The peptide profiles of the 40 strains isolated from the Atlantic Forest (CENA350-389) were investigated using LC/DAD/ESI/QTOF/MS/MS. As an example,
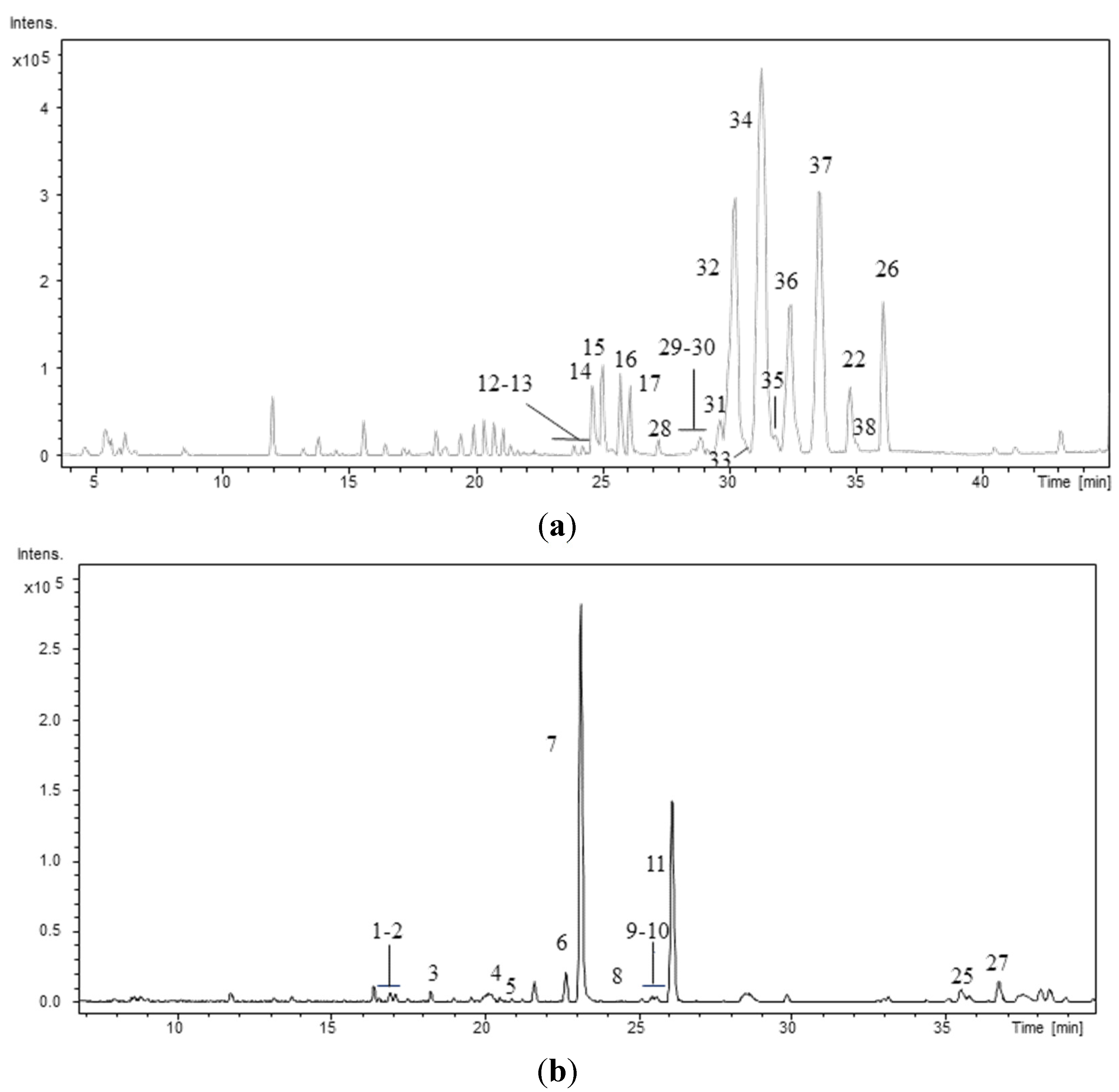
Figure 1 shows the chromatographic profiles of two strains: one
Brasilonema sp. and one
Desmonostoc sp. The retention times (RT), protonated molecules ([M + H]
+), molecular formula provided for the experimental
m/z, and error and millisigma (mSigma) values for the major peaks are summarized in
Table 2. Mass accuracy was below 5 ppm for all the detected compounds. Peak identification was performed based on the data presented in
Table 2,
Table 3,
Table 4,
Table 5,
Table 6,
Table 7,
Table 8,
Table 9,
Table 10,
Table 11,
Table 12,
Table 13 and
Table 14 and based on previously published data [
23,
24,
26,
31,
32,
33,
34,
35,
36,
37,
38,
39,
40]. This approach allowed the elucidation of the planar structures of 38 peptides, including 10 new aeruginosins, 16 new anabaenopeptins, and 11 new cyanopeptolins. Among the surveyed strains the heterocytous strains
Nostoc sp. CENA352, CENA358, and CENA369,
Brasilonema sp. CENA360, CENA361, CENA381, and CENA382 and
Desmonostoc sp. CENA386 and CENA371 were identified as producers of cyanopeptides. Cyanopeptides were not detected in the extracts of the remaining 31 cyanobacterial strains under our experimental conditions.
Figure 1.
Base Peak Chromatogram of the hydromethanolic extract of the strains (a) Desmonostoc sp. CENA386 and (b) Brasilonema sp. CENA358. Conditions as described in experimental section.
2.1. Aeruginosins
Seven strains belonging to the orders Nostocales (
Nostoc sp. CENA352, CENA358, and CENA369; and
Brasilonema sp. CENA360, CENA361, CENA381, and CENA382) were found to produce aeruginosins (
1–11). Aeruginosins are linear tetrapeptides that contain the unusual amino acid 2-carboxy-6-hydroxyoctahydroindol (Choi) in the central position and typically contain an arginine derivative at the
C-terminus [
3]. The
N-terminal position is occupied by a 3-(4-hydroxyphenyl) lactic or phenyl lactic acid, which can be acetylated, brominated, chlorinated, or sulfonated [
41,
42,
43,
44]. A small hydrophobic amino acid (leucine or isoleucine) is typically present in position 2. Additionally, hydroxylation, sulfation, and chlorination of the Choi moiety have also been observed [
45]. These peptides are potent inhibitors of serine proteases, and this bioactivity is largely related to
C-terminal modifications [
46]. This family of compounds has been reported to be produced by cyanobacteria of the genera
Microcystis [
41,
42,
43,
44,
47],
Planktothrix [
24],
Nodularia [
48], and
Nostoc [
37].
The aeruginosins found in these extracts were characterized by closely related structures, most of which were common to both
Nostoc and
Brasilonema producer species. The most prominent peak detected in the MS chromatogram of these extracts (
7) was assigned to the aeruginosin 865 (
m/z 865.4565 [M + H]
+) [
37]. This compound, which was recently isolated from a terrestrial cyanobacterium belonging to
Nostoc sp., was structurally characterized as containing both a fatty acid and a carbohydrate attached to the Choi moiety [
37].
