Interaction of Intercellular Adhesion Molecule 1 (ICAM1) Polymorphisms and Environmental Tobacco Smoke on Childhood Asthma
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Ascertainment of the Presence of Asthma and Exposure to ETS
2.3. DNA Collection and Genotyping
2.4. Statistical Analyses
3. Results
| Variables | Asthmatics (n = 218) | Nonasthmatics (n = 877) | p-value | |
|---|---|---|---|---|
| n (%) | n (%) | |||
| Demographic characteristics | ||||
| Gender | ||||
| Boys | 137 (62.84) | 427 (48.69) | 0.0002 | |
| Girls | 81 (37.16) | 450 (51.31) | ||
| Grade | ||||
| 1–2 | 85 (39.00) | 281 (32.05) | 0.1367 | |
| 3–4 | 88 (40.36) | 382 (43.55) | ||
| 5–6 | 45 (20.64) | 214 (24.40) | ||
| Parental education level | ||||
| Graduate school and above | 38 (17.43) | 99 (11.29) | 0.0001 | |
| University or college | 126 (57.80) | 420 (47.89) | ||
| Senior high school or less | 52 (23.85) | 353 (40.25) | ||
| Missing values | 2 (0.92) | 5 (0.57) | ||
| Exposure Status | ||||
| Maternal smoking during pregnancy | ||||
| No | 213 (97.71) | 858 (97.83) | 0.1379 | |
| Yes | 1 (0.46) | 13 (1.48) | ||
| Missing values | 4 (1.83) | 6 (0.68) | ||
| Numbers of household smokers | ||||
| None | 131 (60.09) | 426 (48.57) | 0.0058 | |
| One | 60 (27.52) | 332 (37.86) | ||
| Two and above | 25 (11.47) | 117 (13.34) | ||
| Missing values | 2 (0.92) | 2 (0.23) | ||
| Breastfed | ||||
| No | 89 (40.83) | 400 (45.61) | 0.4225 | |
| Yes | 126 (57.80) | 468 (53.36) | ||
| Missing values | 3 (1.38) | 9 (1.03) | ||
| Allergic diseases | ||||
| Lifetime allergic rhinitis | ||||
| No | 43 (19.72) | 457 (52.11) | <0.0001 | |
| Yes | 171 (78.44) | 402 (45.84) | ||
| Missing values | 4 (1.83) | 18 (2.05) | ||
| Lifetime eczema | ||||
| No | 122 (55.96) | 658 (75.03) | <0.0001 | |
| Yes | 86 (39.45) | 184 (20.98) | ||
| Missing values | 10 (4.59) | 35 (3.99) | ||
| Family history of allergic diseases | ||||
| Family history of eczema | ||||
| No | 132 (60.55) | 614 (70.01) | 0.0012 | |
| Yes | 77 (35.32) | 206 (23.49) | ||
| Missing values | 9 (4.13) | 57 (6.50) | ||
| Family history of asthma | ||||
| No | 158 (72.48) | 757 (86.32) | <0.0001 | |
| Yes | 52 (23.85) | 77 (8.78) | ||
| Missing values | 8 (3.67) | 43 (8.78) | ||
| Family history of allergic rhinitis | ||||
| No | 78 (35.78) | 402 (45.84) | 0.0021 | |
| Yes | 133 (61.01) | 422 (48.12) | ||
| Missing values | 7 (3.21) | 53 (6.04) | ||
| ICAM1 polymorphisms * | ||||
| rs5491 (A→T) genotype | ||||
| AA | 194 (88.99) | 794 (90.54) | 0.5724 | |
| AT | 24 (11.01) | 81 (9.24) | ||
| TT | 0 | 2 (0.23) | ||
| rs5498 (A→G) genotype | ||||
| AA | 122 (55.96) | 466 (53.14) | 0.6876 | |
