Oleosome-Associated Protein of the Oleaginous Diatom Fistulifera solaris Contains an Endoplasmic Reticulum-Targeting Signal Sequence
Abstract
:1. Introduction
2. Results
2.1. Characterization of Doap1 Gene Structure
2.2. GFP Expression in the Transformants
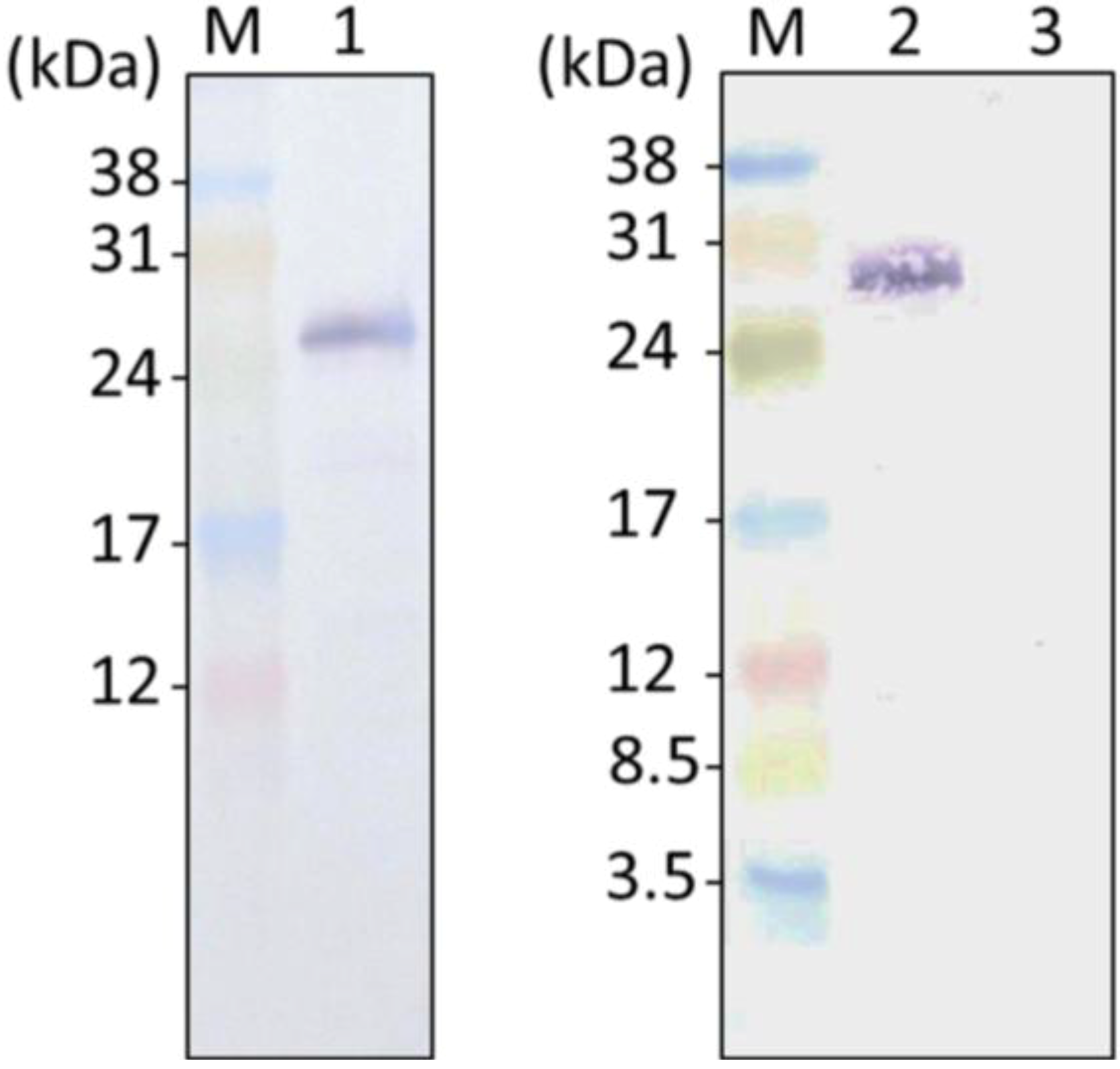

2.3. ER-Targeting of SDOAP1-GFP
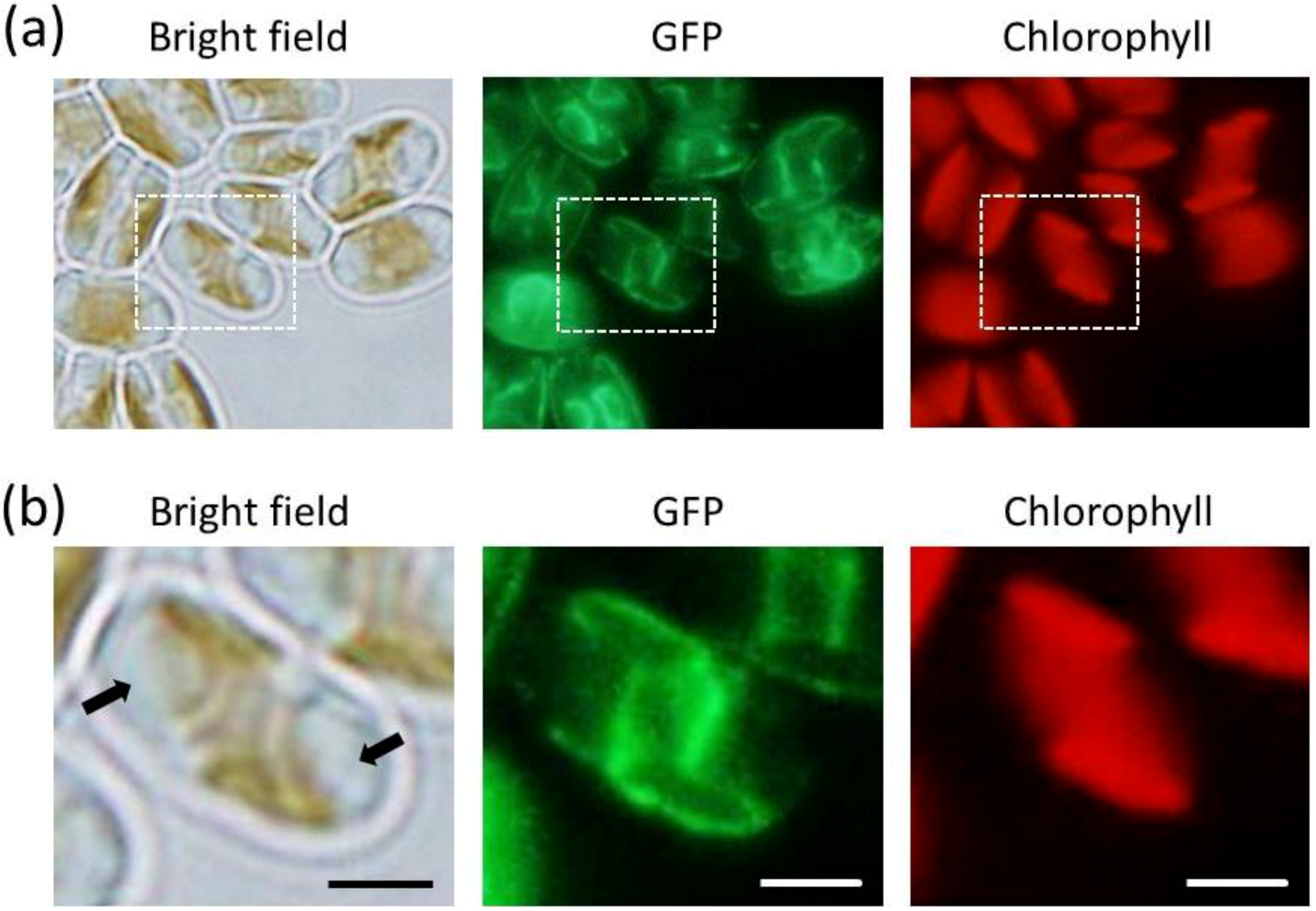
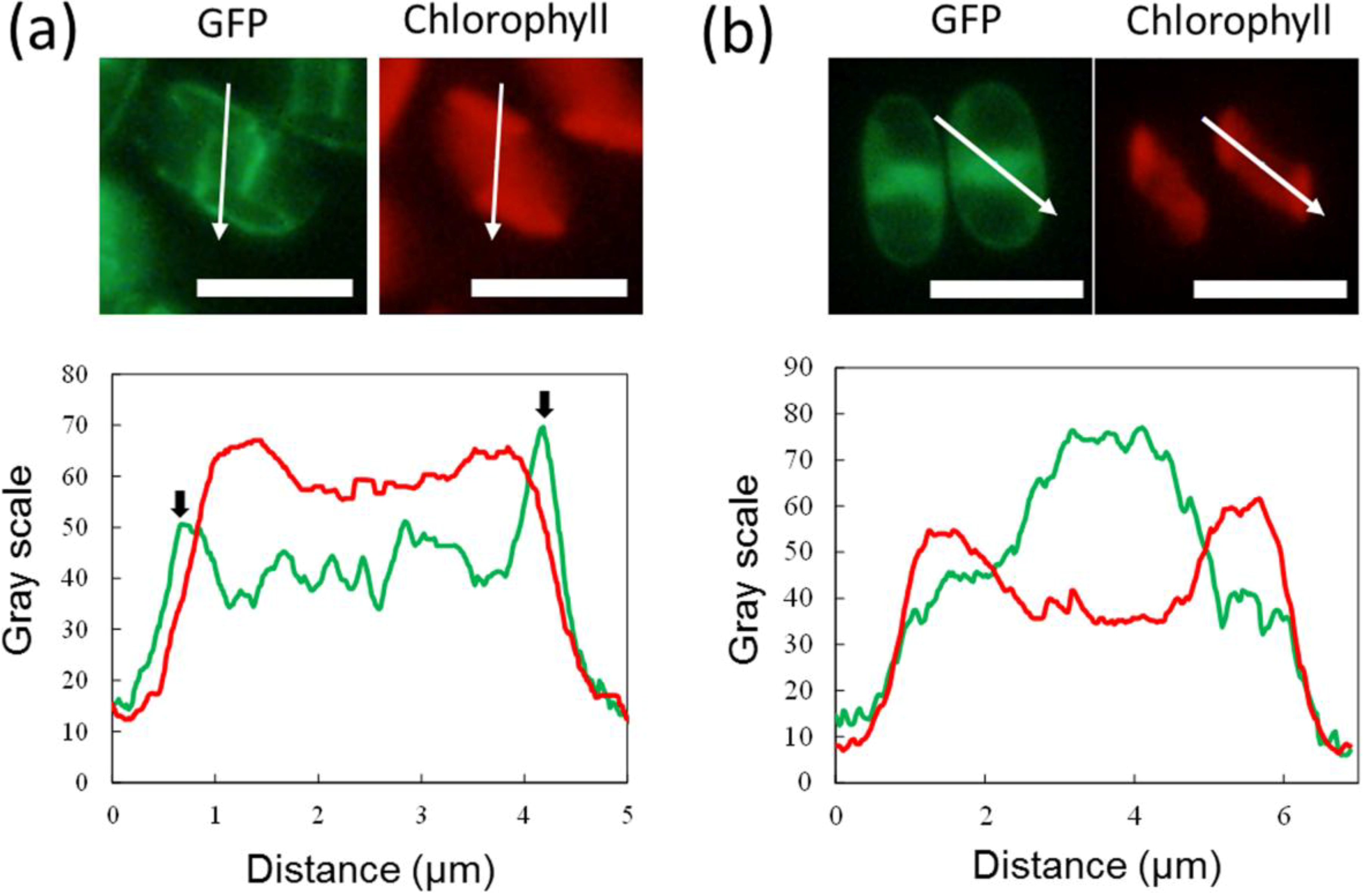
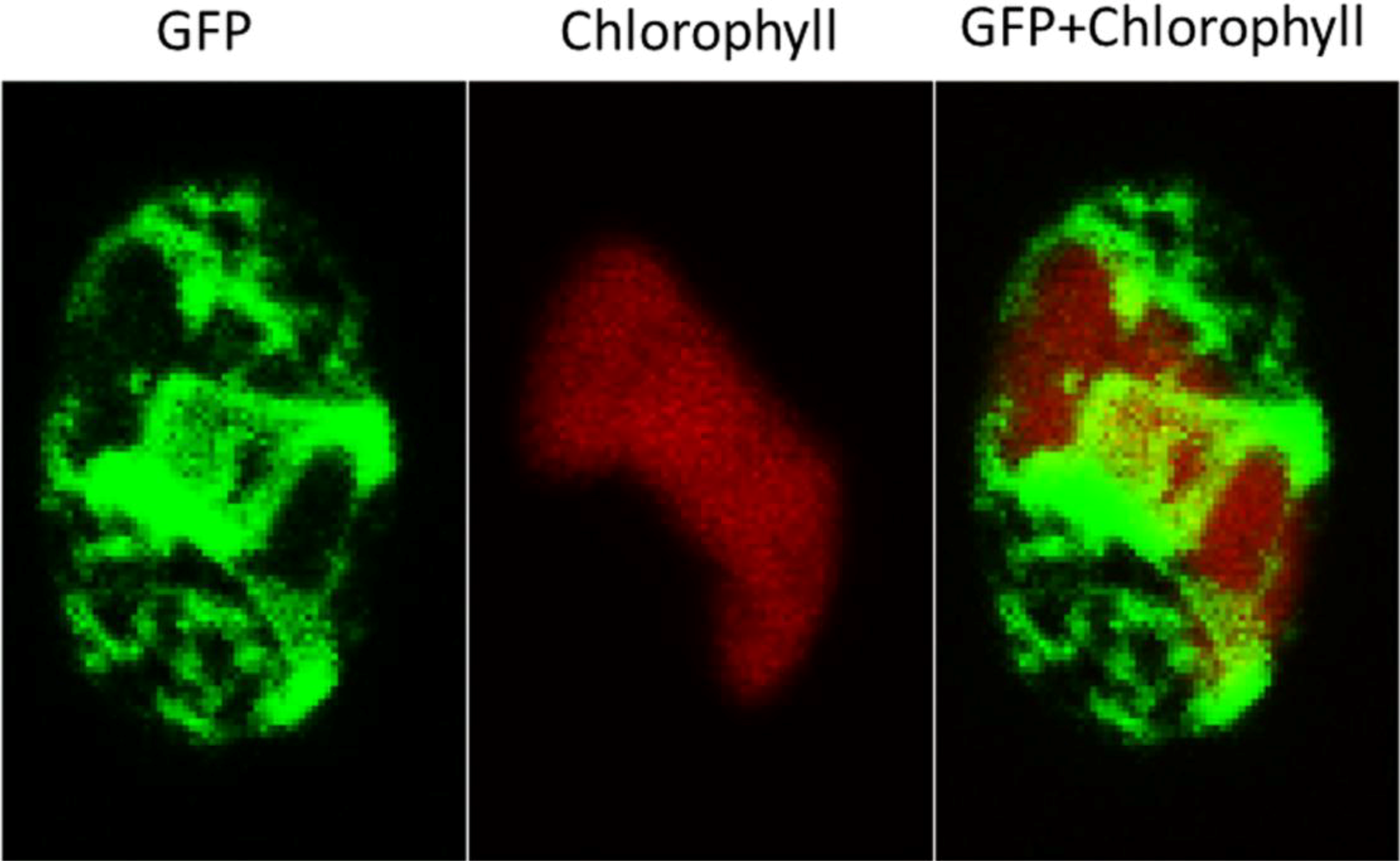
3. Discussion
4. Experimental Section
4.1. Culture Conditions
4.2. Characterization of Nucleotide and Protein Sequences
4.3. Vector Construction and Transformation
4.4. Western Blotting
4.5. Fluorescent Microscopy and Image Analysis
5. Conclusions
Supplementary Files
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Smith, V.H.; Sturm, B.S.; Denoyelles, F.J.; Billings, S.A. The ecology of algal biodiesel production. Trends Ecol. Evol. 2010, 25, 301–309. [Google Scholar]
- Liang, Y.; Sarkany, N.; Cui, Y. Biomass and lipid productivities of Chlorella vulgaris under autotrophic, heterotrophic and mixotrophic growth conditions. Biotechnol. Lett. 2009, 31, 1043–1049. [Google Scholar] [CrossRef]
