Silaffins in Silica Biomineralization and Biomimetic Silica Precipitation
Abstract
:1. Silicon in Nature
2. Silica Biomineralization in Diatoms
2.1. Diatom Biology and Cell Cycle
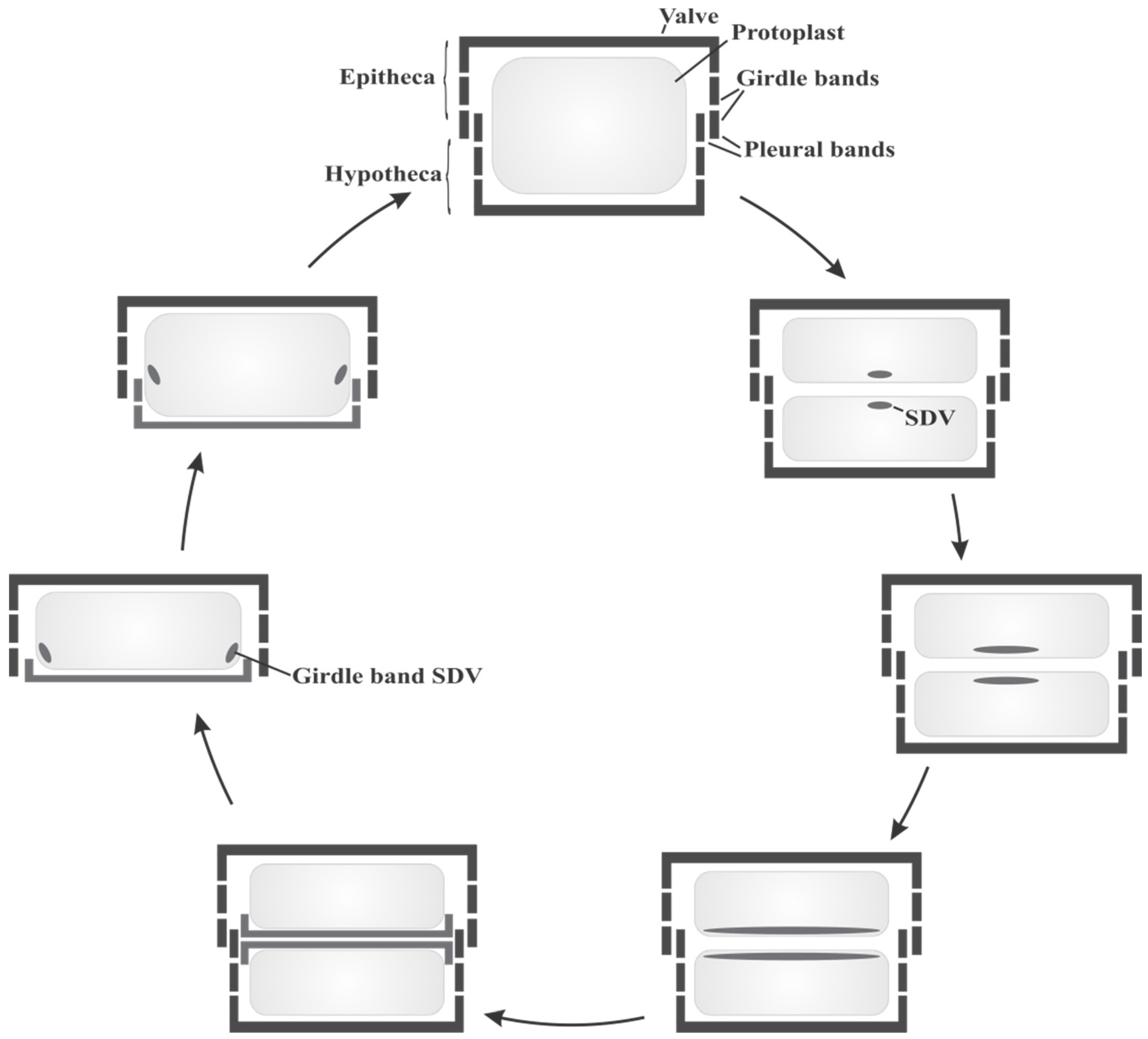
2.2. Organic Constituents of Diatom Cell Walls and Their Role in Silica Formation
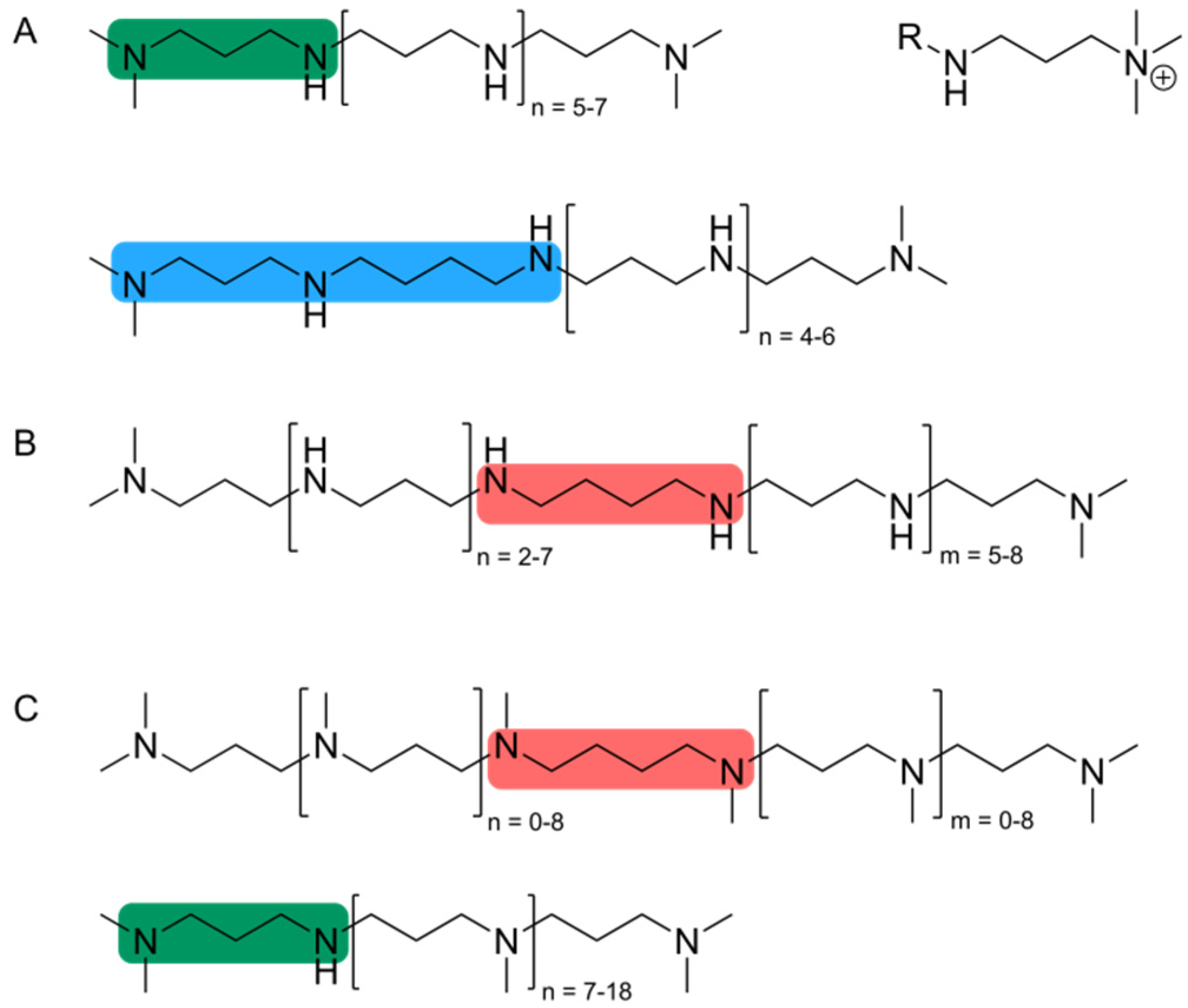
3. Silaffin Proteins and Peptides

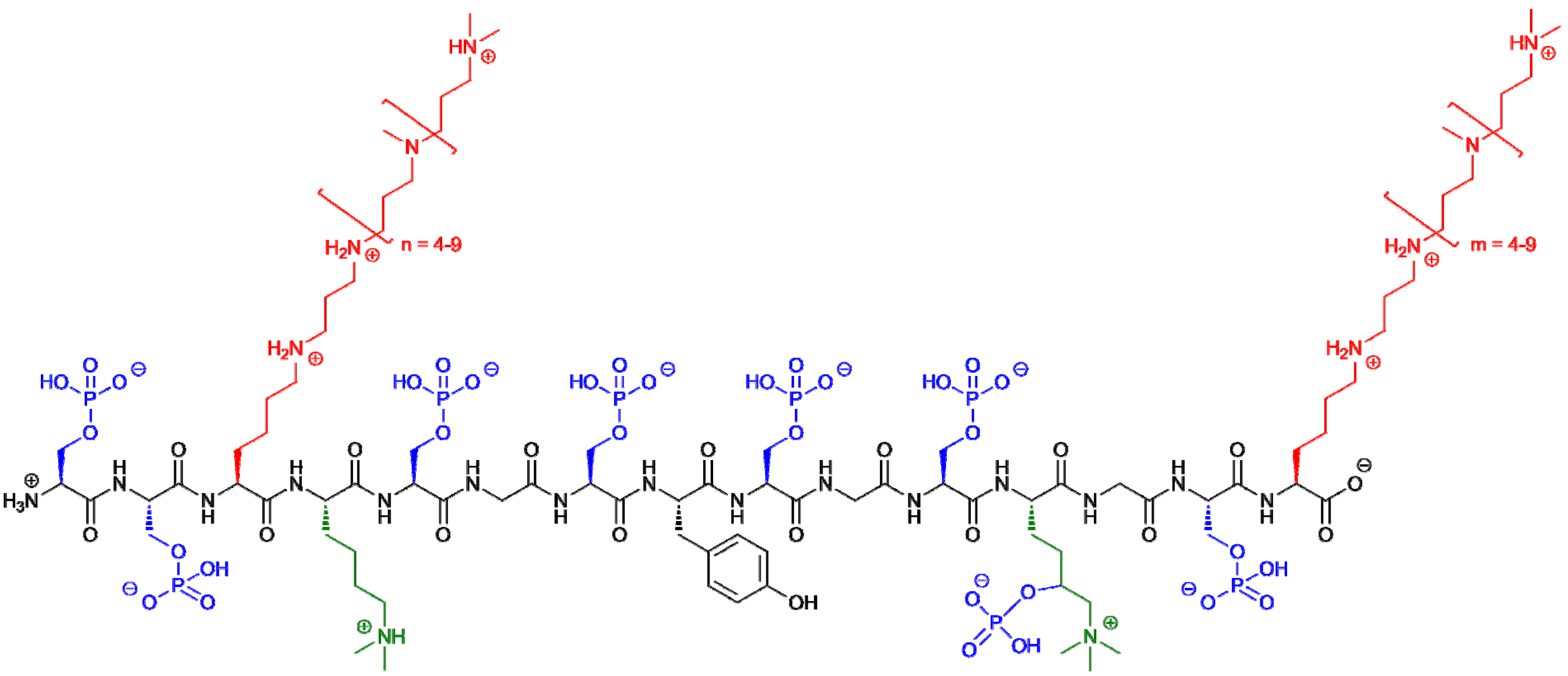
| Diatom Species | Silaffin | Posttranslational Modifications | Silica Precipitation Activity | References | |
|---|---|---|---|---|---|
| at Lysine | at Hydroxyl Amino Acids | ||||
| C. fusiformis | silaffin-1A and silaffin-1B | methylations and polyamine modification at ε-amino group; hydroxylation and phosphorylation at δ-position | phosphorylation | yes | [57,58,59] |
| yes | |||||
| silaffin-2 | sulfation, glycosylation and phosphorylation | no | [61] | ||
| T. pseudonana | tpSil1p | methylations and polyamine modification at ε-amino group; hydroxylation and phosphorylation at δ-position | sulfation, glycosylation and phosphorylation | no | [62,63] |
| tpSil2p | |||||
| tpSil3p | |||||
| tpSil4p | |||||
| E. zodiacus | methylations and polyamine modification at ε-amino group | not analyzed | not analyzed | [64] | |
| C. gracilis | not analyzed | not analyzed | yes | [65] | |
4. Silaffin- and LCPA-Mediated Silica Formation in Diatoms
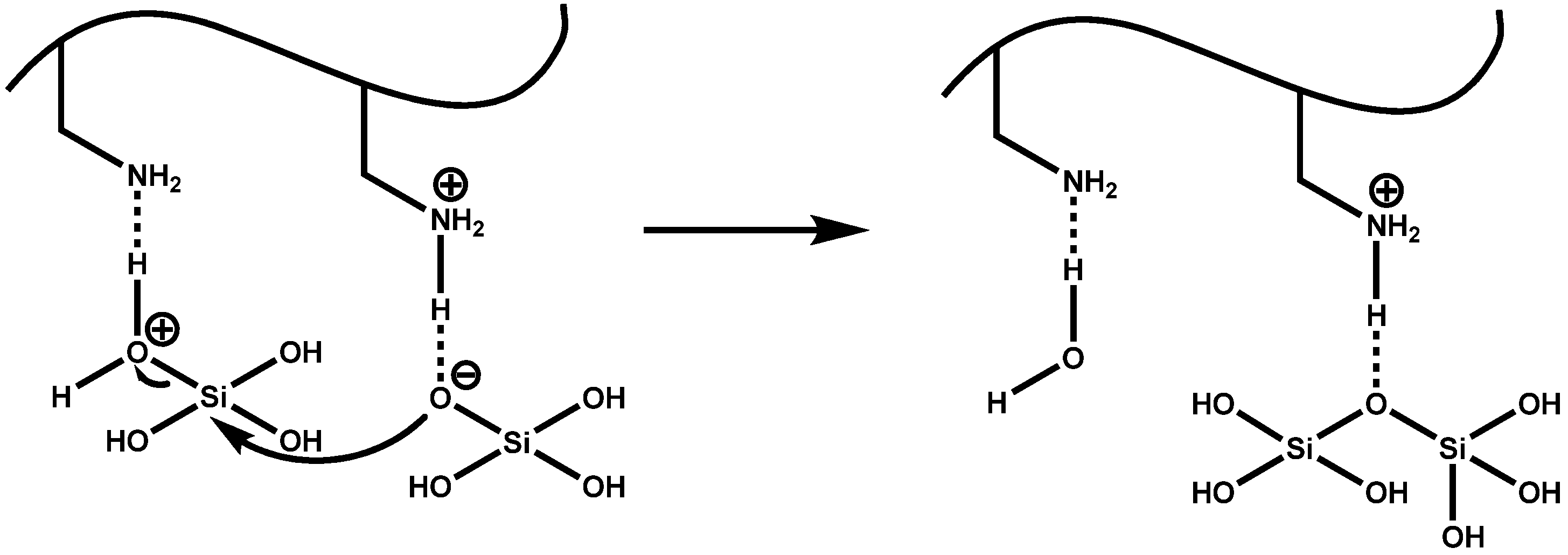
4.1. Chemical and Mechanistic Aspects of Silica Formation
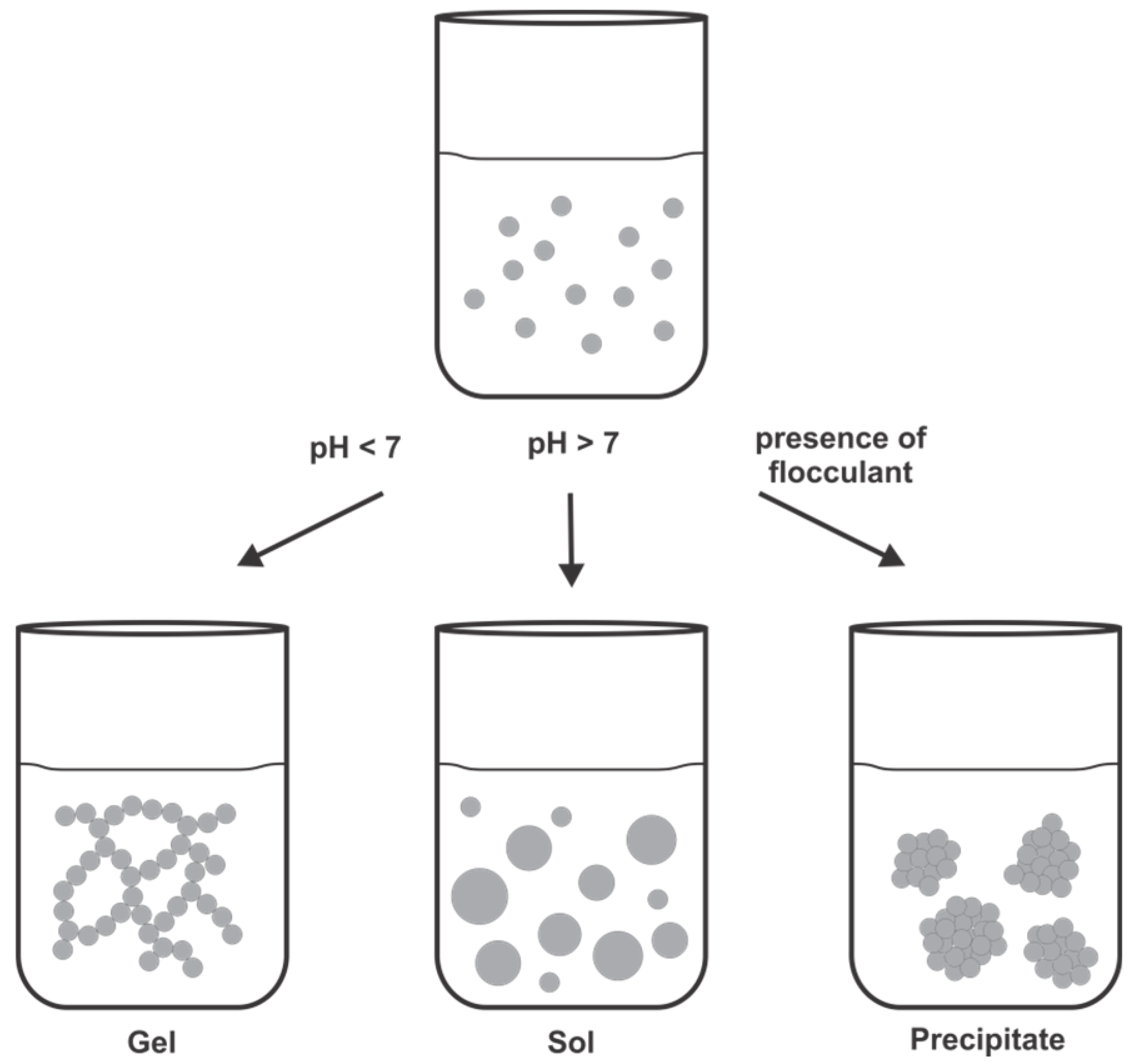
4.2. LCPA-Induced Silica Formation Based on Phase Separation
4.3. Silaffin-Induced Silica Precipitation
5. Biotechnological Applications of Silica
5.1. Synthetic Silica Materials
5.2. Diatomaceous Earth and Biogenic Diatom Silica
5.3. Biomimetic Silica
5.3.1. Formation of Biomimetic Silica with Different Silica-Precipitating Agents
| Silica Morphology | Conditions | References | ||
|---|---|---|---|---|
| silaffin-1A | spherical particles |  | pH 5–5.5 | [57] |
| mixture of native silaffins | cluster of small spheres |  | pH 5–5.5 | [57] |
| R5 peptide | spherical particles |  | phosphate buffered solution, neutral pH, static conditions | [60,81,82,169,170] |
| arch-shaped |  | nitrogen stream bubbling through reaction mixture | [169] | |
| fibrillar |  | mechanical shear force; electrostatic/hydrodynamic force | [169] | |
| sheet-like |  | presence of polyhydroxy compounds (e.g., glycerol) | [170] | |
| poly-l-lysine (PLL) | spherical particles |  | static conditions | [171,172] |
| hexagons |  | phosphate induced self-assembly of long-chain PLL | [173,174] | |
| fibrous |  | electrostatic field, long-chain PLL | [170] | |
| dendrite-like |  | hydrodynamic field | [170] | |
| poly(l-cysteine30-b-l-lysine200) | spheres |  | nitrogen atmosphere | [178] |
| packed columns |  | air-oxidation | ||
| lanreotide | double-walled nanotubes |  | calcination of peptide template after silica formation | [180] |
| A6K, V6K | fibers |  | electrostatic field, flow field | [181] |
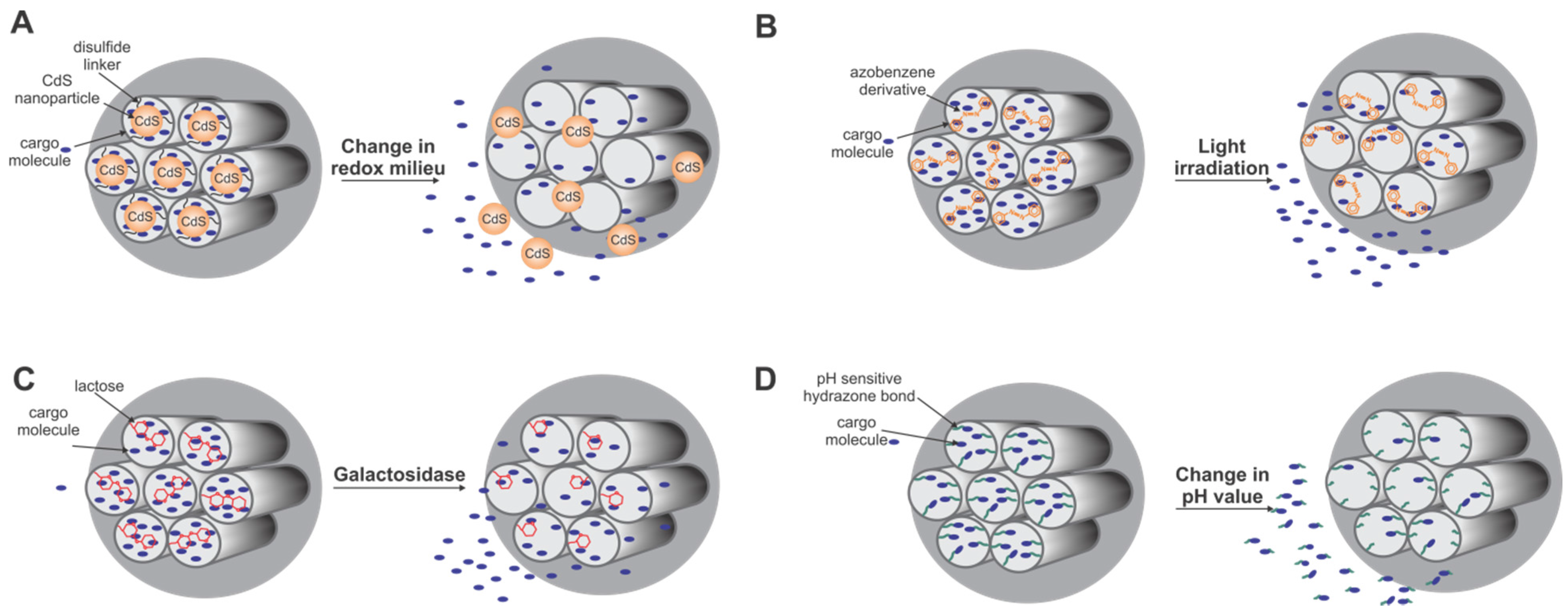
5.3.2. Biotechnological Applications Based on Biomimetic Silica

6. Outlook
Acknowledgments
Conflicts of Interest
References
- Wedepohl, K.H. The composition of the continental crust. Geochim. Cosmochim. Acta 1995, 59, 1217–1232. [Google Scholar] [CrossRef]
- Exley, C. Silicon in life: A bioinorganic solution to bioorganic essentiality. J. Inorg. Biochem. 1998, 69, 139–144. [Google Scholar] [CrossRef]
- Birchall, J.D. The essentiality of silicon in biology. Chem. Soc. Rev. 1995, 24, 351–357. [Google Scholar] [CrossRef]
- Nielsen, F.H. Ultratrace Elements in Nutrition. Ann. Rev. Nutr. 1984, 4, 21–41. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, K.; Milne, D.B. Growth-promoting effects of silicon in rats. Nature 1972, 239, 333–334. [Google Scholar] [CrossRef] [PubMed]
- Carlisle, E.M. Silicon: An essential element for the chick. Science 1972, 178, 619–621. [Google Scholar] [CrossRef] [PubMed]
- Hodson, M.J.; White, P.J.; Mead, A.; Broadley, M.R. Phylogenetic variation in the silicon composition of plants. Ann. Bot. 2005, 96, 1027–1046. [Google Scholar] [CrossRef] [PubMed]
- Mitani, N.; Ma, J.F. Uptake system of silicon in different plant species. J. Exp. Bot. 2005, 56, 1255–1261. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.F.; Yamaji, N. Functions and transport of silicon in plants. Cell. Mol. Life Sci. 2008, 65, 3049–3057. [Google Scholar] [CrossRef] [PubMed]
- Raven, J.A. The transport and function of silicon in plants. Biol. Rev. 1983, 58, 179–207. [Google Scholar] [CrossRef]
- Epstein, E. The anomaly of silicon in plant biology. Proc. Natl. Acad. Sci. USA 1994, 91, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Epstein, E. Silicon. Annu. Rev. Plant. Physiol. Plant Mol. Biol. 1999, 50, 641–664. [Google Scholar] [CrossRef] [PubMed]