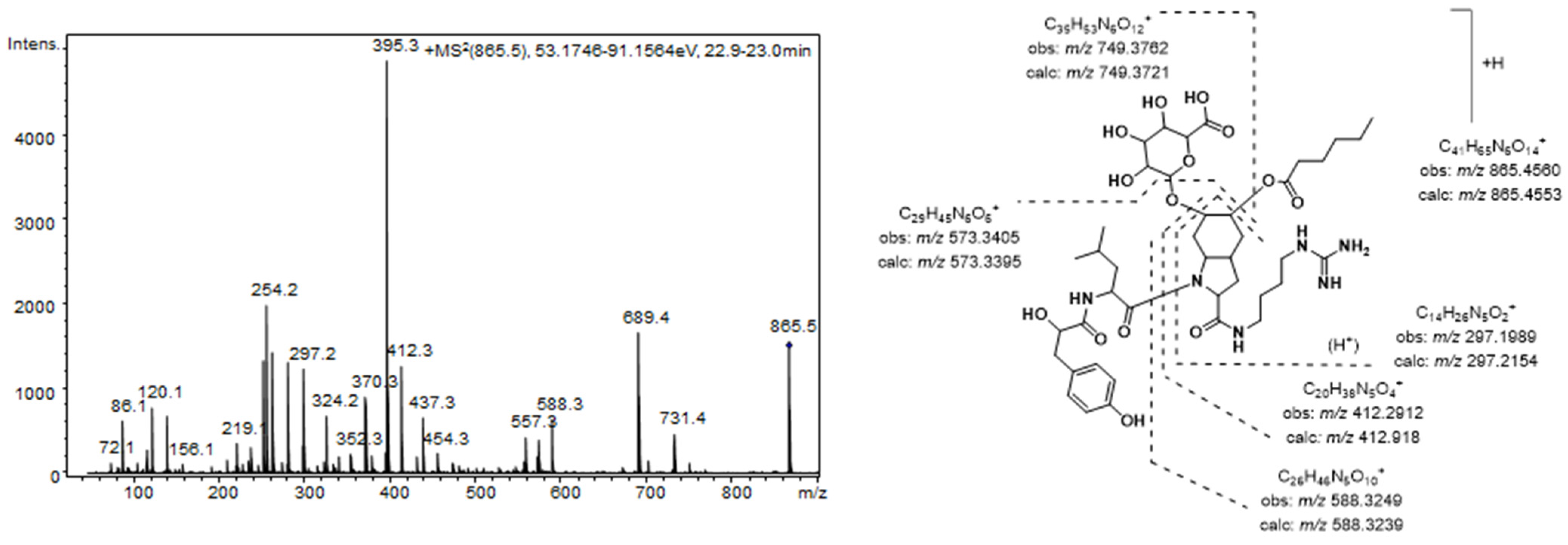
Figure 2 shows the product ion spectra of this aeruginosin. A collision energy of 70 eV was necessary to obtain a spectrum with abundant and intense product ions. Consistent with the existence of an agmatine (Agma) residue in the molecule, the
C-terminal ions (
m/z 588.3249, 412.2912, and 297.1989) and the corresponding satellite ions, which were produced via ammonia or water loss (
m/z 571.2985, 395.2657, 279.1820, and 261.1717) dominated the spectrum. Additionally, the ions generated by the cleavage of the glycosidic acid and/or the ester bond established the sugar and the lipid acids as glucuronic acid and hexanoic acid, respectively. The presence of ions at
m/z 18 u higher (
m/z 156.1035 and 138.0866) than the diagnostic ions that are typically generated from the Choi residue (
m/z 140 and 122) indicated the dihydroxylation of the indole ring in this amino acid and were key fragments for the detection of other aeruginosin congeners.
Figure 2.
Product ion spectrum for [M + H]+ of compound 7 and its predicted fragmentation pattern. Conditions as mentioned in the experimental section.
Figure 2.
Product ion spectrum for [M + H]+ of compound 7 and its predicted fragmentation pattern. Conditions as mentioned in the experimental section.
A total of 10 additional aeruginosins that shared conserved substructures (
1–
6,
8–
11) and yielded [M + H]
+ at
m/z 753.4042 (
1), 767.3828 (
2), 737.4044 (
3), 823.4470 (
4), 837.4253 (
5), 851.4780 (
6), 879.5078 (
8), 879.5078 (
9), 849.4596 (
10), and 893.4872 (
11) were identified as new congeners. The product ion spectra of these aeruginosins were similar to that of compound
7, allowing us to elucidate their structures based on a comparison of spectra (
Table 3,
Table 4 and
Table 5).
Table 3.
Product ion spectra data for compounds 4–7, 9, and 11.
Table 3.
Product ion spectra data for compounds 4–7, 9, and 11.
| Product Ion Assignment a | 4 (m/z) | 5 (m/z) | 6 (m/z) | 7 (m/z) | 9 (m/z) | 11 (m/z) |
|---|
| Leu immonium | 86.0967 | 86.0947 | 86.0907 | 86.0945 | 86.0906 | 86.0932 |
| OH-Choi immonium ion − H2O | 138.0791 | 138.0866 | 138.0910 | 138.0912 | 138.0938 | 138.0903 |
| OH-Choi immonium ion | 156.1035 | 156.1035 | 156.1033 | 156.1026 | - | 156.1031 |
| OH-Choi-Agma − NH3 − H2O + H | 261.1693 | 261.1707 | 261.1799 | 261.1717 | 261.1791 | 261.1709 |
| OH-Choi-Agma − H2O + H | 279.1802 | 279.1801 | 279.1799 | 279.1820 | 279.1796 | 279.1828 |
| OH-Choi-Agma + H | 297.1925 | 297.1885 | 297.1882 | 297.1989 | 297.1880 | 297.1878 |
| (R1)-O-Choi-Agma + H − NH3 | 367.2348 | 367.2342 | 395.2672 | 395.2657 | 409.2803 | 423.2969 |
| (R1)-O-Choi-Agma + H | 384.2618 | 384.2617 | 412.2803 | 412.2912 | 426.3113 | 440.3226 |
| (R1, R2)-OH-Choi-Agma+ H − NH3 | - b | 543.2599 | - | 571.2985 | - | 599.3216 |
| (R1, R2)-OH-Choi-Agma + H | 546.3176 | 560.2818 | 574.3454 | 588.3249 | - | 616.3618 |
| Hpla-Leu-OHChoi-Agma + H − H2O | 573.3423 | 573.3422 | 573.3417 | 573.3405 | - | 573.3409 |
| Hpla-Leu-(R1)OH-Choi-Agma | 661.3949 | 661.3909 | 689.4266 | 689.4250 | 703.4423 | 717.4543 |
| Hpla-Leu-(R2)OH-Choi-Agma | 735.3952 | 749.3722 | 735.3945 | 749.3762 | - | - |
| Hpla-Leu-(R1, R2)-OH-Choi-Agma | 823.4470 | 837.4253 | 851.4780 | 865.4560 | 879.5078 | 893.4872 |
Table 4.
Product ion spectra data for compounds 1–3.
Table 4.
Product ion spectra data for compounds 1–3.
| Product Ion Assignment a | 1 (m/z) | 3 (m/z) | 2 * (m/z) |
|---|
| Leu immonium | 86.0908 | 86.0908 | 86.0967 |
| OH-Choi immonium ion − H2O | 138.0867 | 138.0941 | 122.1066 * |
| OH-Choi immonium ion | 156.0957 | 156.1036 | 140.1034 * |
| OH-Choi-Agma − NH3 − H2O | 261.1592 | - b | - |
| OH-Choi-Agma − NH3 | 297.1920 | 297.1928 | 263.1868 * |
| OH-Choi-Agma | 314.2143 | 314.2255 | 281.1914 * |
| R2-OH-Choi-Agma − NH3 | - | 473.2107 | 443.2511 * |
| R2-OH-Choi-Agma | 476.2717 | 490.2454 | 460.2740 * |
| Hpla-Leu-OHChoi-Agma | 591.3360 | 591.3204 | 575.3841 * |
| Hpla-Leu-(R2)-OH-Choi-Agma | 753.4042 | 767.3828 | 737.4044 * |
Table 5.
Product ion spectra data for compounds 8 and 10.
Table 5.