| AG | 83 (38.07) | 348 (39.68) | ||
| GG | 13 (5.96) | 63 (7.18) | ||
| rs5491 | n | aOR | 95% CI |
| AA | 988 | ref | ref |
| AT + TT | 107 | 1.12 | 0.89–1.41 |
| rs5498 | n | aOR | 95% CI |
| AA+AG | 1019 | ref | ref |
| GG | 76 | 1.08 | 0.77–1.52 |
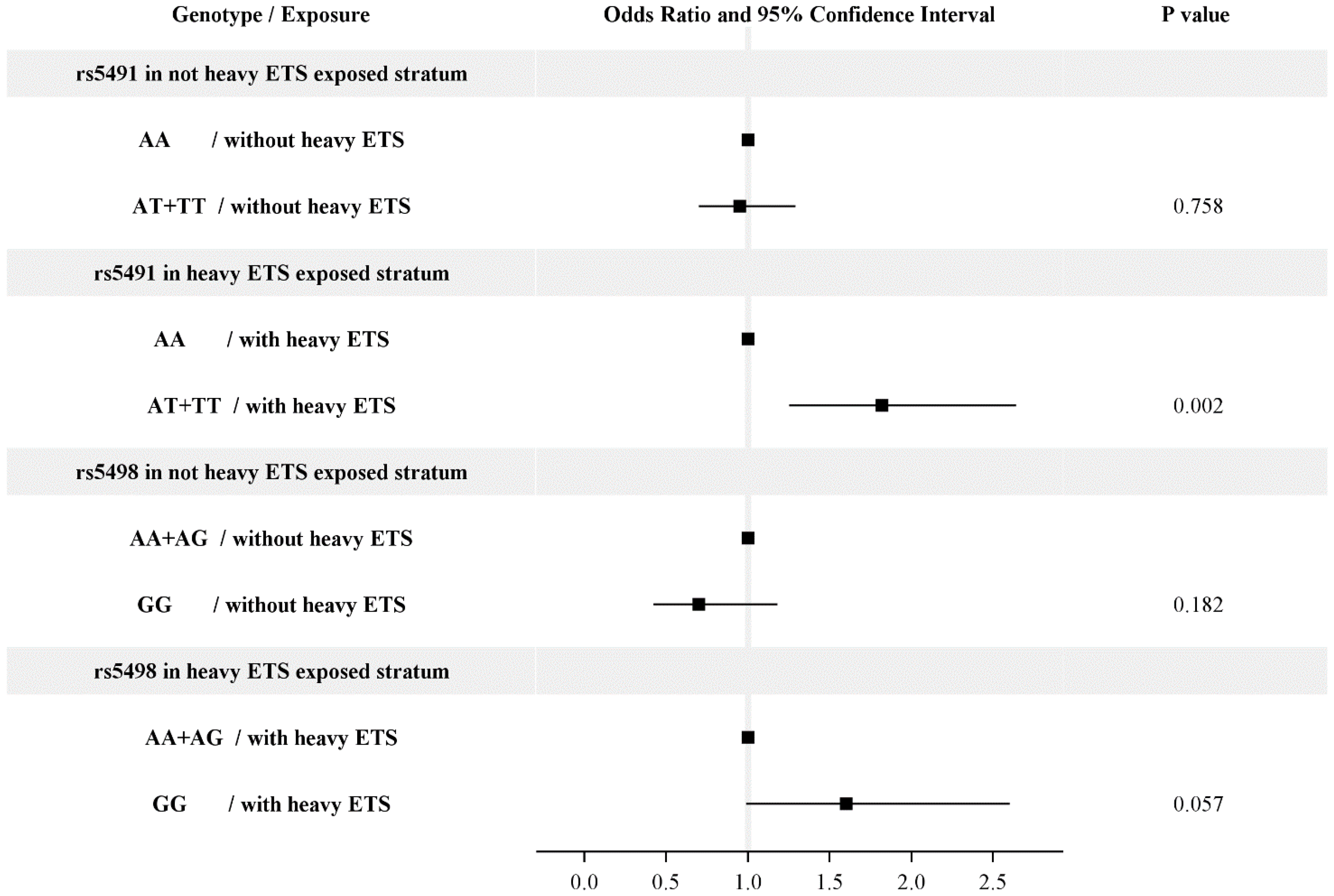
| rs5491 | n | aOR | 95% CI |
| AA | 938 | ref | ref |
| AA | 50 | 0.61 | 0.35–1.07 |
| AT + TT | 81 | 0.97 | 0.74–1.27 |
| AT + TT | 26 | 1.68 | 1.09–2.59 |
| p for the interaction = 0.0063 | |||
| Haplotype * | Frequency | aOR | 95% CI |
| A-G | 0.218 | ref | ref |
| A-A | 0.732 | 0.96 | 0.84–1.11 |
| T-A | 0.002 | 1.38 | 0.56–3.44 |
| T-G | 0.048 | 1.03 | 0.82–1.30 |
4. Discussion and Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Akinbami, L.J.; Moorman, J.E.; Bailey, C.; Zahran, H.S.; King, M.; Johnson, C.A.; Liu, X. Trends in asthma prevalence, health care use, and mortality in the United States, 2001–2010. NCHS Data Brief. 2012, 94, 1–8. [Google Scholar]
- Eder, W.; Ege, M.J.; von Mutius, E. The asthma epidemic. N. Engl. J. Med. 2006, 355, 2226–2235. [Google Scholar] [CrossRef]
- Von Mutius, E. The rising trends in asthma and allergic disease. Clin. Experiment. Allergy 1998, 28, 45–49. [Google Scholar] [CrossRef]
- Lee, Y.L.; Hwang, B.F.; Lin, Y.C.; Guo, Y.L. Time trend of asthma prevalence among school children in Taiwan: ISAAC phase I and III surveys. Pediatr. Allergy Immunol. 2007, 18, 188–195. [Google Scholar] [CrossRef]
- Kabir, Z.; Manning, P.J.; Holohan, J.; Goodman, P.G.; Clancy, L. Prevalence of symptoms of severe asthma and allergies in Irish school children: An ISAAC protocol study, 1995–2007. Int. J. Environ. Res. Public Health 2011, 8, 3192–3201. [Google Scholar] [CrossRef]
- U.S. Centers for Disease Control. CDC Vital Signs. 2011. Available online: http://www.cdc.gov/italsigns/pdf/2011-05-vitalsigns.pdf (accessed on 15 August 2013). [Google Scholar]
- Arruda, L.K.; Sole, D.; Baena-Cagnani, C.E.; Naspitz, C.K. Risk factors for asthma and atopy. Curr. Opin. Allergy Clin. Immuno. 2005, 5, 153–159. [Google Scholar] [CrossRef]
- Gold, D.R. Environmental tobacco smoke, indoor allergens, and childhood asthma. Environ. Health Perspect. 2000, 108, 643–651. [Google Scholar] [CrossRef]
- Li, Y.F.; Langholz, B.; Salam, M.T.; Gilliland, F.D. Maternal and grandmaternal smoking patterns are associated with early childhood asthma. Chest 2005, 127, 1232–1241. [Google Scholar] [CrossRef]
- Li, Y.F.; Gauderman, W.J.; Avol, E.; Dubeau, L.; Gilliland, F.D. Associations of tumor necrosis factor G-308A with childhood asthma and wheezing. Amer. J. Respir. Crit. Care Med. 2006, 173, 970–976. [Google Scholar] [CrossRef]