- Currie, H.A.; Perry, C.C. Silica in Plants: Biological, Biochemical and Chemical Studies. Ann. Bot. 2007, 100, 1383–1389. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, H.; Demadis, K.D.; Pokrovsky, O.S.; Koutsoukos, P.G. Modern views on desilicification: Biosilica and abiotic silica dissolution in natural and artificial environments. Chem. Rev. 2010, 110, 4656–4689. [Google Scholar] [CrossRef] [PubMed]
- Lewin, J.; Reimann, B.E. Silicon and plant growth. Ann. Rev. Plant. Physiol. 1969, 20, 289–304. [Google Scholar] [CrossRef]
- Norton, T.A.; Melkonian, M.; Andersen, R.A. Algal biodiversity. Phycologia 1996, 35, 353–365. [Google Scholar] [CrossRef]
- Mann, D.G.; Droop, S.J.M. 3. Biodiversity, biogeography and conservation of diatoms. Hydrobiologia 1996, 336, 19–32. [Google Scholar] [CrossRef]
- Round, F.E.; Crawford, R.M.; Mann, D.G. The Diatoms: Biology and Morphology of the Genera; Cambridge University Press: Cambridge, UK, 1990; p. 747. [Google Scholar]
- Kröger, N.; Poulsen, N. Diatoms—From Cell Wall Biogenesis to Nanotechnology. Annu. Rev. Genet. 2008, 42, 83–107. [Google Scholar] [CrossRef] [PubMed]
- Del Amo, Y.; Brzezinski, M.A. The chemical form of dissolved Si taken up by marine diatoms. J. Phycol. 1999, 35, 1162–1170. [Google Scholar]
- Tréguer, P.; Nelson, D.M.; van Bennekom, A.J.; DeMaster, D.J.; Leynaert, A.; Quéquiner, B. The silica balance in the world ocean: A reestimate. Science 1995, 268, 375–379. [Google Scholar] [CrossRef] [PubMed]
- Martin-Jézéquel, V.; Hildebrand, M.; Brzezinski, M.A. Silicon metabolism in diatoms: Implications for growth. J. Phycol. 2000, 36, 821–840. [Google Scholar] [CrossRef]
- Hildebrand, M.; Volcani, B.E.; Gassmann, W.; Schroeder, J.I. A gene family of silicon transporters. Nature 1997, 20, 688–689. [Google Scholar] [CrossRef] [PubMed]
- Thamatrakoln, K.; Alverson, A.J.; Hildebrand, M. Comparative sequence analysis of diatom silicon transporters: Toward a mechanistic model of silicon transport. J. Phycol. 2006, 42, 822–834. [Google Scholar] [CrossRef]
- Marron, A.O.; Alston, M.J.; Heavens, D.; Akam, M.; Caccamo, M.; Holland, P.W.H.; Walker, G. A family of diatom-like silicon transporters in the siliceous loricate choanoflagellates. Proc. R. Soc. 2013, 280. [Google Scholar] [CrossRef] [PubMed]
- Simpson, T.L.; Volcani, B.E. Silicon and Siliceous Structures in Biological Systems; Springer-Verlag: New York, NY, USA, 1981. [Google Scholar]
- Li, C.W.; Chu, S.; Lee, M. Characterizing the silica deposition vesicle of diatoms. Protoplasma 1989, 151, 158–163. [Google Scholar] [CrossRef]
- Vrieling, E.G.; Gieskes, W.W.C.; Beelen, T.P.M. Silicon deposition in diatoms: Control by the pH inside the silicon deposition vesicle. J. Phycol. 1999, 35, 548–559. [Google Scholar] [CrossRef]
- Tesson, B.; Hildebrand, M. Extensive and intimate association of the cytoskeleton with forming silica in diatoms: Control over patterning on the meso-and micro-scale. PLoS ONE 2010, 5. [Google Scholar] [CrossRef] [PubMed]
- Chiappino, M.L.; Volcani, B.E. Studies on the biochemistry and fine structure of silica shell formation in diatoms. Protoplasma 1977, 93, 205–221. [Google Scholar] [CrossRef]
- Kröger, N.; Wetherbee, R. Pleuralins are involved in theca differentiation in the diatom Cylindrotheca fusiformis. Protist 2000, 151, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Volcani, B.E. Biochemistry of Silicon and Related Problems; Bendz, G., Lindquist, I., Eds.; Plenum Publishing: New York, NY, USA, 1978; pp. 177–204. [Google Scholar]
- Nakajima, T.; Volcani, B.E. 3,4-Dihydroxyproline—A new amino acid in diatom cell wall. Science 1969, 164, 1400–1401. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, T.; Volcani, B.E. ε-N-Trimethyl-L-δ-hydroxysine phosphate and its nonphosphorylated compound in diatom cell walls. Biochem. Biophys. Res. Commun. 1970, 39, 28–33. [Google Scholar] [CrossRef]
- Hecky, R.E.; Mopper, K.; Kilham, P.; Degens, T.E. The amino acid and sugar composition of diatom cell-walls. Mar. Biol. 1973, 19, 323–331. [Google Scholar] [CrossRef]
- Kröger, N.; Bergsdorf, C.; Sumper, M. A new calcium-binding glycoprotein family constitutes a major diatom cell wall component. EMBO J. 1994, 13, 4676–4683. [Google Scholar] [PubMed]
- Kröger, N.; Bergsdorf, C.; Sumper, M. Frustulins: Domain conservation in a protein family associated with diatom cell walls. Eur. J. Biochem. 1996, 239, 259–264. [Google Scholar] [CrossRef] [PubMed]
- Kröger, N.; Lehmann, G.; Rachel, R.; Sumper, M. Characterization of a 200-kDa diatom protein that is specifically associated with a silica-based substructure of the cell wall. Eur. J. Biochem. 1997, 250, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Van de Poll, W.H.; Vrieling, E.G.; Gieskes, W.W.C. Location and expression of frustulins in the pennate diatoms Cylindrotheca fusiformis, Navicula pelliculosa, and Navicula salinarum (Bacillariophyceae). J. Phycol. 1999, 35, 1044–1053. [Google Scholar] [CrossRef]