Product ion spectra data for compounds 8 and 10.
| Product Ion Assignment a | 8 (m/z) | 10 (m/z) |
|---|
| X immonium | 100.1148 | 86.0906 |
| OH-Choi immonium ion − H2O | 138.1248 | 138.0863 |
| OH-Choi immonium ion | - b | 156.0952 |
| OH-Choi-Agma − NH3 − H2O | 261.1689 | 261.1687 |
| OH-Choi-Agma − H2O | 279.1797 | 279.1796 |
| OH-Choi-Agma − NH3 | 297.1771 | - |
| OH-Choi-Agma + H | - | - |
| (R1)-OH-Choi-Agma − NH3 | 395.2544 | 395.2659 |
| (R1)-OH-Choi-Agma + H | 412.2930 | 412.2928 |
| (R1, R2)-OH-Choi-Agma − NH3 | 571.3340 | 571.2881 |
| (R1, R2)-OH-Choi-Agma + H | 588.3205 | 588.3238 |
| Z-X-(R1)OH-Choi-Agma + H | - | 673.4238 |
| Z-X-(R1)-OH-Choi-Agma + H | 703.4398 | - |
| Z-X-(R1, R2)-OH-Choi-Agma + H | 879.4717 | 849.4596 |
In this sense, a structure similar to that of aeruginosin 865 was proposed for compounds 5, 9, and 11 except for the fatty acid esterifying position 5 of the Choi moiety. In this position, butanoic, heptanoic, and octanoic acid were proposed for each compound, respectively. These changes were clearly evidenced by the sequence of ions containing the aforementioned fatty acids (m/z 384.2617, 543.2599, 560.2818, and 661.3909 for compound 5; m/z 409.2803 and 703.4423 for compound 9; and m/z 423.2969, 599.3216, 616.3618, and 717.4543 for compound 11). On the other hand, structural differences between the pairs of compounds 4, 5 and 6, 7 were attributed to Choi-glycosylation.A similar product ion spectra, which differed only in the ions generated by the cleavage of the glycosidic bond (m/z 546.3176, 574.3454, and 735.3952/735.3945) suggested that the glucuronic acid in compounds 5 and 7 was replaced by a hexose in compounds 4 and 6.
Compounds
1 and
3, which also showed structures similar to compounds
4 and
6 and
5,
7,
9, and
11, respectively, were distinguished by the lack of fatty acids in their structures (
Table 4). These peptides could be biosynthetic intermediates of the respective fatty acid-containing aeruginosins. Along with these compounds, an oxygen-deficient variant of compound
1 (
2) was also detected; this structural difference was likely due to the absence of the Choi 5-hydroxylation.
Finally, two other structural variants of compound
7 were also detected (
8 and
10). For compound
8, methylation of the amino acid in the second position was suggested based on the mostly conserved product ion spectrum and the presence of a fragment ion at
m/
z 100.1148 (
NMeLeu immonium). Similarly, for compound
10, a phenyl lactic acid was proposed for the
N-terminus instead of a hydroxyl-phenyl lactic acid (
Table 5).
From a biomedical point of view, the pharmacological potential of these new aeruginosin variants must be evaluated. As mentioned above, aeruginosins typically exhibit antithrombotic activity, making these compounds interesting candidates for the development of anticoagulant drugs [
46]. Additionally, all of these compounds are structurally similar to aeruginosin-865, which has exhibited remarkable anti-inflammatory activity [
37]. Evaluations of the bioactivity of these compounds could provide insights into the structure-activity relationship of this class of aeruginosins.
2.2. Anabaenopeptins
Fifteen compounds produced by
Nostoc sp. CENA352,
Brasilonema sp. CENA360, and
Desmonostoc sp. CENA386 were characterized as new anabaenopeptin analogs (
12–
27). Protonated molecules in the mass range of anabaenopeptins and an important loss of the amino acid in the side-chain position were the diagnostic criteria used to classify these peptides [
23]. These new anabaenopeptins were characterized based on empirical formulae, product ion spectra, and previously described sequences [
23,
24,
26,
34,
36].
Anabaenopeptins are hexapeptides that contain a ring of five amino acids. Position 2 is always occupied by
d-Lys, which both closes the ring with the amino acid at position 6 and establishes a ureido link with the amino acid in position 1, giving rise to a side chain. Positions 4 and 5 are typically occupied by aromatic and methylated amino acids, respectively. Position 3 has been reported to be occupied primarily by valine or isoleucine/leucine and less frequently by methionine [
3]. Various biological activities have been described for these structures, including inhibition of protein phosphatase [
49], carboxypeptidases A [
50,
51] and U [
52], and other protease inhibitory activity [
53]. To date, 30 anabaenopeptins have been isolated from many different cyanobacteria genera [
8,
54] (
Anabaena [
55,
56],
Aphanizomenon [
51],
Lyngbya [
57],
Microcystis [
58],
Oscillatoria [
53,
59],
Planktothrix [
60], and
Schizothrix [
61]) and also from marine sponges [
62,
63].
The detected anabaenopeptins could be grouped according to their structural features. Among these compounds, 10 anabaenopeptins that shared similar structural characteristics were detected in
Desmonostoc sp. CENA360 and
Brasilonema sp. CENA386 (
12–
17,
22,
25–
27). The representative fragmentation pathways and spectra of these compounds are described in
Figure 3 and
Figure 4, and the assignments of the principal ions are shown in
Table 6,
Table 7,
Table 8 and
Table 9. Since the nature of the exocyclic amino acid varies among them, two different fragmentation patterns were observed for these compounds depending on the nature of the amino acid side chain. Extensive fragmentation was observed due to the absence of polar and basic residues in this chain. These anabaenopeptins commonly incorporate the amino acid
N-methyl asparagine (
N-MeAsn) in position 5, and on several occasions, either methylation or ethylation was postulated for the homovariant amino acid in position 4.
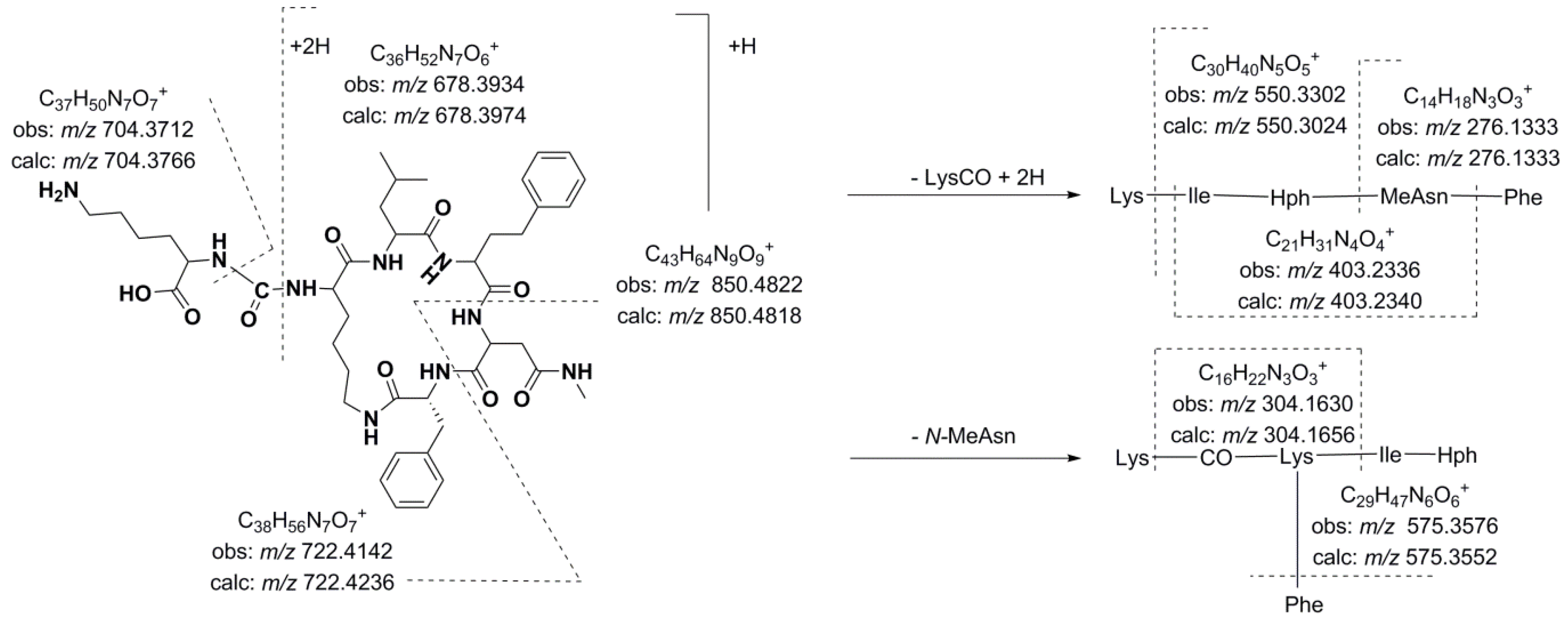
Figure 3.
Predicted fragmentation pattern for compound 12.
Figure 3.
Predicted fragmentation pattern for compound 12.
