATR-HSA system greatly changed after the addition of FA. From the above conclusions, we know that the binding ratio of ATR to HSA is 1:1; therefore, we can speculate that the binding site of ATR on HSA is the same as that of fluoride on HSA. These results suggest that the binding site of ATR is located within site II (subdomain IIIA).

Table 3.
Effects of the site probe on the binding constants of ATR to HSA.
3.4. Combination Distance between ATR and HSA
The fluorescence intensity of HSA also decreased with the addition of ATR, indicating an energy transfer between the amino acid residues in HSA and ATR. The binding distance of the ATR-HSA system can be calculated by separately overlapping the UV-Vis spectra of ATR with the HSA fluorescence spectrum. The overlapping plots at different excitation wavelengths are shown in Figure 5.
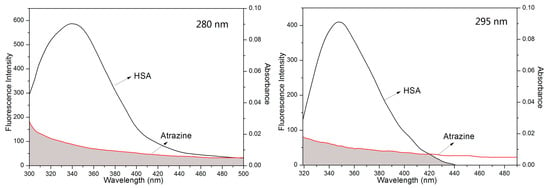
Figure 5.
Overlap of the fluorescence spectrum of HSA and the absorption spectra of the ATR. C (HSA) = c (ATR) = 3.00 × 10−6 mol/L; T = 298 K.
According to Förster’s theory of nonradiative energy transfer [53], the energy transfer rate depends on three factors including the relative orientation of the donor and acceptor dipole, the overlap of the fluorescence emission spectrum of the donor with the absorption spectrum, and the distance between the donor and acceptor. The energy transfer effect is related not only to the distance between the receptor and the donor but also to the critical energy transfer distance R0, as described in the following equation:
where F and F0 are the fluorescence intensities of HSA in the presence and absence of the drug, respectively, r is the binding distance between HSA and ATR, and R0 is the critical distance when the energy transfer is 50%. R0 can be calculated using the following formula:
where K2 is the spatial orientation factor between the emission dipole of HSA and the absorption dipole of the ATR. In general, the value of K2 is 2/3 N and is the refractive index of the medium, with a value of 1.336. ϕ is the quantum yield of HSA in the presence of ATR, and J is the overlap integral of the fluorescence emission spectra of HSA with the absorption spectra of the ATR. J can be calculated from the following formula:
where F(λ) is the fluorescence intensity of the HSA solution at the specified wavelength, and ε(λ) is the molar absorption coefficient of the solution of ATR. J can be evaluated from the spectrum in the plot. The E, J, R and r values of the ATR-HSA system at different excitation wavelengths are listed in Table 4. The value of r in the presence of the drugs was less than 8.0 nm at different excitation wavelengths, which indicates that the quenching of HSA by ATR is consistent with the nonradiative energy transfer theory and potentially indicates that the combination of ATR effectively quenches the endogenous fluorescence of HSA through energy transfer. Through the overlapping points of results from different excitation wavelengths, it is apparent that the largest contribution of endogenous fluorescence of HSA is Trp residues.

Table 4.
Parameters of E, J, R0 and r of ATR-HSA system.
3.5. Thermodynamic Parameters and Binding Modes
The interactions between small molecules and proteins are weak and include four different forces and modes of action: hydrogen bonds, electrostatic interactions, hydrophobic effects and van der Waals forces. Based on a large number of experimental results, Ross [54,55] and others summed the parameters of the interactions of proteins with small molecules, such as the binding force properties and the thermodynamics of the interaction. Their results indicated that hydrophobic forces play a major role when ΔS > 0 and ΔH > 0, and hydrogen bonds and van der Waals forces play a dominant role in the binding process when ΔS < 0 and ΔH < 0. The condition of ΔH < 0 and ΔS > 0 corresponds to electrostatic interactions. Using the van’t Hoff equation, the thermodynamics of the interactions between small molecules and proteins can be determined according to the change in the binding constant at different temperatures, and the type of interactions between small molecules and proteins can be assessed:
where Ka is the binding constant calculated from Equation (2), T is the absolute temperature, and R is the gas constant. According to the van’t Hoff equation, the values of ΔH and ΔS can be obtained from the graphs of 1/T versus ln Ka (Figure 6), where ΔH and ΔS are the slope and intercept in the van’t Hoff formula, respectively. The free energy change (ΔG) is then determined according to the following equation:
where ΔG is negative, indicating that the binding of ATR to HSA in solution is spontaneous (Table 2). The ΔS of the three ligands is positive, which indicates that the main interaction pattern between ATR and HSA is hydrophobic, whereas ΔH is negative, indicating that hydrogen plays a key role in the interaction between ATR and HSA. Based on the above results, hydrophobic forces and hydrogen bonding are the main modes of interaction between ATR and HSA.