- Bidle, K.D.; Azam, F. Accelerated dissolution of diatom silica by marine bacterial assemblages. Nature 1999, 397, 508–512. [Google Scholar]
- Santos, J.; Almeida, S.F.P.; Figueira, E. Cadmium chelation by frustulins: A novel metal tolerance mechanism in Nitzschia palea (Kützing) W. Smith. Ecotoxicology 2013, 22, 166–173. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.K.; Hildebrand, M.; Palenik, B. A stress-induced protein associated with the girdle band region of the diatom Thalassiosira pseudonana (Bacillariophyta). J. Phycol. 2005, 41, 577–589. [Google Scholar] [CrossRef]
- Davis, A.K.; Hildebrand, M.; Palenik, B. Gene expression induced by copper stress in the diatom Thalassiosira pseudonana. Eukaryot. Cell 2006, 5, 1157–1168. [Google Scholar] [CrossRef] [PubMed]
- Hildebrand, M.; Kim, S.; Shi, D.; Scott, K.; Subramaniam, S. 3D imaging of diatoms with ion-abrasion scanning electron microscopy. J. Struct. Biol. 2009, 166, 316–328. [Google Scholar] [CrossRef] [PubMed]
- Hildebrand, M.; Holton, G.; Joy, D.C.; Doktycz, M.J.; Allison, D.P. Diverse and conserved nano- and mesoscale structures of diatom silica revealed by atomic force microscopy. J. Microsc. 2009, 235, 172–187. [Google Scholar] [CrossRef] [PubMed]
- Brunner, E.; Richthammer, P.; Ehrlich, H.; Paasch, S.; Ueberlein, S.; van Pee, K.H. Chitin-based organic networks: An integral part of cell wall biosilica from the diatom Thalassiosira pseudonana. Angew. Chem. Int. Ed. 2009, 48, 9724–9727. [Google Scholar] [CrossRef] [PubMed]
- Spinde, K.; Kammer, M.; Freyer, K.; Ehrlich, H.; Vournakis, J.N.; Brunner, E. Biomimetic silicification of fibrous chitin from diatoms. Chem. Mater. 2011, 23, 2973–2978. [Google Scholar] [CrossRef]
- Scheffel, A.; Poulsen, N.; Shian, S.; Kröger, N. Nanopatterned protein microrings from a diatom that direct silica morphogenesis. Proc. Natl. Acad. Sci. USA 2011, 108, 3175–3180. [Google Scholar] [CrossRef] [PubMed]
- Kröger, N.; Deutzmann, R.; Bergsdorf, C.; Sumper, M. Species-specific polyamines from diatoms control silica morphology. Proc. Natl. Acad. Sci. USA 2000, 97, 14133–14138. [Google Scholar] [CrossRef] [PubMed]
- Sumper, M.; Brunner, E.; Lehmann, G. Biomineralization in diatoms: Characterization of novel polyamines associated with silica. FEBS Lett. 2005, 579, 3765–3769. [Google Scholar] [CrossRef] [PubMed]
- Sumper, M.; Lehmann, G. Silica pattern formation in diatoms: Species-specific polyamine biosynthesis. ChemBioChem 2006, 9, 1419–1427. [Google Scholar] [CrossRef] [PubMed]
- Sumper, M.; Brunner, E. Learning from diatoms: Nature’s tools for the production of nanostructured silica. Adv. Funct. Mat. 2006, 16, 17–26. [Google Scholar] [CrossRef]
- Frigeri, L.G.; Radabaugh, T.R.; Haynes, P.A.; Hildebrand, M. Identification of proteins from a cell wall fraction of the diatom Thalassiosira pseudonana—Insights into silica structure formation. Mol. Cell. Proteomics 2006, 5, 182–193. [Google Scholar] [CrossRef] [PubMed]
- Sumper, M.; Lorenz, S.; Brunner, E. Biomimetic control of size in the polyamine-directed formation of silica nanospheres. Angew. Chem. Int. Ed. 2003, 42, 5192–5195. [Google Scholar] [CrossRef] [PubMed]
- Wenzl, S.; Hett, R.; Richthammer, P.; Sumper, M. Silacidins: Highly acidic phosphopeptides from diatom shells assist in silica precipitation in vitro. Angew. Chem. Int. Ed. 2008, 47, 1729–1732. [Google Scholar] [CrossRef] [PubMed]
- Richthammer, P.; Börmel, M.; Brunner, E.; van Pée, K.H. Biomineralization in diatoms: The role of silacidins. ChemBioChem 2011, 12, 1362–1366. [Google Scholar] [CrossRef] [PubMed]
- Kröger, N.; Deutzmann, R.; Sumper, M. Polycationic peptides from diatom biosilica that direct silica nanosphere formation. Science 1999, 286, 1129–1132. [Google Scholar] [PubMed]
- Kröger, N.; Deutzmann, R.; Sumper, M. Silica-precipitating peptides from diatoms. The chemical structure of silaffin-A from Cylindrotheca fusiformis. J. Biol. Chem. 2001, 276, 26066–26070. [Google Scholar] [CrossRef] [PubMed]
- Kröger, N.; Lorenz, S.; Brunner, E.; Sumper, M. Self-assembly of highly phosphorylated silaffins and their function in biosilica morphogenesis. Science 2002, 29, 584–586. [Google Scholar] [CrossRef] [PubMed]
- Senior, L.; Crump, M.P.; Williams, C.; Booth, P.J.; Mann, S.; Perriman, A.W.; Curnow, P. Structure and function of the silicifying peptide R5. J. Mater. Chem. B 2015, 3, 2607–2614. [Google Scholar] [CrossRef]
- Poulsen, N.; Sumper, M.; Kröger, N. Biosilica formation in diatoms: Characterization of native silaffin-2 and its role in silica morphogenesis. Proc. Natl. Acad. Sci. USA 2003, 100, 12075–12080. [Google Scholar] [CrossRef] [PubMed]
- Poulsen, N.; Kröger, N. Silica morphogenesis by alternative processing of silaffins in the diatom Thalassiosira pseudonana. J. Biol. Chem. 2004, 279, 42993–42999. [Google Scholar] [CrossRef] [PubMed]
