MicroRNAs in Breastmilk and the Lactating Breast: Potential Immunoprotectors and Developmental Regulators for the Infant and the Mother
Abstract
:1. Introduction
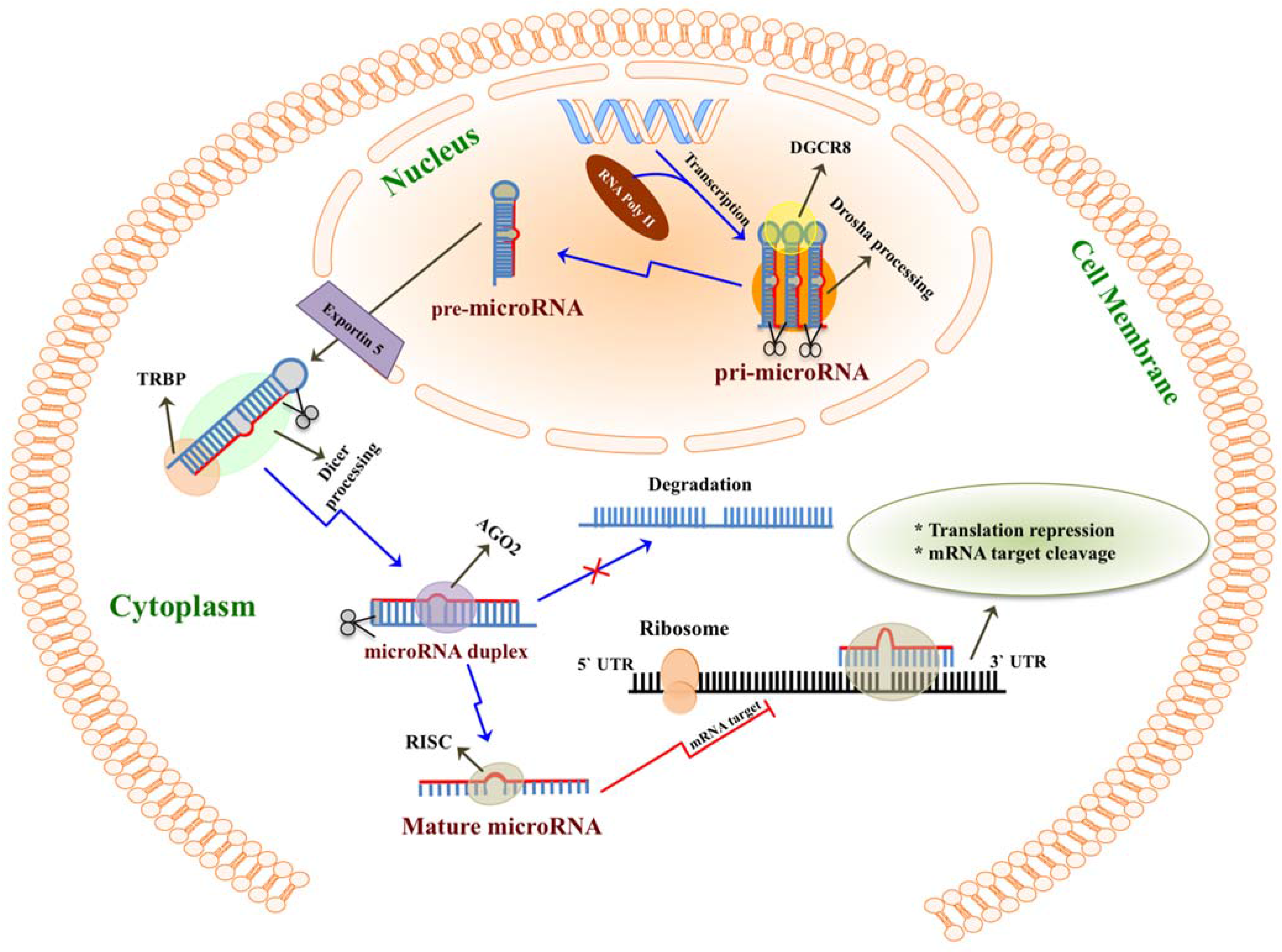
2. MicroRNAs Are Highly Enriched in Milk
2.1. microRNAs in Mammalian Milk
2.2. microRNAs in Different Milk Fractions
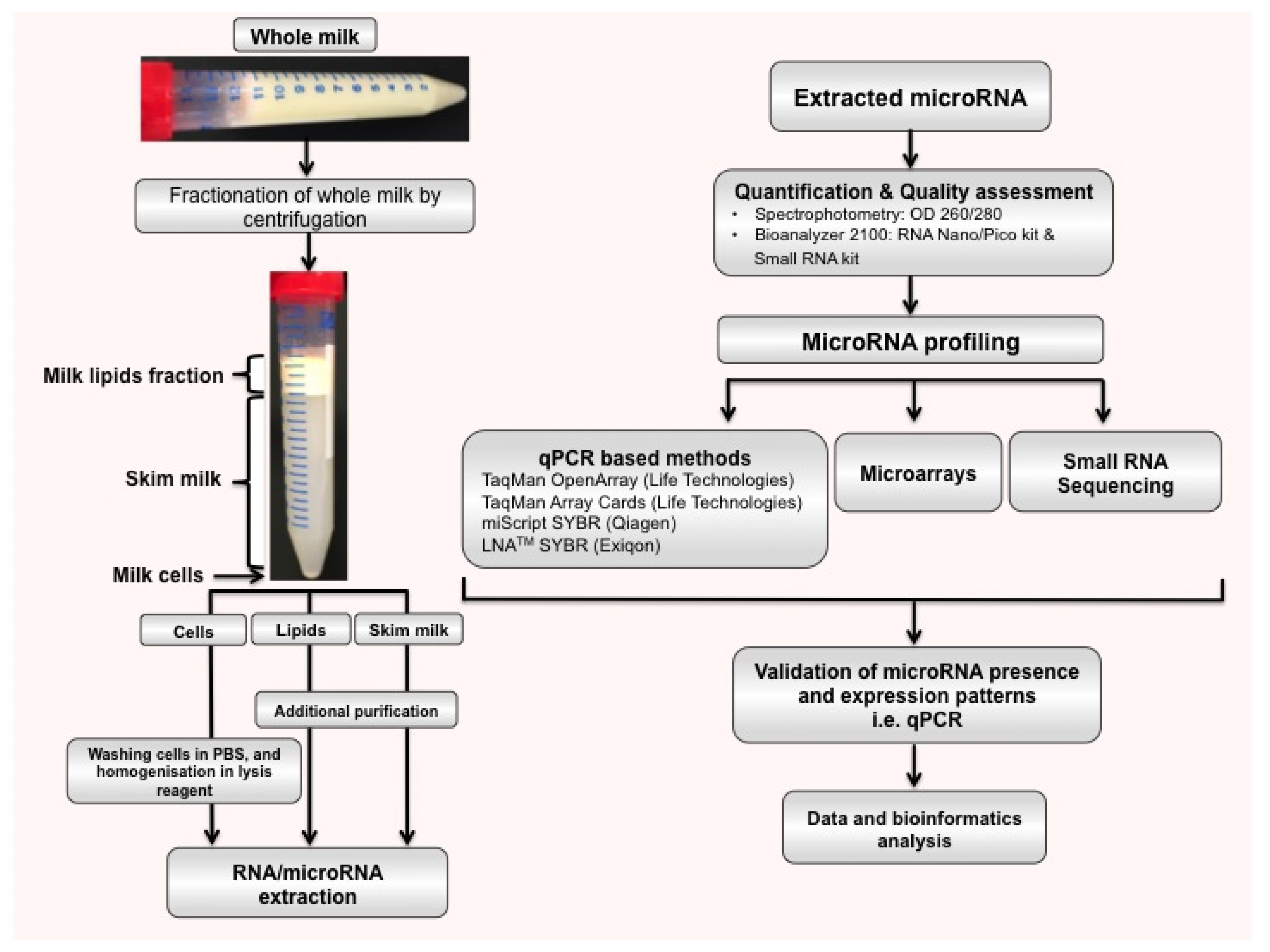
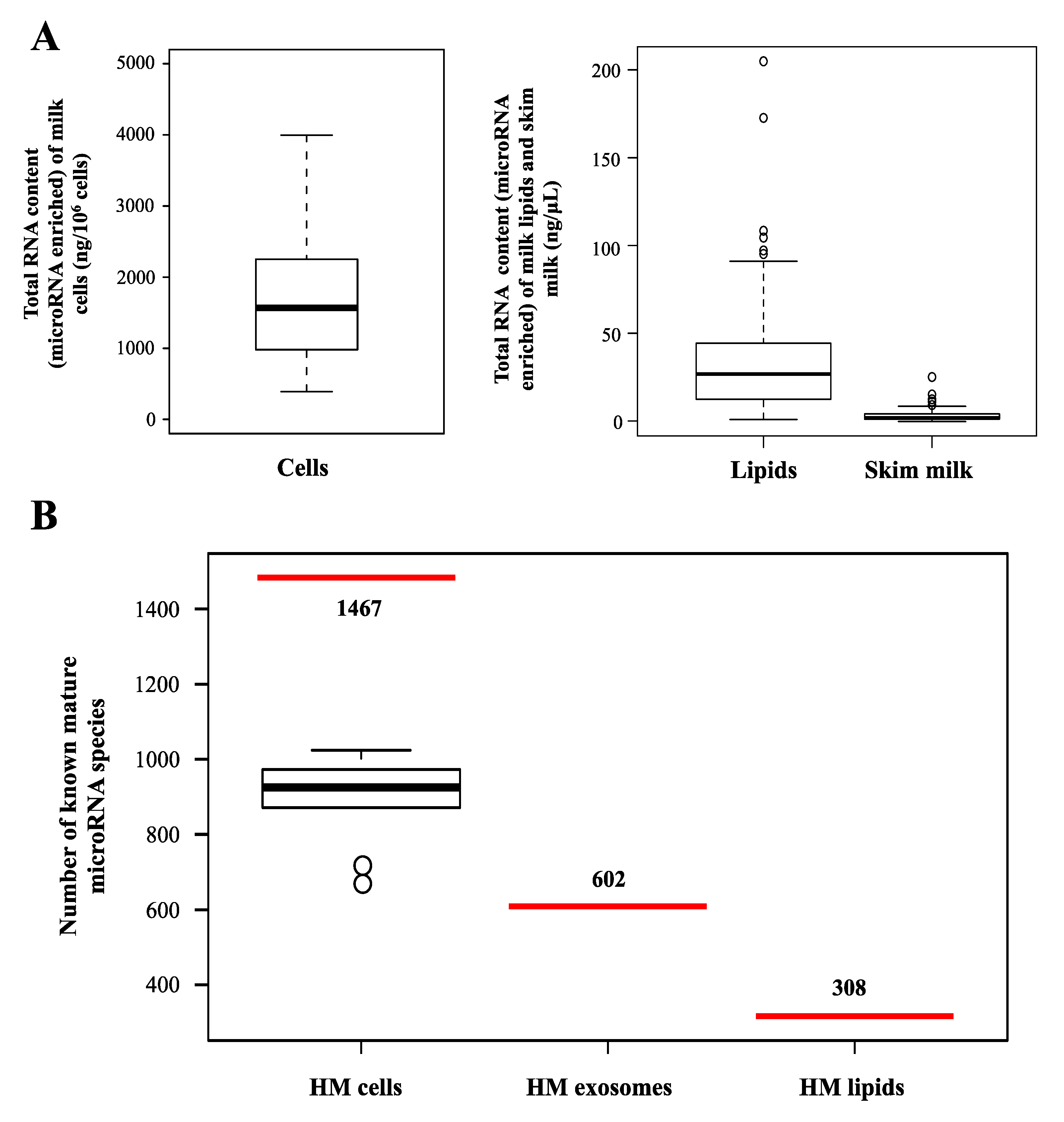
| Species | Milk Fraction | microRNAs * | Profiling Method | Reference |
|---|---|---|---|---|
| Human | Skim milk (mature) | 429 | qPCR | [30] |
| Skim milk (colostrum) | 386 | qPCR | [30] | |
| Skim milk (mature) | 281 | Microarray | [48] | |
| Milk exosomes | 639 ** | Solexa deep sequencing | [36] | |
| Milk lipids | 308 | Solexa deep sequencing | [44] | |
| Milk cells | 450 *** | TaqMan OpenArray | [45,46] | |
| Milk lipids | 337 *** | TaqMan OpenArray | [45,46] | |
| Milk cell pre-feed | 1287 | Solexa deep sequencing | [35] | |
| Milk cell post-feed | 1308 | Solexa deep sequencing | [35] | |
| Bovine | Skim milk (colostrum) | 230 | Solexa deep sequencing | [52] |
| Skim milk (mature) | 213 | Solexa deep sequencing | [52] | |
| Skim milk (colostrum) | 100 | Microarray | [54] | |
| Skim milk (mature) | 53 | Microarray | [54] | |
| Porcine | Milk exosomes | 180 ** | Solexa deep sequencing | [51] |
| Milk exosomes (colostrum) | 491 | Solexa deep sequencing | [55] | |
| Murine (rat) | Skim milk (colostrum) | 128 | Microarray | [56] |
| Skim milk (mature) | 144 | Microarray | [56] |
2.3. Origin of Milk microRNAs
2.4. Milk microRNAs as Diagnostic Tools
3. Functions of Milk microRNAs
3.1. Stability and Uptake of Food-Derived microRNAs
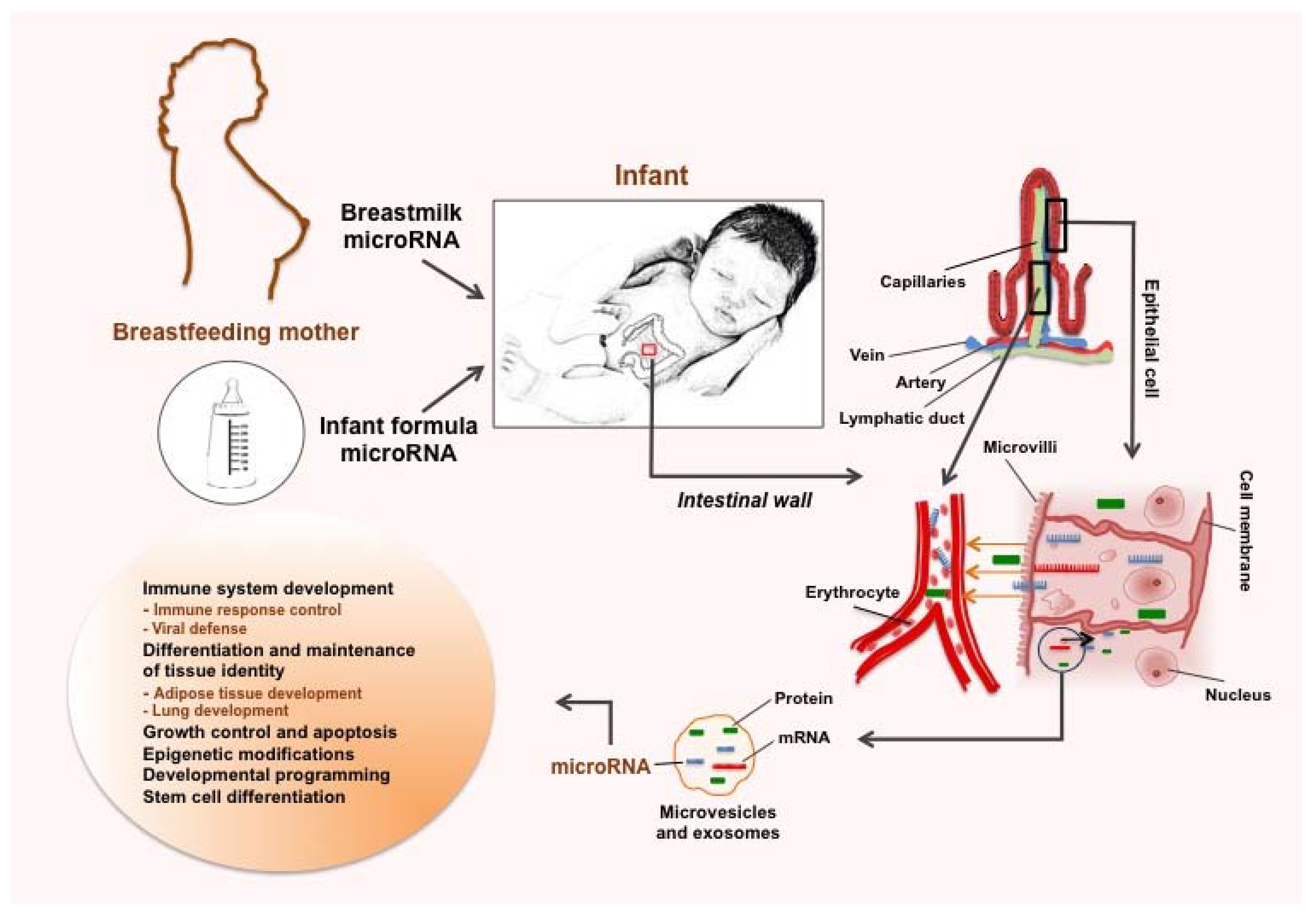
3.2. microRNAs Act as Immune Regulators
| MicroRNA | Regulatory Function(s) | References | Presence in HM | References | Presence in Animal Milk | References |
|---|---|---|---|---|---|---|
| miR-181a | Cell signaling. Development of B cells. | [105,107] | Skim milk. Milk lipids. | [30,44,48] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [52,55,56] |
| miR-181b | Switch recombination in activated B cells. Increased activity of NF-κB. | [105,108,109] | Skim milk (colostrum and mature milk). Milk lipids. | [30,44,48] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [52,55,56] |
| miR-155 | B and T cell differentiation. Innate/adaptive immune response. | [106,110,111] | Skim milk (colostrum and mature milk). Milk lipids. Milk exosomes. | [30,36,44,48] | Bovine skim milk (colostrum and mature milk). | [52,54] |
| miR-17 | B and T cells. Monocyte development. | [112,113] | Skim milk. Milk lipids. Milk exosomes. | [36,44,48] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [52,55,56] |
| miR-92a | B and T cells. Monocyte development. Downregulated in lymphoma. | [114,115] | Skim milk (colostrum and mature milk). Milk lipids. | [30,44,48] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [52,54,55,56] |
| miR-125b | Tumor necrosis factor-α production. Innate immune response. TLR signaling. | [111] | Skim milk (colostrum and mature milk). Milk lipids. | [30,44,48] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [52,54,55,56] |
| miR-146a | Innate immune response. TLR signaling. | [116,117] | Skim milk (colostrum and mature milk). Milk lipids. Milk exosomes. | [30,36,44,48] | Bovine skim milk (colostrum and mature milk). | [52,55] |
| miR-223 | Neutrophil proliferation and activation. Granulopoiesis. | [118,119,120] | Skim milk. Milk lipids. Milk exosomes. | [36,44,48] | Bovine skim milk (colostrum and mature milk). Rat milk whey. | [52,54,56] |
| miR-150 | B and T cells. Suppresses B cell differentiation. | [121,122] | Skim milk. Milk lipids. Milk exosomes. | [30,36,44,48] | Bovine skim milk (colostrum and mature milk). Rat milk whey. | [52,56] |
| miR-30b | Promotes induced cellular invasion. Immune suppression. | [123] | Skim milk (colostrum and mature milk). Milk lipid. Milk exosomes. | [30,36,44] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [51,52,56] |
| miR-182 | Promotes IL-2 (interleukin-2). Induces T cell-mediated immune responses. | [124] | Milk lipids. Milk exosomes. | [36,44] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [51,52,56] |
| miR-200a | Associated with Hodgkin lymphoma. | [125] | Skim milk (colostrum and mature milk). Milk lipids. Milk exosomes. | [30,36,44] | Bovine skim milk (colostrum and mature milk). Rat milk whey.Porcine milk exosomes. | [51,52,54,56] |
| miR-29a | Suppresses immune responses to intracellular pathogens. Downregulated in B cell chronic lymphocytic leukemia. | [126,127] | Skim milk (colostrum and mature milk). Milk lipids. Milk exosomes. | [30,36,44] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [52,54,55,56] |
