Investigation of Marine-Derived Fungal Diversity and Their Exploitable Biological Activities
Abstract
:1. Introduction
2. Results and Discussion
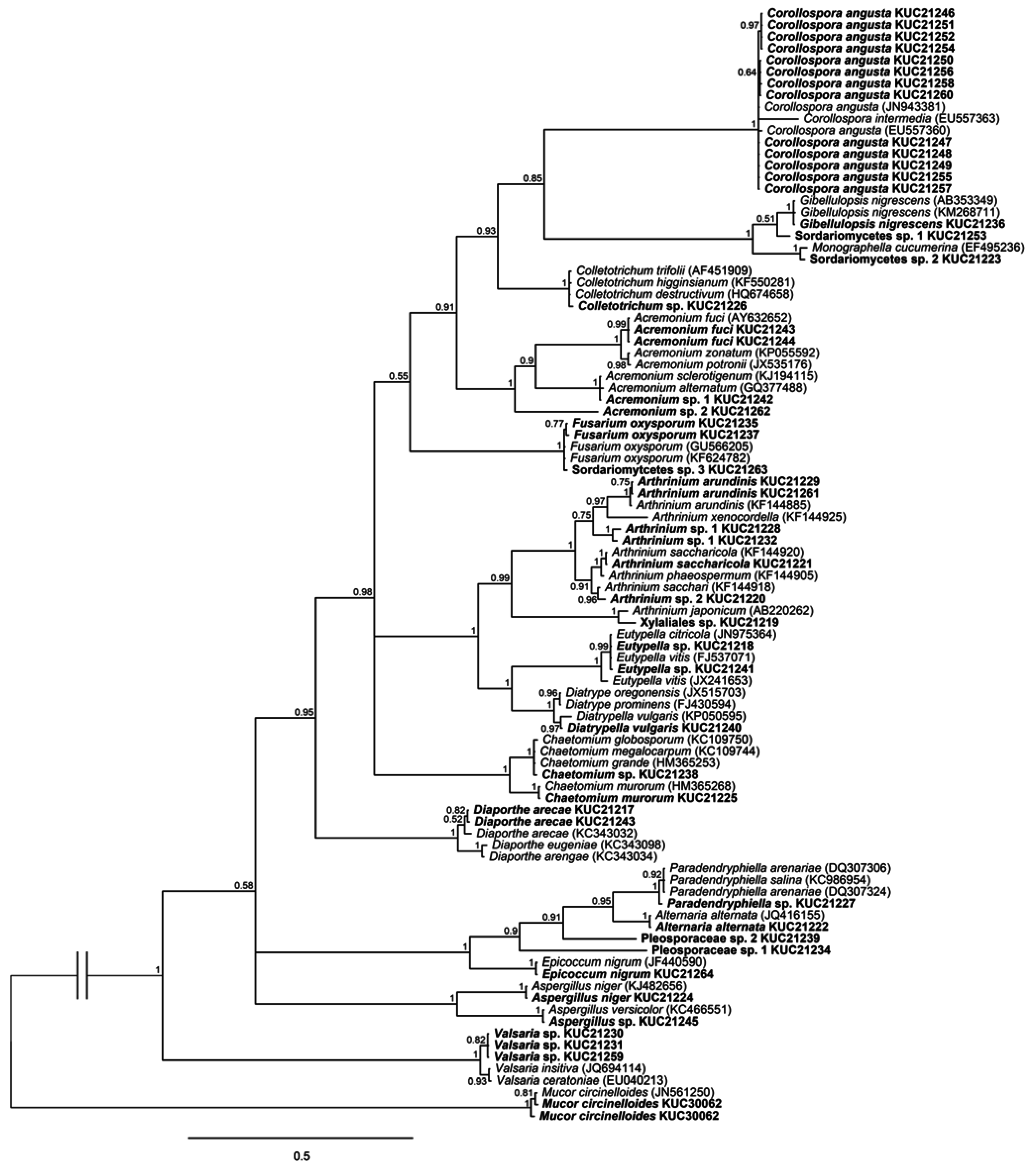
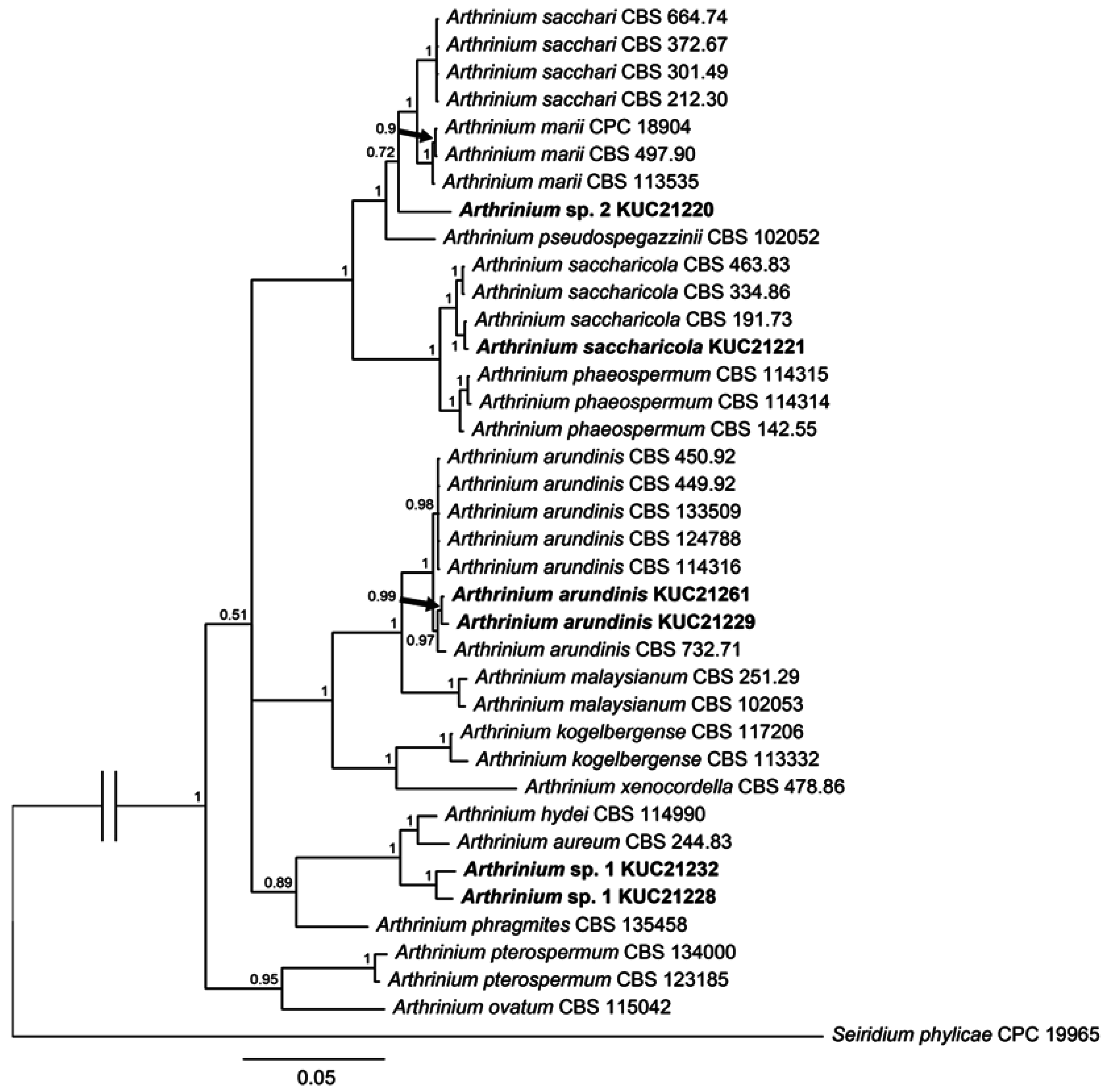
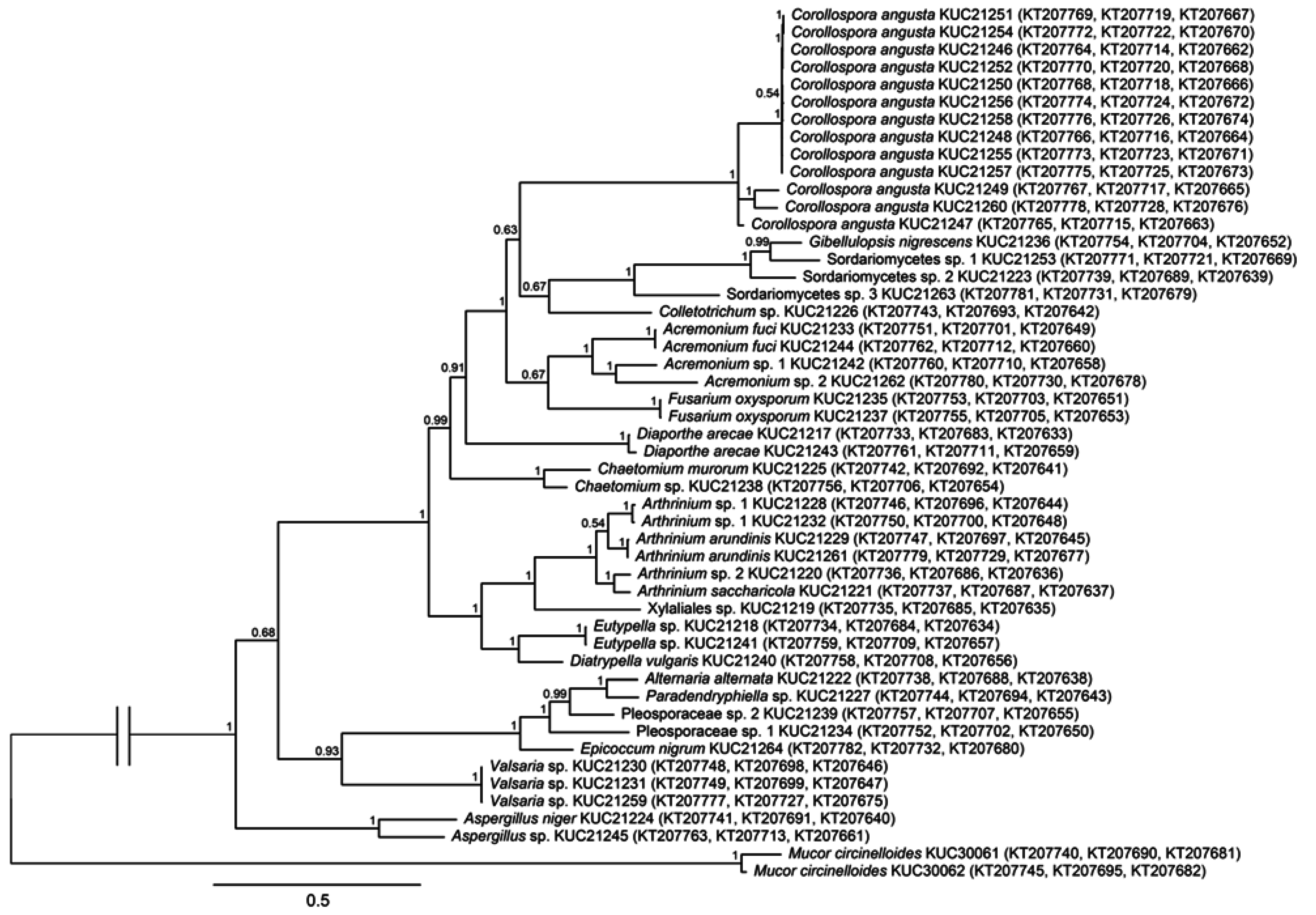
2.1. Fungal Phylogeny and Identification
| Fungal Identity | ID | Antifungal Activity IC50 (μg/mL) | Antioxidant Activity IC50 (μg/mL) | Cellulolytic Enzyme Activity (U/mL) | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Botrytis cinerea | Collectotrichum gloeosporioides | Fusarium oxysporum | ABTS a Radical Scavenging Activity | DPPH b Radical Scavenging Activity | FPU c | EG d | BGL e | BXL f | ||
| Ascomycota | ||||||||||
| Dothideomycetes | ||||||||||
| Alternaria alternata | KUC21222 | N.D. g | N.D. | >1000 | >1000 | N.D. | N.D. g | N.D. | 0.07 ± 0 | N.D. |
| Epicoccum nigrum | KUC21264 | >1000 | >1000 | N.D. | N.D. | N.D. | 0.05 ± 0.09 | N.D. | 0.05 ± 0.01 | 0.02 ± 0 |
| Paradendryphiella sp. | KUC21227 | >1000 | N.D. | >1000 | N.D. | N.D. | 0.15 ± 0 | 0.03 ± 0.06 | 0.04 ± 0 | N.D. |
| Pleosporaceae sp. 1 | KUC21234 | N.D. | >1000 | N.D. | N.D. | N.D. | 0.1 ± 0.09 | N.D. | 0.03 ± 0.02 | N.D. |
| Pleosporaceae sp. 2 | KUC21239 | N.D. | N.D. | >1000 | N.D. | N.D. | N.D. | 0.03 ± 0.06 | 0.05 ± 0 | N.D. |
| Valsaria sp. | KUC21230 | >1000 | N.D. | >1000 | >1000 | N.D. | 0.05 ± 0.08 | 0.04 ± 0.07 | 0.01 ± 0.02 | N.D. |
| KUC21231 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | 0.03 ± 0.02 | N.D. | |
| KUC21259 | N.D. | N.D. | 559.0 | >1000 | N.D. | N.D. | 0.08 ± 0.08 | 0.02 ± 0.03 | N.D. | |
| Eurotiomycetes | ||||||||||
| Aspergillus niger. | KUC21224 | >1000 | >1000 | 72.5 | 272.2 | >1000 | 0.15 ± 0 | 0.04 ± 0.08 | 0.05 ± 0 | 0.02 ± 0 |
| Aspergillus sp. | KUC21245 | >1000 | N.D. | >1000 | >1000 | N.D. | N.D. | 0.14 ± 0.02 | 0.11 ± 0.01 | 0.02 ± 0 |
| Sordariomycetes | ||||||||||
| Acremonium fuci | KUC21233 | N.D. | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. |
| KUC21244 | 523.8 | >1000 | N.D. | N.D. | N.D. | N.D. | 0.03 ± 0.06 | 0.01 ± 0.02 | N.D. | |
| Acremonium sp. 1 | KUC21242 | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. |
| Acremonium sp. 2 | KUC21262 | N.D. | N.D. | 670.7 | >1000 | N.D. | 0.1 ± 0.09 | 0.11 ± 0 | 0.07 ± 0.01 | N.D. |
| Arthrinium arundinis | KUC21229 | N.D. | N.D. | 193.1 | N.D. | N.D. | 0.1 ± 0.09 | 0.17 ± 0.04 | 0.05 ± 0 | N.D. |
| KUC21261 | >1000 | 590.3 | 225.9 | N.D. | N.D. | 0.1 ± 0.09 | 0.18 ± 0.07 | 0.04 ± 0 | N.D. | |
| Arthrinium saccharicola | KUC21221 | 210.5 | N.D. | >1000 | 46 | 88.4 | 0.39 ± 0.13 | 0.38 ± 0 | 1.04 ± 0.03 | 0.02 ± 0 |
| Arthrinium sp. 1 | KUC21228 | 574.8 | N.D. | 391.8 | N.D. | N.D. | 0.15 ± 0 | 0.26 ± 0.04 | 0.1 ± 0.01 | 0.02 ± 0 |
| KUC21232 | 172.8 | >1000 | 139.9 | N.D. | N.D. | 0.15 ± 0 | 0.14 ± 0.02 | 0.06 ± 0 | 0.01 ± 0.01 | |
| Arthrinium sp. 2 | KUC21220 | 179.9 | >1000 | >1000 | >1000 | >1000 | 0.15 ± 0 | 0.14 ± 0.02 | 0.16 ± 0.04 | 0.02 ± 0 |
| Chaetomium murorum | KUC21225 | N.D. | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | 0.04 ± 0 | N.D. |