- Sumper, M.; Hett, R.; Lehmann, G.; Wenzl, S. A code for lysine modifications of a silica biomineralizing silaffin protein. Ang. Chem. Int. Ed. 2007, 46, 8405–8408. [Google Scholar] [CrossRef] [PubMed]
- Wenzl, S.; Deutzmann, R.; Hett, R.; Hochmuth, E.; Sumper, M. Quaternary ammonium groups in silica-associated proteins. Angew. Chem. Int. Ed. 2004, 43, 5933–5936. [Google Scholar] [CrossRef] [PubMed]
- Manurung, A.I.; Pratiwi, A.R.; Syah, D.; Suhartono, M.T. Isolation and characterization of silaffin that catalyze biosilica formation from marine diatom Chaetoceros gracilis. HAYATI J. Biosci. 2007, 14, 119–122. [Google Scholar]
- Sumper, M.; Brunner, E. Silica biomineralisation in diatoms: The model organism Thalassiosira pseudonana. ChemBioChem 2008, 9, 1187–1194. [Google Scholar] [CrossRef] [PubMed]
- Poulsen, N.; Scheffel, A.; Sheppard, V.C.; Chesley, P.M.; Kröger, N. Pentalysine clusters mediate silica targeting of silaffins in Thalassiosira pseudonana. J. Biol. Chem. 2013, 288, 20100–20109. [Google Scholar] [CrossRef] [PubMed]
- Sheppard, V.; Poulsen, N.; Kröger, N. Characterization of an endoplasmic reticulum-associated silaffin kinase from the diatom Thalassiosira pseudonana. J. Biol. Chem. 2010, 285, 1166–1176. [Google Scholar] [CrossRef] [PubMed]
- Iler, R.K. The Chemistry of Silica; Wiley: New York, NY, USA, 1979. [Google Scholar]
- Ostwald, W. Studien über die Bildung und Umwandlung fester Körper. Z. Phys. Chem. 1897, 22, 289–330. [Google Scholar]
- Mizutani, T.; Nagase, H.; Fujiwara, N.; Ogoshi, H. Silicic acid polymerization catalyzed by amines and polyamines. Bull. Chem. Soc. Jpn. 1998, 71, 2017–2022. [Google Scholar] [CrossRef]
- Coradin, T.; Durupthy, O.; Livage, J. Interactions of amino-containing peptides with sodium silicate and colloidal silica: A biomimetic approach of silicification. Langmuir 2002, 18, 2331–2336. [Google Scholar] [CrossRef]
- Delak, K.M.; Sahai, N. Amine-catalyzed biomimetic hydrolysis and condensation of organosilicate. Chem. Mater. 2005, 17, 3221–3227. [Google Scholar] [CrossRef]
- Borowitzka, M.A.; Volcani, B.E. The polymorphic diatom Phaeodactylum tricornutum: Ultrastructure of its morphotypes. J. Phycol. 1978, 14, 10–21. [Google Scholar] [CrossRef]
- Schmid, A.M.M.; Schulz, D. Wall morphogenesis in diatoms: Deposition of silica by cytoplasmic vesicles. Protoplasma 1979, 100, 267–288. [Google Scholar] [CrossRef]
- Sumper, M. A phase separation model for the nanopatterning of diatom biosilica. Science 2002, 295, 2430–2433. [Google Scholar] [CrossRef] [PubMed]
- Brunner, E.; Lutz, K.; Sumper, M. Biomimetic synthesis of silica nanospheres depends on the aggregation and phase separation of polyamines in aqueous solution. Phys. Chem. Chem. Phys. 2004, 6, 854–857. [Google Scholar] [CrossRef]
- Lutz, K.; Gröger, C.; Sumper, M.; Brunner, E. Biomimetic silica formation: Analysis of the phosphate-induced self-assembly of polyamines. Phys. Chem. Chem. Phys. 2005, 7, 2812–2815. [Google Scholar] [CrossRef] [PubMed]
- Bernecker, A.; Wieneke, R.; Riedel, R.; Seibt, M.; Geyer, A.; Steinem, C. Tailored synthetic polyamines for controlled biomimetic silica formation. J. Am. Chem. Soc. 2010, 132, 1023–1031. [Google Scholar] [CrossRef] [PubMed]
- Kröger, N.; Sumper, M. The biochemistry of silica formation in diatoms. In Biomineralization; Baeuerlein, E., Ed.; Wiley-VCH: Weinheim, Germany, 2000; p. 168. [Google Scholar]
- Knecht, M.R.; Wright, D.W. Functional analysis of the biomimetic silica precipitating activity of the R5 peptide from Cylindrotheca fusiformis. Chem. Commun. 2003, 3038–3039. [Google Scholar] [CrossRef]
- Lechner, C.C.; Becker, C.F.W. A sequence-function analysis of the silica precipitating silaffin R5 peptide. J. Pept. Sci. 2014, 20, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Lechner, C.C.; Becker, C.F.W. Exploring the effect of native and artificial peptide modifications on silaffin induced silica precipitation. Chem. Sci. 2012, 3, 3500–3504. [Google Scholar] [CrossRef]
- Berthod, A. Silica: Backbone material of liquid chromatographic column packings. J. Chromatogr. 1991, 549, 1–28. [Google Scholar] [CrossRef]
- Chaudhry, Q.; Scotter, M.; Blackburn, J.; Ross, B.; Boxall, A.; Castle, L.; Aitken, R.; Watkins, R. Applications and implications of nanotechnologies for the food sector. Food Addit. Contam. 2008, 25, 241–258. [Google Scholar] [CrossRef] [PubMed]
- Patwardhan, S.V. Biomimetic and bioinspired silica: Recent developments and applications. Chem. Commun. 2011, 47, 7567–7582. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Barnes, J.C.; Bosoy, A.; Stoddart, J.F.; Zink, J.I. Mesoporous silica nanoparticles in biomedical applications. Chem. Soc. Rev. 2012, 41, 2590–2605. [Google Scholar] [CrossRef] [PubMed]
- United States Environmental Protection Agency. R.E.D. FACTS: Silicon dioxide and Silica Gel; EPA/540/FS-92/173; U.S. Environmental Protection Agency, Office of Prevention, Pesticides and Toxic Substances: Washington, DC, USA, 1991.
