Association of LPP and TAGAP Polymorphisms with Celiac Disease Risk: A Meta-Analysis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Search Strategy
2.2. Inclusion and Exclusion Criteria
2.3. Data Extraction
2.4. Risk of Bias Assessment
2.5. Statistical Analysis
3. Results
3.1. Identifying Relevant Studies
3.2. Risk of Bias Assessment
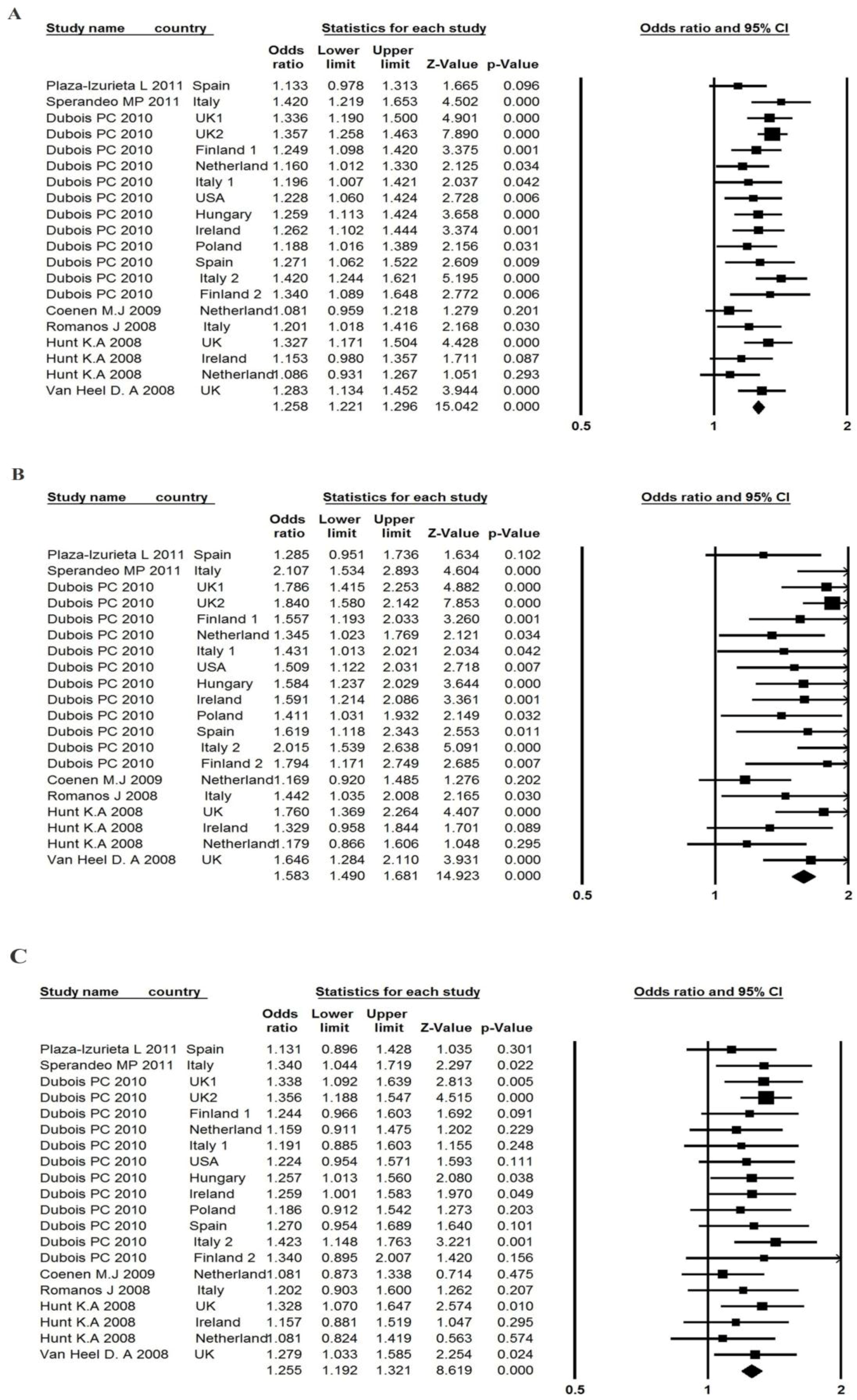
3.3. Association between the LPP rs1464510 Polymorphism and CD Risk
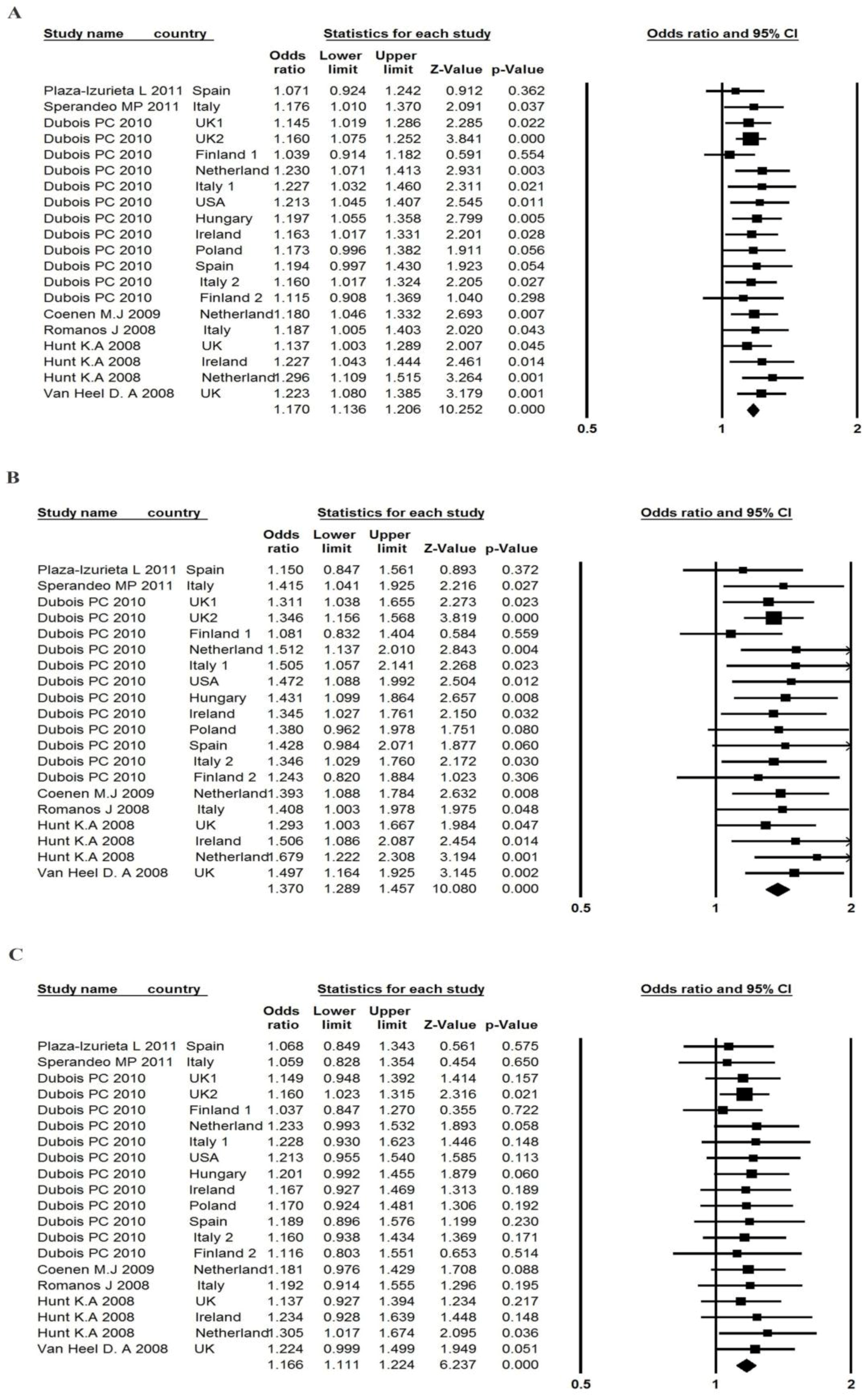
3.4. Association between the TAGAP rs1738074 Polymorphism and CD Risk
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ludvigsson, J.F.; Leffler, D.A.; Bai, J.C.; Biagi, F.; Fasano, A.; Green, P.H.; Hadjivassiliou, M.; Kaukinen, K.; Kelly, C.P.; Leonard, J.N.; et al. The Oslo definitions for coeliac disease and related terms. Gut 2013, 62, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Catassi, C.; Gatti, S.; Fasano, A. The new epidemiology of celiac disease. J. Pediatr. Gastroenterol. Nutr. 2014, 59, S7–S9. [Google Scholar] [CrossRef] [PubMed]
- Van De Kamer, J.H.; Weijers, H.A. Coeliac disease. V. Some experiments on the cause of the harmful effect of wheat gliadin. Acta Paediatr. 1955, 44, 465–469. [Google Scholar] [CrossRef] [PubMed]
- Sollid, L.M. Coeliac disease: Dissecting a complex inflammatory disorder. Nat. Rev. Immunol. 2002, 2, 647–655. [Google Scholar] [CrossRef] [PubMed]
- Castellanos-Rubio, A.; Martin-Pagola, A.; Santín, I.; Hualde, I.; Aransay, A.M.; Castaño, L.; Vitoria, J.C.; Bilbao, J.R. Combined functional and positional gene information for the identification of susceptibility variants in celiac disease. Gastroenterology 2008, 134, 738–746. [Google Scholar] [CrossRef] [PubMed]
- Van Heel, D.A.; Hunt, K.; Greco, L.; Wijmenga, C. Genetics in coeliac disease. Best Pract. Res. Clin. Gastroenterol. 2005, 19, 323–339. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Izurieta, L.; Castellanos-Rubio, A.; Irastorza, I.; Fernandez-Jimenez, N.; Gutierrez, G.; Bilbao, J.R. Revisiting genome wide association studies (GWAS) in coeliac disease: Replication study in spanish population and expression analysis of candidate genes. J. Med. Genet. 2011, 48, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Dubois, P.C.A.; Trynka, G.; Franke, L.; Hunt, K.A.; Romanos, J.; Curtotti, A.; Zhernakova, A.; Heap, G.A.R.; Ádány, R.; Aromaa, A.; et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 2010, 42, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Hunt, K.A.; Zhernakova, A.; Turner, G.; Heap, G.A.; Franke, L.; Bruinenberg, M.; Romanos, J.; Dinesen, L.C.; Ryan, A.W.; Panesar, D.; et al. Newly identified genetic risk variants for celiac disease related to the immune response. Nat. Genet. 