| Chaetomium sp. | KUC21238 | N.D. | N.D. | >1000 | N.D. | N.D. | N.D. | 0.07 ± 0.06 | 0.04 ± 0 | N.D. |
| Colletotrichum sp. | KUC21226 | >1000 | N.D. | >1000 | N.D. | N.D. | 0.15 ± 0 | 0.13 ± 0.01 | 0.04 ± 0 | 0.01 ± 0.01 |
| Corollospora angusta | KUC21246 | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. |
| KUC21247 | N.D. | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | |
| KUC21248 | >1000 | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | |
| KUC21249 | N.D. | N.D. | N.D. | N.D. | N.D. | 0.05 ± 0.09 | N.D. | N.D. | N.D. | |
| KUC21250 | N.D. | N.D. | N.D. | N.D. | N.D. | 0.1 ± 0.09 | N.D. | N.D. | N.D. | |
| KUC21251 | N.D. | N.D. | N.D. | N.D. | N.D. | 0.1 ± 0.09 | N.D. | N.D. | N.D. | |
| KUC21252 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | |
| KUC21254 | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | |
| KUC21255 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | 0.04 ± 0 | N.D. | |
| KUC21256 | N.D. | >1000 | >1000 | N.D. | N.D. | N.D. | N.D. | 0.01 ± 0.02 | N.D. | |
| KUC21257 | >1000 | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | |
| KUC21258 | >1000 | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | |
| KUC21260 | >1000 | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | |
| Diaporthe arecae | KUC21217 | N.D. | N.D. | N.D. | N.D. | N.D. | 0.16 ± 0.01 | 0.14 ± 0.03 | 0.29 ± 0.08 | 0.01 ± 0.01 |
| KUC21243 | >1000 | N.D. | N.D. | N.D. | N.D. | 0.1 ± 0.09 | 0.13 ± 0.02 | 0.13 ± 0.07 | 0.01 ± 0.01 | |
| Diatrypella vulgaris | KUC21240 | N.D. | N.D. | 758 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. |
| Eutypella sp. | KUC21218 | >1000 | N.D. | N.D. | N.D. | N.D. | 0.05 ± 0.09 | 0.14 ± 0.01 | 0.18 ± 0.04 | 0.02 ± 0 |
| KUC21241 | N.D. | >1000 | >1000 | N.D. | N.D. | N.D. | 0.16 ± 0.05 | 0.17 ± 0.09 | 0.01 ± 0.01 | |
| Fusarium oxysporum | KUC21235 | N.D. | N.D. | >1000 | N.D. | N.D. | 0.15 ± 0 | N.D. | 0.04 ± 0 | 0.01 ± 0.01 |
| KUC21237 | N.D. | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | 0.04 ± 0 | N.D. | |
| Gibellulopsis nigrescens | KUC21236 | N.D. | N.D. | >1000 | N.D. | N.D. | 0.05 ± 0.09 | 0.04 ± 0.06 | 0.04 ± 0 | N.D. |
| Sordariomycetes sp. 1 | KUC21253 | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. |
| Sordariomycetes sp. 2 | KUC21223 | >1000 | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. | N.D. |
| Sordariomycetes sp. 3 | KUC21263 | N.D. | N.D. | >1000 | N.D. | N.D. | N.D. | N.D. | 0.04 ± 0 | N.D. |
| Xylariales sp. | KUC21219 | >1000 | >1000 | 97.9 | >1000 | >1000 | 0.05 ± 0.09 | N.D. | 0.05 ± 0.01 | 0.01 ± 0.01 |
| Zygomycota | ||||||||||
| Mucoromyciotina | ||||||||||
| Mucor circinelloides | KUC30061 | >1000 | N.D. | >1000 | >1000 | N.D. | 0.05 ± 0.09 | N.D. | 0.13 ± 0.03 | 0.01 ± 0.01 |
| KUC30062 | N.D. | N.D. | N.D. | N.D. | N.D. | 0.15 ± 0 | 0.04 ± 0.06 | 0.05 ± 0.01 | N.D. | |


2.2. Biological Activities of the Strains
2.2.1. Antifungal Activity and Antioxidant Activity
2.2.2. Antioxidant Activity
2.2.3. Cellulolytic Enzymes