| miR-15a | Downregulated in chronic lymphocytic leukemia. | [128,129] | Skim milk (colostrum and mature milk). Milk lipid. Milk exosomes. | [30,36,44] | Bovine skim milk (colostrum and mature milk). Porcine milk exosomes. | [52,54,55] |
| miR-16 | Induces TNF mRNA degradation. Upregulated in rheumatoid arthritis. | [20,130] | Skim milk. Milk lipids. | [30,44] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [52,55,56] |
| miR-21 | Up-regulated in B-cell lymphoma and chronic lymphocytic leukemia. | [131] | Skim milk (colostrum and mature milk). Milk lipids. Milk exosomes. | [30,36,44] | Bovine skim milk (colostrum and mature milk). Rat milk whey. Porcine milk exosomes. | [51,52,55,56] |
| miR-20a | Inhibits monocyte proliferation, differentiation and maturation. | [112] | Skim milk (colostrum and mature milk). Milk lipids. Milk exosomes. | [30,36,44] | Bovine skim milk (colostrum and mature milk). Rat milk whey. | [52,54,56] |
| miR-106a | Inhibits monocyte proliferation, differentiation and maturation. | [112] | Milk lipids. Milk exosomes. | [36,44] | Bovine skim milk (colostrum and mature milk). Porcine milk exosomes. | [52,54,55] |
3.3. microRNAs Are Key Regulators of Milk Lipid Metabolism
3.4. Various Potential Benefits of Human Milk microRNAs
4. Infant Formula is Poor in microRNAs Compared to Human Milk
| MicroRNA | Mature Sequence (miRBase 20.0) | Existence | Expression Level | References |
|---|---|---|---|---|
| bta-miR-26a hsa-miR-26a-5p | UUCAAGUAAUCCAGGAUAGGCU UUCAAGUAAUCCAGGAUAGGCU | Bovine milk. Infant formula. HM. | Low in formulae compared to raw bovine milk. Highly expressed in HM lipid fraction. | [44,52] |
| bta-miR-26b hsa-miR-26b-5p | UUCAAGUAAUUCAGGAUAGGUU UUCAAGUAAUUCAGGAUAGGU | Bovine milk. Infant formula. HM. | Low in formulae compared to raw bovine milk. Highly expressed in HM lipids and exosomes. | [36,44,52] |
| bta-miR-200c hsa-miR-200c-3p | UAAUACUGCCGGGUAAUGAUGGA UAAUACUGCCGGGUAAUGAUGGA | Bovine milk. Infant formula. HM. | Low in formulae compared to raw bovine milk. Highly expressed in HM lipid fraction. | [44,52,54] |
| bta-miR-21-5p hsa-miR-21-5p | UAGCUUAUCAGACUGAUGUUGACU UAGCUUAUCAGACUGAUGUUGA | Bovine milk. Infant formula. HM. | Low in formulae compared to raw bovine milk. Highly expressed in HM lipids, skim milk and exosomes. | [30,36,44,48,52] |
| bta-miR-30d hsa-miR-30d-5p | UGUAAACAUCCCCGACUGGAAGCU UGUAAACAUCCCCGACUGGAAG | Bovine milk. Infant formula. HM. | Low in formulae compared to raw bovine milk. Highly expressed in HM lipids and skim milk. | [30,44,52] |
| bta-miR-99a-5p hsa-miR-99a-5p | AACCCGUAGAUCCGAUCUUGU AACCCGUAGAUCCGAUCUUGUG | Bovine milk. Infant formula. HM. | Low in formulae compared to raw bovine milk. Highly expressed in HM lipids and skim milk. | [30,44,52] |
| bta-miR-148 hsa-miR-148a-3p | UCAGUGCACUACAGAACUUUGU UCAGUGCACUACAGAACUUUGU | Bovine milk. Infant formula. HM. | Low in formula compared to raw bovine milk. Highly expressed in HM lipid, skim milk and exosomes. | [30,36,44,52,54] |
5. Conclusions and Outlook
Acknowledgements
Author Contributions
Conflicts of Interest
References
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- He, L.; Hannon, G.J. MicroRNAs: Small RNAs with a big role in gene regulation. Nat. Rev. Genet. 2004, 5, 522–531. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, C.C.; Cheng, H.H.; Tewari, M. MicroRNA profiling: Approaches and considerations. Nat. Rev. Genet. 2012, 13, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Fabian, M.R.; Sonenberg, N.; Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 2010, 79, 351–379. [Google Scholar] [CrossRef] [PubMed]
- Krol, J.; Loedige, I.; Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 2010, 11, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Kim, V.N.; Han, J.; Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 2009, 10, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Du, T.; Zamore, P.D. microPrimer: The biogenesis and function of microRNA. Development 2005, 132, 4645–4652. [Google Scholar] [CrossRef] [PubMed]
- International Human Genome Sequencing, C. Finishing the euchromatic sequence of the human genome. Nature 2004, 431, 931–945. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.E. Functional aspects of animal microRNAs. Cell. Mol. Life Sci. 2008, 65, 545–562. [Google Scholar] [CrossRef] [PubMed]
- O’Driscoll, L. The emerging world of microRNAs. Anticancer Res. 2006, 26, 4271–4278. [Google Scholar] [PubMed]
- Houbaviy, H.B.; Murray, M.F.; Sharp, P.A. Embryonic stem cell-specific MicroRNAs. Dev. Cell 2003, 5, 351–358. [Google Scholar] [CrossRef]