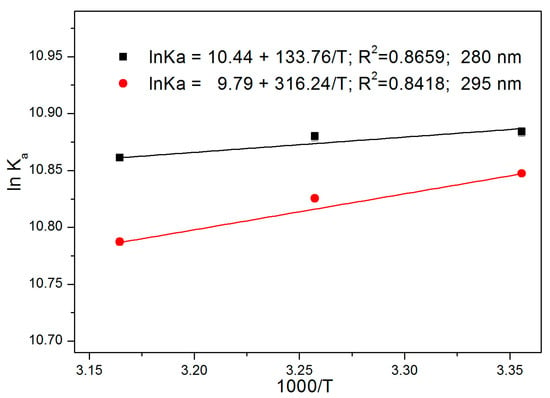
Figure 6.
Van’t Hoff plots for the ATR-HSA systems.
3.6. Study of the Synchronous Fluorescence Spectra
Synchronous fluorescence spectroscopy has many advantages and is often used to characterize complex mixtures; this technique can also be used to investigate changes in protein conformation [56]. The intrinsic fluorescence of HSA is mainly derived from Tyr and Trp residues [57]. In this experiment, synchronous fluorescence spectroscopy was used to assess the effect of metsulfuron-methyl, prometryn and ATR on the molecular conformation of HSA. When ∆λ was set to 15 nm, the synchronous fluorescence spectrum contained features characteristic of Tyr residues in HSA. When the distance between the excitation and emission wavelengths was set to 60 nm, the resulting synchronous fluorescence spectrum contained characteristics of Trp residues in HSA.
Figure 7 shows the change in the HSA synchronic fluorescence spectrum after the addition of ATR. When ∆λ = 60 nm, the synchronous fluorescence intensity decreased with increasing ATR concentration, and the maximum emission wavelength was a 4-nm blueshift; these results indicate that the histidine residues underwent conformational changes, polarity decreased, and hydrophobicity increased. When ∆λ = 15 nm, although the fluorescence intensity decreased with the increase in the ATR concentration, there was no obvious blueshift or redshift, and the quenching at ∆λ = 60 nm was better than at ∆λ = 15 nm. The above results indicate that ATR has no effect on the microenvironment of the Tyr residues but has an effect on the microenvironment of the Trp residues in HSA. In addition, these data indicate that ATR is bound closer to the Trp residue than to the Tyr residue and that the binding site is predominantly distributed over the Trp moiety.
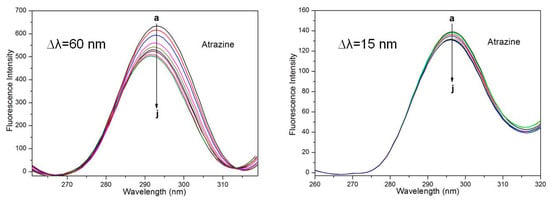
Figure 7.
Synchronous fluorescence spectra of HSA with various amounts of ATR.