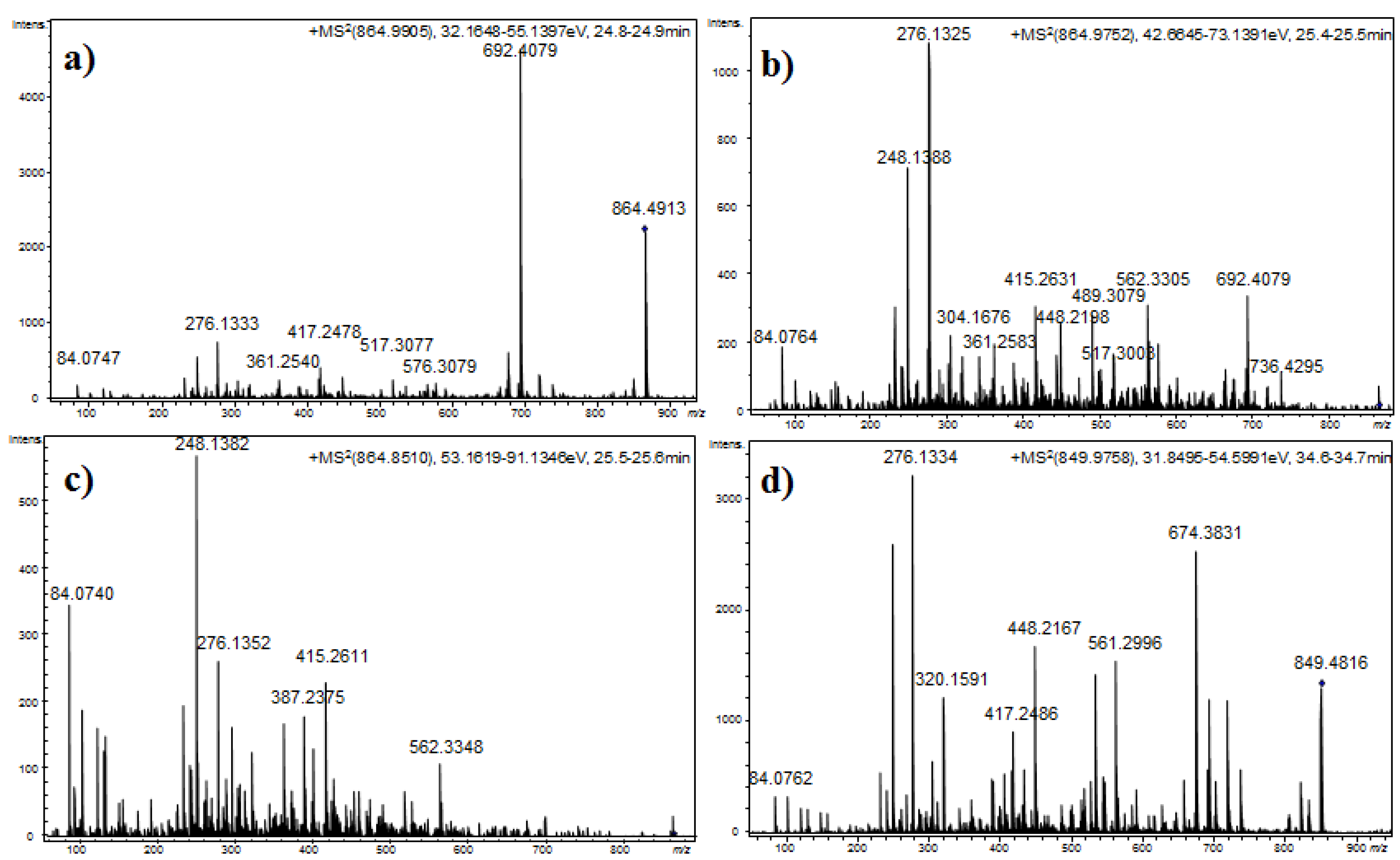
Figure 4.
Product ion spectra for [M + H]+ of compound 14 at (a) CE: 35 eV; (b) CE: 50 eV; and (c) CE: 70 eV; and (d) compound 21 at CE 35 eV.
Figure 4.
Product ion spectra for [M + H]+ of compound 14 at (a) CE: 35 eV; (b) CE: 50 eV; and (c) CE: 70 eV; and (d) compound 21 at CE 35 eV.
Table 6.
Product ion spectra data for compounds 12, 14, and 16.
Table 6.
Product ion spectra data for compounds 12, 14, and 16.
| Product Ion Assignment a | 12 (m/z) | 14 (m/z) | 16 (m/z) |
|---|
| Lys fragment | 84.0768 | 84.0748 | 84.0760 |
| MeAsn immonium ion | 101.0702 | 101.0670 | 101.0649 |
| Phe immonium ion | 120.0738 | 120.0780 | 120.0793 |
| Z immonium ion | - b | 148.1125 | 162.1259 |
| MeAsn-Phe − CO + H | 248.1406 | 248.1400 | 248.1355 |
| MeAsn-Phe + H | 276.1333 | 276.1350 | 276.1320 |
| CO-Lys-Phe + H | 304.1630 | 304.1642 | - |
| Z-MeAsn + H | - | - | 318.1750 |
| Phe-Lys-Ile + H | - | 387.2375 | 387.2269 |
| Lys-Ile-Z + H | - | - | 429.2842 |
| Z-Ile-Lys + H | 401.2661 | 415.2674 | 429.2808 |
| Ile-Z-MeAsn + H | 403.2336 | 417.2517 | 431.2620 |
| Phe-Lys-Ile-Z + H | - | 562.3374 | 576.3330 |
| Ile-Z-MeAsn-Phe + H | 550.3302 | 564.3783 | 578.3556 |
| Lys-CO-Lys-(Ile-Z) + H | 575.3576 | 589.3531 | 603.3697 |
| Lys-Ile-Z-MeAsn-Phe − NH3 + 2H | 661.3682 | 675.3851 | 689.4008 |
| Lys-Ile-Z-MeAsn-Phe + 2H | 678.3934 | 692.4075 | 706.4232 |
| CO-[Lys-Ile-Z-MeAsn-Phe] + H | 704.3712 | 718.3935 | 732.4006 |
| Lys-CO-Lys-(Phe)-(Ile-Z) + H | 722.4142 | 736.4312 | 750.4436 |
| Lys-CO-[Lys-Ile-Z-MeAsn-Phe] + H | 850.4818 | 864.4972 | 878.5122 |
Table 7.
Product ion spectra data for compounds 13, 15, and 17.
Table 7.
Product ion spectra data for compounds 13, 15, and 17.
| Product Ion Assignment a | 13 (m/z) | 15 (m/z) | 17 (m/z) |
|---|
| Lys related fragment | 70.0544 | 70.0619 | 70.0552 |
| Lys fragment | 84.0706 | 84.0731 | 84.0350 |
| MeAsn immonium ion | 101.0617 | 101.068 | - |
| Phe immonium ion | 120.0679 | 120.0748 | - |
| Arg + H | 175.1205 | 175.1184 | 175.1171 |
| CO-Arg | 201.0964 | 201.0966 | 201.0970 |
| MeAsn-Phe − CO + H | -b | 248.1435 | 248.1286 |
| MeAsn-Phe + H | - | 276.1298 | 276.1337 |
| Phe-Lys-Ile + H | - | 387.2358 | 387.2356 |
| Ile-Z-MeAsn + H | 403.7866 | 417.2410 | - |
| Ile-Z-MeAsn-Phe + 2H | - | 564.3384 | - |
| Lys-Ile-Z-MeAsn-Phe − NH3 + 2H | - | - | 689.3691 |
| Lys-Ile-Z-MeAsn-Phe + 2H | 678.4045 | 692.4084 | 706.4222 |
| Arg-CO-Lys-(Phe)-(Ile-Z) + H | - | - | 778.4472 |
| Arg-CO-[Lys-Ile-Z-MeAsn-Phe] + H | 878.4864 | 892.5023 | 906.5196 |
Table 8.
Product ion spectra data for compounds 22 and 26.
Table 8.