- Judson, R.L.; Babiarz, J.E.; Venere, M.; Blelloch, R. Embryonic stem cell-specific microRNAs promote induced pluripotency. Nat. Biotechnol. 2009, 27, 459–461. [Google Scholar] [CrossRef] [PubMed]
- Xu, N.; Papagiannakopoulos, T.; Pan, G.; Thomson, J.A.; Kosik, K.S. MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 2009, 137, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Huangfu, D.; Maehr, R.; Guo, W.; Eijkelenboom, A.; Snitow, M.; Chen, A.E.; Melton, D.A. Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nat. Biotechnol. 2008, 26, 795–797. [Google Scholar] [CrossRef]
- Yoshida, Y.; Takahashi, K.; Okita, K.; Ichisaka, T.; Yamanaka, S. Hypoxia enhances the generation of induced pluripotent stem cells. Cell Stem Cell 2009, 5, 237–241. [Google Scholar] [CrossRef] [PubMed]
- Ventura, A.; Young, A.G.; Winslow, M.M.; Lintault, L.; Meissner, A.; Erkeland, S.J.; Newman, J.; Bronson, R.T.; Crowley, D.; Stone, J.R.; et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 2008, 132, 875–886. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Srinivasan, L.; Calado, D.P.; Patterson, H.C.; Zhang, B.; Wang, J.; Henderson, J.M.; Kutok, J.L.; Rajewsky, K. Lymphoproliferative disease and autoimmunity in mice with increased miR-17–92 expression in lymphocytes. Nat. Immunol. 2008, 9, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Jing, Q.; Huang, S.; Guth, S.; Zarubin, T.; Motoyama, A.; Chen, J.; Di Padova, F.; Lin, S.C.; Gram, H.; Han, J. Involvement of microRNA in AU-rich element-mediated mRNA instability. Cell 2005, 120, 623–634. [Google Scholar] [CrossRef] [PubMed]
- Landgraf, P.; Rusu, M.; Sheridan, R.; Sewer, A.; Iovino, N.; Aravin, A.; Pfeffer, S.; Rice, A.; Kamphorst, A.O.; Landthaler, M.; et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007, 129, 1401–1414. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, R.M.; Taganov, K.D.; Boldin, M.P.; Cheng, G.; Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 2007, 104, 1604–1609. [Google Scholar] [CrossRef] [PubMed]
- Monticelli, S.; Ansel, K.M.; Xiao, C.; Socci, N.D.; Krichevsky, A.M.; Thai, T.H.; Rajewsky, N.; Marks, D.S.; Sander, C.; Rajewsky, K.; et al. MicroRNA profiling of the murine hematopoietic system. Genome Biol. 2005. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Lin, H.; Xiao, J.; Lu, Y.; Luo, X.; Li, B.; Zhang, Y.; Xu, C.; Bai, Y.; Wang, H.; et al. The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat. Med. 2007, 13, 486–491. [Google Scholar] [CrossRef]
- Poy, M.N.; Eliasson, L.; Krutzfeldt, J.; Kuwajima, S.; Ma, X.; Macdonald, P.E.; Pfeffer, S.; Tuschl, T.; Rajewsky, N.; Rorsman, P.; et al. A pancreatic islet-specific microRNA regulates insulin secretion. Nature 2004, 432, 226–230. [Google Scholar] [CrossRef] [PubMed]
- Giraldez, A.J.; Cinalli, R.M.; Glasner, M.E.; Enright, A.J.; Thomson, J.M.; Baskerville, S.; Hammond, S.M.; Bartel, D.P.; Schier, A.F. MicroRNAs regulate brain morphogenesis in zebrafish. Science 2005, 308, 833–838. [Google Scholar] [CrossRef] [PubMed]
- Mahn, R.; Heukamp, L.C.; Rogenhofer, S.; von Ruecker, A.; Muller, S.C.; Ellinger, J. Circulating microRNAs (miRNA) in serum of patients with prostate cancer. Urology 2011. [Google Scholar] [CrossRef] [PubMed]
- Gotte, M. MicroRNAs in breast cancer pathogenesis. Minerva Ginecol. 2010, 62, 559–571. [Google Scholar] [PubMed]
- Si, M.L.; Zhu, S.; Wu, H.; Lu, Z.; Wu, F.; Mo, Y.Y. miR-21-mediated tumor growth. Oncogene 2007, 26, 2799–2803. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.H.; Lee, M.J.; Galas, D.J.; Wang, K. The microRNA spectrum in 12 body fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef] [PubMed]
- Silveri, L.; Tilly, G.; Vilotte, J.L.; Le Provost, F. MicroRNA involvement in mammary gland development and breast cancer. Reprod. Nutr. Dev. 2006, 46, 549–556. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, L.; Hou, D.; Chen, X.; Li, D.; Zhu, L.; Zhang, Y.; Li, J.; Bian, Z.; Liang, X.; Cai, X.; et al. Exogenous plant MIR168a specifically targets mammalian LDLRAP1: Evidence of cross-kingdom regulation by microRNA. Cell Res. 2012, 22, 107–126. [Google Scholar] [PubMed]
- Palmer, J.D.; Soule, B.P.; Simone, B.A.; Zaorsky, N.G.; Jin, L.; Simone, N.L. Dietary alterations caused by microRNA: Can food be medicinal? Ageing Res. Rev. 2014. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Sang, X.; Hong, Z. Beyond nutrients: Food-derived microRNAs provide cross-kingdom regulation. Bioessays 2012, 34, 280–284. [Google Scholar] [CrossRef] [PubMed]
- Alsaweed, M.; Tat-Lai, C.; Newnham, J.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Small RNA Sequencing of Human Milk Cells reveals Numerous Known and Novel miRNAs that may be affected by Milk Removal. In Proceedings of the Combined Biological Sciences Meeting, Perth, Australia, 28 August 2015.