3.7. Conformational Changes of HSA Detected by CD and 3D Fluorescence Spectroscopy
CD spectroscopy is widely used to study the interactions between small molecules and proteins because of its high sensitivity and because it is a very effective method to analyse the secondary structure of proteins [58,59,60]. Figure 8 shows CD spectra over an interval of 200–260 nm with and without ATR. The CD spectra of HSA have negative peaks at 208 nm and 220 nm, which are characteristic of an α-helix in the protein. With the continuous addition of ATR, the values of the CD spectra increased; that is, after the addition of ATR, the HSA structure changed. Table 5 lists the percentages of α-helixes, β-sheets, β-turns and random coils that were calculated using the CDSSTR algorithm in the CDPro package. α-helixes decreased as the ATR concentration increased, which indicates that the ATR has a direct effect on the amino acid residues of HSA, which leads to the destabilization of the spatial protein structure and to spiral structure extension. Agreeing with the FT-IR spectroscopy results of the Purcell’s study, it was shown that the binding of ATR to HSA results in a change in the conformation of HSA [61].
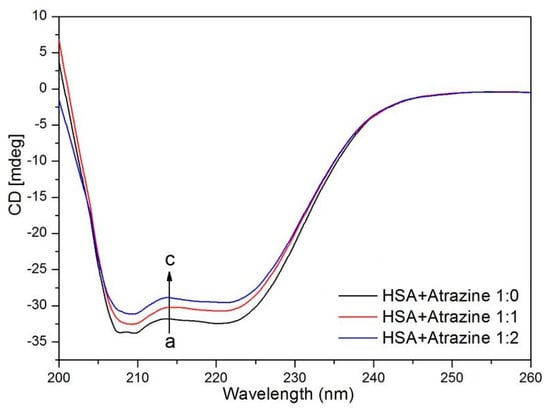
Figure 8.
Far-UV CD spectra of HSA in the presence of different concentrations of ATR.

Table 5.
Conformational changes in the secondary structure of HSA in the presence and absence of ATR.
Three-dimensional fluorescence spectroscopy is an effective method to comprehensively investigate protein structure [62]. The 3D fluorescence spectra are used to investigate the interference of ATR with the HSA spatial structure, which can reflect the structural changes of HSA. Table 6 lists 3D fluorescence characteristics. Figure 9 shows 3D fluorescence spectra of the ATR-HSA system. (λex = λem) is the Rayleigh scattering peak, peak b (λem = 2λex) is the secondary scattering peak, and peak 1 mainly expresses the spectral characteristics of the Tyr and Trp residues. Because the excitation wavelength was λex = 280 nm, the main excitation was related to the fluorescence of the Tyr and Trp residues, and the Phe residue fluorescence was negligible. Peak 2 mainly reflects the fluorescence signal of the polypeptide chain skeleton structure of the protein (for example, C = O). When the ATR are present, the fluorescence intensity of peak 2 decreases, indicating that the HSA peptide chain structure changed, which is consistent with the CD results. The fluorescence intensities of peaks 1 and 2 decreased, verifying the results of synchrotron fluorescence and CD. These results show that the addition of ATR exposed some hydrophobic residues to the water environment, resulting in a conformational transformation of the protein, i.e., a transition from a compact structure to a loose state in which the active portion of the protein was lost.

Table 6.
Intensity ratios of peaks 1 and 2.
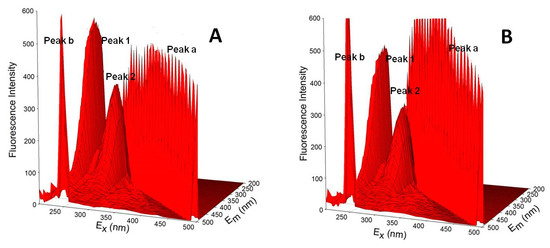
Figure 9.
Three-dimensional fluorescence spectra of free HSA (A) and HSA complexed with the ATR (B).