Product ion spectra data for compounds 22 and 26.
| Product Ion Assignment a | 22 (m/z) | 26 (m/z) |
|---|
| Lys fragment | 84.0753 | 84.0768 |
| Leu immonium ion | 86.0924 | 86.0924 |
| MeAsn immonium ion | 101.0695 | 101.0677 |
| Phe immonium ion | 120.0810 | 120.0724 |
| Z immonium ion | 148.1124 | 162.1273 |
| MeAsn-Phe − CO + H | 248.1381 | 248.1369 |
| MeAsn-Phe + H | 276.1335 | 276.1326 |
| CO-Lys-Phe + H | 304.1667 | - b |
| Phe-Lys-Ile | 387.2406 | 387.2370 |
| Ile-Z-MeAsn + H | 417.2486 | 431.2661 |
| Z-Ile-Lys + H | 415.2558 | - |
| Ile-Z-MeAsn-Phe + 2H | 564.3281 | - |
| Leu-CO-Lys-Phe | 433.2435 | - |
| CO-Lys-Phe-NMeAsn | 431.1872 | - |
| Leu-CO-Lys-NMeAsn-Phe − CO + H | 533.2997 | 533.2996 |
| Leu-CO-Lys-NMeAsn-Phe | 561.2995 | 561.2967 |
| Leu-CO-Lys-(Ile)-(NMeAsn-Phe) − NH3+ H | - | 657.3648 |
| Leu-CO-Lys-(Ile)-(NMeAsn-Phe) | 674.3823 | 674.3810 |
| Leu-CO-Lys-Z-NMeAsn-Phe | 736.3966 | 750.4096 |
| [Lys-Ile-Z-MeAsn-Phe] + 2H | 692.4077 | 706.4201 |
| Leu-CO[Lys-Ile-Z-NMeAsn-Phe] | 849.4872 | 863.5028 |
Table 9.
Product ion spectra data for compounds 25 and 27.
Table 9.
Product ion spectra data for compounds 25 and 27.
| Product Ion Assignment a | 25 (m/z) | 27 (m/z) |
|---|
| Lys fragment | 84.0714 | 84.0714 |
| MeAsn immonium ion | 101.0633 | 101.057 |
| Phe immonium ion | 120.0701 | 120.077 |
| Z immonium ion | 148.1074 | 162.1336 |
| MeAsn-Phe − CO + H | 248.1425 | 248.1362 |
| MeAsn-Phe + H | 276.1338 | 276.1333 |
| CO-Lys-Phe + H | 304.1570 | - b |
| Phe-CO-Lys + H | 320.1601 | 320.1534 |
| Phe-Lys-Ile + H | - | 387.2388 |
| Ile-Z-MeAsn + H | 417.2582 | - |
| Z-Ile-Lys + H | 415.2709 | - |
| Phe-CO-Lys-Phe + H | 467.2262 | - |
| Phe-CO-Lys-Phe-MeAsn + H | 595.2844 | 595.2842 |
| Phe-CO-Lys-Phe-MeAsn-Z + H | 770.3866 | - |
| Phe-CO-Lys-(Ile-Z)-(MeAsn) + H | 736.3827 | - |
| Phe-CO-Lys-Ile-Z-MeAsn + H | 736.3827 | - |
| Lys-Ile-Z-MeAsn-Phe | - | 706.4299 |
| Lys-Ile-Z-MeAsn-Phe + 2H | 708.3705 | 708.3720 |
| CO-[Lys-Ile-Z-MeAsn-Phe] + H | 718.3895 | 732.4082 |
| Phe-CO-(Lys-Ile-Z)-(Phe) + H | 755.3925 | 767.3730 |
| Phe-CO-[Lys-Ile-Z-MeAsn-Phe] + H | 883.4695 | 897.4858 |
Thus, compound
12 (
m/z 850.4818 [M + H]
+) was characterized as Lys-CO[Lys-Ile-Hph-MeAsn-Phe]. As expected, the presence of Lys in the side chain generated a simple fragmentation pattern. Consequently, just two low intensity series of fragment ions that were assigned to two preferential primary cleavages were observed in the spectrum (
Figure 4). However, enough information was generated to allow a tentative interpretation of the spectrum. The preferential fragmentation observed was attributed to the cleavage of the ureido linkage and the opening of the ring and indicated the presence of lysine in the side chain (
m/z 678.3934). Further amino acid residue losses from this lineal ion (
i.e., acylium or similar) yielded a series of ions at
m/z 550.3302, 403.2336, 304.1630, and 276.1333 (loss of Lys, Phe, Ile/Leu, and Hph, respectively). Another series of less abundant ions that were generated by the loss of
N-methyl asparagine and phenylalanine was also observed at
m/z 722.4142 (loss of MeAsn) and 575.3576 (loss of Phe). The dipeptide fragments at
m/z 304.1630 and 276.1333 determined the partial sequence Hph-MeAsn-Phe and established the isoleucine/leucine at position 3 (
m/z 403.2336) and the lysine that closed the ring (
m/z 678.3934). Ions that are typically observed in the low
m/z region were also present (Lys at
m/z 84.0768, MeAsn at
m/z 101.0702, Phe at
m/z 120.0738).
Compounds 14 and 16, which exhibited molecular mass increases of 14 and 28 Da, respectively, when compared to the compound at m/z 850.4818 [M + H]+, exhibited similar product ion spectra. Differences that were observed in the fragments containing the amino acid at position 4 suggested that this residue was either methylated (m/z 417.2517, 564.3783, 692.4075, and 736.4312) or ethylated (m/z 431.2620, 576.3330, 689.4008, and 750.4436). The existence of ions at m/z 148.1125 and 162.1213 that were attributed to the N-methylhomophenylalanine (N-MeHph) and N-ethylhomophenylalanine (N-EtHph) immonium ions, respectively, also supported these assignments.
Using similar reasoning, the remaining anabaenopeptins were subsequently characterized. Losses of 157 u that were observed for compounds
22 and
26, losses of 191 u that were observed for compounds
25 and
27, and losses of 200 u that were observed for compounds
13,
15, and
17 were used to identify the amino acid side chain as leucine, phenylalanine, or arginine, respectively. A fragmentation pattern similar to that described in the preceding paragraphs characterized the cyclic structures. As mentioned above, compounds with [M + H]
+ at
m/z 878.5122, 892.5023, and 906.5196 exhibited low efficiency fragmentations, which are characteristic of oligopeptides containing strongly basic residues. When the collision energy used for fragmentation was increased to improve efficiency, the abundance of the fragment ions was compromised (
Figure 4).
Four additional anabaenopeptins (
18,
19,
23,
24) were found in
Nostoc sp. CENA352. Accurate mass measurement, isotopic profiles, and MS/MS spectra were conclusive to distinguish these compounds from previously described anabaenopeptins and characterize these compounds as new variants [
36,
53]. Structurally, all of these anabaenopeptins incorporated phenylalanine in the side chain position, as evidenced by the loss of a 191-u fragment from the protonated molecules (
Table 10). Positions 5 and 6 of these four peptides were also conserved and were occupied by
N-methyl-alanine and homotyrosine respectively. For the homovariant amino acids in position 4, either homophenylalanine (
18,
19) or homotyrosine (
23,
24) was proposed; for position 3, the commonly reported valine (
18 and
23) or leucine/isoleucine (
19 and
24) was proposed. For example, diagnostic low mass fragment ions that were observed in the product ion spectrum of the compound at
m/z 842.4416 [M + H]
+ (
23) provided information about the amino acid residues, indicating the presence of lysine (
m/z 84.0744),
N-methyl-alanine (
m/z 114.0515), phenylalanine (
m/z 120.0736), homophenylalanine (
m/z 134.0921), and homotyrosine (150.0887), while higher product ions suggested their sequence. Fragment ions at
m/z 247.1440 and 263.1399 indicated the attachment of
N-methyl-alanine to both homotyrosine and homophenylalanine, and subsequently established the sequence Val-Hph-MeAla for
m/z 346.2124 and Lys-Val-Hph for
m/z 387.2384. Taking into account the reported anabaenopeptins structures, the Phe-CO-[Lys-Val-Hph-MeAla-Hty] sequence was reasonably postulated. The intense fragment ion generated by the loss of a homophenylalanine residue at
m/z 681.3591 and the less intense series of b-ions (
m/z 596.3041, 497.2381, 320.1602, and 419.2283) were thus comprehensively attributed (
Table 10).