2008, 40, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Van Heel, D.A.; Franke, L.; Hunt, K.A.; Gwilliam, R.; Zhernakova, A.; Inouye, M.; Wapenaar, M.C.; Barnardo, M.C.N.M.; Bethel, G.; Holmes, G.K.T.; et al. A genome-wide association study for celiac disease identifies risk variants in the region harboring IL2 and IL21. Nat. Genet. 2007, 39, 827–829. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Stahl, E.A.; Kurreeman, F.A.S.; Gregersen, P.K.; Siminovitch, K.A.; Worthington, J.; Padyukov, L.; Raychaudhuri, S.; Plenge, R.M. Fine mapping the TAGAP risk locus in rheumatoid arthritis. Genes Immun. 2011, 12, 314–318. [Google Scholar] [CrossRef] [PubMed]
- Festen, E.A.; Goyette, P.; Green, T.; Boucher, G.; Beauchamp, C.; Trynka, G.; Dubois, P.C.; Lagace, C.; Stokkers, P.C.; Hommes, D.W.; et al. A meta-analysis of genome-wide association scans identifies IL18RAP, PTPN2, TAGAP, and PUS10 as shared risk loci for Crohn’s disease and celiac disease. PLoS Genet. 2011, 7, e1001283. [Google Scholar] [CrossRef] [PubMed]
- Smyth, D.J.; Plagnol, V.; Walker, N.M.; Cooper, J.D.; Downes, K.; Yang, J.H.M.; Howson, J.M.M.; Stevens, H.; McManus, R.; Wijmenga, C.; et al. Shared and distinct genetic variants in type 1 diabetes and celiac disease. N. Engl. J. Med. 2008, 359, 2767–2777. [Google Scholar] [CrossRef] [PubMed]
- Eyre, S.; Hinks, A.; Bowes, J.; Flynn, E.; Martin, P.; Wilson, A.G.; Morgan, A.W.; Emery, P.; Steer, S.; Hocking, L.J.; et al. Overlapping genetic susceptibility variants between three autoimmune disorders: Rheumatoid arthritis, type 1 diabetes and coeliac disease. Arthritis Res. Ther. 2010. [Google Scholar] [CrossRef] [PubMed]
- Hinks, A.; Martin, P.; Flynn, E.; Eyre, S.; Packham, J.; Barton, A.; Worthington, J.; Thomson, W. Investigation of type 1 diabetes and coeliac disease susceptibility loci for association with juvenile idiopathic arthritis. Ann. Rheum. Dis. 2010, 69, 2169–2172. [Google Scholar] [CrossRef] [PubMed]
- Hue, S.; Mention, J.J.; Monteiro, R.C.; Zhang, S.; Cellier, C.; Schmitz, J.; Verkarre, V.; Fodil, N.; Bahram, S.; Cerf-Bensussan, N.; et al. A direct role for NKG2D/MICA interaction in villous atrophy during celiac disease. Immunity 2004, 21, 367–377. [Google Scholar] [CrossRef] [PubMed]