- Zhou, Q.; Li, M.; Wang, X.; Li, Q.; Wang, T.; Zhu, Q.; Zhou, X.; Wang, X.; Gao, X.; Li, X. Immune-related microRNAs are abundant in breast milk exosomes. Int. J. Biol. Sci. 2012, 8, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Baier, S.R.; Nguyen, C.; Xie, F.; Wood, J.R.; Zempleni, J. MicroRNAs are absorbed in biologically meaningful amounts from nutritionally relevant doses of cow milk and affect gene expression in peripheral blood mononuclear cells, HEK-293 kidney cell cultures, and mouse livers. J. Nutr. 2014, 144, 1495–1500. [Google Scholar] [CrossRef] [PubMed]
- Wolf, T.; Baier, S.R.; Zempleni, J. The intestinal transport of bovine milk exosomes is mediated by endocytosis in human colon carcinoma Caco-2 cells and rat small intestinal IEC-6 cells. J. Nutr. 2015. [Google Scholar] [CrossRef] [PubMed]
- Pieters, B.C.; Arntz, O.J.; Bennink, M.B.; Broeren, M.G.; van Caam, A.P.; Koenders, M.I.; van Lent, P.L.; van den Berg, W.B.; de Vries, M.; van der Kraan, P.M.; et al. Commercial cow milk contains physically stable extracellular vesicles expressing immunoregulatory TGF-beta. PLoS ONE 2015. [Google Scholar] [CrossRef] [PubMed]
- Kramer, M.S. “Breast is best”: The evidence. Early Hum. Dev. 2010, 86, 729–732. [Google Scholar] [CrossRef] [PubMed]
- Kramer, M.S.; Kakuma, R. The optimal duration of exclusive breastfeeding. J. Adv. Nurs. 2001, 35, 313–315. [Google Scholar] [PubMed]
- Wang, K.; Zhang, S.; Weber, J.; Baxter, D.; Galas, D.J. Export of microRNAs and microRNA-protective protein by mammalian cells. Nucleic Acids Res. 2010, 38, 7248–7259. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Iguchi, H.; Ochiya, T. Circulating microRNA in body fluid: A new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010, 101, 2087–2092. [Google Scholar] [CrossRef] [PubMed]
- Munch, E.M.; Harris, R.A.; Mohammad, M.; Benham, A.L.; Pejerrey, S.M.; Showalter, L.; Hu, M.; Shope, C.D.; Maningat, P.D.; Gunaratne, P.H.; et al. Transcriptome profiling of microRNA by Next-Gen deep sequencing reveals known and novel miRNA species in the lipid fraction of human breast milk. PLoS ONE 2013. [Google Scholar] [CrossRef] [PubMed]
- Alsaweed, M.; Tat-Lai, C.; Newnham, J.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Breastmilk miRNA Primarily Originate from the Mammary Gland. In Proceedings of Combined Biological Sciences Meeting, Perth, Australia, 28 August 2015.
- Alsaweed, M.; Tat-Lai, C.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Human milk miRNAs primarily originate from the mammary gland resulting in unique miRNA profiles of fractionated milk. Scientific Reports 2015. under review. [Google Scholar]
- Alsaweed, M.; Hepworth, A.R.; Lefevre, C.; Hartmann, P.E.; Geddes, D.T.; Hassiotou, F. Human milk microRNA and total RNA differ depending on milk fractionation. J. Cell Biochem. 2015. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Izumi, H.; Sekine, K.; Ochiya, T. microRNA as a new immune-regulatory agent in breast milk. Silence 2010. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Liu, H.; Jin, X.; Lo, L.; Liu, J. Expression profiles of microRNAs from lactating and non-lactating bovine mammary glands and identification of miRNA related to lactation. BMC Genomics 2012. [Google Scholar] [CrossRef] [PubMed]
- Avril-Sassen, S.; Goldstein, L.D.; Stingl, J.; Blenkiron, C.; Le Quesne, J.; Spiteri, I.; Karagavriilidou, K.; Watson, C.J.; Tavare, S.; Miska, E.A.; et al. Characterisation of microRNA expression in post-natal mouse mammary gland development. BMC Genomics 2009. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Li, M.; Wang, T.; Liang, Y.; Zhong, Z.; Wang, X.; Zhou, Q.; Chen, L.; Lang, Q.; He, Z.; et al. Lactation-related microRNA expression profiles of porcine breast milk exosomes. PLoS ONE 2012. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Gao, C.; Li, H.; Huang, L.; Sun, Q.; Dong, Y.; Tian, C.; Gao, S.; Dong, H.; Guan, D.; et al. Identification and characterization of microRNAs in raw milk during different periods of lactation, commercial fluid, and powdered milk products. Cell Res. 2010, 20, 1128–1137. [Google Scholar] [CrossRef] [PubMed]
- Hata, T.; Murakami, K.; Nakatani, H.; Yamamoto, Y.; Matsuda, T.; Aoki, N. Isolation of bovine milk-derived microvesicles carrying mRNAs and microRNAs. Biochem. Biophys. Res. Commun. 2010, 396, 528–533. [Google Scholar] [CrossRef] [PubMed]
- Izumi, H.; Kosaka, N.; Shimizu, T.; Sekine, K.; Ochiya, T.; Takase, M. Bovine milk contains microRNA and messenger RNA that are stable under degradative conditions. J. Dairy Sci. 2012, 95, 4831–4841. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Xi, Q.Y.; Ye, R.S.; Cheng, X.; Qi, Q.E.; Wang, S.B.; Shu, G.; Wang, L.N.; Zhu, X.T.; Jiang, Q.Y.; et al. Exploration of microRNAs in porcine milk exosomes. BMC Genomics 2014, 15. [Google Scholar] [CrossRef] [PubMed]
- Izumi, H.; Kosaka, N.; Shimizu, T.; Sekine, K.; Ochiya, T.; Takase, M. Time-dependent expression profiles of microRNAs and mRNAs in rat milk whey. PLoS ONE 2014. [Google Scholar] [CrossRef] [PubMed]
- Modepalli, V.; Kumar, A.; Hinds, L.A.; Sharp, J.A.; Nicholas, K.R.; Lefevre, C. Differential temporal expression of milk miRNA during the lactation cycle of the marsupial tammar wallaby (Macropus eugenii). BMC Genomics 2014. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Geddes, D.T.; Hartmann, P.E. Cells in human milk: State of the science. J. Hum. Lact. 2013, 29, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Molinari, C.E.; Casadio, Y.S.; Hartmann, B.T.; Arthur, P.G.; Hartmann, P.E. Longitudinal analysis of protein glycosylation and beta-casein phosphorylation in term and preterm human milk during the first 2 months of lactation. Br. J. Nutr. 2013, 110, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Mitoulas, L.R.; Kent, J.C.; Cox, D.B.; Owens, R.A.; Sherriff, J.L.; Hartmann, P.E. Variation in fat, lactose and protein in human milk over 24 h and throughout the first year of lactation. Br. J. Nutr. 2002, 88, 29–37. [Google Scholar] [CrossRef]
- Hassiotou, F.; Hepworth, A.R.; Williams, T.M.; Twigger, A.J.; Perrella, S.; Lai, C.T.; Filgueira, L.; Geddes, D.T.; Hartmann, P.E. Breastmilk cell and fat contents respond similarly to removal of breastmilk by the infant. PLoS ONE 2013. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Hepworth, A.R.; Metzger, P.; Tat Lai, C.; Trengove, N.; Hartmann, P.E.; Filgueira, L. Maternal and infant infections stimulate a rapid leukocyte response in breastmilk. Clin. Transl. Immunol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Powe, C.E.; Knott, C.D.; Conklin-Brittain, N. Infant sex predicts breast milk energy content. Am. J. Hum. Biol. 2010, 22, 50–54. [Google Scholar] [CrossRef] [PubMed]
- Bachour, P.; Yafawi, R.; Jaber, F.; Choueiri, E.; Abdel-Razzak, Z. Effects of smoking, mother’s age, body mass index, and parity number on lipid, protein, and secretory immunoglobulin a concentrations of human milk. Breastfeed. Med. 2012, 7, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Bauer, J.; Gerss, J. Longitudinal analysis of macronutrients and minerals in human milk produced by mothers of preterm infants. Clin. Nutr. 2011, 30, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.W.; Jin, M.J.; Yu, Y.X.; Zhang, S.C.; Liu, B.; Jiang, X.; Pan, Y.F.; Li, Q.I.; Ma, S.Y.; Chen, K. Associations of lifestyle-related factors, hsa-miR-149 and hsa-miR-605 gene polymorphisms with gastrointestinal cancer risk. Mol. Carcinog. 2012. [Google Scholar] [CrossRef]
- Makrides, M.; Neumann, M.A.; Gibson, R.A. Effect of maternal docosahexaenoic acid (DHA) supplementation on breast milk composition. Eur. J. Clin. Nutr. 1996, 50, 352–357. [Google Scholar] [PubMed]
- Melnik, B.C.; John, S.M.; Schmitz, G. Milk consumption during pregnancy increases birth weight, a risk factor for the development of diseases of civilization. J. Transl. Med. 2015. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Wu, W.; Zhang, M.; Li, J.; Peng, Y.; Miao, T.T.; Zhu, H.; Xu, G. Aberrant upregulation of miR-21 in placental tissues of macrosomia. J. Perinatol. 2014, 34, 658–663. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Geddes, D.T. Immune cell-mediated protection of the mammary gland and the infant during breastfeeding. Adv. Nutr. 2015, 6, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Lawless, N.; Reinhardt, T.A.; Bryan, K.; Baker, M.; Pesch, B.; Zimmerman, D.; Zuelke, K.; Sonstegard, T.; O’Farrelly, C.; Lippolis, J.D.; et al. MicroRNA regulation of bovine monocyte inflammatory and metabolic networks in an in vivo infection model. G3 (Bethesda) 2014, 4, 957–971. [Google Scholar] [CrossRef] [PubMed]
- Sourvinou, I.S.; Markou, A.; Lianidou, E.S. Quantification of circulating miRNAs in plasma: Effect of preanalytical and analytical parameters on their isolation and stability. J. Mol. Diagn. 2013, 15, 827–834. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Moisa, S.; Khan, M.J.; Wang, J.; Bu, D.; Loor, J.J. MicroRNA expression patterns in the bovine mammary gland are affected by stage of lactation. J. Dairy Sci. 2012, 95, 6529–6535. [Google Scholar] [CrossRef] [PubMed]