- Mahapatra, D.M.; Chanakya, H.; Ramachandra, T. Euglena sp. as a suitable source of lipids for potential use as biofuel and sustainable wastewater treatment. J. Appl. Phycol. 2013, 25, 855–865. [Google Scholar] [CrossRef]
- Matsunaga, T.; Matsumoto, M.; Maeda, Y.; Sugiyama, H.; Sato, R.; Tanaka, T. Characterization of marine microalga, Scenedesmus sp. strain JPCC GA0024 toward biofuel production. Biotechnol. Lett. 2009, 31, 1367–1372. [Google Scholar] [CrossRef]
- Oh, S.H.; Han, J.G.; Kim, Y.; Ha, J.H.; Kim, S.S.; Jeong, M.H.; Jeong, H.S.; Kim, N.Y.; Cho, J.S.; Yoon, W.B.; et al. Lipid production in Porphyridium cruentum grown under different culture conditions. J. Biosci. Bioeng. 2009, 108, 429–434. [Google Scholar] [CrossRef]
- Rodolfi, L.; Chini Zittelli, G.; Bassi, N.; Padovani, G.; Biondi, N.; Bonini, G.; Tredici, M.R. Microalgae for oil: Strain selection, induction of lipid synthesis and outdoor mass cultivation in a low-cost photobioreactor. Biotechnol. Bioeng. 2009, 102, 100–112. [Google Scholar] [CrossRef]