Table 10.
Product ion spectra data for compounds 18, 19, 23, and 24.
Table 10.
Product ion spectra data for compounds 18, 19, 23, and 24.
| Product Ion Assignment a | 18 (m/z) | 19 (m/z) | 23 (m/z) | 24 (m/z) |
|---|
| Lys fragment | 84.0734 | 84.0757 | 84.0744 | 84.0749 |
| MeAla + H − CO | 114.0499 | 114.0557 | 114.0515 | 114.0514 |
| Phe immonium ion | - b | 120.0736 | 120.0736 | 120.0742 |
| Hph immonium | - | - | 134.0921 | 134.0929 |
| Htyr immonium | 150.0907 | 150.0907 | 150.0887 | 150.0873 |
| Hph-MeAla +H | - | - | 247.1440 | 247.1485 |
| HTyr-MeAla + H | 263.1381 | 263.1394 | 263.1399 | 263.1391 |
| Y-Z-MeAla + H | 362.2138 | 376.2236 | 346.2124 | 360.2270 |
| Lys-Y-Z + H | 403.2366 | 417.2447 | 387.2384 | 401.2544 |
| MeAla-Hty-Lys-Y + H | 490.2942 | - | 490.3023 | 514.3045 |
| Phe-CO-Lys-Y + H | - | - | 419.2283 | - |
| Phe-CO-Lys + 2H | - | - | 320.1602 | 320.1605 |
| Phe-CO-Lys-Hty + 2H | 497.2336 | 497.2298 | 497.2381 | 497.2387 |
| Phe-CO-Lys-Y-Z + H | - | 610.3218 | 596.3041 | 610.3266 |
| Phe-CO-Lys-Hty-MeAla + 2H | 582.2993 | 582.2962 | 582.2918 | 582.2915 |
| Lys-Y-Z-MeAla-Hty | 665.3655 | 679.3793 | 649.3720 | 663.3819 |
| Lys-Y-Z-MeAla-Hty + 2H | 667.3786 | 681.3953 | 651.3873 | 665.4013 |
| Phe-CO-Lys-(Y)-(Hty-MeAla) + 2H | 681.3604 | 695.3756 | 681.3597 | 695.3756 |
| CO-[Lys-Y-Z-MeAla-Hty] + H | 693.3629 | - | - | - |
| Phe-CO-[Lys-Y-Z-MeAla-Hty] + H − CO | 830.4384 | 844.4575 | 814.4502 | 828.4634 |
| Phe-CO-[Lys-Y-Z-MeAla-Hty] + H − H2O | 840.4385 | 854.4465 | 824.4377 | 838.4398 |
| Phe-CO-[Lys-Y-Z-MeAla-Hty] + H | 858.4405 | 872.4544 | 842.4416 | 856.4561 |
In addition, a pair of unusual tryptophan-containing anabaenopeptins (
20–
21) was also detected at
m/z 803.4417 [M + H]
+ and 803.4425 [M + H]
+, exclusively in the genus
Brasilonema sp. (CENA360 and CENA382). As this pair of compounds exhibited similar protonated molecules and product ion spectra but differed in retention times, these compounds were classified as diastereoisomers. The structures of these compounds were postulated in accordance with the fragment ions listed in
Table 11. According to our literature search, prior to our work, the occurrence of tryptophan-containing anabaenopeptins in cyanobacteria was limited to the genus
Tychonema sp. [
64]. The structurally related compounds isolated from this genus, the brunsvicamides A–C, inhibit tyrosine phosphatase B of
Mycobacterium tuberculosis (MptpB) [
64] and are highly selectivity inhibitors for human leukocyte elastase (HLE) [
65]. However, those peptides and the compounds reported here differ in their amino acid sequences. Unlike the brunsvicamides, the postulated anabaenopeptins contain tryptophan in position 4 and
N-methyl-alanine in the methylated amino acid position. The structures of the compounds described in this study are more closely related to a synthetic brunsvicamide analog described by Walther
et al. [
66], which was found to be an inhibitor of carboxypeptidase A. Thus, further biological tests of these compounds are warranted.
Table 11.
Product ion spectra data for compounds 20 and 21.
Table 11.
Product ion spectra data for compounds 20 and 21.
| Product Ion Assignment | 20 (m/z) | 21 (m/z) |
|---|
| Lys fragment | 84.0730 | 84.0721 |
| MeAla + H − CO | 114.0519 | 114.0504 |
| Phe immonium ion | 120.0833 | 120.0747 |
| Trp fragment | 130.0645 | 130.0607 |
| Lys-Ile + H | 240.1738 | 240.1752 |
| Trp-MeAla + H | 272.1384 | 272.1429 |
| Ile-Trp-MeAla + H | 385.2229 | 385.2238 |
| CO-Lys-Phe + H | 320.1599 | 320.1602 |
| Val-CO-Lys-Ile + H | 385.2229 | 385.2211 |
| CO-Lys-Phe-MeAla + H | 405.2128 | 405.2084 |
| Val-CO-Lys-Ile + H | 419.2214 | 419.2240 |
| Val-CO-Lys-Phe-MeAla + H | 504.2829 | 504.2806 |
| Val-CO-Lys-Phe-Ile + H | 532.3069 | 532.3204 |
| Val-CO-Lys-(Ile)-(Phe-MeAla) + H | 617.3706 | 617.3686 |
| Lys-Ile-Trp-MeAla-Phe + H | 658.3680 | 658.3493 |
| Lys-Ile-Trp-MeAla-Phe + 2H | 660.3879 | 660.3833 |
| CO-Lys-Ile-Trp-MeAla-Phe + H | 686.3690 | 686.3681 |
| Val-CO-Lys-(Ile-Trp)-(Phe) + H–CO2 | 674.3749 | 674.3970 |
| Val-CO-[Lys-Ile-Trp-MeAla-Phe] + H − CO2 | 759.4538 | 759.4521 |
| Val-CO-[Lys-Ile-Trp-MeAla-Phe] + H − CO | 775.4563 | 775.4595 |
| Val-CO-[Lys-Ile-Trp-MeAla-Phe] + H − H2O | 785.4408 | 785.4278 |
| Val-CO-[Lys-Ile-Trp-MeAla-Phe] + H | 803.4417 | 803.4425 |
2.3. Cyanopeptolins
Eleven new cyanopeptolins with protonated molecules at
m/z 984.5381 (
29), 984.5398 (
33), 998.5531 (
31), 998.5549 (
35), 998.5564 (
36), 1002.5031 (
28), 1004.4831 (
30), 1012.5737 (
37), 1018.5011 (
32), 1026.5858 (
38), and 1032.5170 (
34) were present in the extracts from
Desmonostoc strains CENA371 and CENA386. Although no single diagnostic fragment ion can be used to identify this family of peptides, series of fragments related to the conserved position 4 (3-amino-6-hydroxy-2-piperidone amino acid (Ahp) could be used to identify these compounds [
23]. Under our experimental conditions, doubly charged ions ([M − H
2O + 2H]
2+) with an abundance comparable to that of protonated molecules were also observed in the mass spectra of all detected cyanopeptolins. MS and MS/MS analyses suggested closely related structures for all of these cyanopeptides (
Table 12,
Table 13 and
Table 14). As the most common structural feature, these cyanopeptolins contained
N-acetyl-proline-glutamine as side chain, with the fifth position occupied by either a dimethylated tyrosine or chlorinated-methylated tyrosine. Valine and leucine alternated in positions 4 and 6.
Table 12.
Product ion spectra data for compounds 30, 32, 34, and 37.
Table 12.