| Enzyme | Strains | Substrate | Cultivation Conditions | Activity (U/mL) | References |
|---|---|---|---|---|---|
| FPU a | A. saccharicola KUC 21221 | Cellulose | Shake flask, 25 °C, 168 h | 0.39 | This study |
| Penicillium echinulatum | Cellulose | Stir fermenter, 37 °C, 192 h | 0.27 | [56] | |
| EG b | A. saccharicola KUC 21221 | Cellulose | Shake flask, 25 °C, 168 h | 0.39 | This study |
| Hypoxylon oceanicum | Cellulose + Filter paper | Shake flask, 25 °C, 360 h | 0.04 | [57] | |
| BGL c | A. saccharicola KUC 21221 | Cellulose | Shake flask, 25 °C, 168 h | 1.04 | This study |
| Aspergillus niger | Cellulose | Shake flask, 28 °C, 168 h | 1.02 | [58] |
3. Experimental Section
3.1. Sampling and Isolation of Fungi
3.2. DNA Extraction, PCR and Identification
3.3. Phylogenetic Analysis
3.4. Biological Activity
3.4.1. Preparation of Extracts
3.4.2. Antifungal Activity
3.4.3. Antioxidant Activity
ABTS Radical-Scavenging Assay
DPPH Radical-Scavenging Assay
3.4.4. Cellulolytic Enzyme
Enzyme Preparation
Enzyme Assays
3.5. Statistical Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Abdel-Wahab, M.A.; Pang, K.L.; Nagahama, T.; Abdel-Aziz, F.A.; Jones, E.G. Phylogenetic evaluation of anamorphic species of Cirrenalia and Cumulospora with the description of eight new genera and four new species. Mycol. Prog. 2010, 9, 537–558. [Google Scholar] [CrossRef]
- Manimegalai, K.; Devi, N.A.; Padmavathy, S. Marine fungi as a source of secondary metabolites of antibiotics. Int. J. Biotechnol. Bioeng. Res. 2013, 4, 275–282. [Google Scholar]
- Park, M.S.; Fong, J.J.; Oh, S.Y.; Kwon, K.K.; Sohn, J.H.; Lim, Y.W. Marine-derived Penicillium in Korea: Diversity, enzyme activity, and antifungal properties. Antonie Leeuwenhoek 2014, 106, 331–345. [Google Scholar] [CrossRef] [PubMed]
- Jones, E.B. Are there more marine fungi to be described? Bot. Mar. 2011, 54, 343–354. [Google Scholar] [CrossRef]
- Suryanarayanan, T.S.; Venkatachalam, A.; Thirunavukkarasu, N.; Ravishankar, J.P.; Doble, M.; Geetha, V. Internal mycobiota of marine macroalgae from the Tamilnadu coast: Distribution, diversity and biotechnological potential. Bot. Mar. 2010, 53, 457–468. [Google Scholar] [CrossRef]
- Zuccaro, A.; Schoch, C.L.; Spatafora, J.W.; Kohlmeyer, J.; Draeger, S.; Mitchell, J.I. Detection and identification of fungi intimately associated with the brown seaweed Fucus serratus. Appl. Environ. Microbiol. 2008, 74, 931–941. [Google Scholar] [CrossRef] [PubMed]
- Suryanarayanan, T.S. Fungal endosymbionts of seaweeds. In Biology of Marine Fungi; Springer: Berlin, Germany, 2012. [Google Scholar]
- White, T.J.; Bruns, T.D.; Lee, S.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Griffith, G.W. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [PubMed]
- Huh, N.; Jang, Y.; Lee, J.; Kim, G.-H.; Kim, J.-J. Phylogenetic analysis of major molds inhabiting woods and their discoloration characteristics. Part 1. Genus Trichoderma. Holzforschung 2011, 65, 257–263. [Google Scholar] [CrossRef]