- Hwang, B.F.; Young, L.H.; Tsai, C.H.; Tung, K.Y.; Wang, P.C.; Su, M.W. Fine particle, ozone exposure, and asthma/wheezing: Effect modification by glutathione S-transferase P1 polymorphisms. PLoS One 2013, 8. [Google Scholar] [CrossRef]
- Tung, K.Y.; Tsai, C.H.; Lee, Y.L. Microsomal epoxide hydroxylase genotypes/diplotypes, traffic air pollution, and childhood asthma. Chest 2011, 139, 839–848. [Google Scholar] [CrossRef]
- Dong, G.H.; Ma, Y.N.; Ding, H.L.; Jin, J.; Cao, Y.; Zhao, Y.D. Housing characteristics, home environmental factors and respiratory health in 3945 pre-school children in China. Int. J. Environ. Health Res. 2008, 18, 267–282. [Google Scholar] [CrossRef]
- Peden, D.B. The epidemiology and genetics of asthma risk associated with air pollution. J. Allergy Clin. Immunol. 2005, 115, 213–219. [Google Scholar] [CrossRef]
- Li, Y.F.; Gauderman, W.J.; Conti, D.V.; Lin, P.C.; Avol, E.; Gilliland, F.D. Glutathione S-transferase P1, maternal smoking, and asthma in children: A haplotype-based analysis. Environ. Health Perspect. 2008, 116, 409–415. [Google Scholar]
- Li, Y.F.; Tseng, P.J.; Lin, C.C.; Hung, C.L.; Lin, S.C.; Su, W.C. NAD(P)H: Quinone oxidoreductase 1, glutathione S-transferase M1, environmental tobacco smoke exposure, and childhood asthma. Mutat. Res. 2009, 678, 53–58. [Google Scholar] [CrossRef]
- Su, M.W.; Tsai, C.H.; Tung, K.Y.; Hwang, B.F.; Liang, P.H.; Chiang, B.L. GSTP1 is a hub gene for gene-air pollution interactions on childhood asthma. Allergy 2013, 68, 1614–1617. [Google Scholar] [CrossRef]
- Greve, J.M.; Davis, G.; Meyer, A.M.; Forte, C.P.; Yost, S.C.; Marlor, C.W. The major human rhinovirus receptor is ICAM-1. Cell 1989, 56, 839–847. [Google Scholar] [CrossRef]
- Majoor, C.J.; van de Pol, M.A.; Kamphuisen, P.W.; Meijers, J.C.; Molenkamp, R.; Wolthers, K.C. Evaluation of coagulation activation after Rhinovirus infection in patients with asthma and healthy control subjects: An observational study. Respir. Res. 2014, 15. [Google Scholar] [CrossRef]
- Xatzipsalti, M.; Papadopoulos, N.G. Cellular and animals models for rhinovirus infection in asthma. Contrib. Microbiol. 2007, 14, 33–41. [Google Scholar] [CrossRef]
- Rawlinson, W.D.; Waliuzzaman, Z.; Carter, I.W.; Belessis, Y.C.; Gilbert, K.M.; Morton, J.R. Asthma exacerbations in children associated with rhinovirus but not human metapneumovirus infection. J. Infect. Dis. 2003, 187, 1314–1318. [Google Scholar] [CrossRef]
- Li, Y.F.; Tsao, Y.H.; Gauderman, W.J.; Conti, D.V.; Avol, E.; Dubeau, L. Intercellular adhesion molecule-1 and childhood asthma. Hum. Genet. 2005, 117, 476–484. [Google Scholar] [CrossRef]