Product ion spectra data for compounds 30, 32, 34, and 37.
| Product Ion Assignment a | 30 (m/z) | 32 (m/z) | 34 (m/z) | 37 (m/z) |
|---|
| Pro immonium | 70.0542 | 70.0595 | 70.0571 | 70.0541 |
| Val immonium | -b | 72.0692 | - | - |
| Leu immonium | - | - | 86.0900 | 86.0935 |
| NAcPro immonium | 112.0695 | 112.0732 | 112.0679 | 112.0679 |
| NAcPro + H | 140.0623 | 140.0773 | 140.0678 | 140.0706 |
| NMe-R1-Tyr immonium | 184.0572 | 184.0486 | 184.0523 | 164.1052 |
| Leu-Ahp − H2O − CO + H | - | 181.1263 | 181.1290 | 181.1300 |
| Leu-Ahp − H2O + H | 209.1239 | - | 209.1257 | 209.1286 |
| Ac-Pro-Gln + H | 268.1207 | 268.1276 | 268.1270 | 268.1271 |
| Ac-Pro-Gln-Thr − H2O + H | 351.1656 | 351.1645 | 351.1638 | 351.1633 |
| NMe-R1-Tyr-X-Ahp − H2O + H | 406.1509 | 406.1515 | 420.1648 | 400.2181 |
| AcPro-Gln-Thr-Leu − H2O + H | 464.2465 | 464.2447 | 464.2467 | 464.2468 |
| AcPro-Gln-Thr-Leu-Ahp-X − 2H2O + H | - | - | - | 672.3922 |
| AcPro-Gln-Thr-Leu-Ahp-X − H2O + H | - | - | - | 690.3777 |
| Thr-Leu-Ahp-X-NMe-R1-Tyr-Z − H2O + 2H | 727.0780 | 733.3705 | 747.3811 | 727.4318 |
| AcPro-Gln-Thr-(Z-NMe-R1-Tyr)-(Leu) + H | - | - | 806.3871 | 786.4305 |
| AcPro-Gln-Thr-Leu-Ahp-Z-NMe-R1Tyr − H2O + H | - | - | 901.4207 | 881.4576 |
| AcPro-Gln-[Thr-Leu-Ahp-X-NMe-R1-Tyr-Z] − H2O +H | 986.4693 | 1000.4815 | 1014.5018 | 994.5553 |
| AcPro-Gln-[Thr-Leu-Ahp-X-NMe-R1-Tyr-Z] + H | 1004.4831 | 1018.5011 | 1032.5170 | 1012.5737 |
Table 13.
Product ion spectra data for compounds 29, 31, 33, and 35–36.
Table 13.
Product ion spectra data for compounds 29, 31, 33, and 35–36.
| Product Ion Assignment a | 29 (m/z) | 31 (m/z) | 33 (m/z) | 35 (m/z) | 36 (m/z) |
|---|
| Pro immonium | -b | - | - | - | 70.0563 |
| Val immonium | - | - | - | - | - |
| Leu immonium | 86.0924 | - | - | - | 86.0876 |
| NAcPro immonium | 112.0755 | 112.0755 | 112.0755 | 112.0688 | 112.0679 |
| NAcPro + H | 140.0618 | 140.0693 | 140.0693 | 140.0693 | 140.0696 |
| NMe-R1-Tyr immonium | 164.1024 | 150.0907 | 164.1024 | 164.1024 | 164.1052 |
| Leu-Ahp − H2O − CO + H | 181.1353 | 181.1267 | - | - | 181.1283 |
| Leu-Ahp − H2O + H | 209.1243 | 209.1335 | - | 209.1244 | 209.1331 |
| Ac-Pro-Gln + H | 268.1258 | 268.1258 | - | 268.1258 | 268.1207 |
| Ac-Pro-Gln-Thr − H2O + H | 351.1645 | 351.1637 | 351.1638 | 351.1658 | 351.1642 |
| NMe-R1-Tyr-X-Ahp − H2O + H | 386.2025 | 386.2008 | 386.2064 | 386.2034 | 400.2250 |
| AcPro-Gln-Thr-Leu − H2O + H | 464.2439 | 464.2464 | 464.2493 | 464.2495 | 464.2549 |
| AcPro-Gln-Thr-Leu-Ahp-X − 2H2O + H | - | - | - | 658.3446 | - |
| AcPro-Gln-Thr-Leu-Ahp-X − H2O + H | - | - | - | - | - |
| Thr-Leu-Ahp-X-NMe-R1-Tyr-Z − H2O + 2H | 699.4064 | 713.4158 | 699.4065 | 713.4195 | 713.3971 |
| AcPro-Gln-Thr-(Z-NMe-R1-Tyr)-(Leu) + H | - | 772.3942 | - | 786.4170 | - |
| AcPro-Gln-Thr-Leu-Ahp-X-NMe-R1Tyr − H2O + H | - | - | - | 867.4460 | 867.4647 |
| AcPro-Gln-[Thr-Leu-Ahp-X-NMe-R1-Tyr-Z] − H2O +H | 966.5061 | 980.5489 | 966.5144 | 980.5472 | 980.5382 |
| Ac-Pro-Gln-[Thr-Leu-Ahp-X-NMe-R1-Tyr-Z] + H | 984.5381 | 998.5531 | 984.5398 | 998.5550 | 998.5564 |
Table 14.
Product ion spectra data for compound 28 and 38.
Table 14.
Product ion spectra data for compound 28 and 38.
| Product Ion Assignment a | 28 (m/z) | 38 (m/z) |
|---|
| Pro immonium | - b | - |
| Leu immonium | 86.08763 | - |
| Y immonium | - | 126.0845 |
| Y + H | - | 154.0743 |
| NMe-R1-Tyr immonium | 184.0486 | 164.1060 |
| Leu-Ahp − H2O − CO + H | - | - |
| Leu-Ahp − H2O + H | 209.1239 | - |
| Y-Gln + H | 268.1207 | 288.1510 |
| Ac-Pro-Gln-Thr − H2O + H | 321.1521 | 365.1662 |
| NMe-R1-Tyr-X-Ahp − H2O + H | 420.1585 | 400.2211 |
| Y-Gln-Thr-Leu − H2O + H | 434.2416 | 478.2652 |
| Y-Gln-Thr-Leu-Ahp-X − 2H2O + H | - | 686.5626 |
| Y-Gln-Thr-Leu-Ahp-X − H2O + H | - | - |
| Thr-Leu-Ahp-X-NMe-R1-Tyr-Z − H2O + 2H | - | 727.4402 |
| Y-Gln-Thr-(Z-NMe-R1-Tyr)-(Leu) + H | - | 800.4147 |
| Y-Gln-Thr-Leu-Ahp-X-NMe-R1Tyr − H2O + H | - | - |
| Y-Gln-[Thr-Leu-Ahp-X-NMe-R1-Tyr-Z] − H2O +H | 984.4914 | 1008.5481 |
| Y-Gln-[Thr-Leu-Ahp-X-NMe-R1-Tyr-Z] + H | 1002.5031 | 1026.5858 |
This family of cyclic peptides with high structural variability featured a ring formed by six amino acids and a side chain of different lengths and composition [
3]. An ester bond between the hydroxyl group of the threonine in position one and the carboxyl group of the terminal amino acid cyclizes the ring. The threonine amino acid in position 1 is occasionally replaced by 3-hydroxy-4-methylproline [
67,
68]. The 3-amino-6-hydroxy-2-piperidone amino acid (Ahp) always occupied position 3, while a methylated aromatic amino acid and other neutral amino acids are found in positions 5 and 6, respectively. The highly variable side chain may contain an aliphatic fatty acid or a glyceric acid, which is attached either directly to the threonine in position 1 or through one or two amino acids [
3]. This family of peptides is often described as protease inhibitors [
69,
70,
71,
72,
73]. Cyanopeptolins have been isolated mostly from
Microcystis [
71,
74,
75] but also from other genera such as
Lyngbya [
76,
77],
Nostoc [
67,
78],
Oscillatoria [
79,
80],
Planktothrix [
60,
81],
Scytonema [
82], or
Symploca [
72].