- Crous, P.W.; Groenewald, J.Z. A phylogenetic re-evaluation of Arthrinium. IMA Fungus 2013, 4, 133–154. [Google Scholar] [CrossRef] [PubMed]
- James, T.Y.; Kauff, F.; Schoch, C.L.; Matheny, P.B.; Hofstetter, V.; Cox, C.J.; Celio, G.; Gueidan, C.; Fraker, E.; Miadlikowska, J.; et al. Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature 2006, 443, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Sakayaroj, J.; Pang, K.L.; Jones, E.G. Multi-gene phylogeny of the Halosphaeriaceae: Its ordinal status, relationships between genera and morphological character evolution. Fungal Divers. 2011, 46, 87–109. [Google Scholar] [CrossRef]
- Mayer, A.M.; Rodríguez, A.D.; Berlinck, R.G.; Fusetani, N. Marine pharmacology in 2007–2008: Marine compounds with antibacterial, anticoagulant, antifungal, anti-inflammatory, antimalarial, antiprotozoal, antituberculosis, and antiviral activities; affecting the immune and nervous system, and other miscellaneous mechanisms of action. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2011, 153, 191–222. [Google Scholar] [PubMed]
- Kelecom, A. Secondary metabolites from marine microorganisms. An. Acad. Bras. Cienc. 2002, 74, 151–170. [Google Scholar] [CrossRef] [PubMed]
- Bugni, T.S.; Ireland, C.M. Marine-derived fungi: A chemically and biologically diverse group of microorganisms. Nat. Prod. Rep. 2004, 21, 143–163. [Google Scholar] [CrossRef] [PubMed]
- Wei, M.Y.; Wang, C.Y.; Liu, Q.A.; Shao, C.L.; She, Z.G.; Lin, Y.C. Five sesquiterpenoids from a marine-derived fungus Aspergillus sp. isolated from a gorgonian Dichotella gemmacea. Mar. Drugs 2010, 8, 941–949. [Google Scholar] [CrossRef] [PubMed]
- Bai, Z.Q.; Lin, X.; Wang, J.; Zhou, X.; Liu, J.; Yang, B.; Yang, X.; Liao, S.; Wang, L.; Liu, Y. New Meroterpenoids from the Endophytic Fungus Aspergillus flavipes AIL8 Derived from the Mangrove Plant Acanthus ilicifolius. Mar. Drugs 2015, 13, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Kito, K.; Ookura, R.; Yoshida, S.; Namikoshi, M.; Ooi, T.; Kusumi, T. New cytotoxic 14-membered macrolides from marine-derived fungus Aspergillus ostianus. Org. Lett. 2008, 10, 225–228. [Google Scholar] [CrossRef] [PubMed]
- Fenical, W.; Jensen, P.R.; Cheng, X.C. Avraincillamide, a Cytotoxic Marine Natural Product, and Derivatives Thereof. U.S. Patent 6,066,635, 23 May 2000. [Google Scholar]
- Balboa, E.M.; Conde, E.; Soto, M.L.; Pérez-Armada, L.; Domínguez, H. Cosmetics from Marine Sources. In Hb25_Springer Handbook of Marine Biotechnology; Springer: Berlin, Germany, 2015; pp. 1015–1042. [Google Scholar]
- Veiga, M.; Esparis, A.; Fabregas, J. Isolation of cellulolytic actinomycetes from marine sediments. Appl. Environ. Microbiol. 1983, 46, 286. [Google Scholar] [PubMed]
- Cowie, G.L.; Hedges, J.I. Carbohydrate sources in a coastal marine environment. Geochim. Cosmochim. Acta 1984, 48, 2075–2087. [Google Scholar] [CrossRef]
- Rohrmann, S.; Molitoris, H.P. Screening for wood-degrading enzymes in marine fungi. Can. J. Bot. 1992, 70, 2116–2123. [Google Scholar] [CrossRef]