- Pamirsky, I.E.; Golokhvast, K.S. Silaffins of diatoms: From applied biotechnology to biomedicine. Mar. Drugs 2013, 11, 3155–3167. [Google Scholar] [CrossRef] [PubMed]
- Kresge, C.T.; Leonowicz, M.E.; Roth, W.J.; Vartuli, J.C.; Beck, J.S. Ordered mesoporous molecular sieves synthesized by a liquid-crystal template mechanism. Nature 1992, 359, 710–712. [Google Scholar] [CrossRef]
- Beck, J.S.; Vartuli, J.C.; Roth, W.J.; Leonowicz, M.E.; Kresge, C.T.; Schmitt, K.D.; Chu, C.T.W.; Olson, D.H.; Sheppard, E.W. A new family of mesoporous molecular sieves prepared with liquid crystal templates. J. Am. Chem. Soc. 1992, 114, 10834–10843. [Google Scholar] [CrossRef]
- Wan, Y.; Zhao, D. On the controllable soft-templating approach to mesoporous silicates. Chem. Rev. 2007, 107, 2821–2860. [Google Scholar] [CrossRef] [PubMed]
- Giraldo, L.F.; López, B.L.; Pérez, L.; Urrego, S.; Sierra, L.; Mesa, M. Mesoporous Silica Applications. Macromol. Symp. 2007, 258, 129–141. [Google Scholar] [CrossRef]
- Vallet-Regi, M.; Ramila, A.; del Real, R.P.; Perez-Pariente, J. A new property of MCM-41: Drug delivery system. J. Chem. Mater. 2001, 13, 308–311. [Google Scholar] [CrossRef]
- Radu, D.R.; Lai, C.Y.; Jeftinija, K.; Rowe, E.W.; Jeftinija, S.; Lin, V.S.Y. A polyamidoamine dendrimer-capped mesoporous silica nanosphere-based gene transfection reagent. J. Am. Chem. Soc. 2004, 126, 13216–13217. [Google Scholar] [CrossRef] [PubMed]
- Slowing, I.; Trewyn, B.G.; Lin, V.S.Y. Effect of surface functionalization of MCM-41-type mesoporous silica nanoparticles on the endocytosis by human cancer cells. J. Am. Chem. Soc. 2006, 128, 14792–14793. [Google Scholar] [CrossRef] [PubMed]
- Trewyn, B.G.; Nieweg, J.A.; Zhao, Y.; Lin, V.S.Y. Biocompatible mesoprous silica nanoparticles with different morphologies for animal cell membrane penetration. Chem. Eng. J. 2008, 137, 23–29. [Google Scholar] [CrossRef]
- Mortera, R.; Vivero-Escoto, J.; Slowing, I.I.; Garrone, E.; Onida, B.; Lin, V.S. Cell-induced intracellular controlled release of membrane impermeable cysteine from a mesoporous silica nanoparticle-based drug delivery system. Chem. Commun. 2009, 3219–3221. [Google Scholar] [CrossRef] [PubMed]
- Schlossbauer, A.; Schaffert, D.; Kecht, J.; Wagner, E.; Bein, T. Click chemistry for highdensity biofunctionalization of mesoporous silica. J. Am. Chem. Soc. 2008, 130, 12558–12559. [Google Scholar] [CrossRef] [PubMed]
- Charnay, C.; Bégu, S.; Tourné-Péteilh, C.; Nicole, L.; Lerner, D.A.; Devoisselle, J.M. Inclusion of ibuprofen in mesoporous templated silica: Drug loading and release property. Eur. J. Pharm. Biopharm. 2004, 57, 533–540. [Google Scholar] [CrossRef] [PubMed]
- Nieto, A.; Colilla, M.; Balas, F.; Vallet-Regi, M. Surface electrochemistry of mesoporous silicas as a key factor in the design of tailored delivery devices. Langmuir 2010, 26, 5038–5049. [Google Scholar] [CrossRef] [PubMed]
- Tourne-Peteilh, C.; Lerner, D.A.; Charnay, C.; Nicole, L.; Bégu, S.; Devoisselle, J.M. The potential of ordered mesoporous silica for the storage of drugs: The example of a pentapeptide encapsulated in a MSU-Tween 80. ChemPhysChem 2003, 4, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Slowing, I.I.; Trewyn, B.G.; Lin, V.S.Y. Mesoporous silica nanoparticles for intracellular delivery of membrane-impermeable proteins. J. Am. Chem. Soc. 2007, 129, 8845–8849. [Google Scholar] [CrossRef] [PubMed]
- Lei, C.; Liu, P.; Chen, B.; Mao, Y.; Engelmann, H.; Shin, Y.; Jaffar, J.; Hellstrom, I.; Liu, J.; Hellstrom, K.E. Local Release of Highly Loaded Antibodies from Functionalized Nanoporous Support for Cancer Immunotherapy. J. Am. Chem. Soc. 2010, 132, 6906–6907. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.M.; Lin, H.A.; Zibrowius, B.; Spliethoff, B.; Schueth, F.; Liou, S.C.; Chu, M.W.; Chen, C.H. Selective surface functionalization and metal deposition in the micropores of mesoporous silica SBA-15. Chem. Mater. 2007, 19, 3205–3211. [Google Scholar] [CrossRef]
- Kecht, J.; Schlossbauer, A.; Bein, T. Selective functionalization of the outer and inner surfaces in mesoporous silica nanoparticles. Chem. Mater. 2008, 20, 7207–7214. [Google Scholar] [CrossRef]
- Doadrio, J.C.; Sousa, E.M.B.; Izquierdo-Barba, I.; Doadrio, A.L.; Perez-Pariente, J.; Vallet-Regi, M. Functionalization of mesoporous materials with long alkyl chains as a strategy for controlling drug delivery pattern. J. Mater. Chem. 2006, 16, 462–466. [Google Scholar] [CrossRef]
- Popova, M.D.; Szegedi, A.; Kolev, I.N.; Mihaly, J.; Tzanikov, B.S.; Momekov, G.T.; Lambov, N.G.; Yoncheva, K.P. Carboxilic modified spherical mesoporous silica as drug delivery carriers. Int. J. Pharm. 2012, 436, 778–785. [Google Scholar] [CrossRef] [PubMed]
- Vallet-Regí, M.; Balas, F.; Arcos, D. Mesoporous Materials for Drug Delivery. Angew. Chem. Int. Ed. 2007, 46, 7548–7558. [Google Scholar] [CrossRef] [PubMed]
- Liong, M.; Lu, J.; Kovochich, M.; Xia, T.; Ruehm, S.G.; Nel, A.E.; Tamanoi, F.; Zink, J.I. Multifunctional inorganic nanoparticles for imaging, targeting, and drug delivery. ACS Nano 2008, 2, 889–896. [Google Scholar] [CrossRef] [PubMed]
- Rosenholm, J.M.; Meinander, A.; Peuhu, E.; Niemi, R.; Eriksson, J.E.; Sahlgren, C.; Linden, M. Targeting of porous hybrid silica nanoparticles to cancer cells. ACS Nano 2009, 3, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Brevet, D.; Gary-Bobo, M.; Raehm, L.; Richeter, S.; Hocine, O.; Amro, K.; Loock, B.; Couleaud, P.; Frochot, C.; Morere, A.; et al. Mannose-targeted mesoporous silica nanoparticles for photodynamic therapy. Chem. Commun. 2009, 1475–1477. [Google Scholar] [CrossRef] [PubMed]
- Luo, Z.; Cai, K.; Hu, Y.; Zhao, L.; Liu, P.; Duan, L.; Yang, W. Mesoporous silica nanoparticles end-capped with collagen: Redox-responsive nanoreservoirs for targeted drug delivery. Angew. Chem. Int. Ed. 2010, 50, 640–643. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.H.; Lee, C.H.; Chen, M.C.; Souris, J.S.; Tseng, F.G.; Yang, C.S.; Mou, C.Y.; Chen, C.T.; Lo, L.W. Trifunctionalization of mesoporous silica nanoparticles for comprehensive cancer theranostics-the trio of imaging, targeting and therapy. J. Mater. Chem. 2010, 20, 6149–6157. [Google Scholar] [CrossRef]
- Ferris, D.P.; Lu, J.; Gothard, C.; Yanes, R.; Thomas, C.R.; Olsen, J.C.; Stoddart, J.F.; Tamanoi, F.; Zink, J.I. Synthesis of biomolecule-modified mesoporous silica nanoparticles for targeted hydrophobic drug delivery to cancer cells. Small 2011, 7, 1816–1826. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.P.; Chen, C.Y.; Hung, Y.; Chang, F.H.; Mou, C.Y. Monoclonal antibody-functionalized mesoporous silica nanoparticles (MSN) for selective targeting breast cancer cells. J. Mater. Chem. 2009, 19, 5737–5743. [Google Scholar] [CrossRef]
- Cauda, V.; Engelke, H.; Sauer, A.; Arcizet, D.; Braeuchle, C.; Raedler, J.; Bein, T. Colchicine-loaded lipid bilayer-coated 50 nm mesoporous nanoparticles efficiently induce microtubule depolymerization upon cell uptake. Nano Lett. 2010, 10, 2484–2492. [Google Scholar] [CrossRef] [PubMed]
- Mackowiak, S.A.; Schmidt, A.; Weiss, V.; Argyo, C.; von Schirnding, C.; Bein, T.; Bräuchle, C. Targeted drug delivery in cancer cells with red light photoactivated mesoporous silica nanoparticles. Nano Lett. 2013, 13, 2576–2583. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.Y.; Trewyn, B.G.; Jeftinija, D.M.; Jeftinija, K.; Xu, S.; Jeftinija, S.; Lin, V.S. A mesoporous silica nanosphere-based carrier system with chemically removable CdS nanoparticle caps for stimuli-responsive controlled release of neurotransmitters and drug molecules. J. Am. Chem. Soc. 2003, 125, 4451–4459. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Zhao, X.; Wu, T.; Feng, P. Tunable redox-responsive hybrid nanogated ensembles. J. Am. Chem. Soc. 2008, 130, 14418–14419. [Google Scholar] [CrossRef] [PubMed]
- Bernardos, A.; Aznar, E.; Marcos, M.D.; Martinez-Manez, R.; Sancenon, F.; Soto, J.; Barat, J.M.; Amoros, P. Enzyme-responsive controlled release using mesoporous silica supports capped with lactose. Angew. Chem. Int. Ed. 2009, 48, 5884–5887. [Google Scholar] [CrossRef] [PubMed]
- Coll, C.; Mondragon, L.; Martinez-Manez, R.; Sancenon, F.; Marcos, M.D.; Soto, J.; Amoros, P.; Perez-Paya, E. Enzyme-mediated controlled release systems by anchoring peptide sequences on mesoporous silica supports. Angew. Chem. Int. Ed. 2011, 50, 2138–2140. [Google Scholar] [CrossRef] [PubMed]