4. Conclusions
The interactions between ATR and HSA were studied using steady-state fluorescence spectroscopy at different excitation wavelengths, and the fluorescence quenching mechanism was determined for static quenching. We verified that the fluorescence quenched during the binding of ATR to HSA was mainly contributed to by Trp residues in HSA. The changes in the HSA conformation and secondary structure were verified using UV-Vis, 3D fluorescence, synchronous fluorescence, and CD spectroscopy. Based on the above data, ATR changes the conformation and secondary structure of HSA by combining at site II on HSA, and ATR and HSA (Trp) undergo energy transfer during binding, based on the calculations of the bond distance. The thermodynamic parameters indicated spontaneous combinations of ATR and HSA. ATR and HSA form a complex in a ratio of 1:1, primarily through electrostatic interactions. As ATR and HSA form a complex, fluorescence quenching occurs. The results of this study provide useful information for the use of ATR while minimizing its potential effects on humans.
Acknowledgments
The authors gratefully acknowledge financial support from the National Natural Science Foundation of China (Nos. 31601657 and 31701808), the Anhui Agricultural University Youth Fund Project (2014zr003 and yj2015-25), the Provincial Training Programs of Innovation and Entrepreneurship for Undergraduates (No. 201610364024), special funds of agro-product quality safety risk assessment of the Ministry of Agriculture of the People’s Republic of China (GJFP201600501, GJFP201700501 and GJFP201800501) and Open Funding of Key Laboratory of Agri-food Safety of Anhui Province.
Author Contributions
This research was carried on by all authors. Yi Wang and Meiqing Zhu designed the theme of the study and prepared the manuscript. Lijun Wang, Yi Wang, Jie Zhou, Jie Ding, Wei Li, Yue Xin, Shisuo Fan and Zhen Wang participated in the fluorescence experiments.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Broszat, M.; Ernst, H.; Spangenberg, B. A simple method for quantifying triazine herbicides using thin-layer chromatography and a CCD camera. J. Liquid Chromatogr. Relat. Technol. 2010, 33, 948–956. [Google Scholar] [CrossRef]
- Fernandez, M.V.; Gardinali, P.R. Risk assessment of triazine herbicides in surface waters and bioaccumulation of Irgarol and M1 by submerged aquatic vegetation in Southeast Florida. Sci. Total Environ. 2016, 541, 1556–1571. [Google Scholar] [CrossRef] [PubMed]
- Frias, S.; Sanchez, M.J.; Rodriguez, M.A. Determination of triazine compounds in ground water samples by micellar electrokinetic capillary chromatography. Anal. Chim. Acta 2004, 503, 271–278. [Google Scholar] [CrossRef]
- Alfonso, M.; Collados, R.; Yruela, I.; Picorel, R. Photoinhibition and recovery in a herbicide-resistant mutant from Glycine max (L.) Merr. cell cultures deficient in fatty acid unsaturation. Planta 2004, 219, 428–439. [Google Scholar] [CrossRef] [PubMed]
- Buonasera, K.; Pezzotti, G.; Scognamiglio, V.; Tibuzzi, A.; Giardi, M.T. New platform of biosensors for prescreening of pesticide residues to support laboratory analyses. J. Agric. Food Chem. 2010, 58, 5982–5990. [Google Scholar] [CrossRef] [PubMed]
- Lambreva, M.D.; Giardi, M.T.; Rambaldi, I.; Antonacci, A.; Pastorelli, S.; Bertalan, I.; Husu, I.; Johanningmeier, U.; Rea, G. A powerful molecular engineering tool provided efficient chlamydomonas mutants as bio-sensing elements for herbicides detection. PLoS ONE 2013, 8, e061851. [Google Scholar] [CrossRef] [PubMed]