- Meyers, S.P.; Prindle, B.; Reynolds, E.S. Cellulolytic activity of marine fungi. Degradation of lignocellulose material. Tappi 1960, 43, 534–538. [Google Scholar]
- Nakagiri, A.; Tokura, R. Taxonomic studies of the genus Corollospora (Halosphaeriaceae, Ascomycotina) with descriptions of seven new species. Trans. Mycol. Soc. Jpn. 1987, 28, 413–436. [Google Scholar]
- Prasannarai, K.; Sridhar, K.R. Abundance and diversity of marine fungi on intertidal woody litter of the west coast of India on prolonged incubation. Fungal Divers. 2003, 14, 127–141. [Google Scholar]
- Luo, W.; Vrijmoed, L.L.; Jones, E.B. Screening of marine fungi for lignocellulose-degrading enzyme activities. Bot. Mar. 2005, 48, 379–386. [Google Scholar] [CrossRef]
- Khan, S.A.; Hamayun, M.; Kim, H.-Y.; Yoon, H.-J.; Seo, J.-C.; Choo, Y.-S.; Lee, I.-J.; Kim, S.D.; Rhee, I.-K.; Kim, J.-G. A new strain of Arthrinium phaeospermum isolated from Carex kobomugi Ohwi is capable of gibberellin production. Biotechnol. Lett. 2009, 31, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-J.; Lee, S.-S.; Ra, J.-B.; Lee, H.; Huh, N.; Kim, G.-H. Fungi associated with bamboo and their decay capabilities. Holzforschung 2011, 65, 271–275. [Google Scholar] [CrossRef]
- Martínez-Cano, C.; Grey, W.E.; Sands, D.C. First report of Arthrinium arundinis causing kernel blight on barley. Plant Dis. 1992, 76, 1077. [Google Scholar] [CrossRef]
- Saccardo, P.A. Sylloge Fungorum; Sumptibus Auctoris: Pavia, Italy, 1931; Volume 25, p. 771. [Google Scholar]
- Miao, L.I.; Kwong, T.F.; Qian, P.Y. Effect of culture conditions on mycelial growth, antibacterial activity, and metabolite profiles of the marine-derived fungus Arthrinium cf saccharicola. Appl. Microbiol. Biotechnol. 2006, 72, 1063–1073. [Google Scholar] [CrossRef] [PubMed]
- Glenn, A.E.; Bacon, C.W.; Price, R.; Hanlin, R.T. Molecular phylogeny of Acremonium and its taxonomic implications. Mycologia 1996, 88, 369–383. [Google Scholar] [CrossRef]
- Samson, R.A.; Hoekstra, E.S.; Frisvad, J.C. Introduction to Food-And Airborne Fungi, 7th ed.; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 2004. [Google Scholar]
- Zuccaro, A.; Summerbell, R.C.; Gams, W.; Schroers, H.J.; Mitchell, J.I. A new Acremonium species associated with Fucus spp., and its affinity with a phylogenetically distinct marine Emericellopsis clade. Stud. Mycol. 2004, 50, 283–297. [Google Scholar]
- Oren, A.; Gunde-Cimerman, N. Fungal Life in the Dead Sea. In Biology of Marine Fungi; Springer: Berlin, Germany, 2012. [Google Scholar]
- Khambhaty, Y.; Mody, K.; Basha, S.; Jha, B. Kinetics, equilibrium and thermodynamic studies on biosorption of hexavalent chromium by dead fungal biomass of marine Aspergillus niger. Chem. Eng. J. 2009, 145, 489–495. [Google Scholar] [CrossRef]
- Xue, D.S.; Chen, H.Y.; Ren, Y.R.; Yao, S.J. Enhancing the activity and thermostability of thermostable β-glucosidase from a marine Aspergillus niger at high salinity. Process Biochem. 2012, 47, 606–611. [Google Scholar] [CrossRef]