- Holmgren Peterson, K.; Magnusson, K.E.; Stenhammar, L.; Falth-Magnusson, K. Confocal laser scanning microscopy of small-intestinal mucosa in celiac disease. Scand. J. Gastroenterol. 1995, 30, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Goldmann, W.H. Mechanical aspects of cell shape regulation and signaling. Cell Biol. Int. 2002, 26, 313–317. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, S.; Nanayakkara, M.; Kosova, R.; Lania, G.; Sarno, M.; Gaito, A.; Galatola, M.; Greco, L.; Cuomo, M.; Troncone, R.; et al. A celiac cellular phenotype, with altered LPP sub-cellular distribution, is inducible in controls by the toxic gliadin peptide p31–43. PLoS ONE 2013, 8, e79763. [Google Scholar]
- Mertins, P.; Eberl, H.C.; Renkawitz, J.; Olsen, J.V.; Tremblay, M.L.; Mann, M.; Ullrich, A.; Daub, H. Investigation of protein-tyrosine phosphatase 1B function by quantitative proteomics. Mol. Cell. Proteom. 2008, 7, 1763–1777. [Google Scholar] [CrossRef] [PubMed]
- Dube, N.; Cheng, A.; Tremblay, M.L. The role of protein tyrosine phosphatase 1B in ras signaling. Proc. Natl. Acad. Sci. USA. 2004, 101, 1834–1839. [Google Scholar] [CrossRef] [PubMed]
- Trackman, P.C.; Nanayakkara, M.; Lania, G.; Maglio, M.; Kosova, R.; Sarno, M.; Gaito, A.; Discepolo, V.; Troncone, R.; Auricchio, S.; et al. Enterocyte proliferation and signaling are constitutively altered in celiac disease. PLoS ONE 2013, 8, e76006. [Google Scholar]
- Connelly, T.M.; Berg, A.S.; Harris, L.R.; Hegarty, J.P.; Ruggiero, F.M.; Deiling, S.M.; Brinton, D.L.; Koltun, W.A. T-cell activation Rho GTPase–activating protein expression varies with inflammation location and severity in Crohn’s disease. J. Surg. Res. 2014, 190, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Connelly, T.M.; Sehgal, R.; Berg, A.S.; Hegarty, J.P.; Deiling, S.; Stewart, D.B.; Poritz, L.S.; Koltun, W.A. Mutation in tagap is protective of anal sepsis in ileocolic crohn’s disease. Dis. Colon Rectum 2012, 55, 1145–1152. [Google Scholar] [CrossRef] [PubMed]
- Moon, S.Y.; Zheng, Y. Rho GTPase-activating proteins in cell regulation. Trends Cell Biol. 2003, 13, 13–22. [Google Scholar] [CrossRef]
- Zheng, Y. Dbl family guanine nucleotide exchange factors. Trends Biochem. Sci. 2001, 26, 724–732. [Google Scholar] [CrossRef]
- Lamarche, N.; Hall, A. Gaps for rho-related GTPases. Trends Genet. 1994, 10, 436–440. [Google Scholar] [CrossRef]
- Mardilovich, K.; Olson, M.F.; Baugh, M. Targeting rho GTPase signaling for cancer therapy. Future Oncol. 2012, 8, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Ligeti, E.; Welti, S.; Scheffzek, K. Inhibition and termination of physiological responses by GTPase activating proteins. Physiol. Rev. 2012, 92, 237–272. [Google Scholar] [CrossRef] [PubMed]
- Coenen, M.J.H.; Trynka, G.; Heskamp, S.; Franke, B.; van Diemen, C.C.; Smolonska, J.; van Leeuwen, M.; Brouwer, E.; Boezen, M.H.; Postma, D.S.; et al. Common and different genetic background for rheumatoid arthritis and coeliac disease. Hum. Mol. Genet. 2009, 18, 4195–4203. [Google Scholar] [CrossRef] [PubMed]