- Thiel, H.; Varrelmann, M. Identification of a new PSII target site psbA mutation leading to D1 amino acid Leu (218) Val exchange in the Chenopodium album D1 protein and comparison to cross-resistance profiles of known modifications at positions 251 and 264. Pest Manag. Sci. 2014, 70, 278–285. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Liu, Z.F.; Deng, G.G.; Chen, W.; Li, M.Y.; Yang, L.J.; Li, Y.; Yang, X.D.; Zhang, H.B. Synthesis and antitumor activity of novel N-substituted tetrahydro-beta-carboline-imidazolium salt derivatives. Org. Biomol. Chem. 2016, 14, 9423–9430. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.J.; Xia, S.; Ma, S.X.; Zhou, S.Y.; Zhao, X.Q.; Wang, S.H.; Li, M.Y.; Yang, X.D. Host-guest system of hesperetin and beta-cyclodextrin or its derivatives: Preparation, characterization, inclusion mode, solubilization and stability. Mater. Sci. Eng. 2016, 59, 1016–1024. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Gonzalez-Esguevillas, M.; Berritt, S.; Yang, X.; Bellomo, A.; Walsh, P.J. Palladium-catalyzed C-H arylation of alpha, beta-unsaturated imines: Catalyst-controlled synthesis of enamine and allylic amine derivatives. Angew. Chem. 2016, 55, 2825–2829. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Li, Y.; Li, A.M. Treatment of actual chemical wastewater by a heterogeneous fenton process using natural pyrite. Int. J. Environ. Res. Public Health 2015, 12, 13762–13778. [Google Scholar] [CrossRef] [PubMed]
- Brvar, M.; Okrajsek, R.; Kosmina, P.; Staric, F.; Kaps, R.; Kozelj, G.; Bunc, M. Metabolic acidosis in prometryn (triazine herbicide) self-poisoning. Clin. Toxicol. 2008, 46, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Zhong, J.; Li, S.; Li, M.; Wu, L.; Wang, Q.; Mao, J.; Liu, S.; Zheng, B.; Wang, M.; et al. Total syntheses of (R)-Strongylodiols C and D. J. Nat. Prod. 2016, 79, 244–247. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.S.; Zhao, Q.; Yan, X.; Li, H.Y.; Zhang, P.; Wang, L.; Zhou, T.Y.; Li, Y.; Ding, L. Rapid preparation of expanded graphite by microwave irradiation for the extraction of triazine herbicides in milk samples. Food Chem. 2016, 197, 943–949. [Google Scholar] [CrossRef] [PubMed]
- Boffetta, P.; Adami, H.O.; Berry, C.; Mandel, J.S. Atrazine and cancer: A review of the epidemiologic evidence. Eur. J. Cancer Prev. 2013, 22, 169–180. [Google Scholar] [CrossRef] [PubMed]
- Gammon, D.W.; Aldous, C.N.; Carr, W.C.; Sanborn, J.R.; Pfeifer, K.F. A risk assessment of atrazine use in California: Human health and ecological aspects. Pest Manag. Sci. 2005, 61, 331–355. [Google Scholar] [CrossRef] [PubMed]
- Kuhnisch, J.; Goddon, I.; Berger, S.; Senkel, H.; Bucher, K.; Oehme, T.; Hickel, R.; Heinrich-Weltzien, R. Development, methodology and potential of the new universal visual scoring system (UniViSS) for caries detection and diagnosis. Int. J. Environ. Res. Public Health 2009, 6, 2500–2509. [Google Scholar] [CrossRef] [PubMed]
- Cavas, T. In vivo genotoxicity evaluation of atrazine and atrazine-based herbicide on fish Carassius auratus using the micronucleus test and the comet assay. Food Chem. Toxicol. 2011, 49, 1431–1435. [Google Scholar] [CrossRef] [PubMed]