- Gordon, T.R.; Martyn, R.D. The evolutionary biology of Fusarium oxysporum. Annu. Rev. Phytopathol. 1997, 35, 111–128. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Mu, J.; Feng, Y.; Kang, Y.; Zhang, J.; Gu, P.J.; Wang, Y.; Ma, L.-F.; Zhu, Y.H. Broad-spectrum antimicrobial epiphytic and endophytic fungi from marine organisms: Isolation, bioassay and taxonomy. Mar. Drugs 2009, 7, 97–112. [Google Scholar] [CrossRef] [PubMed]
- Khallil, A.R.; El-Hissy, F.T.; Bagy, M.M.K. Mycoflora of mangroves of Red Sea in Egypt. Folia Microbiol. 1991, 36, 456–464. [Google Scholar] [CrossRef]
- Hong, S.B.; Kim, D.H.; Lee, M.; Baek, S.Y.; Kwon, S.W.; Houbraken, J.; Samson, R.A. Zygomycota associated with traditional meju, a fermented soybean starting material for soy sauce and soybean paste. J. Microbiol. 2012, 50, 386–393. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Lee, J.; Kwak, J.; Kim, K.; Seo, M.; Lee, Y.W. Fungi associated with the traditional starter cultures used for rice wine in Korea. J. Kor. Soc. Appl. Biol. Chem. 2011, 54, 933–943. [Google Scholar] [CrossRef]
- Vicente, G.; Bautista, L.F.; Rodríguez, R.; Gutiérrez, F.J.; Sádaba, I.; Ruiz-Vázquez, R.M.; Torres-Martínezb, S.; Garre, V. Biodiesel production from biomass of an oleaginous fungus. Biochem. Eng. J. 2009, 48, 22–27. [Google Scholar] [CrossRef]
- Lübbehüsen, T.L.; Nielsen, J.; Mcintyre, M. Aerobic and anaerobic ethanol production by Mucor circinelloides during submerged growth. Appl. Microbiol. Biotechnol. 2004, 63, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Jiménez, E.; Dorta, F.; Medina, C.; Ramírez, A.; Ramírez, I.; Peña-Cortés, H. Anti-phytopathogenic activities of macro-algae extracts. Mar. Drugs 2011, 9, 739–756. [Google Scholar] [CrossRef] [PubMed]
- Ebada, S.S.; Schulz, B.; Wray, V.; Totzke, F.; Kubbutat, M.H.; Müller, W.E.; Hamacherf, A.; Kassackf, M.U.; Ling, W.; Proksch, P. Arthrinins A–D: Novel diterpenoids and further constituents from the sponge derived fungus Arthrinium sp. Bioorg. Med. Chem. 2011, 19, 4644–4651. [Google Scholar] [CrossRef] [PubMed]
- Tsukamoto, S.; Yoshida, T.; Hosono, H.; Ohta, T.; Yokosawa, H. Hexylitaconic acid: A new inhibitor of p53-HDM2 interaction isolated from a marine-derived fungus, Arthrinium sp. Bioorg. Med. Chem. Lett. 2006, 16, 69–71. [Google Scholar] [CrossRef] [PubMed]
- Tsukada, M.; Fukai, M.; Miki, K.; Shiraishi, T.; Suzuki, T.; Nishio, K.; Sugita, T.; Ishino, M.; Kinoshita, K.; Takahashi, K.; et al. Chemical constituents of a marine fungus, Arthrinium sacchari. J. Nat. Prod. 2011, 74, 1645–1649. [Google Scholar] [CrossRef] [PubMed]
- Bloor, S. Arthrinic acid, a novel antifungal polyhydroxyacid from Arthrinium phaeospermum. J. Antibiot. 2008, 61, 515–517. [Google Scholar] [CrossRef] [PubMed]
- Amens, B.N. Dietary carcinogens and anticarcinogens. Science 1983, 221, 1256. [Google Scholar] [CrossRef]
- Li, X.; Li, X.M.; Xu, G.M.; Li, C.S.; Wang, B.G. Antioxidant metabolites from marine alga-derived fungus Aspergillus wentii EN-48. Phytochem. Lett. 2014, 7, 120–123. [Google Scholar] [CrossRef]