- Bian, Y.; Lei, Y.; Wang, C.; Wang, J.; Wang, L.; Liu, L.; Liu, L.; Gao, X.; Li, Q. Epigenetic regulation of miR-29s affects the lactation activity of dairy cow mammary epithelial cells. J. Cell. Physiol. 2015, 230, 2152–2163. [Google Scholar] [CrossRef] [PubMed]
- Dickinson, B.; Zhang, Y.; Petrick, J.S.; Heck, G.; Ivashuta, S.; Marshall, W.S. Lack of detectable oral bioavailability of plant microRNAs after feeding in mice. Nat. Biotechnol. 2013, 31, 965–967. [Google Scholar] [CrossRef] [PubMed]
- Mu, J.; Zhuang, X.; Wang, Q.; Jiang, H.; Deng, Z.B.; Wang, B.; Zhang, L.; Kakar, S.; Jun, Y.; Miller, D.; et al. Interspecies communication between plant and mouse gut host cells through edible plant derived exosome-like nanoparticles. Mol. Nutr. Food Res. 2014, 58, 1561–1573. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Segura, L.; Perez-Andrade, M.; Miranda-Rios, J. The emerging role of MicroRNAs in the regulation of gene expression by nutrients. J. Nutrigenet. Nutrigenomics 2013, 6, 16–31. [Google Scholar] [CrossRef] [PubMed]
- Tian, T.; Wang, Y.; Wang, H.; Zhu, Z.; Xiao, Z. Visualizing of the cellular uptake and intracellular trafficking of exosomes by live-cell microscopy. J. Cell Biochem. 2010, 111, 488–496. [Google Scholar] [CrossRef] [PubMed]
- Tian, T.; Zhu, Y.L.; Hu, F.H.; Wang, Y.Y.; Huang, N.P.; Xiao, Z.D. Dynamics of exosome internalization and trafficking. J. Cell. Physiol. 2013, 228, 1487–1495. [Google Scholar] [CrossRef] [PubMed]
- Tian, T.; Zhu, Y.L.; Zhou, Y.Y.; Liang, G.F.; Wang, Y.Y.; Hu, F.H.; Xiao, Z.D. Exosome uptake through clathrin-mediated endocytosis and macropinocytosis and mediating miR-21 delivery. J. Biol. Chem. 2014, 289, 22258–22267. [Google Scholar] [CrossRef] [PubMed]
- Arntz, O.J.; Pieters, B.C.; Oliveira, M.C.; Broeren, M.G.; Bennink, M.B.; de Vries, M.; van Lent, P.L.; Koenders, M.I.; van den Berg, W.B.; et al. Oral administration of bovine milk derived extracellular vesicles attenuates arthritis in two mouse models. Mol. Nutr. Food Res. 2015, 59, 1701–1712. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Mobley, A.; Geddes, D.T.; Hartmann, P.E.; Wilkie, T.M. Breastmilk Imparts the Mother’s Stem Cells to the Infant; Experimental Biology: Boston, MA, USA, 2015. [Google Scholar]
- Hassiotou, F.; Mobley, A.; Ocal, O.; Filgueira, L.; Geddes, D.T.; Hartmann, P.E.; Wilkie, T.M. Breastmilk Stem Cell Transfer from the Mother to Neonatal Organs: A Route of Migration and Integration; International Society for Research in Human Milk and Lactation: Charleston, SC, USA, 2014. [Google Scholar]
- Sun, Q.; Chen, X.; Yu, J.; Zen, K.; Zhang, C.Y.; Li, L. Immune modulatory function of abundant immune-related microRNAs in microvesicles from bovine colostrum. Protein Cell 2013, 4, 197–210. [Google Scholar] [CrossRef] [PubMed]
- Ji, L.; Chen, X. Regulation of small RNA stability: Methylation and beyond. Cell Res. 2012, 22, 624–636. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Ba, Y.; Ma, L.; Cai, X.; Yin, Y.; Wang, K.; Guo, J.; Zhang, Y.; Chen, J.; Guo, X.; et al. Characterization of microRNAs in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008, 18, 997–1006. [Google Scholar] [CrossRef] [PubMed]
- Bai, W.L.; Yin, R.H.; Yang, R.J.; Khan, W.A.; Ma, Z.J.; Zhao, S.J.; Jiang, W.Q.; Wang, Z.Y.; Zhu, Y.B.; Luo, G.B.; et al. Technical note: Identification of suitable normalizers for microRNA expression analysis in milk somatic cells of the yak (Bos grunniens). J. Dairy Sci. 2013, 96, 4529–4534. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Gilad, S.; Meiri, E.; Yogev, Y.; Benjamin, S.; Lebanony, D.; Yerushalmi, N.; Benjamin, H.; Kushnir, M.; Cholakh, H.; Melamed, N.; et al. Serum microRNAs are promising novel biomarkers. PLoS ONE 2008. [Google Scholar] [CrossRef] [PubMed]
- Fallingborg, J. Intraluminal pH of the human gastrointestinal tract. Dan. Med. Bull. 1999, 46, 183–196. [Google Scholar] [PubMed]
- Title, A.C.; Denzler, R.; Stoffel, M. Uptake and function studies of maternal milk-derived microRNAs. J. Biol. Chem. 2015. [Google Scholar] [CrossRef] [PubMed]
- Bar-Sagi, D.; Fernandez, A.; Feramisco, J.R. Regulation of membrane turnover by ras proteins. Biosci. Rep. 1987, 7, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Salunkhe, V.A.; Esguerra, J.L.; Ofori, J.K.; Mollet, I.G.; Braun, M.; Stoffel, M.; Wendt, A.; Eliasson, L. Modulation of microRNA-375 expression alters voltage-gated Na(+) channel properties and exocytosis in insulin-secreting cells. Acta Physiol. (Oxf.) 2015, 213, 882–892. [Google Scholar] [CrossRef]
- Yang, J.; Farmer, L.M.; Agyekum, A.A.; Hirschi, K.D. Detection of dietary plant-based small RNAs in animals. Cell Res. 2015, 25, 517–520. [Google Scholar] [CrossRef] [PubMed]
- Pothof, J.; Verkaik, N.S.; van, I.W.; Wiemer, E.A.; Ta, V.T.; van der Horst, G.T.; Jaspers, N.G.; van Gent, D.C.; Hoeijmakers, J.H.; Persengiev, S.P. MicroRNA-mediated gene silencing modulates the UV-induced DNA-damage response. EMBO J. 2009, 28, 2090–2099. [Google Scholar] [CrossRef] [PubMed]
- Kagias, K.; Podolska, A.; Pocock, R. Reliable reference miRNAs for quantitative gene expression analysis of stress responses in Caenorhabditis elegans. BMC Genomics 2014. [Google Scholar] [CrossRef] [PubMed]
- Kraemer, A.; Chen, I.P.; Henning, S.; Faust, A.; Volkmer, B.; Atkinson, M.J.; Moertl, S.; Greinert, R. UVA and UVB irradiation differentially regulate microRNA expression in human primary keratinocytes. PLoS ONE 2013. [Google Scholar] [CrossRef] [PubMed]
- Syed, D.N.; Khan, M.I.; Shabbir, M.; Mukhtar, H. MicroRNAs in skin response to UV radiation. Curr. Drug Targets 2013, 14, 1128–1134. [Google Scholar] [CrossRef] [PubMed]
- Grignol, V.; Fairchild, E.T.; Zimmerer, J.M.; Lesinski, G.B.; Walker, M.J.; Magro, C.M.; Kacher, J.E.; Karpa, V.I.; Clark, J.; Nuovo, G.; et al. MiR-21 and miR-155 are associated with mitotic activity and lesion depth of borderline melanocytic lesions. Br. J. Cancer 2011, 105, 1023–1029. [Google Scholar] [CrossRef] [PubMed]
- Lindsay, M.A. microRNAs and the immune response. Trends Immunol 2008, 29, 343–351. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.F.; Liston, A. MicroRNA in the immune system, microRNA as an immune system. Immunology 2009, 127, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Pauley, K.M.; Cha, S.; Chan, E.K. MicroRNA in autoimmunity and autoimmune diseases. J. Autoimmun. 2009, 32, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.X.; Munitz, A.; Rothenberg, M.E. MicroRNA-21 is up-regulated in allergic airway inflammation and regulates IL-12p35 expression. J. Immunol. 2009, 182, 4994–5002. [Google Scholar] [CrossRef] [PubMed]
- Oddy, W.H. The long-term effects of breastfeeding on asthma and atopic disease. Adv. Exp. Med. Biol. 2009, 639, 237–251. [Google Scholar] [PubMed]
- Chen, C.Z.; Li, L.; Lodish, H.F.; Bartel, D.P. MicroRNAs modulate hematopoietic lineage differentiation. Science 2004, 303, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Vigorito, E.; Perks, K.L.; Abreu-Goodger, C.; Bunting, S.; Xiang, Z.; Kohlhaas, S.; Das, P.P.; Miska, E.A.; Rodriguez, A.; Bradley, A.; et al. microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity 2007, 27, 847–859. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.J.; Chau, J.; Ebert, P.J.; Sylvester, G.; Min, H.; Liu, G.; Braich, R.; Manoharan, M.; Soutschek, J.; Skare, P.; et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 2007, 129, 147–161. [Google Scholar] [CrossRef] [PubMed]
- De Yebenes, V.G.; Belver, L.; Pisano, D.G.; Gonzalez, S.; Villasante, A.; Croce, C.; He, L.; Ramiro, A.R. miR-181b negatively regulates activation-induced cytidine deaminase in B cells. J. Exp. Med. 2008, 205, 2199–2206. [Google Scholar] [CrossRef] [PubMed]
- Iliopoulos, D.; Jaeger, S.A.; Hirsch, H.A.; Bulyk, M.L.; Struhl, K. STAT3 activation of miR-21 and miR-181b-1 via PTEN and CYLD are part of the epigenetic switch linking inflammation to cancer. Mol. Cell 2010, 39, 493–506. [Google Scholar] [CrossRef] [PubMed]
- Quinn, S.R.; Mangan, N.E.; Caffrey, B.E.; Gantier, M.P.; Williams, B.R.; Hertzog, P.J.; McCoy, C.E.; O’Neill, L.A. The role of Ets2 transcription factor in the induction of microRNA-155 (miR-155) by lipopolysaccharide and its targeting by interleukin-10. J. Biol. Chem. 2014, 289, 4316–4325. [Google Scholar] [CrossRef] [PubMed]
- Tili, E.; Michaille, J.J.; Cimino, A.; Costinean, S.; Dumitru, C.D.; Adair, B.; Fabbri, M.; Alder, H.; Liu, C.G.; Calin, G.A.; et al. Modulation of miR-155 and miR-125b levels following lipopolysaccharide/TNF-alpha stimulation and their possible roles in regulating the response to endotoxin shock. J. Immunol. 2007, 179, 5082–5089. [Google Scholar] [CrossRef] [PubMed]