- Stara, A.; Kristan, J.; Zuskova, E.; Velisek, J. Effect of chronic exposure to prometryne on oxidative stress and antioxidant response in common carp (Cyprinus carpio L.). Pest Biochem. Phys. 2013, 105, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Vieira, C.E.D.; Costa, P.G.; Lunardelli, B.; de Oliveira, L.F.; Cabrera, L.D.; Risso, W.E.; Primel, E.G.; Meletti, P.C.; Fillmann, G.; dos Reis Martinez, C.B. Multiple biomarker responses in Prochilodus lineatus subjected to short-term in situ exposure to streams from agricultural areas in Southern Brazil. Sci. Total Environ. 2016, 542, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Sha, S.C.; Li, M.; Kim, B.S.; Morgan, C.; Huang, R.; Yang, X.; Walsh, P.J. Nickel-catalyzed arylation of heteroaryl-containing diarylmethanes: Exceptional reactivity of the Ni (NIXANTPHOS)-based catalyst. Chem. Sci. 2016, 7, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Yucel, B.; Jiménez, J.; Rotella, M.; Fu, Y.; Walsh, P.J. Umpolung synthesis of diarylmethylaminesviapalladium—Catalyzed arylation ofn-benzyl aldimines. Adv. Synth. Catal. 2016, 358, 1910–1915. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Zhang, X. Spectrophotometric studies on the interaction between pazufloxacin mesilate and human serum albumin or lysozyme. J. Lumin. 2008, 128, 81–86. [Google Scholar] [CrossRef]
- Yue, Y.Y.; Zhang, Y.H.; Li, Y.; Zhu, J.H.; Qin, J.; Chen, X.G. Interaction of nobiletin with human serum albumin studied using optical spectroscopy and molecular modeling methods. J. Lumin. 2008, 128, 513–520. [Google Scholar] [CrossRef]
- Chen, C.L.; Capanema, E.A.; Gracz, H.S. Comparative studies on the delignification of pine kraft-anthraquinone pulp with hydrogen peroxide by binucleus Mn (IV) complex catalysis. J. Agric. Food Chem. 2003, 51, 6223–6232. [Google Scholar] [CrossRef] [PubMed]
- Liitia, T.M.; Maunu, S.L.; Hortling, B.; Toikka, M.; Kilpelainen, I. Analysis of technical lignins by two- and three-dimensional NMR spectroscopy. J. Agric. Food Chem. 2003, 51, 2136–2143. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, Y.; Jiang, J.; Liu, R.; Li, M.; Wang, Y.; Su, Y.; Zhu, B.; Zhang, S.; Huang, W. A DRIFTS study of low-temperature CO oxidation over Au/SnO2 catalyst prepared by co-precipitation method. Catal. Commun. 2009, 10, 640–644. [Google Scholar] [CrossRef]
- Wang, S.; Zhang, J.; Jiang, J.; Liu, R.; Zhu, B.; Xu, M.; Wang, Y.; Cao, J.; Li, M.; Yuan, Z.; et al. Porous ceria hollow microspheres: Synthesis and characterization. Microporous Mesoporous Mater. 2009, 123, 349–353. [Google Scholar] [CrossRef]
- Makarska-Bialokoz, M. Investigation of the binding affinity in vitamin B12-Bovine serum albumin system using various spectroscopic methods. Spectrochim. Acta A 2017, 184, 262–269. [Google Scholar] [CrossRef] [PubMed]
- Cysewski, P. A post-SCF complete basis set study on the recognition patterns of uracil and cytosine by aromatic and pi-aromatic stacking interactions with amino acid residues. Phys. Chem. Chem. Phys. 2008, 10, 2636–2645. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Yucel, B.; Adrio, J.; Bellomo, A.; Walsh, P.J. Synthesis of diarylmethylamines via palladium-catalyzed regioselective arylation of 1,1,3-triaryl-2-azaallyl anions. Chem. Sci. 2014, 5, 2383–2391. [Google Scholar] [CrossRef] [PubMed]