- Sun, H.H.; Mao, W.J.; Chen, Y.; Guo, S.D.; Li, H.Y.; Qi, X.H.; Chen, Y.L.; Xu, J. Isolation, chemical characteristics and antioxidant properties of the polysaccharides from marine fungus Penicillium sp. F23-2. Carbohydr. Polym. 2009, 78, 117–124. [Google Scholar] [CrossRef]
- Parra, R.; Aldred, D.; Magan, N. Medium optimization for the production of the secondary metabolite squalestatin S1 by a Phoma sp. combining orthogonal design and response surface methodology. Enzym. Microb. Technol. 2005, 37, 704–711. [Google Scholar] [CrossRef]
- Gogoi, D.K.; Boruah, H.P.D.; Saikia, R.; Bora, T.C. Optimization of process parameters for improved production of bioactive metabolite by a novel endophytic fungus Fusarium sp. DF2 isolated from Taxus wallichiana of North East India. World J. Microbiol. Biotechnol. 2008, 24, 79–87. [Google Scholar] [CrossRef]
- Sun, Y.; Cheng, J. Hydrolysis of lignocellulosic materials for ethanol production: A review. Bioresour. Technol. 2002, 83, 1–11. [Google Scholar] [CrossRef]
- Martins, L.F.; Kolling, D.; Camassola, M.; Dillon, A.J.P.; Ramos, L.P. Comparison of Penicillium echinulatum and Trichoderma reesei cellulases in relation to their activity against various cellulosic substrates. Bioresour. Technol. 2008, 99, 1417–1424. [Google Scholar] [CrossRef] [PubMed]
- Pointing, S.B.; Buswell, J.A.; Jones, E.G.; Vrijmoed, L.L.P. Extracellular cellulolytic enzyme profiles of five lignicolous mangrove fungi. Mycol. Res. 1999, 103, 696–700. [Google Scholar] [CrossRef]
- Narasimha, G.; Sridevi, A.; Buddolla, V.; Subhosh, C.M.; Rajasekhar, R.B. Nutrient effects on production of cellulolytic enzymes by Aspergillus niger. Afr. J. Biotechnol. 2006, 5, 472–476. [Google Scholar]
- Gao, L.; Gao, F.; Jiang, X.; Zhang, C.; Zhang, D.; Wang, L.; Wu, G.; Chen, S. Biochemical characterization of a new β-glucosidase (Cel3E) from Penicillium piceum and its application in boosting lignocelluloses bioconversion and forming disaccharide inducers: New insights into the role of β-glucosidase. Process Biochem. 2014, 49, 768–774. [Google Scholar] [CrossRef]
- Jones, E.G.; Jennings, D.H. The effect of salinity on the growth of marine fungi in comparison with non-marine species. Trans. Br. Mycol. Soc. 1964, 47, 619–625. [Google Scholar] [CrossRef]
- Borines, M.G.; de Leon, R.L.; McHenry, M.P. Bioethanol production from farming non-food macroalgae in Pacific island nations: Chemical constituents, bioethanol yields, and prospective species in the Philippines. Renew. Sustain. Energy Rev. 2011, 15, 4432–4435. [Google Scholar] [CrossRef]
- Kjer, J.; Debbab, A.; Aly, A.H.; Prokasc, P. Methods for isolation of marine-deried endophytic fungi and their bioactive secondary products. Nat. Protoc. 2010, 5, 479–490. [Google Scholar] [CrossRef] [PubMed]
- Gardes, M.; Bruns, T.D. ITS primers with enhanced specificity for basidiomycetes: Application to the identification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef] [PubMed]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [PubMed]