- Corteggiani Carpinelli, E.; Telatin, A.; Vitulo, N.; Forcato, C.; D’Angelo, M.; Schiavon, R.; Vezzi, A.; Giacometti, G.M.; Morosinotto, T.; Valle, G. Chromosome scale genome assembly and transcriptome profiling of Nannochloropsis gaditana in nitrogen depletion. Mol. Plant 2014, 7, 323–325. [Google Scholar] [CrossRef]
- Guarnieri, M.T.; Nag, A.; Smolinski, S.L.; Darzins, A.; Seibert, M.; Pienkos, P.T. Examination of triacylglycerol biosynthetic pathways via de novo transcriptomic and proteomic analyses in an unsequenced microalga. PLoS One 2011, 6, e25851. [Google Scholar]
- Radakovits, R.; Jinkerson, R.E.; Fuerstenberg, S.I.; Tae, H.; Settlage, R.E.; Boore, J.L.; Posewitz, M.C. Draft genome sequence and genetic transformation of the oleaginous alga Nannochloropis gaditana. Nat. Commun. 2012, 3, 686. [Google Scholar] [CrossRef]
- Rismani-Yazdi, H.; Haznedaroglu, B.Z.; Hsin, C.; Peccia, J. Transcriptomic analysis of the oleaginous microalga Neochloris oleoabundans reveals metabolic insights into triacylglyceride accumulation. Biotechnol. Biofuels 2012, 5, 74. [Google Scholar] [CrossRef]