- Makarska-Bialokoz, M. Analysis of the binding interaction in uric acid—Human hemoglobin system by spectroscopic techniques. Spectrochim. Acta A 2017, 178, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.Y.; Zhang, Y.P.; Chen, X.Q.; Peng, M.J. Investigation of flavonoids bearing different substituents on ring c and their Cu2+ complex binding with bovine serum albumin: Structure-affinity relationship aspects. J. Agric. Food Chem. 2011, 59, 10761–10769. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.Q.; Bao, Y.H.; Chen, Y. Characteristics and antioxidant activity of water-soluble mail lard reaction products from interactions in a whey protein isolate and sugars system. Food Chem. 2013, 139, 355–361. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.S.; Al-Lohedan, H.A. Interaction of human serum albumin with sulfadiazine. J. Mol. Liq. 2014, 197, 124–130. [Google Scholar] [CrossRef]
- Li, M.; Berritt, S.; Walsh, P.J. Palladium-catalyzed regioselective arylation of 1,1,3-triaryl-2-azaallyl anions with aryl chlorides. Org. Lett. 2014, 16, 4312–4315. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhu, M.Q.; Liu, F.; Wu, X.W.; Pan, D.D.; Liu, J.; Fan, S.S.; Wang, Z.; Tang, J.; Na, R.S.; et al. Comparative studies of interactions between fluorodihydroquinazolin derivatives and human serum albumin with fluorescence spectroscopy. Molecules 2016, 21, 1373. [Google Scholar] [CrossRef] [PubMed]
- Soares, S.; Mateus, N.; de Freitas, V. Interaction of different polyphenols with bovine serum albumin (BSA) and human salivary alpha-amylase (HSA) by fluorescence quenching. J. Agric. Food Chem. 2007, 55, 6726–6735. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.X.; Fan, X.Y.; Jiang, F.L.; Liu, Y.; Lei, K.L. Resonance energy transfer, pH-induced folded states and the molecular interaction of human serum albumin and icariin. Luminescence 2015, 30, 1026–1033. [Google Scholar] [CrossRef] [PubMed]
- Mueller, D.R.; Krenos, J. Reaction of metastable Ar* (P-3 (2)) and Kr* (P-3 (2)) atoms with water vapor: Excitation functions for electronic quenching collisions. J. Phys. Chem. B 2002, 106, 8142–8147. [Google Scholar] [CrossRef]
- Liu, F.P.; Zhong, J.C.; Zheng, B.; Li, S.N.; Gao, G.; Wang, Z.Y.; Li, M.Y.; Hou, S.C.; Wang, M.; Bian, Q.H. Catalytic asymmetric synthesis of (S,4E,15Z)-docosa-4,15-dien-1-yn-3-ol, an antitumor marine natural product. Tetrahedron 2015, 26, 961–965. [Google Scholar] [CrossRef]
- Sohn, Y. Structural and spectroscopic characteristics of terbium hydroxide/oxide nanorods and plates. Ceram. Int. 2014, 40, 13803–13811. [Google Scholar] [CrossRef]
- Yang, R.H.; Zhu, J.Z.; Li, Y.L.; Zhang, H. A study on the preparation of regular multiple micro-electrolysis filler and the application in pretreatment of oil refinery wastewater. Int. J. Environ. Res. Public Health 2016, 13, 457. [Google Scholar] [CrossRef] [PubMed]
- Zheng, B.; Li, M.; Gao, G.; He, Y.; Walsh, P.J. Palladium-catalyzed α-arylation of methyl sulfonamides with aryl chlorides. Adv. Synth. Catal. 2016, 358, 2156–2162. [Google Scholar] [CrossRef] [PubMed]
- Ding, F.; Wang, M.L.; Zhang, S.X.; Ai, X.Z. Changes in SBPase activity influence photosynthetic capacity, growth, and tolerance to chilling stress in transgenic tomato plants. Sci. Rep. 2016, 6, 32741–32755. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Kim, B.S.; Li, M.; Walsh, P.J. Palladium-catalyzed selective alpha-alkenylation of pyridylmethyl ethers with vinyl bromides. Org. Lett. 2016, 18, 2371–2374. [Google Scholar] [CrossRef] [PubMed]
- Bessonneau, V.; Thomas, O. Assessment of exposure to alcohol vapor from alcohol-based hand rubs. Int. J. Environ. Res. Public Health 2012, 9, 868–879. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Wang, Q.Y.; Yang, T.; Cao, J.Z.; Lin, Q.L.; Yuan, Z.Q.; Li, L. Polyethyleneimine capped silver nanoclusters as efficient antibacterial agents. Int. J. Environ. Res. Public Health 2016, 13, 334. [Google Scholar] [CrossRef] [PubMed]