- O’Donnell, K.; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol. Phylogenet. Evol. 1997, 7, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Glass, N.; Donaldson, G. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Maddison, W.P.; Maddison, D.R. MacClade 4: Analysis of Phylogeny and Character Evolution; Version 4.08; Sinauer Associates: Sunderland, MA, USA, 2005. [Google Scholar]
- Nylander, J.A.A. MrModeltest v2. Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Hong, J.H.; Lee, J.; Min, M.; Ryu, S.M.; Lee, D.; Kim, G.-H.; Kim, J.-J. 6-Pentyl-α-pyrone as an anti-sapstain compound produced by Trichoderma gamsii KUC1747 inhibits the germination of ophiostomatoid fungi. Holzforschung 2014, 68, 769–774. [Google Scholar] [CrossRef]
- Lee, Y.M.; Lee, H.; Kim, G.-H.; Kim, J.-J. Miniaturized enzyme production and development of micro-assays for cellulolytic and xylanolytic enzymes. J. Microbiol. Methods 2011, 86, 124–127. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Z.; Storms, R.; Tsang, A. Microplate-based filter paper assay to measure total cellulase activity. Biotechnol. Bioeng. 2004, 88, 832–837. [Google Scholar] [CrossRef] [PubMed]
- Ghose, T.K. Measurement of cellulase activities. Pure Appl. Chem. 1987, 59, 257–268. [Google Scholar] [CrossRef]
- Bailey, M.J.; Biely, P.; Poutanen, K. Interlaboratory testing of methods for assay of xylanase activity. J. Biotechnol. 1992, 23, 257–270. [Google Scholar] [CrossRef]
- Valásková, V.; Baldrian, P. Degradation of cellulose and hemicelluloses by the brown rot fungus Piptoporus betulinus: Production of extracellular enzymes and characterization of the major cellulases. Microbiology 2006, 152, 3613–3622. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Jang, Y.; Lee, H.; Lee, S.; Kim, G.-H.; Kim, J.-J. Screening for xylanase and β-xylosidase production from wood-inhabiting Penicillium strains for potential use in biotechnological applications. Holzforschung 2012, 66, 267–271. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hong, J.-H.; Jang, S.; Heo, Y.M.; Min, M.; Lee, H.; Lee, Y.M.; Lee, H.; Kim, J.-J. Investigation of Marine-Derived Fungal Diversity and Their Exploitable Biological Activities. Mar. Drugs 2015, 13, 4137-4155. https://doi.org/10.3390/md13074137
Hong J-H, Jang S, Heo YM, Min M, Lee H, Lee YM, Lee H, Kim J-J. Investigation of Marine-Derived Fungal Diversity and Their Exploitable Biological Activities. Marine Drugs. 2015; 13(7):4137-4155. https://doi.org/10.3390/md13074137
Chicago/Turabian StyleHong, Joo-Hyun, Seokyoon Jang, Young Mok Heo, Mihee Min, Hwanhwi Lee, Young Min Lee, Hanbyul Lee, and Jae-Jin Kim. 2015. "Investigation of Marine-Derived Fungal Diversity and Their Exploitable Biological Activities" Marine Drugs 13, no. 7: 4137-4155. https://doi.org/10.3390/md13074137
APA StyleHong, J.-H., Jang, S., Heo, Y. M., Min, M., Lee, H., Lee, Y. M., Lee, H., & Kim, J.-J. (2015). Investigation of Marine-Derived Fungal Diversity and Their Exploitable Biological Activities. Marine Drugs, 13(7), 4137-4155. https://doi.org/10.3390/md13074137