- Wang, H.; Alvarez, S.; Hicks, L.M. Comprehensive comparison of iTRAQ and label-free LC-based quantitative proteomics approaches using two Chlamydomonas reinhardtii strains of interest for biofuels engineering. J. Proteome Res. 2012, 11, 487–501. [Google Scholar] [CrossRef]
- Davidi, L.; Katz, A.; Pick, U. Characterization of major lipid droplet proteins from Dunaliella. Planta 2012, 236, 19–33. [Google Scholar] [CrossRef]
- Frandsen, G.I.; Mundy, J.; Tzen, J.T. Oil bodies and their associated proteins, oleosin and caleosin. Physiol. Plant. 2001, 112, 301–307. [Google Scholar] [CrossRef]
- Lin, I.; Jiang, P.-L.; Chen, C.-S.; Tzen, J.T. A unique caleosin serving as the major integral protein in oil bodies isolated from Chlorella sp. cells cultured with limited nitrogen. Plant Physiol. Biochem. 2012, 61, 80–87. [Google Scholar] [CrossRef]
- Moellering, E.R.; Benning, C. RNA interference silencing of a major lipid droplet protein affects lipid droplet size in Chlamydomonas reinhardtii. Eukaryot. Cell 2010, 9, 97–106. [Google Scholar] [CrossRef]