- Sperandeo, M.P.; Tosco, A.; Izzo, V.; Tucci, F.; Troncone, R.; Auricchio, R.; Romanos, J.; Trynka, G.; Auricchio, S.; Jabri, B.; et al. Potential celiac patients: A model of celiac disease pathogenesis. PLoS ONE 2011, 6, e21281. [Google Scholar] [CrossRef] [PubMed]
- Thakkinstian, A.; McKay, G.J.; McEvoy, M.; Chakravarthy, U.; Chakrabarti, S.; Silvestri, G.; Kaur, I.; Li, X.; Attia, J. Systematic review and meta-analysis of the association between complement component 3 and age-related macular degeneration: A huge review and meta-analysis. Am. J. Epidemiol. 2011, 173, 1365–1379. [Google Scholar] [CrossRef] [PubMed]
- Romanos, J.; Barisani, D.; Trynka, G.; Zhernakova, A.; Bardella, M.T.; Wijmenga, C. Six new coeliac disease loci replicated in an italian population confirm association with coeliac disease. J. Med. Genet. 2008, 46, 60–63. [Google Scholar] [CrossRef] [PubMed]
- Munafò, M.R.; Flint, J. Meta-analysis of genetic association studies. Trends Genet. 2004, 20, 439–444. [Google Scholar] [CrossRef] [PubMed]
- Petit, M.M.R. The focal adhesion and nuclear targeting capacity of the LIM-containing lipoma-preferred partner (LPP) protein. J. Biol. Chem. 2002, 278, 2157–2168. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Kern, M.J.; Otey, C.A.; Wamhoff, B.R.; Somlyo, A.V. Angiotensin II, focal adhesion kinase, and PRX1 enhance smooth muscle expression of lipoma preferred partner and its newly identified binding partner palladin to promote cell migration. Circ. Res. 2007, 100, 817–825. [Google Scholar] [CrossRef] [PubMed]
- Feighery, C.F.; McManus, R. Session 3: Joint nutrition society and irish nutrition and dietetic institute symposium on “nutrition and autoimmune disease”. Proc. Nutr. Soc. 2009, 68, 122. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Wijmenga, C.; Withoff, S. From genome-wide association studies to disease mechanisms: Celiac disease as a model for autoimmune diseases. Semin. Immunopathol. 2012, 34, 567–580. [Google Scholar] [CrossRef] [PubMed]

| Authors, Year (Ref.) | Ethnicity | Genotype Method | Gene | Type of SNP | MAF | Sample Size | ||
|---|---|---|---|---|---|---|---|---|
| Case | Control | Case | Control | |||||
| Plaza-Izurieta et al., 2011 [7] | Spanish | RT-PCR | LPP | rs1464510 | 0.450 | 0.419 | 1094 | 540 |
| TAGAP | rs1738074 | 0.423 | 0.406 | |||||
| Sperandeo et al., 2011 [31] | Italian | TaqMan | LPP | rs1464510 | 0.493 | 0.406 | 637 | 711 |
| TAGAP | rs1738074 | 0.465 | 0.425 | |||||
| Dubois et al., 2010 [8] | British | Illumina Hap300v1-1 + IlluminaHap550-2v3 | LPP | rs1464510 | 0.522 | 0.450 | 737 | 2596 |