- Fontana, L.; Pelosi, E.; Greco, P.; Racanicchi, S.; Testa, U.; Liuzzi, F.; Croce, C.M.; Brunetti, E.; Grignani, F.; Peschle, C. MicroRNAs 17-5p-20a-106a control monocytopoiesis through AML1 targeting and M-CSF receptor upregulation. Nat. Cell Biol. 2007, 9, 775–787. [Google Scholar] [CrossRef] [PubMed]
- Admyre, C.; Johansson, S.M.; Qazi, K.R.; Filen, J.J.; Lahesmaa, R.; Norman, M.; Neve, E.P.; Scheynius, A.; Gabrielsson, S. Exosomes with immune modulatory features are present in human breast milk. J. Immunol. 2007, 179, 1969–1978. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Oikawa, K.; Takanashi, M.; Kudo, M.; Ohyashiki, J.; Ohyashiki, K.; Kuroda, M. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS ONE 2009. [Google Scholar] [CrossRef]
- Ohyashiki, K.; Umezu, T.; Yoshizawa, S.; Ito, Y.; Ohyashiki, M.; Kawashima, H.; Tanaka, M.; Kuroda, M.; Ohyashiki, J.H. Clinical impact of down-regulated plasma miR-92a levels in non-Hodgkin’s lymphoma. PLoS ONE 2011. [Google Scholar] [CrossRef] [PubMed]
- Taganov, K.D.; Boldin, M.P.; Chang, K.J.; Baltimore, D. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 2006, 103, 12481–12486. [Google Scholar] [CrossRef] [PubMed]
- Perry, M.M.; Moschos, S.A.; Williams, A.E.; Shepherd, N.J.; Larner-Svensson, H.M.; Lindsay, M.A. Rapid changes in microRNA-146a expression negatively regulate the IL-1beta-induced inflammatory response in human lung alveolar epithelial cells. J. Immunol. 2008, 180, 5689–5698. [Google Scholar] [CrossRef]
- Fukao, T.; Fukuda, Y.; Kiga, K.; Sharif, J.; Hino, K.; Enomoto, Y.; Kawamura, A.; Nakamura, K.; Takeuchi, T.; Tanabe, M. An evolutionarily conserved mechanism for microRNA-223 expression revealed by microRNA gene profiling. Cell 2007, 129, 617–631. [Google Scholar] [CrossRef]
- Johnnidis, J.B.; Harris, M.H.; Wheeler, R.T.; Stehling-Sun, S.; Lam, M.H.; Kirak, O.; Brummelkamp, T.R.; Fleming, M.D.; Camargo, F.D. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature 2008, 451, 1125–1129. [Google Scholar] [CrossRef] [PubMed]
- Fazi, F.; Rosa, A.; Fatica, A.; Gelmetti, V.; de Marchis, M.L.; Nervi, C.; Bozzoni, I. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell 2005, 123, 819–831. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Wang, S.; Mayr, C.; Bartel, D.P.; Lodish, H.F. miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc. Natl. Acad. Sci. USA 2007, 104, 7080–7085. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Calado, D.P.; Galler, G.; Thai, T.H.; Patterson, H.C.; Wang, J.; Rajewsky, N.; Bender, T.P.; Rajewsky, K. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 2007, 131, 146–159. [Google Scholar] [CrossRef] [PubMed]
- Gaziel-Sovran, A.; Segura, M.F.; Di Micco, R.; Collins, M.K.; Hanniford, D.; Vega-Saenz de Miera, E.; Rakus, J.F.; Dankert, J.F.; Shang, S.; Kerbel, R.S.; et al. miR-30b/30d regulation of GalNAc transferases enhances invasion and immunosuppression during metastasis. Cancer Cell 2011, 20, 104–118. [Google Scholar] [CrossRef] [PubMed]
- Stittrich, A.B.; Haftmann, C.; Sgouroudis, E.; Kuhl, A.A.; Hegazy, A.N.; Panse, I.; Riedel, R.; Flossdorf, M.; Dong, J.; Fuhrmann, F.; et al. The microRNA miR-182 is induced by IL-2 and promotes clonal expansion of activated helper T lymphocytes. Nat. Immunol. 2010, 11, 1057–1062. [Google Scholar] [CrossRef] [PubMed]
- Navarro, A.; Gaya, A.; Martinez, A.; Urbano-Ispizua, A.; Pons, A.; Balague, O.; Gel, B.; Abrisqueta, P.; Lopez-Guillermo, A.; Artells, R.; et al. MicroRNA expression profiling in classic Hodgkin lymphoma. Blood 2008, 111, 2825–2832. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Xu, S.; Liu, X.; Zhang, Q.; Xu, X.; Liu, M.; Hua, M.; Li, N.; Yao, H.; Cao, X. The microRNA miR-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-gamma. Nat. Immunol. 2011, 12, 861–869. [Google Scholar] [CrossRef] [PubMed]
- Pekarsky, Y.; Santanam, U.; Cimmino, A.; Palamarchuk, A.; Efanov, A.; Maximov, V.; Volinia, S.; Alder, H.; Liu, C.G.; Rassenti, L.; et al. Tcl1 expression in chronic lymphocytic leukemia is regulated by miR-29 and miR-181. Cancer Res. 2006, 66, 11590–11593. [Google Scholar] [CrossRef] [PubMed]
- Fulci, V.; Chiaretti, S.; Goldoni, M.; Azzalin, G.; Carucci, N.; Tavolaro, S.; Castellano, L.; Magrelli, A.; Citarella, F.; Messina, M.; et al. Quantitative technologies establish a novel microRNA profile of chronic lymphocytic leukemia. Blood 2007, 109, 4944–4951. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef] [PubMed]
- Pauley, K.M.; Satoh, M.; Chan, A.L.; Bubb, M.R.; Reeves, W.H.; Chan, E.K. Upregulated miR-146a expression in peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Res. Ther. 2008. [Google Scholar] [CrossRef] [PubMed]
- Lawrie, C.H.; Soneji, S.; Marafioti, T.; Cooper, C.D.; Palazzo, S.; Paterson, J.C.; Cattan, H.; Enver, T.; Mager, R.; Boultwood, J.; et al. MicroRNA expression distinguishes between germinal center B cell-like and activated B cell-like subtypes of diffuse large B cell lymphoma. Int. J. Cancer 2007, 121, 1156–1161. [Google Scholar] [CrossRef] [PubMed]
- Koralov, S.B.; Muljo, S.A.; Galler, G.R.; Krek, A.; Chakraborty, T.; Kanellopoulou, C.; Jensen, K.; Cobb, B.S.; Merkenschlager, M.; Rajewsky, N.; et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell 2008, 132, 860–874. [Google Scholar] [CrossRef] [PubMed]
- Mendell, J.T. miRiad roles for the miR-17–92 cluster in development and disease. Cell 2008, 133, 217–222. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Beltran, A.; Chetwynd, E.; Stuebe, A.M.; Twigger, A.J.; Metzger, P.; Trengove, N.; Lai, C.T.; Filgueira, L.; Blancafort, P.; et al. Breastmilk is a novel source of stem cells with multilineage differentiation potential. Stem Cells 2012, 30, 2164–2174. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Hepworth, A.R.; Beltran, A.S.; Mathews, M.M.; Stuebe, A.M.; Hartmann, P.E.; Filgueira, L.; Blancafort, P. Expression of the pluripotency transcription Factor OCT4 in the normal and aberrant mammary gland. Front. Oncol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Sonkoly, E.; Stahle, M.; Pivarcsi, A. MicroRNAs and immunity: Novel players in the regulation of normal immune function and inflammation. Semin. Cancer Biol. 2008, 18, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Keller, M.A.; Gendreau-Reid, L.; Heiner, D.C.; Rodriguez, A.; Short, J.A. IgG4 in human colostrum and human milk: Continued local production or selective transport from serum. Acta Paediatr. Scand. 1988, 77, 24–29. [Google Scholar] [CrossRef] [PubMed]
- Peitersen, B.; Bohn, L.; Andersen, H. Quantitative determination of immunoglobulins, lysozyme, and certain electrolytes in breast milk during the entire period of lactation, during a 24-h period, and in milk from the individual mammary gland. Acta Paediatr. Scand. 1975, 64, 709–717. [Google Scholar] [CrossRef] [PubMed]
- Goldman, A.S.; Garza, C.; Nichols, B.L.; Goldblum, R.M. Immunologic factors in human milk during the first year of lactation. J. Pediatr. 1982, 100, 563–567. [Google Scholar] [CrossRef]
- Xiao, C.; Rajewsky, K. MicroRNA control in the immune system: Basic principles. Cell 2009, 136, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Smith, H.; Ueno, Y.; Patel, T. The MicroRNA let-7a modulates interleukin-6-dependent STAT-3 survival signaling in malignant human cholangiocytes. J. Biol. Chem. 2007, 282, 8256–8264. [Google Scholar] [CrossRef]
- Sheedy, F.J.; Palsson-McDermott, E.; Hennessy, E.J.; Martin, C.; O’Leary, J.J.; Ruan, Q.; Johnson, D.S.; Chen, Y.; O’Neill, L.A. Negative regulation of TLR4 via targeting of the proinflammatory tumor suppressor PDCD4 by the microRNA miR-21. Nat. Immunol. 2010, 11, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Manetti, R.; Parronchi, P.; Giudizi, M.G.; Piccinni, M.P.; Maggi, E.; Trinchieri, G.; Romagnani, S. Natural killer cell stimulatory factor (interleukin 12 [IL-12]) induces T helper type 1 (Th1)-specific immune responses and inhibits the development of IL-4-producing Th cells. J. Exp. Med. 1993, 177, 1199–1204. [Google Scholar] [CrossRef] [PubMed]
- Jennewein, C.; von Knethen, A.; Schmid, T.; Brune, B. MicroRNA-27b contributes to lipopolysaccharide-mediated peroxisome proliferator-activated receptor gamma (PPARgamma) mRNA destabilization. J. Biol. Chem. 2010, 285, 11846–11853. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Eleswarapu, S.; Jiang, H. Identification and characterization of microRNAs from the bovine adipose tissue and mammary gland. FEBS Lett. 2007, 581, 981–988. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.; Tan, A.H.; Hu, X.; Athanasopoulos, V.; Simpson, N.; Silva, D.G.; Hutloff, A.; Giles, K.M.; Leedman, P.J.; Lam, K.P.; et al. Roquin represses autoimmunity by limiting inducible T-cell co-stimulator messenger RNA. Nature 2007, 450, 299–303. [Google Scholar] [CrossRef] [PubMed]