- Puthothu, B.; Krueger, M.; Bernhardt, M.; Heinzmann, A. ICAM1 amino-acid variant K469E is associated with paediatric bronchial asthma and elevated sICAM1 levels. Genes Immun. 2006, 7, 322–326. [Google Scholar] [CrossRef]
- Klaassen, E.M.; van de Kant, K.D.; Jobsis, Q.; Penders, J.; van Schooten, F.J.; Quaak, M. Integrative genomic analysis identifies a role for intercellular adhesion molecule 1 in childhood asthma. Pediatr. Allergy Immunol. 2014, 25, 166–172. [Google Scholar] [CrossRef]
- Gilliland, F.D.; Li, Y.F.; Dubeau, L.; Berhane, K.; Avol, E.; McConnell, R. Effects of glutathione S-transferase M1, maternal smoking during pregnancy, and environmental tobacco smoke on asthma and wheezing in children. Amer. J. Respir. Crit. Care Med. 2002, 166, 457–463. [Google Scholar]
- McKeever, T.M.; Lewis, S.A.; Smith, C.; Collins, J.; Heatlie, H.; Frischer, M. Siblings, multiple births, and the incidence of allergic disease: A birth cohort study using the West Midlands general practice research database. Thorax 2001, 56, 758–762. [Google Scholar] [CrossRef]
- Liao, M.F.; Huang, J.L.; Chiang, L.C.; Wang, F.Y.; Chen, C.Y. Prevalence of asthma, rhinitis, and eczema from ISAAC survey of schoolchildren in central Taiwan. J. Asthma 2005, 42, 833–837. [Google Scholar] [CrossRef]
- Tsuang, H.C.; Su, H.J.; Kao, F.F.; Shih, H.C. Effects of changing risk factors on increasing asthma prevalence in southern Taiwan. Paediatr. Perinat. Epidemiol. 2003, 17, 3–9. [Google Scholar] [CrossRef]
- Gilliland, F.D.; Li, Y.F.; Peters, J.M. Effects of maternal smoking during pregnancy and environmental tobacco smoke on asthma and wheezing in children. Amer. J. Respir. Crit. Care Med. 2001, 163, 429–436. [Google Scholar] [CrossRef]
- Zhao, J.; Shen, K.; Xiang, L.; Zhang, G.; Xie, M.; Bai, J. The knowledge, attitudes and practices of parents of children with asthma in 29 cities of China: A multi-center study. BMC Pediatr. 2013, 13. [Google Scholar] [CrossRef]
- Lee, K.M.; Shen, M.; Chapman, R.S.; Yeager, M.; Welch, R.; He, X. Polymorphisms in immunoregulatory genes, smoky coal exposure and lung cancer risk in Xuan Wei, China. Carcinogenesis 2007, 28, 1437–1441. [Google Scholar] [CrossRef]
- Li, X.X.; Liu, J.P.; Cheng, J.Q.; Han, S.H.; Geng, Y.J.; Wei, S. Intercellular adhesion molecule-1 gene K469E polymorphism and ischemic stroke: A case-control study in a Chinese population. Mol. Biol. Rep. 2009, 36, 1565–1571. [Google Scholar]
- Bielinski, S.J.; Reiner, A.P.; Nickerson, D.; Carlson, C.; Bailey, K.R.; Thyagarajan, B. Polymorphisms in the ICAM1 gene predict circulating soluble intercellular adhesion molecule-1(sICAM-1). Atherosclerosis 2011, 216, 390–394. [Google Scholar]