- Nguyen, H.M.; Baudet, M.; Cuiné, S.; Adriano, J.M.; Barthe, D.; Billon, E.; Bruley, C.; Beisson, F.; Peltier, G.; Ferro, M. Proteomic profiling of oil bodies isolated from the unicellular green microalga Chlamydomonas reinhardtii: With focus on proteins involved in lipid metabolism. Proteomics 2011, 11, 4266–4273. [Google Scholar] [CrossRef]
- Nojima, D.; Yoshino, T.; Maeda, Y.; Tanaka, M.; Nemoto, M.; Tanaka, T. Proteomics analysis of oil body-associated proteins in the oleaginous diatom. J. Proteome Res. 2013, 12, 5293–5301. [Google Scholar] [CrossRef]
- Peled, E.; Leu, S.; Zarka, A.; Weiss, M.; Pick, U.; Khozin-Goldberg, I.; Boussiba, S. Isolation of a novel oil globule protein from the green alga Haematococcus pluvialis (Chlorophyceae). Lipids 2011, 46, 851–861. [Google Scholar] [CrossRef]
- Vieler, A.; Brubaker, S.B.; Vick, B.; Benning, C. A lipid droplet protein of Nannochloropsis with functions partially analogous to plant oleosins. Plant Physiol. 2012, 158, 1562–1569. [Google Scholar] [CrossRef]
- Liang, Y.; Maeda, Y.; Yoshino, T.; Matsumoto, M.; Tanaka, T. Profiling of fatty acid methyl esters from the oleaginous diatom Fistulifera sp. strain JPCC DA0580 under nutrition-sufficient and -deficient conditions. J. Appl. Phycol. 2014. [Google Scholar] [CrossRef]