| TAGAP | rs1738074 | 0.472 | 0.438 | |||||
| British | Illumina 670-QuadCustom_v1 + Illumina 1.2MDuoCustom_v1 | LPP | rs1464510 | 0.524 | 0.448 | 1849 | 4936 | |
| TAGAP | rs1738074 | 0.475 | 0.438 | |||||
| Finnish | Illumina 670-QuadCustom_v1 + Illumina610-Quad | LPP | rs1464510 | 0.601 | 0.547 | 647 | 1829 | |
| TAGAP | rs1738074 | 0.430 | 0.421 | |||||
| Dutch | Illumina 670-QuadCustom_v1 | LPP | rs1464510 | 0.531 | 0.493 | 803 | 846 | |
| TAGAP | rs1738074 | 0.445 | 0.395 | |||||
| Italian | Illumina 670-QuadCustom_v1 | LPP | rs1464510 | 0.517 | 0.472 | 497 | 543 | |
| TAGAP | rs1738074 | 0.464 | 0.413 | |||||
| American | IlluminaGoldenGate | LPP | rs1464510 | 0.511 | 0.459 | 973 | 555 | |
| TAGAP | rs1738074 | 0.470 | 0.423 | |||||
| Hungarian | IlluminaGoldenGate | LPP | rs1464510 | 0.533 | 0.475 | 965 | 1067 | |
| TAGAP | rs1738074 | 0.415 | 0.372 | |||||
| Irish | IlluminaGoldenGate | LPP | rs1464510 | 0.501 | 0.443 | 597 | 1456 | |
| TAGAP | rs1738074 | 0.500 | 0.462 | |||||
| Polish | IlluminaGoldenGate | LPP | rs1464510 | 0.495 | 0.452 | 564 | 716 | |
| TAGAP | rs1738074 | 0.364 | 0.328 | |||||
| Spanish | IlluminaGoldenGate | LPP | rs1464510 | 0.462 | 0.403 | 550 | 433 | |
| TAGAP | rs1738074 | 0.443 | 0.400 | |||||
| Italian | IlluminaGoldenGate | LPP | rs1464510 | 0.495 | 0.408 | 1010 | 804 | |
| TAGAP | rs1738074 | 0.461 | 0.425 | |||||
| Finnish | IlluminaGoldenGate + Illumina610-Quad | LPP | rs1464510 | 0.602 | 0.531 | 259 | 653 | |
| TAGAP | rs1738074 | 0.448 | 0.421 | |||||
| Coenen et al., 2009 [30] | Dutch | Illumina HAP550 | LPP | rs1464510 | 0.530 | 0.510 | 795 | 1683 |
| TAGAP | rs1738074 | 0.440 | 0.400 | |||||
| Romanos et al., 2008 [33] | Italian | TaqMan technology | LPP | rs1464510 | 0.520 | 0.474 | 538 | 593 |
| TAGAP | rs1738074 | 0.454 | 0.412 | |||||
| Hunt et al., 2008 [9] | British | IlluminaGoldenGate | LPP | rs1464510 | 0.517 | 0.446 | 719 | 1561 |
| TAGAP | rs1738074 | 0.460 | 0.428 | |||||
| Irish | IlluminaGoldenGate | LPP | rs1464510 | 0.483 | 0.448 | 416 | 957 | |
| TAGAP | rs1738074 | 0.519 | 0.468 | |||||
| Dutch | IlluminaGoldenGate | LPP | rs1464510 | 0.521 | 0.500 | 508 | 888 | |
| TAGAP | rs1738074 | 0.459 | 0.395 | |||||
| Van Heel et al., 2008 [10] | British | Illumina Hap300 | LPP | rs1464510 | 0.519 | 0.457 | 778 | 1422 |
| TAGAP | rs1738074 | 0.472 | 0.422 | |||||
| Author, Year (Ref.) | Ascertainment of Celiac Disease | Ascertainment of Control | Quality Control for Genotyping | Population Stratification | Confounding Bias | Selective Outcome Report | HWE |
|---|---|---|---|---|---|---|---|