- Lakowicz, J.R. On spectral relaxation in proteins. Photochem. Photobiol. 2000, 72, 421–437. [Google Scholar] [CrossRef]
- Li, M.; Gutierrez, O.; Berritt, S.; Pascual-Escudero, A.; Yeşilçimen, A.; Yang, X.; Adrio, J.; Huang, G.; Nakamaru-Ogiso, E.; Kozlowski, M.C.; et al. Transition-metal-free chemo- and regioselective vinylation of azaallyls. Nat. Chem. 2017, 9, 997–1004. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.C.; Wang, H.M.; Niu, Q.X.; Ye, D.Y.; Liang, G.W. Binding between saikosaponin C and human serum albumin by fluorescence spectroscopy and molecular docking. Molecules 2016, 21, 153. [Google Scholar] [CrossRef] [PubMed]
- Handing, K.B.; Shabalin, I.G.; Szlachta, K.; Majorek, K.A.; Minor, W. Crystal structure of equine serum albumin in complex with cetirizine reveals a novel drug binding site. Mol. Immunol. 2016, 71, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Forster, M.T.; Hoecker, A.C.; Kang, J.S.; Quick, J.; Seifert, V.; Hattingen, E.; Hilker, R.; Weise, L.M. Does navigated transcranial stimulation increase the accuracy of tractography? A prospective clinical trial based on intraoperative motor evoked potential monitoring during deep brain stimulation. Neurosurgery 2015, 76, 766–775. [Google Scholar] [CrossRef] [PubMed]
- Ross, P.D.; Conway, J.F.; Cheng, N.Q.; Dierkes, L.; Firek, B.A.; Hendrix, R.W.; Steven, A.C.; Duda, R.L. A free energy cascade with locks drives assembly and maturation of bacteriophage HK97 capsid. J. Mol. Biol. 2006, 364, 512–525. [Google Scholar] [CrossRef] [PubMed]
- Sessions, R.B.; Thomas, G.L.; Parker, M.J. Water as a conformational editor in protein folding. J. Mol. Biol. 2004, 343, 1125–1133. [Google Scholar] [CrossRef] [PubMed]
- Peng, W.; Ding, F.; Peng, Y.K.; Jiang, Y.T.; Zhang, L. Binding patterns and structure-affinity relationships of food azo dyes with lysozyme: A multitechnique approach. J. Agric. Food Chem. 2013, 61, 12415–12428. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.Q.; Lei, H.G.; Min, S.G.; Duan, Z.Q. Spectroscopic study on interaction of rodenticide brodifacoum with bovine serum albumin. Spectrosc. Spect. Anal. 2009, 29, 2998–3002. [Google Scholar]
- Ding, F.; Huang, J.L.; Lin, J.; Li, Z.Y.; Liu, F.; Jiang, Z.Q.; Sun, Y. A study of the binding of CI Mordant Red 3 with bovine serum albumin using fluorescence spectroscopy. Dyes Pigment. 2009, 82, 65–70. [Google Scholar] [CrossRef]
- Dong, J.H.; Dai, W.T.; Xu, J.R.; Li, S.N. Spectral estimation model construction of heavy metals in mining reclamation areas. Int. J. Environ. Res. Public Health 2016, 13, 640. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.X.; Zhao, W.J.; Yan, X.; Shu, T.T.; Xiong, Q.L.; Chen, F.T. Pollution characteristics and health risk assessment of airborne heavy metals collected from Beijing bus stations. Int. J. Environ. Res. Public Health 2015, 12, 9658–9671. [Google Scholar] [CrossRef] [PubMed]
- Purcell, M.; Neault, J.F.; Malonga, H.; Arakawa, H.; Carpentier, R.; Tajmir-Riahi, H.A. Interactions of atrazine and 2,4-D with human serum albumin studied by gel and capillary electrophoresis, and FTIR spectroscopy. Biochim. Biophys. Acta 2001, 1548, 129–138. [Google Scholar] [CrossRef]
- Zhou, X.M.; Lu, W.J.; Su, L.; Shan, Z.J.; Chen, X.G. Binding of phthalate plasticizers to human serum albumin in vitro: A multispectroscopic approach and molecular modeling. J. Agric. Food Chem. 2012, 60, 1135–1145. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).