- Schlossbauer, A.; Kecht, J.; Bein, T. Biotin-avidin as a protease-responsive cap system for controlled guest release from colloidal mesoporous silica. Angew. Chem. Int. Ed. 2009, 48, 3092–3095. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Geng, J.; Pu, F.; Yang, X.; Ren, J.; Qu, X. Polyvalent nucleic acid/mesoporous silica nanoparticle conjugates: Dual stimuli-responsive vehicles for intracellular drug delivery. Angew. Chem. Int. Ed. 2011, 50, 882–886. [Google Scholar] [CrossRef] [PubMed]
- Angelos, S.; Choi, E.; Vogtle, F.; de Cola, L.; Zink, J.I. Photo-driven expulsion of molecules from mesostructured silica nanoparticles. J. Phys. Chem. C 2007, 111, 6589–6592. [Google Scholar] [CrossRef]
- Ferris, D.P.; Zhao, Y.L.; Khashab, N.M.; Khatib, H.A.; Stoddart, J.F.; Zink, J.I. Light-operated mechanized nanoparticles. J. Am. Chem. Soc. 2009, 131, 1686–1688. [Google Scholar] [CrossRef] [PubMed]
- Schloßbauer, A.; Sauer, A.M.; Cauda, V.; Schmidt, A.; Engelke, H.; Rothbauer, U.; Zolghadr, K.; Leonhardt, H.; Bräuchle, C.; Bein, T. Cascaded photoinduced drug delivery to cells from multifunctional core-shell mesoporous silica. Adv. Healthc. Mater. 2012, 1, 316–320. [Google Scholar] [CrossRef] [PubMed]
- Climent, E.; Bernardos, A.; Martinez-Manez, R.; Maquieira, A.; Marcos, M.D.; Pastor-Navarro, N.; Puchades, R.; Sancenon, F.; Soto, J.; Amoros, P. Controlled delivery systems using antibody-capped mesoporous nanocontainers. J. Am. Chem. Soc. 2009, 131, 14075–14080. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.H.; Cheng, S.H.; Huang, I.P.; Souris, J.S.; Yang, C.S.; Mou, C.Y.; Lo, L.W. Intracellular pH-responsive mesoporous silica nanoparticles for the controlled release of anticancer chemotherapeutics. Angew. Chem. Int. Ed. 2010, 49, 8214–8219. [Google Scholar] [CrossRef] [PubMed]
- Muhammad, F.; Guo, M.; Qi, W.; Sun, F.; Wang, A.; Guo, Y.; Zhu, G. pH-triggered controlled drug release from mesoporous silica nanoparticles via intracelluar dissolution of ZnO nanolids. J. Am. Chem. Soc. 2011, 133, 8778–8781. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Dong, H.; Xu, J.; Feng, Q. Simultaneous immobilization of horseradish peroxidase and glucose oxidase in mesoporous sol-gel host materials. Chem. Phys. Chem. 2002, 3, 802–808. [Google Scholar] [PubMed]
- Dai, Z.; Liu, S.; Ju, H.; Chen, H. Direct electron transfer and enzymatic activity of hemoglobin in a hexagonal mesoporous silica matrix. Biosens. Bioelectron. 2004, 19, 861–867. [Google Scholar] [CrossRef] [PubMed]
- Dai, Z.; Xu, X.; Ju, H. Direct electrochemistry and electrocatalysis of myoglobin immobilized on a hexagonal mesoporous silica matrix. Anal. Biochem. 2004, 332, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Descalzo, A.B.; Marcos, M.D.; Martinez-Manez, R.; Soto, J.; Beltran, D.; Amoros, P. Anthrylmethylamine functionalised mesoporous silica-based materials as hybrid fluorescent chemosensors for ATP. J. Mater. Chem. 2005, 15, 2721–2731. [Google Scholar] [CrossRef]
- Radu, D.R.; Lai, C.Y.; Wiench, J.W.; Pruski, M.; Lin, V.S.Y. Gatekeeping layer effect: A poly(lactic acid)-coated mesoporous silica nanosphere-based fluorescence probe for detection of amino-containing neurotransmitters. J. Am. Chem. Soc. 2004, 126, 1640–1641. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Li, H.; Javadi, A.; Gong, S. Multifunctional mesoporous silica nanoparticles as labels for the preparation of ultrasensitive electrochemical immunosensors. Biomaterials 2010, 31, 3281–3286. [Google Scholar] [CrossRef] [PubMed]
- Hurley, M.T.; Wang, Z.; Mahle, A.; Rabin, D.; Liu, Q.; English, D.S.; Zachariah, M.R.; Stein, D.; DeShong, P. Synthesis, characterization, and application of antibody functionalized fluorescent silica nanoparticles. Adv. Funct. Mater. 2013, 23, 3335–3343. [Google Scholar] [CrossRef]
- Calvert, R. Diatomaceous earth. J. Chem. Educ. 1930, 7, 2829. [Google Scholar] [CrossRef]
- Xiaohua, Q.; Mingzhu, L.; Zhenbin, C.; Rui, L. Preparation and properties of diatomite composite superabsorbent. Polym. Adv. Technol. 2007, 18, 184–193. [Google Scholar]
- Korunic, Z. Diatomaceous Earths, a Group of Natural Insecticides. J. Stored Prod. Res. 1998, 34, 87–97. [Google Scholar] [CrossRef]
- Ivanov, S.E.; Belyakov, A.V. Diatomite and its applications. Glass Ceram. 2008, 65, 48–51. [Google Scholar] [CrossRef]
- Osmanlioglu, A.E. Natural diatomite process for removal of radioactivity from liquid waste. Appl. Radiat. Isot. 2007, 65, 17–20. [Google Scholar] [CrossRef] [PubMed]
- Al-Ghouti, M.A.; Khraisheh, M.A.M.; Allen, S.J.; Ahmad, M.N. The removal of dyes from textile wastewater: A study of the physical characteristics and adsorption mechanisms of diatomaceous earth. J. Environ. Manag. 2003, 69, 229–238. [Google Scholar] [CrossRef]
- Hsien, K.J.; Tsai, W.T.; Su, T.Y. Preparation of diatomite-TiO2 composite for photodegradation of bisphenol-A in water. J. Sol. Gel. Sci. Technol. 2009, 51, 63–69. [Google Scholar] [CrossRef]
- Zhu, Q.W.; Zhang, Y.H.; Zhou, F.S.; Lv, F.Z.; Ye, Z.F.; Fan, F.D.; Chu, P.K. Preparation and characterization of Cu2O-ZnO immobilized on diatomite for photocatalytic treatment of red water produced from manufacturing of TNT. Chem. Eng. J. 2011, 171, 61–68. [Google Scholar] [CrossRef]
- Hamm, C.E.; Merkel, R.; Springer, O.; Jurkojc, P.; Maier, C.; Prechtel, K.; Smetacek, V. Architecture and material properties of diatom shells provide effective mechanical protection. Nature 2003, 421, 841–843. [Google Scholar] [CrossRef] [PubMed]
- Butcher, K.S.A.; Ferris, J.M.; Philips, M.R.; Wintrebertt-Foquet, M.; Wah, J.W.J.; Jovanovic, N.; Vyverman, W.; Chepurnov, V. A luminescence study of porous diatoms. Mater. Sci. Eng. C 2005, 25, 658–663. [Google Scholar] [CrossRef]
- Fuhrmann, T.; Landwehr, S.; El Rharbi-Kucki, M.; Sumper, M. Diatoms as living photonic crystals. Appl. Phys. B 2004, 78, 257–260. [Google Scholar] [CrossRef]
- Payne, E.K.; Rosi, N.L.; Xue, C.; Mirkin, C.A. Sacrificial biological templates for the formation of nanostructured metallic microshells. Angew. Chem. 2005, 117, 5192–5195. [Google Scholar] [CrossRef]
- Sandhage, K.H. Materials “Alchemy”: Shape-preserving chemical transformation of micro-to-macroscopic 3-D structures. JOM 2010, 62, 32–43. [Google Scholar] [CrossRef]
- Bao, Z.H.; Song, M.K.; Davis, S.C.; Cai, Y.; Liu, M.; Sandhage, K.H. High surface area, micro/mesoporous carbon particles with selecTable 3-D biogenic morphologies for tailored catalysis, filtration, or adsorption. Energy Environ. Sci. 2011, 4, 3980–3984. [Google Scholar] [CrossRef]
- Jeffryes, C.; Campbell, J.; Li, H.Y.; Jiao, J.; Rorrer, G. The potential of diatom nanobiotechnology for applications in solar cells, batteries, and electroluminescent devices. Energy Environ. Sci. 2011, 4, 3930–3941. [Google Scholar] [CrossRef]
- De Stefano, L.; Rendina, I.; De Stefano, M.; Bismuto, A.; Maddalena, P. Marine diatoms as optical chemical sensors. Appl. Phys. Lett. 2005, 87, 233902. [Google Scholar] [CrossRef]
- Lettieri, S.; Setaro, A.; De Stefano, L.; De Stefano, M.; Maddalena, P. The gas-detection properties of light-emitting diatoms. Adv. Funct. Mater. 2008, 8, 1257–1264. [Google Scholar] [CrossRef]
- Gale, D.K.; Gutu, T.; Jiao, J.; Chang, C.H.; Rorrer, G.L. Photoluminescence detection of biomolecules by antibody-functionalized diatom biosilica. Adv. Funct. Mater. 2009, 19, 926–933. [Google Scholar] [CrossRef]
- De Stefano, L.; Larnberti, A.; Rotiroti, L.; De Stefano, M. Interfacing the nanostructured biosilica microshells of the marine diatom Coscinodiscus wailesii with biological matter. Acta Biomater. 2008, 4, 126–130. [Google Scholar] [CrossRef] [PubMed]
- Townley, H.E.; Parker, A.R.; White-Cooper, H. Exploitation of diatom frustules for nanotechnology: Tethering active biomolecules. Adv. Funct. Mater. 2008, 18, 369–374. [Google Scholar] [CrossRef]
- Lin, K.C.; Kunduru, V.; Bothara, M.; Rege, K.; Prasad, S.; Ramakrishna, B.L. Biogenic nanoporous silica-based sensor for enhanced electrochemical detection of cardiovascular biomarkers proteins. Biosens. Bioelectron. 2010, 25, 2336–2342. [Google Scholar] [CrossRef] [PubMed]
- Aw, M.S.; Simovic, S.; Addai-Mensah, J.; Losic, D. Silica microcapsules from diatoms as new carrier for delivery of therapeutics. Nanomedicine 2011, 6, 1159–1173. [Google Scholar] [CrossRef] [PubMed]