| Plaza-Izurieta et al., 2011 [7] | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| Sperandeo et al., 2011 [31] | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| Dubois et al., 2010 [8] | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| Coenen et al., 2009 [30] | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| Romanos et al., 2008 [33] | Yes | Yes | Unclear | Yes | Unclear | Yes | Yes |
| Hunt et al., 2008 [9] | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| Van Heel et al., 2008 [10] | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| Author (Ref.) | Country | Case Genotype | Control Genotype | A vs. C | AA vs. CC | AC vs. CC | HWE | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AA | AC | CC | AA | AC | CC | OR | 95% CI | OR | 95% CI | OR | 95% CI | |||
| Plaza-Izurieta et al. [7] | Spain | 222 | 541 | 331 | 95 | 263 | 182 | 1.133 | 0.978–1.313 | 1.258 | 0.951–1.736 | 1.131 | 0.896–1.428 | 0.999 |
| Sperandeo et al. [31] | Italy | 152 | 324 | 161 | 108 | 362 | 241 | 1.420 | 1.219–1.653 | 2.107 | 1.534–2.893 | 1.340 | 1.044–1.719 | 0.141 |
| Dubois et al. [8] | UK1 | 201 | 368 | 168 | 526 | 1285 | 785 | 1.336 | 1.190–1.500 | 1.786 | 1.415–2.253 | 1.338 | 1.092–1.639 | 0.997 |
| UK2 | 508 | 922 | 419 | 991 | 2441 | 1504 | 1.357 | 1.258–1.463 | 1.840 | 1.580–2.142 | 1.356 | 1.188–1.547 | 0.992 | |
| Finland 1 | 234 | 310 | 103 | 547 | 907 | 375 | 1.249 | 1.098–1.420 | 1.557 | 1.193–2.033 | 1.244 | 0.966–1.603 | 0.978 | |
| The Netherlands | 226 | 400 | 177 | 206 | 423 | 217 | 1.160 | 1.012–1.330 | 1.345 | 1.023–1.769 | 1.159 | 0.911–1.475 | 0.996 | |
| Italy 1 | 133 | 248 | 116 | 121 | 271 | 151 | 1.196 | 1.007–1.421 | 1.431 | 1.013–2.021 | 1.191 | 0.885–1.603 | 0.977 | |
| USA | 254 | 486 | 233 | 117 | 276 | 162 | 1.228 | 1.060–1.424 | 1.509 | 1.122–2.031 | 1.224 | 0.954–1.571 | 0.978 | |
| Hungary | 274 | 480 | 211 | 241 | 532 | 294 | 1.259 | 1.113–1.424 | 1.584 | 1.237–2.029 | 1.257 | 1.013–1.560 | 0.991 | |
| Ireland | 150 | 298 | 149 | 286 | 718 | 452 | 1.262 | 1.102–1.444 | 1.591 | 1.214–2.086 | 1.259 | 1.001–1.583 | 0.977 | |
| Poland | 138 | 282 | 144 | 146 | 355 | 215 | 1.188 | 1.016–1.389 | 1.411 | 1.031–1.932 | 1.186 | 0.912–1.542 | 0.980 | |
| Spain | 117 | 274 | 159 | 70 | 209 | 154 | 1.271 | 1.062–1.522 | 1.619 | 1.118–2.343 | 1.270 | 0.954–1.689 | 0.948 | |
| Italy 2 | 247 | 505 | 258 | 134 | 388 | 282 | 1.420 | 1.244–1.621 | 2.015 | 1.539–2.638 | 1.423 | 1.148–1.763 | 0.978 | |
| Finland 2 | 94 | 124 | 41 | 184 | 325 | 144 | 1.340 | 1.089–1.648 | 1.794 | 1.171–2.749 | 1.340 | 0.895–2.007 | 0.983 | |