- Chen, C.F.; Wu, K.G.; Hsu, M.C.; Tang, R.B. Prevalence and relationship between allergic diseases and infectious diseases. J. Microbiol. Immunol. Infect. 2001, 34, 57–62. [Google Scholar]
- Stanciu, L.A.; Djukanovic, R. The role of ICAM-1 on T-cells in the pathogenesis of asthma. Eur. Respir. J. 1998, 11, 949–957. [Google Scholar] [CrossRef]
- Craig, A.; Fernandez-Reyes, D.; Mesri, M.; McDowall, A.; Altieri, D.C.; Hogg, N. A functional analysis of a natural variant of intercellular adhesion molecule-1 (ICAM-1Kilifi). Hum. Mol. Genet. 2000, 9, 525–530. [Google Scholar] [CrossRef]
- Grigg, J.; Riedler, J.; Robertson, C.F. Soluble intercellular adhesion molecule-1 in the bronchoalveolar lavage fluid of normal children exposed to parental cigarette smoke. Eur. Respir. J. 1999, 13, 810–813. [Google Scholar] [CrossRef]
- Sarecka-Hujar, B.; Zak, I.; Krauze, J. Interactions between rs5498 polymorphism in the ICAM1 gene and traditional risk factors influence susceptibility to coronary artery disease. Clin. Exp. Med. 2009, 9, 117–124. [Google Scholar] [CrossRef]
- Cardon, L.R.; Palmer, L.J. Population stratification and spurious allelic association. Lancet 2003, 361, 598–604. [Google Scholar] [CrossRef]
- Yang, H.C.; Lin, C.H.; Hsu, C.L.; Hung, S.I.; Wu, J.Y.; Pan, W.H. A comparison of major histocompatibility complex SNPs in Han Chinese residing in Taiwan and Caucasians. J. Biomed. Sci. 2006, 13, 489–498. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, Y.-F.; Lin, C.-C.; Tai, C.-K. Interaction of Intercellular Adhesion Molecule 1 (ICAM1) Polymorphisms and Environmental Tobacco Smoke on Childhood Asthma. Int. J. Environ. Res. Public Health 2014, 11, 6504-6516. https://doi.org/10.3390/ijerph110606504
Li Y-F, Lin C-C, Tai C-K. Interaction of Intercellular Adhesion Molecule 1 (ICAM1) Polymorphisms and Environmental Tobacco Smoke on Childhood Asthma. International Journal of Environmental Research and Public Health. 2014; 11(6):6504-6516. https://doi.org/10.3390/ijerph110606504
Chicago/Turabian StyleLi, Yu-Fen, Che-Chen Lin, and Chien-Kuo Tai. 2014. "Interaction of Intercellular Adhesion Molecule 1 (ICAM1) Polymorphisms and Environmental Tobacco Smoke on Childhood Asthma" International Journal of Environmental Research and Public Health 11, no. 6: 6504-6516. https://doi.org/10.3390/ijerph110606504
APA StyleLi, Y.-F., Lin, C.-C., & Tai, C.-K. (2014). Interaction of Intercellular Adhesion Molecule 1 (ICAM1) Polymorphisms and Environmental Tobacco Smoke on Childhood Asthma. International Journal of Environmental Research and Public Health, 11(6), 6504-6516. https://doi.org/10.3390/ijerph110606504