- Aw, M.S.; Simovic, S.; Yu, Y.; Addai-Mensah, J.; Losic, D. Porous silica microshells from diatoms as biocarrier for drug delivery applications. Powder Technol. 2012, 223, 52–58. [Google Scholar] [CrossRef]
- Aw, M.S.; Bariana, M.; Yu, Y.; Addai-Mensah, J.; Losic, D. Surface functionalized diatom microcapsules for drug delivery of water-insoluble drugs. J. Biomater. Appl. 2013, 28, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Bariana, M.; Aw, M.S.; Kurkuri, M.; Losic, D. Tuning drug loading and release properties of diatom silica microparticles by surface modifications. Int. J. Pharm. 2013, 443, 230–241. [Google Scholar] [CrossRef] [PubMed]
- Poulsen, N.; Berne, C.; Spain, J.; Kröger, N. Silica immobilization of an enzyme through genetic engineering of the diatom Thalassiosira pseudonana. Angew. Chem. Int. Ed. 2007, 46, 1843–1846. [Google Scholar] [CrossRef] [PubMed]
- Sheppard, V.C.; Scheffel, A.; Poulsen, N.; Kröger, N. Live diatom silica immobilization of multimeric and redox-active enzymes. Appl. Environ. Microbiol. 2012, 78, 211–218. [Google Scholar] [CrossRef] [PubMed]
- Annenkov, V.V.; Patwardhan, S.V.; Belton, D.; Danilovtseva, E.N.; Perry, C.C. A new stepwise synthesis of a family of propylamines derived from diatom silaffins and their activity in silicification. Chem. Commun. 2006, 1521–1523. [Google Scholar] [CrossRef] [PubMed]
- Belton, D.J.; Patwardhan, S.V.; Annenkov, V.V.; Danilovtseva, E.N.; Perry, C.C. From biosilicification to tailored materials: Optimizing hydrophobic domains and resistance to protonation of polyamines. Proc. Natl. Acad. Sci. USA 2008, 105, 5963–5968. [Google Scholar] [CrossRef] [PubMed]
- Masse, S.; Laurent, G.; Chuburu, F.; Cadiou, C.; Dechamps, I.; Coradin, T. Modification of the Stöber process by a polyazamacrocycle leading to unusual core-shell silica nanoparticles. Langmuir 2008, 24, 4026–4031. [Google Scholar] [CrossRef] [PubMed]
- Masse, S.; Laurent, G.; Coradin, T. Influence of cyclic polyamines on silica formation during the Stöber process. Phys. Chem. Chem. Phys. 2009, 11, 10204–10210. [Google Scholar] [CrossRef] [PubMed]
- Naik, R.R.; Whitlock, P.W.; Rodriguez, F.; Brott, L.L.; Glawe, D.D.; Clarson, S.J.; Stone, M.O. Controlled formation of biosilica structures in vitro. Chem. Commun. 2003, 238–239. [Google Scholar] [CrossRef]
- Rodríguez, F.; Glawe, D.D.; Naik, R.R.; Hallinan, K.P.; Stone, M.O. Study of the chemical and physical influences upon in vitro peptide-mediated silica formation. Biomacromolecules 2004, 5, 261–265. [Google Scholar] [CrossRef] [PubMed]
- Patwardhan, S.V.; Mukherjee, N.; Clarson, S.J. The use of poly-l-lysine to form novel silica morphologies and the role of polypeptides in biosilicification. J. Inorg. Organomet. Polym. 2001, 11, 193–198. [Google Scholar] [CrossRef]
- Patwardhan, S.V.; Mukherjee, N.; Steinitz-Kannan, M.; Clarson, S.J. Bioinspired synthesis of new silica structures. Chem. Commun. 2003, 1122–1123. [Google Scholar] [CrossRef]
- Tomczak, M.M.; Glawe, D.D.; Drummy, L.F.; Lawrence, C.G.; Stone, M.O.; Perry, C.C.; Pochan, D.J.; Deming, T.J.; Naik, R.R. Polypeptide-templated synthesis of hexagonal silica platelets. J. Am. Chem. Soc. 2005, 127, 12577–12582. [Google Scholar] [CrossRef] [PubMed]
- Patwardhan, S.V.; Maheshwari, R.; Mukherjee, N.; Kiick, K.L.; Clarson, S.J. Conformation and assembly of polypeptide scaffolds in templating the synthesis of silica: An example of a polylysine macromolecular “Switch”. Biomacromolecules 2006, 7, 491–497. [Google Scholar] [CrossRef] [PubMed]
- McKenna, B.J.; Birkedal, H.; Bartl, M.H.; Deming, T.J.; Stucky, G.D. Micrometer-sized spherical assemblies of polypeptides and small molecules by acid-base chemistry. Angew. Chem. Int. Ed. 2004, 43, 5652–5655. [Google Scholar] [CrossRef] [PubMed]
- Pires, J.; Fernandes, A.C.; Avó, R. Bio-inspired synthesis of mesoporous silicas using large molecular weight poly-l-lysine at neutral pH. J. Mater. Sci. 2014, 49, 6087–6092. [Google Scholar] [CrossRef]
- Coradin, T.; Roux, C.; Livage, J. Biomimetic self-activated formation of multi-scale porous silica in the presence of arginine-based surfactants. J. Mater. Chem. 2002, 12, 1242–1244. [Google Scholar] [CrossRef]
- Cha, J.N.; Stucky, G.D.; Morse, D.E.; Deming, T.J. Biomimetic synthesis of ordered silica structures mediated by block copolypeptides. Nature 2000, 403, 289–292. [Google Scholar] [PubMed]
- Valéry, C.; Paternostre, M.; Robert, B.; Gulik-Krzywicki, T.; Narayanan, T.; Dedieu, J.C.; Keller, G.; Torres, M.L.; Cherif-Cheikh, R.; Calvo, P.; et al. Biomimetic organization: Octapeptide self-assembly into nanotubes of viral capsid-like dimension. Proc. Natl. Acad. Sci. USA 2003, 100, 10258–10262. [Google Scholar] [CrossRef] [PubMed]
- Pouget, E.; Dujardin, E.; Cavalier, A.; Moreac, A.; Valéry, C.; Marchi-Artzner, V.; Weiss, T.; Renault, A.; Paternostre, M.; Artzner, F. Hierarchical architectures by synergy between dynamical template self-assembly and biomineralization. Nat. Mater. 2007, 6, 434–439. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, J.Y.X.; Liu, D.; Zheng, J.; Pan, Y.; Lin, Y. Controlled biosilification using self-assembled short peptides A6K and V6K. RSC Adv. 2013, 3, 2784–2793. [Google Scholar] [CrossRef]
- Han, W.; MacEvan, S.R.; Chilkoti, A.; López, G.P. Bio-inspired synthesis of hybrid silica nanoparticles template from elastin-like polypeptide micelles. Nanoscale 2015, 7, 12038–12044. [Google Scholar] [CrossRef] [PubMed]
- Patwardhan, S.V.; Mukherjee, N.; Clarson, S.J. Effect of process parameters on the polymer mediated synthesis of silica at neutral pH. Silicon Chem. 2002, 1, 47–54. [Google Scholar] [CrossRef]
- Patwardhan, S.V.; Mukherjee, N.; Clarson, S.J. Formation of fiber-like amorphous silica structures by externally applied shear. J. Inorg. Organomet. Polym. 2001, 11, 117–121. [Google Scholar] [CrossRef]
- Patwardhan, S.V.; Clarson, S.J. Silicification and biosilicification. Silicon Chem. 2002, 1, 207–214. [Google Scholar] [CrossRef]
- Jin, R.H.; Yuan, J.J. One-pot and rapid synthesis of uniformed silica spheres via mediation of linear poly(ethyleneimine)s and dyes. Polym. J. 2007, 39, 822–827. [Google Scholar] [CrossRef]
- Jin, R.H.; Yuan, J.J. Synthesis of poly(ethyleneimine)s-silica hybrid particles with complex shapes and hierarchical structures. Chem. Commun. 2005, 1399–1401. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.J.; Jin, R.H. Multiply shaped silica mediated by aggregates of linear poly(ethyleneimine). Adv. Mater. 2005, 17, 885–888. [Google Scholar] [CrossRef]
- Jin, R.H.; Yuan, J.J. Simple synthesis of hierarchically structured silicas by poly(ethyleneimine) aggregates preorganized by media modulation. Macromol. Chem. Phys. 2005, 206, 2160–2170. [Google Scholar] [CrossRef]
- Yuan, J.J.; Zhu, P.X.; Fukazawa, N.; Jin, R.H. Synthesis of nanofiber-based silica networks mediated by organized poly(ethylene imine): Structure, properties, and mechanism. Adv. Funct. Mater. 2006, 16, 2205–2212. [Google Scholar] [CrossRef]
- Knecht, M.R.; Wright, D.W. Amine-terminated dendrimers as biomimetic templates for silica nanosphere formation. Langmuir 2004, 20, 4728–4732. [Google Scholar] [CrossRef] [PubMed]
- Knecht, M.R.; Sewell, S.L.; Wright, D.W. Size control of dendrimer-templated silica. Langmuir 2005, 21, 2058–2061. [Google Scholar] [CrossRef] [PubMed]
- Neville, F.; Broderick, M.J.; Gibson, T.; Millner, P.A. Fabrication and activity of silicate nanoparticles and nanosilicate-entrapped enzymes using polyethyleneimine as a biomimetic polymer. Langmuir 2011, 27, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, J.B.; Meyer, R.L.; Poulsen, C.H.; Kragh, K.M.; Besenbacher, F.; Laursen, B.S. Biomimetic silica encapsulation of enzymes for replacement of biocides in antifouling coatings. Green Chem. 2010, 12, 387–394. [Google Scholar] [CrossRef]
- Berne, C.; Betancor, L.; Luckarift, H.R.; Spain, J.C. Application of a microfluidic reactor for screening cancer prodrug activation using silica-immobilized nitrobenzene nitroreductase. Biomacromolecules 2006, 7, 2631–2636. [Google Scholar] [CrossRef] [PubMed]
- Betancor, L.; Berne, C.; Luckarift, H.R.; Spain, J.C. Coimmobilization of a redox enzyme and a cofactor regeneration system. Chem. Commun. 2006, 3640–3642. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Liang, L.; Han, C.; Liu, R. Stabilization of phenylalanine ammonia lyase from Rhodotorula glutinis by encapsulation in polyethyleneimine-mediated biomimetic silica. Appl. Biochem. Biotechnol. 2015, 176, 999–1011. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.A.; Hong, E.D.; Wright, D. Rapid and efficient enzyme encapsulation in a dendrimer silica nanocomposite. Macromol. Biosci. 2006, 6, 839–845. [Google Scholar] [CrossRef] [PubMed]