- Cobb, B.S.; Hertweck, A.; Smith, J.; O’Connor, E.; Graf, D.; Cook, T.; Smale, S.T.; Sakaguchi, S.; Livesey, F.J.; Fisher, A.G.; Merkenschlager, M. A role for Dicer in immune regulation. J. Exp. Med. 2006, 203, 2519–2527. [Google Scholar] [CrossRef] [PubMed]
- Thai, T.H.; Calado, D.P.; Casola, S.; Ansel, K.M.; Xiao, C.; Xue, Y.; Murphy, A.; Frendewey, D.; Valenzuela, D.; Kutok, J.L.; et al. Regulation of the germinal center response by microRNA-155. Science 2007, 316, 604–608. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, A.R.; Schlauch, K.; Lao, R.; Halayko, A.J.; Gerthoffer, W.T.; Singer, C.A. MicroRNA expression in human airway smooth muscle cells: Role of miR-25 in regulation of airway smooth muscle phenotype. Am. J. Respir. Cell Mol. Biol. 2010, 42, 506–513. [Google Scholar] [CrossRef] [PubMed]
- Lei, B.X.; Liu, Z.H.; Li, Z.J.; Li, C.; Deng, Y.F. miR-21 induces cell proliferation and suppresses the chemosensitivity in glioblastoma cells via downregulation of FOXO1. Int. J. Clin. Exp. Med. 2014, 7, 2060–2066. [Google Scholar] [PubMed]
- Song, W.; Wang, L.; Wang, L.; Li, Q. Interplay of miR-21 and FoxO1 modulates growth of pancreatic ductal adenocarcinoma. Tumour Biol. 2015, 36, 4741–4745. [Google Scholar] [CrossRef]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 2008, 10, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Brabletz, T.; Jung, A.; Hlubek, F.; Lohberg, C.; Meiler, J.; Suchy, U.; Kirchner, T. Negative regulation of CD4 expression in T cells by the transcriptional repressor ZEB. Int. Immunol. 1999, 11, 1701–1708. [Google Scholar] [CrossRef] [PubMed]
- Naeem, A.; Zhong, K.; Moisa, S.J.; Drackley, J.K.; Moyes, K.M.; Loor, J.J. Bioinformatics analysis of microRNA and putative target genes in bovine mammary tissue infected with Streptococcus uberis. J. Dairy Sci. 2012, 95, 6397–6408. [Google Scholar] [CrossRef] [PubMed]
- Vickers, K.C.; Remaley, A.T. Lipid-based carriers of microRNAs and intercellular communication. Curr. Opin. Lipidol. 2012, 23, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Rayner, K.J.; Hennessy, E.J. Extracellular communication via microRNA: Lipid particles have a new message. J. Lipid Res. 2013, 54, 1174–1181. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Hernando, C.; Suarez, Y.; Rayner, K.J.; Moore, K.J. MicroRNAs in lipid metabolism. Curr. Opin. Lipidol. 2011, 22, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Rayner, K.J.; Suarez, Y.; Davalos, A.; Parathath, S.; Fitzgerald, M.L.; Tamehiro, N.; Fisher, E.A.; Moore, K.J.; Fernandez-Hernando, C. MiR-33 contributes to the regulation of cholesterol homeostasis. Science 2010, 328, 1570–1573. [Google Scholar] [CrossRef] [PubMed]
- Najafi-Shoushtari, S.H.; Kristo, F.; Li, Y.; Shioda, T.; Cohen, D.E.; Gerszten, R.E.; Naar, A.M. MicroRNA-33 and the SREBP host genes cooperate to control cholesterol homeostasis. Science 2010, 328, 1566–1569. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, G.; Langmann, T. Structure, function and regulation of the ABC1 gene product. Curr. Opin. Lipidol. 2001, 12, 129–140. [Google Scholar] [CrossRef] [PubMed]
- Yvan-Charvet, L.; Wang, N.; Tall, A.R. Role of HDL, ABCA1, and ABCG1 transporters in cholesterol efflux and immune responses. Arterioscler. Thromb. Vasc. Biol. 2010, 30, 139–143. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Huang, Z.; Wang, L.; Wang, Y.; Wu, F.; Meng, S.; Wang, C. MicroRNA-125a-5p partly regulates the inflammatory response, lipid uptake, and ORP9 expression in oxLDL-stimulated monocyte/macrophages. Cardiovasc. Res. 2009, 83, 131–139. [Google Scholar] [CrossRef] [PubMed]
- Olkkonen, V.M.; Levine, T.P. Oxysterol binding proteins: In more than one place at one time? Biochem. Cell Biol. 2004, 82, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Raychaudhuri, S.; Im, Y.J.; Hurley, J.H.; Prinz, W.A. Nonvesicular sterol movement from plasma membrane to ER requires oxysterol-binding protein-related proteins and phosphoinositides. J. Cell Biol. 2006, 173, 107–119. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Luo, J.; Zhang, L.; Wang, W.; Gou, D. MiR-103 controls milk fat accumulation in goat (Capra hircus) mammary gland during lactation. PLoS ONE 2013. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Xie, H.; Mori, M.A.; Alexander, R.; Yuan, B.; Hattangadi, S.M.; Liu, Q.; Kahn, C.R.; Lodish, H.F. Mir193b-365 is essential for brown fat differentiation. Nat. Cell Biol. 2011, 13, 958–965. [Google Scholar] [CrossRef] [PubMed]
- Thery, C.; Zitvogel, L.; Amigorena, S. Exosomes: Composition, biogenesis and function. Nat. Rev. Immunol. 2002, 2, 569–579. [Google Scholar] [PubMed]
- Subra, C.; Laulagnier, K.; Perret, B.; Record, M. Exosome lipidomics unravels lipid sorting at the level of multivesicular bodies. Biochimie 2007, 89, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Huber, J.; Vales, A.; Mitulovic, G.; Blumer, M.; Schmid, R.; Witztum, J.L.; Binder, B.R.; Leitinger, N. Oxidized membrane vesicles and blebs from apoptotic cells contain biologically active oxidized phospholipids that induce monocyte-endothelial interactions. Arterioscler. Thromb. Vasc. Biol. 2002, 22, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, D.; Chen, X.; Li, J.; Li, L.; Bian, Z.; Sun, F.; Lu, J.; Yin, Y.; Cai, X.; et al. Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol. Cell 2010, 39, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Kopecki, Z.; Luchetti, M.M.; Adams, D.H.; Strudwick, X.; Mantamadiotis, T.; Stoppacciaro, A.; Gabrielli, A.; Ramsay, R.G.; Cowin, A.J. Collagen loss and impaired wound healing is associated with c-Myb deficiency. J. Pathol. 2007, 211, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Ridzon, D.; Wong, L.; Chen, C. Characterization of microRNA expression profiles in normal human tissues. BMC Genomics 2007. [Google Scholar] [CrossRef] [PubMed]
- Wang, V.; Wu, W. MicroRNA-based therapeutics for cancer. BioDrugs 2009, 23, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Bader, A.G.; Brown, D.; Winkler, M. The promise of microRNA replacement therapy. Cancer Res. 2010, 70, 7027–7030. [Google Scholar] [CrossRef] [PubMed]
- Duursma, A.M.; Kedde, M.; Schrier, M.; le Sage, C.; Agami, R. miR-148 targets human DNMT3b protein coding region. RNA 2008, 14, 872–877. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Schug, J.; McKenna, L.B.; Le Lay, J.; Kaestner, K.H.; Greenbaum, L.E. Tissue-specific regulation of mouse microRNA genes in endoderm-derived tissues. Nucleic Acids Res. 2011, 39, 454–463. [Google Scholar] [CrossRef] [PubMed]
- Sempere, L.F.; Freemantle, S.; Pitha-Rowe, I.; Moss, E.; Dmitrovsky, E.; Ambros, V. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 2004. [Google Scholar] [CrossRef] [PubMed]
- Szafranska, A.E.; Davison, T.S.; John, J.; Cannon, T.; Sipos, B.; Maghnouj, A.; Labourier, E.; Hahn, S.A. MicroRNA expression alterations are linked to tumorigenesis and non-neoplastic processes in pancreatic ductal adenocarcinoma. Oncogene 2007, 26, 4442–4452. [Google Scholar] [CrossRef] [PubMed]
- Masaki, S.; Ohtsuka, R.; Abe, Y.; Muta, K.; Umemura, T. Expression patterns of microRNAs 155 and 451 during normal human erythropoiesis. Biochem. Biophys. Res. Commun. 2007, 364, 509–514. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Aurora, A.B.; Johnson, B.A.; Qi, X.; McAnally, J.; Hill, J.A.; Richardson, J.A.; Bassel-Duby, R.; Olson, E.N. The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev. Cell 2008, 15, 261–271. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Fan, G.C. Extracellular/circulating microRNAs and their potential role in cardiovascular disease. Am. J. Cardiovasc. Dis. 2011, 1, 138–149. [Google Scholar] [PubMed]
- Wang, K.; Zhang, S.; Marzolf, B.; Troisch, P.; Brightman, A.; Hu, Z.; Hood, L.E.; Galas, D.J. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc. Natl. Acad. Sci. USA 2009, 106, 4402–4407. [Google Scholar] [CrossRef] [PubMed]
- Neville, M.C.; Anderson, S.M.; McManaman, J.L.; Badger, T.M.; Bunik, M.; Contractor, N.; Crume, T.; Dabelea, D.; Donovan, S.M.; Forman, N.; et al. Lactation and neonatal nutrition: Defining and refining the critical questions. J. Mammary Gland Biol. Neoplasia 2012, 17, 167–188. [Google Scholar] [CrossRef] [PubMed]
- Baumgartner, S.; Martin, D.; Hagios, C.; Chiquet-Ehrismann, R. Tenm, a Drosophila gene related to tenascin, is a new pair-rule gene. EMBO J. 1994, 13, 3728–3740. [Google Scholar] [PubMed]
- Oohashi, T.; Zhou, X.H.; Feng, K.; Richter, B.; Morgelin, M.; Perez, M.T.; Su, W.D.; Chiquet-Ehrismann, R.; Rauch, U.; Fassler, R. Mouse ten-m/Odz is a new family of dimeric type II transmembrane proteins expressed in many tissues. J. Cell Biol. 1999, 145, 563–577. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Michelson, Y.; Freger, V.; Avraham, Z.; Venken, K.J.; Bellen, H.J.; Justice, M.J.; Wides, R. Drosophila Ten-m and filamin affect motor neuron growth cone guidance. PLoS ONE 2011. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Siegel, F.; Kipschull, S.; Haas, B.; Frohlich, H.; Meister, G.; Pfeifer, A. miR-155 regulates differentiation of brown and beige adipocytes via a bistable circuit. Nat. Commun. 2013. [Google Scholar] [CrossRef] [PubMed]