| Coenen et al. [30] | The Netherlands | 223 | 396 | 176 | 438 | 841 | 404 | 1.081 | 0.959–1.218 | 1.169 | 0.920–1.485 | 1.081 | 0.873–1.338 | 0.994 |
| Romanos et al. [33] | Italy | 145 | 269 | 124 | 133 | 296 | 164 | 1.201 | 1.018–1.416 | 1.442 | 1.035–2.008 | 1.202 | 0.903–1.600 | 0.980 |
| Hunt et al. [9] | UK | 192 | 359 | 168 | 311 | 771 | 479 | 1.327 | 1.171–1.504 | 1.760 | 1.369–2.264 | 1.328 | 1.070–1.647 | 0.981 |
| Ireland | 97 | 208 | 111 | 192 | 473 | 292 | 1.153 | 0.980–1.357 | 1.329 | 0.958–1.844 | 1.157 | 0.881–1.519 | 0.986 | |
| The Netherlands | 138 | 253 | 117 | 222 | 444 | 222 | 1.086 | 0.931–1.267 | 1.179 | 0.866–1.606 | 1.081 | 0.824–1.419 | 1.000 | |
| Van Heel et al. [10] | UK | 210 | 388 | 180 | 297 | 706 | 419 | 1.283 | 1.134–1.452 | 1.646 | 1.284–2.110 | 1.279 | 1.033–1.585 | 0.990 |
| Overall odds ratio | - | - | - | - | - | - | - | 1.258 | 1.221–1.296 | 1.583 | 1.490–1.681 | 1.255 | 1.192–1.321 | - |
| Author (Ref.) | Country | Case Genotype | Control Genotype | A vs. G | AA vs. GG | AG vs. GG | HWE | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AA | AG | GG | AA | AG | GG | OR | 95% CI | OR | 95% CI | OR | 95% CI | |||
| Plaza-Izurieta et al. [7] | Spain | 196 | 534 | 364 | 89 | 261 | 190 | 1.071 | 0.924–1.242 | 1.150 | 0.847–1.561 | 1.068 | 0.849–1.343 | 0.968 |
| Sperandeo et al. [31] | Italy | 144 | 305 | 188 | 125 | 354 | 231 | 1.176 | 1.010–1.370 | 1.415 | 1.041–1.925 | 1.059 | 0.828–1.354 | 0.596 |
| Dubois et al. [8] | UK1 | 164 | 367 | 205 | 498 | 1278 | 820 | 1.145 | 1.019–1.286 | 1.311 | 1.038–1.655 | 1.149 | 0.948–1.392 | 0.999 |
| UK2 | 417 | 922 | 510 | 947 | 2430 | 1559 | 1.160 | 1.075–1.252 | 1.346 | 1.156–1.568 | 1.160 | 1.023–1.315 | 0.999 | |
| Finland 1 | 120 | 317 | 210 | 324 | 892 | 613 | 1.039 | 0.914–1.182 | 1.081 | 0.832–1.404 | 1.037 | 0.847–1.270 | 0.987 | |
| The Netherlands | 159 | 397 | 247 | 132 | 404 | 310 | 1.230 | 1.071–1.413 | 1.512 | 1.137-2.010 | 1.233 | 0.993–1.532 | 0.984 | |
| Italy 1 | 107 | 247 | 143 | 93 | 263 | 187 | 1.227 | 1.032–1.460 | 1.505 | 1.057–2.141 | 1.228 | 0.930–1.623 | 0.974 | |
| USA | 215 | 485 | 273 | 99 | 271 | 185 | 1.213 | 1.045–1.407 | 1.472 | 1.088–1.992 | 1.213 | 0.955–1.540 | 0.989 | |
| Hungary | 166 | 469 | 330 | 148 | 498 | 421 | 1.197 | 1.055–1.358 | 1.431 | 1.099–1.864 | 1.201 | 0.992–1.455 | 0.970 | |
| Ireland | 149 | 299 | 149 | 311 | 724 | 421 | 1.163 | 1.017–1.331 | 1.345 | 1.027–1.761 | 1.167 | 0.927–1.469 | 0.993 | |
| Poland | 75 | 261 | 228 | 77 | 316 | 323 | 1.173 | 0.996–1.382 | 1.380 | 0.962–1.978 | 1.170 | 0.924–1.481 | 0.982 | |
| Spain | 108 | 271 | 171 | 69 | 208 | 156 | 1.194 | 0.997–1.430 | 1.428 | 0.984–2.071 | 1.189 | 0.896–1.576 | 0.981 | |