- Forsyth, C.; Yip, T.W.S.; Patwardhan, S.V. CO2 sequestration by enzyme immobilized onto bioinspired silica. Chem. Commun. 2013, 49, 3191–3193. [Google Scholar] [CrossRef] [PubMed]
- Luckarift, H.R.; Dickerson, M.B.; Sandhage, K.H.; Spain, J.C. Rapid, room-temperature synthesis of antibacterial bionanocomposites of lysozyme with amorphous silica or titania. Small 2006, 2, 640–643. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, M.; Luckarift, H.R.; Sarsenova, A.; Wild, J.R.; Ramanculov, E.K.; Olsen, E.V.; Simonian, A.L. Lysozyme-mediated formation of protein-silica nano-composites for biosensing applications. Colloids Surf. 2009, 73, 58–64. [Google Scholar] [CrossRef] [PubMed]
- Luckarift, H.R.; Balasubramanian, S.; Paliwal, S.; Johnson, G.R.; Simonian, A.L. Enzyme-encapsulated silica monolayers for rapid functionalization of a gold surface. Colloids Surf. 2007, 58, 28–33. [Google Scholar] [CrossRef] [PubMed]
- Ivnitski, D.; Artyushkova, K.; Rincon, R.A.; Atanassov, P.; Luckarift, H.R.; Johnson, G.R. Entrapment of enzymes and carbon nanotubes in biologically synthesized silica: Glucose oxidase-catalyzed direct electron transfer. Small 2008, 4, 357–364. [Google Scholar] [CrossRef] [PubMed]
- Luan, P.P.; Jiang, Y.J.; Zhang, S.P.; Gao, J.; Su, Z.G.; Ma, G.H.; Zhang, Y.F. Chitosan-mediated formation of biomimetic silica nanoparticles: An effective method for manganese peroxidise immobilization and stabilization. J. Biosci. Bioeng. 2014, 118, 575–582. [Google Scholar] [CrossRef] [PubMed]
- Luckarift, H.R.; Spain, J.C.; Naik, R.R.; Stone, M.O. Enzyme immobilization in a biomimetic silica support. Nat. Biotechnol. 2004, 22, 211–213. [Google Scholar] [CrossRef] [PubMed]
- Naik, R.R.; Tomczak, M.M.; Luckarift, H.R.; Spain, J.C.; Stone, M.O. Entrapment of enzymes and nanoparticles using biomimetically synthesized silica. Chem. Commun. 2004, 1684–1685. [Google Scholar] [CrossRef] [PubMed]
- Fragai, M.; Luchinat, C.; Martelli, T.; Ravera, E.; Sagi, I.; Solomonov, I.; Udi, Y. SSNMR of biosilica-entrapped enzymes permits an easy assessment of preservation of native conformation in atomic detail. Chem. Comm. 2014, 50, 421–423. [Google Scholar] [CrossRef] [PubMed]
- Coradin, T.; Coupé, A.; Livage, J. Interactions of bovine serum albumin and lysozyme with sodium silicate solutions. Colloids Surf. B 2003, 29, 189–196. [Google Scholar] [CrossRef]
- Cardoso, M.B.; Luckarift, H.R.; Urban, V.S.; O’Neill, H.; Johnson, G.R. Protein localization in silica nanospheres derived via biomimetic mineralization. Adv. Funct. Mater. 2010, 20, 3031–3038. [Google Scholar] [CrossRef]
- Nam, D.H.; Won, K.; Kim, Y.H.; Sang, B.I. A novel route for immobilization of proteins to silica particles incorporating silaffin domains. Biotechnol. Progr. 2009, 25, 1643–1649. [Google Scholar] [CrossRef] [PubMed]
- Marner, W.D.; Shaikh, A.S.; Muller, S.J.; Keasling, J.D. Enzyme immobilization via silaffin-mediated autoencapsulation in a biosilica support. Biotechnol. Progr. 2009, 25, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Jo, B.H.; Seo, J.H.; Yang, Y.J.; Baek, K.; Choi, Y.S.; Pack, S.P.; Oh, S.H.; Cha, H.J. Bioinspired silica nanocomposite with autoencapsulated carbonic anhydrase as a robust biocatalyst for CO2 sequestration. ACS Catal. 2014, 4, 4332–4340. [Google Scholar] [CrossRef]
- Choi, O.; Kim, B.C.; An, J.H.; Min, K.; Kim, Y.H.; Um, Y.; Oh, M.K.; Sang, B.I. A biosensor based on the self-entrapment of glucose oxidase within biomimetic silica nanoparticles induced by a fusion enzyme. Enzyme Microb. Technol. 2011, 49, 441–445. [Google Scholar] [CrossRef] [PubMed]
- Nam, D.H.; Lee, J.O.; Sang, B.I.; Won, K.; Kim, Y.H. Silaffin peptides as a novel signal enhancer for gravimetric biosensors. Appl. Biochem. Biotechnol. 2013, 170, 25–31. [Google Scholar] [CrossRef] [PubMed]
- Muir, T.W.; Sondhi, D.; Cole, P.A. Expressed protein ligation: A general method for protein engineering. Proc. Natl. Acad. Sci. USA 1998, 95, 6705–6710. [Google Scholar] [CrossRef] [PubMed]
- Lechner, C.C.; Becker, C.F.W. Immobilising proteins on silica with site-specifically attached modified silaffin peptides. Biomater. Sci. 2015, 3, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Lechner, C.C.; Becker, C.F.W. Modified silaffin R5 peptides enable encapsulation and release of cargo molecules from biomimetic silica particles. Bioorg. Med. Chem. 2013, 21, 3533–3541. [Google Scholar] [CrossRef] [PubMed]
- Luckarift, H.R.; Johnson, G.R.; Spain, J.C. Silica-immobilized enzyme reactors: Application to cholinesterase-inhibition studies. J. Chromatogr. B 2006, 843, 310–316. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.H.; Zhang, Y.X.; Li, G.H. Recent Research Progress on Magnetic Nanocomposites with Silica Shell Structures Preparation and Nanotheranostic Applications. Recent Pat. Nanotechnol. 2014, 8, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Brott, L.L.; Naik, R.R.; Pikas, D.J.; Kirkpatrick, S.M.; Tomlin, D.W.; Whitlock, P.W.; Clarson, S.J.; Stone, M.O. Ultrafast holographic nanopatterning of biocatalytically formed silica. Nature 2001, 413, 291–293. [Google Scholar] [CrossRef] [PubMed]
- Coffman, E.A.; Melechko, A.V.; Allison, D.P.; Simpson, M.L.; Doktycz, M.J. Surface patterning of silica nanostructures using bio-inspired templates and directed synthesis. Langmuir 2004, 20, 8431–8436. [Google Scholar] [CrossRef] [PubMed]
- Betancor, L.; Luckarift, H.R.; Seo, J.H.; Brand, O.; Spain, J.C. Three dimensional immobilization of β-galactosidase on a silicon surface. Biotechnol. Bioeng. 2008, 99, 261–267. [Google Scholar] [CrossRef] [PubMed]
- Vamvakaki, V.; Hatzimarinaki, M.; Chaniotakis, N. Biomimetically synthesized silica-carbon nanofiber architectures for the development of highly stable electrochemical biosensor systems. Anal. Chem. 2008, 80, 5970–5975. [Google Scholar] [CrossRef] [PubMed]
- Hatzimarinaki, M.; Vamvakaki, V.; Chaniotakis, N. Spectro-electrochemical studies of acetylcholinesterase in carbon nanofiber-bioinspired silica nanocomposites for biosensor development. J. Mater. Chem. 2009, 19, 428–433. [Google Scholar] [CrossRef]
- Kim, I.Y.; Joachim, E.; Choi, H.; Kim, K. Toxicity of silica nanoparticles depends on size, dose, and cell type. Nanomed. Nanotechnol. Biol. Med. 2015, 11, 1407–1416. [Google Scholar] [CrossRef] [PubMed]
- Gehrke, H.; Frühmesser, A.; Pelka, J.; Esselen, M.; Hecht, L.L.; Blank, H.; Schuchmann, H.P.; Gerthsen, D.; Marquardt, C.; Diabaté, S.; et al. In vitro toxicity of amorphous silica nanoparticles in human colon carcinoma cells. Nanotoxicology 2013, 7, 274–293. [Google Scholar] [CrossRef] [PubMed]
- Tenzer, S.; Docter, D.; Kuharev, J.; Musyanovych, A.; Fetz, V.; Hecht, R.; Schlenk, F.; Fischer, D.; Kiouptsi, K.; Reinhardt, C.; et al. Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nat. Nanotechnol. 2013, 8, 772–781. [Google Scholar] [CrossRef] [PubMed]
- Shemetov, A.A.; Nabiev, I.; Sukhanova, A. Molecular interaction of proteins and peptides with nanoparticles. ACS Nano 2012, 6, 4585–4602. [Google Scholar] [CrossRef] [PubMed]
- Cedervall, T.; Lynch, I.; Lindman, S.; Berggård, T.; Thulin, E.; Nilsson, H.; Dawson, K.A.; Linse, S. Probing the interactions of proteins and nanoparticles. Proc. Natl. Acad. Sci. USA 2007, 104, 2050–2055. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lechner, C.C.; Becker, C.F.W. Silaffins in Silica Biomineralization and Biomimetic Silica Precipitation. Mar. Drugs 2015, 13, 5297-5333. https://doi.org/10.3390/md13085297
Lechner CC, Becker CFW. Silaffins in Silica Biomineralization and Biomimetic Silica Precipitation. Marine Drugs. 2015; 13(8):5297-5333. https://doi.org/10.3390/md13085297
Chicago/Turabian StyleLechner, Carolin C., and Christian F. W. Becker. 2015. "Silaffins in Silica Biomineralization and Biomimetic Silica Precipitation" Marine Drugs 13, no. 8: 5297-5333. https://doi.org/10.3390/md13085297
APA StyleLechner, C. C., & Becker, C. F. W. (2015). Silaffins in Silica Biomineralization and Biomimetic Silica Precipitation. Marine Drugs, 13(8), 5297-5333. https://doi.org/10.3390/md13085297