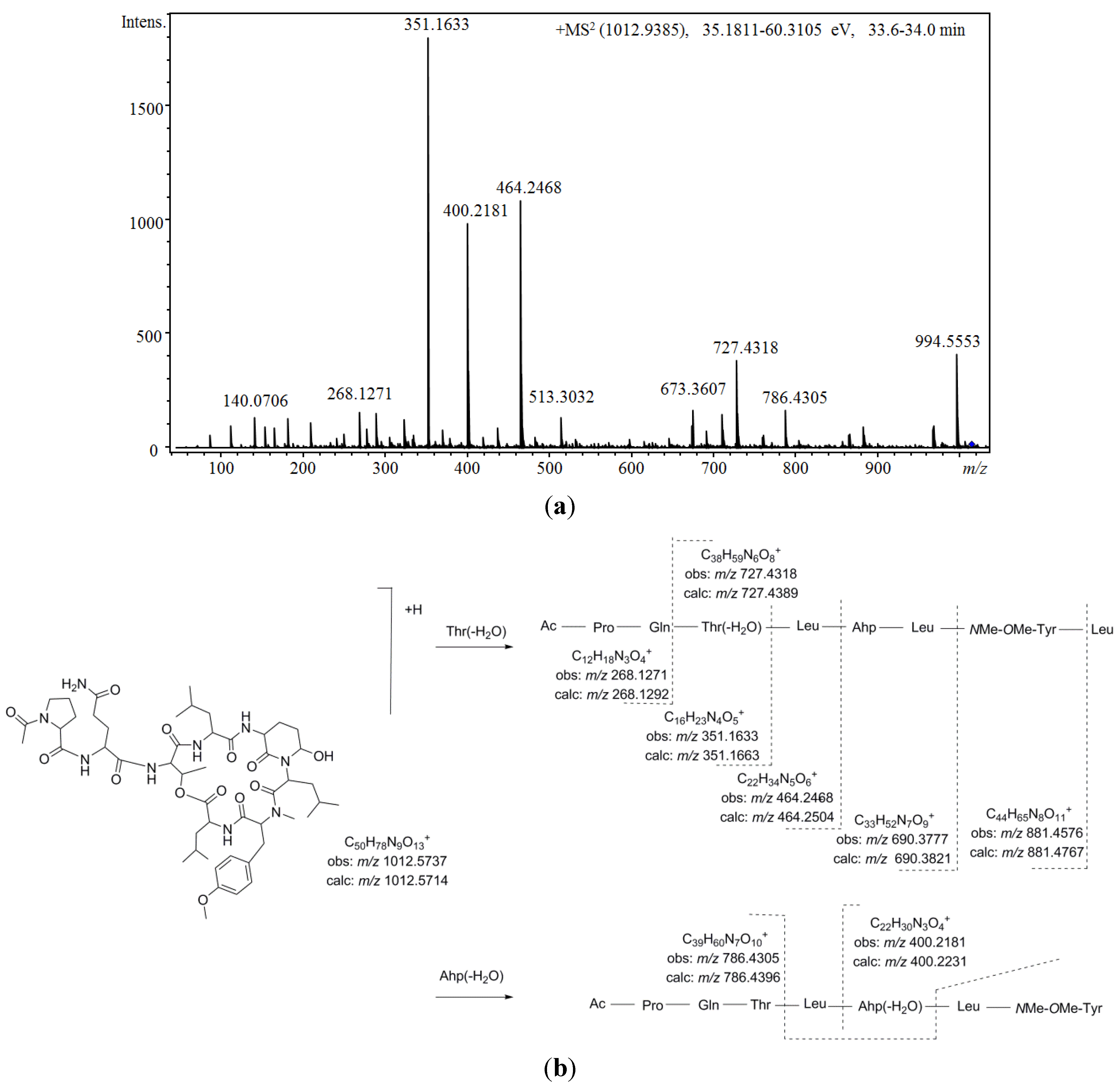
Figure 5 shows the proposed depsipeptide cyclic structure for compound
37 (Ac-Pro-Gln[Thr-Leu-Ahp-Leu-
NMe-OMe-Tyr-Leu]) and its predicted fragmentation pattern. The most abundant ions observed in the product ion spectrum of this compound were attributed to the loss of amino acid residues from the
C-terminus (
m/z 881.4576, 690.3777, 464.2468, 351.1633, and 268.1271) of the dehydrated protonated molecule at
m/z 994.5553. This precursor ion was suggested to be generated by the cleavage of the ester linkage accompanied by the dehydration of the Thr [
34]. Further loss of the acetyl-proline-glutamine side chain from this linear ion was also noted (
m/z 727.4318). Simultaneously, evidence of dehydration at the Ahp position was observed at
m/z 400.2184 and 786.4305. An inspection of the low
m/z region of the spectrum also supported the proposed structure, revealing the presence of ions associated with the amino acids Ahp (
m/z 209.1286 and 181.1300),
NMe
OMeTyr (
m/z 164.1052), AcPro (
m/z 112.0679 and 140.0703), Leu (
m/z 86.0979), and Pro (
m/z 70.0563). To our knowledge, the
N-acetyl-proline-glutamine side chain has not been
Figure 5.
(a) Product ion spectrum for [M + H]+ of compound 37 and (b) its predicted fragmentation pattern. Conditions as described in the experimental section.
Figure 5.
(a) Product ion spectrum for [M + H]+ of compound 37 and (b) its predicted fragmentation pattern. Conditions as described in the experimental section.
Comparing the aforementioned data with those obtained for compound
36, a highly similar structure was deduced. Differences of 14 u in the protonated molecules and a fragment ion at
m/z 713.3971 in combination with the existence of product ions at
m/z 464.2549 and 400.2250 indicated that the leucine attributed to position 6 in compound
37 was replaced by valine. In addition, two other isobaric compounds were observed at earlier retention times (
31 at 30.0 min, and
35 at 31.1 min). Structural differences from compound
36 were proposed based on mass data (
Table 13). Thus, monomethylated tyrosine was suggested for compound
31 (
m/z 772.3942, 713.3942, and 150.0907) and a valine residue occupying the fourth position instead of the sixth position was suggested for
35 (
m/z 867.4460, 786.4170, 713.4195, and 164.1024). Furthermore, two other minor compounds (
29 and
33) with molecular masses 14 Da lower than the compounds mentioned above and with similar product ion spectra were also detected. Based on the mass results (
m/z 966.5061/966.5144 and 699.4064/699.4065), the incorporation of valine at positions 4 and 6 was postulated for this pair of diastereoisomers. Simultaneously with these compounds, another congener was observed at
m/z 1026.5858 [M + H]
+ (
38). Propanoylation of the proline residue instead of the more commonly detected acetylation was suggested for this compound on the basis of its product ion spectrum, which differed in product ions containing the side chain (
m/z 365.1662, 478.2652, and 686.5626) in combination with the existence of product ions at
m/z 126.0845 and 154.0743 (attributed to
N-propanoyl-proline).
Similarly, four additional compounds (
28,
30,
32, and
34) exhibited fragmentation patterns highly similar to that of compound
37. While most of the mass spectrum remained conserved, fragment ions containing the original dimethylated tyrosine amino acid exhibited
m/
z shifts when compared to that of the model compound. Additionally, the isotopic pattern of these fragment ions in conjunction with that of the protonated molecule revealed the presence of a chlorine atom in their structures. For compound
34, these fragment ions shifted 20 u (
m/z 901.4207, 806.3871, 747.3811, 420.1648, and 184.0523), leading us to propose the chlorination of a methylated tyrosine at position 5. For compound
32, shifts of 20 and 14 u were observed (
m/z 733.3705, 406.2447, and 184.0486), suggesting that in addition to the modification mentioned above, a substitution of leucine with valine at position 4 also occurred. Based on the same logic, a structure similar to compound
34, just with the leucine residue at position 6 replaced by valine, was proposed for compound
30. Finally, for compound
28, differences of 30 u in product ions containing the side chain were observed in comparison to compound
34 (
m/z 321.1521 and
m/z 434.2416). Thus, the same cyclic peptide was proposed for compound
28 with the side chain tentatively attributed to methylated-dehydroproline-glutamine (Mdhp-Gln) or other analogs. The chlorinated and methylated tyrosine amino acid proposed for position 5 was quite unusual and has only been observed in a small number of cyanopeptolins [
71,
80].
The structural similarity of these compounds to other cyanopeptolins with observed protease inhibitory activity warrants further bioactivity assays. Trypsin inhibitory selectivity was suggested to be related to the existence of basic residues adjacent to Ahp, while chymotrypsin selectivity was proposed to be related to hydrophobic residues. Additionally, residues in other positions as side chains or in the fifth position appear to influence this activity [
5,
72,
83]. These assays will be able to establish the influence of the particular properties of these compounds on the selectivity and potency of these and other activities.