- He, A.; Zhu, L.; Gupta, N.; Chang, Y.; Fang, F. Overexpression of micro ribonucleic acid 29, highly up-regulated in diabetic rats, leads to insulin resistance in 3T3-L1 adipocytes. Mol. Endocrinol. 2007, 21, 2785–2794. [Google Scholar] [CrossRef] [PubMed]
- Dehwah, M.A.; Xu, A.; Huang, Q. MicroRNAs and type 2 diabetes/obesity. J. Genet. Genomics 2012, 39, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Geddes, D.T. Programming of appetite control during breastfeeding as a preventative strategy against the obesity epidemic. J. Hum. Lact. 2014, 30, 136–142. [Google Scholar] [CrossRef] [PubMed]
- Van Kouwenhove, M.; Kedde, M.; Agami, R. MicroRNA regulation by RNA-binding proteins and its implications for cancer. Nat. Rev. Cancer 2011, 11, 644–656. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Gaard, M.; Tretli, S.; Loken, E.B. Dietary fat and the risk of breast cancer: A prospective study of 25,892 Norwegian women. Int. J. Cancer 1995, 63, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.Q.; Xu, J.Y.; Tezuka, H.; Li, J.; Arita, J.; Hoshi, K.; Sato, A. Consumption of commercial whole and non-fat milk increases the incidence of 7,12-dimethylbenz(a)anthracene-induced mammary tumors in rats. Cancer Detect. Prev. 2007, 31, 339–343. [Google Scholar] [CrossRef] [PubMed]
- Allen, N.E.; Key, T.J.; Appleby, P.N.; Travis, R.C.; Roddam, A.W.; Tjonneland, A.; Johnsen, N.F.; Overvad, K.; Linseisen, J.; Rohrmann, S.; et al. Animal foods, protein, calcium and prostate cancer risk: The European Prospective Investigation into Cancer and Nutrition. Br. J. Cancer 2008, 98, 1574–1581. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Chavarro, J.E.; Cao, Y.; Qiu, W.; Mucci, L.; Sesso, H.D.; Stampfer, M.J.; Giovannucci, E.; Pollak, M.; Liu, S.; et al. Whole milk intake is associated with prostate cancer-specific mortality among U.S. male physicians. J. Nutr. 2013, 143, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Torfadottir, J.E.; Steingrimsdottir, L.; Mucci, L.; Aspelund, T.; Kasperzyk, J.L.; Olafsson, O.; Fall, K.; Tryggvadottir, L.; Harris, T.B.; Launer, L.; et al. Milk intake in early life and risk of advanced prostate cancer. Am. J. Epidemiol. 2012, 175, 144–153. [Google Scholar] [CrossRef] [PubMed]
- Duarte-Salles, T.; Fedirko, V.; Stepien, M.; Trichopoulou, A.; Bamia, C.; Lagiou, P.; Lukanova, A.; Trepo, E.; Overvad, K.; Tjonneland, A.; et al. Dairy products and risk of hepatocellular carcinoma: The european prospective investigation into cancer and nutrition. Int. J. Cancer 2014, 135, 1662–1672. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. MiR-21: An environmental driver of malignant melanoma? J. Transl. Med. 2015. [Google Scholar] [CrossRef]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Olivieri, F.; Spazzafumo, L.; Santini, G.; Lazzarini, R.; Albertini, M.C.; Rippo, M.R.; Galeazzi, R.; Abbatecola, A.M.; Marcheselli, F.; Monti, D.; et al. Age-related differences in the expression of circulating microRNAs: miR-21 as a new circulating marker of inflammaging. Mech. Ageing Dev. 2012, 133, 675–685. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; John, S.M.; Schmitz, G. Milk is not just food but most likely a genetic transfection system activating mTORC1 signaling for postnatal growth. Nutr. J. 2013. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Hwang, S.J.; Bae, Y.C.; Jung, J.S. MiR-21 regulates adipogenic differentiation through the modulation of TGF-beta signaling in mesenchymal stem cells derived from human adipose tissue. Stem Cells 2009, 27, 3093–3102. [Google Scholar] [PubMed]
- Hammond, S.M. MicroRNAs as tumor suppressors. Nat. Genet. 2007, 39, 582–583. [Google Scholar] [CrossRef] [PubMed]
- Johnson, S.M.; Grosshans, H.; Shingara, J.; Byrom, M.; Jarvis, R.; Cheng, A.; Labourier, E.; Reinert, K.L.; Brown, D.; Slack, F.J. RAS is regulated by the let-7 microRNA family. Cell 2005, 120, 635–647. [Google Scholar] [CrossRef] [PubMed]
- Miller, E.M.; Aiello, M.O.; Fujita, M.; Hinde, K.; Milligan, L.; Quinn, E.A. Field and laboratory methods in human milk research. Am. J. Hum. Biol. 2013, 25, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Hartmann, P.E. At the dawn of a new discovery: The potential of breast milk stem cells. Adv. Nutr. 2014, 5, 770–778. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Milk—A Nutrient System of Mammalian Evolution Promoting mTORC1-Dependent Translation. Int. J. Mol. Sci. 2015, 16, 17048–17087. [Google Scholar] [CrossRef] [PubMed]
- Greer, F.R.; Sicherer, S.H.; Burks, A.W.; American Academy of Pediatrics. Effects of early nutritional interventions on the development of atopic disease in infants and children: The role of maternal dietary restriction, breastfeeding, timing of introduction of complementary foods, and hydrolyzed formulas. Pediatrics 2008, 121, 183–191. [Google Scholar] [PubMed]
- Fein, S.B.; Falci, C.D. Infant formula preparation, handling, and related practices in the United States. J. Am. Diet. Assoc. 1999, 99, 1234–1240. [Google Scholar] [CrossRef]
- Cregan, M.D.; Fan, Y.; Appelbee, A.; Brown, M.L.; Klopcic, B.; Koppen, J.; Mitoulas, L.R.; Piper, K.M.; Choolani, M.A.; Chong, Y.S.; et al. Identification of nestin-positive putative mammary stem cells in human breastmilk. Cell Tissue Res. 2007, 329, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Verduci, E.; Banderali, G.; Barberi, S.; Radaelli, G.; Lops, A.; Betti, F.; Riva, E.; Giovannini, M. Epigenetic effects of human breast milk. Nutrients 2014, 6, 1711–1724. [Google Scholar] [CrossRef] [PubMed]
- Bode, L.; McGuire, M.; Rodriguez, J.M.; Geddes, D.T.; Hassiotou, F.; Hartmann, P.E.; McGuire, M.K. It’s alive: Microbes and cells in human milk and their potential benefits to mother and infant. Adv. Nutr. 2014, 5, 571–573. [Google Scholar] [CrossRef] [PubMed]
- Food and Drug Administration, Department of Health & Human Services. Current good manufacturing practices, quality control procedures, quality factors, notification requirements, and records and reports, for infant formula. Fed. Regist. 2014, 79, 33057–33072. [Google Scholar] [PubMed]
- Gigli, I.; Maizon, D.O. microRNAs and the mammary gland: A new understanding of gene expression. Genet. Mol. Biol. 2013, 36, 465–474. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; John, S.M.; Schmitz, G. Milk: An exosomal microRNA transmitter promoting thymic regulatory T cell maturation preventing the development of atopy? J. Transl. Med. 2014. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Zhu, X.; He, B.; Zhang, Y.; Kang, B.; Wang, Z.; Ni, X. MiR-204 regulates cardiomyocyte autophagy induced by ischemia-reperfusion through LC3-II. J. Biomed. Sci. 2011, 18, 35. [Google Scholar] [CrossRef] [PubMed]
- Blenkiron, C.; Miska, E.A. miRNAs in cancer: Approaches, aetiology, diagnostics and therapy. Hum. Mol. Genet. 2007. [Google Scholar] [CrossRef] [PubMed]
- Heneghan, H.M.; Miller, N.; Lowery, A.J.; Sweeney, K.J.; Kerin, M.J. MicroRNAs as novel biomarkers for breast cancer. J. Oncol. 2009. [Google Scholar] [CrossRef] [PubMed]
- Heneghan, H.M.; Miller, N.; Lowery, A.J.; Sweeney, K.J.; Newell, J.; Kerin, M.J. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann. Surg. 2010, 251, 499–505. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alsaweed, M.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. MicroRNAs in Breastmilk and the Lactating Breast: Potential Immunoprotectors and Developmental Regulators for the Infant and the Mother. Int. J. Environ. Res. Public Health 2015, 12, 13981-14020. https://doi.org/10.3390/ijerph121113981
Alsaweed M, Hartmann PE, Geddes DT, Kakulas F. MicroRNAs in Breastmilk and the Lactating Breast: Potential Immunoprotectors and Developmental Regulators for the Infant and the Mother. International Journal of Environmental Research and Public Health. 2015; 12(11):13981-14020. https://doi.org/10.3390/ijerph121113981
Chicago/Turabian StyleAlsaweed, Mohammed, Peter E. Hartmann, Donna T. Geddes, and Foteini Kakulas. 2015. "MicroRNAs in Breastmilk and the Lactating Breast: Potential Immunoprotectors and Developmental Regulators for the Infant and the Mother" International Journal of Environmental Research and Public Health 12, no. 11: 13981-14020. https://doi.org/10.3390/ijerph121113981
APA StyleAlsaweed, M., Hartmann, P. E., Geddes, D. T., & Kakulas, F. (2015). MicroRNAs in Breastmilk and the Lactating Breast: Potential Immunoprotectors and Developmental Regulators for the Infant and the Mother. International Journal of Environmental Research and Public Health, 12(11), 13981-14020. https://doi.org/10.3390/ijerph121113981