| Italy 2 | 215 | 502 | 293 | 145 | 393 | 266 | 1.160 | 1.017–1.324 | 1.346 | 1.029–1.760 | 1.160 | 0.938–1.434 | 0.994 | |
| Finland 2 | 52 | 128 | 79 | 116 | 318 | 219 | 1.115 | 0.908–1.369 | 1.243 | 0.820–1.884 | 1.116 | 0.803–1.551 | 0.976 | |
| Coenen et al. [30] | The Netherlands | 154 | 392 | 249 | 269 | 808 | 606 | 1.180 | 1.046–1.332 | 1.393 | 1.088–1.784 | 1.181 | 0.976–1.429 | 0.990 |
| Romanos et al. [33] | Italy | 111 | 267 | 160 | 101 | 287 | 205 | 1.187 | 1.005–1.403 | 1.408 | 1.003–1.978 | 1.192 | 0.914–1.555 | 0.974 |
| Hunt et al. [9] | UK | 152 | 357 | 210 | 286 | 764 | 511 | 1.137 | 1.003–1.289 | 1.293 | 1.003–1.667 | 1.137 | 0.927–1.394 | 0.988 |
| Ireland | 112 | 208 | 96 | 210 | 476 | 271 | 1.227 | 1.043–1.444 | 1.506 | 1.086–2.087 | 1.234 | 0.928–1.639 | 0.971 | |
| The Netherlands | 107 | 252 | 148 | 139 | 424 | 325 | 1.296 | 1.109–1.515 | 1.679 | 1.222–2.308 | 1.305 | 1.017–1.674 | 0.971 | |
| Van Heel et al. [10] | UK | 173 | 388 | 217 | 253 | 694 | 475 | 1.223 | 1.080–1.385 | 1.497 | 1.164–1.925 | 1.224 | 0.999–1.499 | 0.986 |
| Overall odds ratio | - | - | - | - | - | - | - | 1.170 | 1.136–1.206 | 1.370 | 1.289–1.457 | 1.166 | 1.111–1.224 | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, S.-Q.; Zhang, N.; Zhou, Z.-X.; Huang, C.-C.; Zeng, C.-L.; Xiao, D.; Guo, C.-C.; Han, Y.-J.; Ye, X.-H.; Ye, X.-G.; et al. Association of LPP and TAGAP Polymorphisms with Celiac Disease Risk: A Meta-Analysis. Int. J. Environ. Res. Public Health 2017, 14, 171. https://doi.org/10.3390/ijerph14020171
Huang S-Q, Zhang N, Zhou Z-X, Huang C-C, Zeng C-L, Xiao D, Guo C-C, Han Y-J, Ye X-H, Ye X-G, et al. Association of LPP and TAGAP Polymorphisms with Celiac Disease Risk: A Meta-Analysis. International Journal of Environmental Research and Public Health. 2017; 14(2):171. https://doi.org/10.3390/ijerph14020171
Chicago/Turabian StyleHuang, Shi-Qi, Na Zhang, Zi-Xing Zhou, Chui-Can Huang, Cheng-Li Zeng, Di Xiao, Cong-Cong Guo, Ya-Jing Han, Xiao-Hong Ye, Xing-Guang Ye, and et al. 2017. "Association of LPP and TAGAP Polymorphisms with Celiac Disease Risk: A Meta-Analysis" International Journal of Environmental Research and Public Health 14, no. 2: 171. https://doi.org/10.3390/ijerph14020171
APA StyleHuang, S.-Q., Zhang, N., Zhou, Z.-X., Huang, C.-C., Zeng, C.-L., Xiao, D., Guo, C.-C., Han, Y.-J., Ye, X.-H., Ye, X.-G., Ou, M.-L., Zhang, B.-H., Liu, Y., Zeng, E. Y., Yang, G., & Jing, C.-X. (2017). Association of LPP and TAGAP Polymorphisms with Celiac Disease Risk: A Meta-Analysis. International Journal of Environmental Research and Public Health, 14(2), 171. https://doi.org/10.3390/ijerph14020171