- Matsumoto, M.; Sugiyama, H.; Maeda, Y.; Sato, R.; Tanaka, T.; Matsunaga, T. Marine diatom, Navicula sp. strain JPCC DA0580 and marine green alga, Chlorella sp. strain NKG400014 as potential sources for biodiesel production. Appl. Biochem. Biotechnol. 2010, 161, 483–490. [Google Scholar] [CrossRef]
- Sato, R.; Maeda, Y.; Yoshino, T.; Tanaka, T.; Matsumoto, M. Seasonal variation of biomass and oil production of the oleaginous diatom Fistulifera sp. in outdoor vertical bubble column and raceway-type bioreactors. J. Biosci. Bioeng. 2014, 117, 720–724. [Google Scholar] [CrossRef]
- Satoh, A.; Ichii, K.; Matsumoto, M.; Kubota, C.; Nemoto, M.; Tanaka, M.; Yoshino, T.; Matsunaga, T.; Tanaka, T. A process design and productivity evaluation for oil production by indoor mass cultivation of a marine diatom, Fistulifera sp. JPCC DA0580. Bioresour. Technol. 2013, 137, 132–138. [Google Scholar] [CrossRef]
- Apt, K.E.; Zaslavkaia, L.; Lippmeier, J.C.; Lang, M.; Kilian, O.; Wetherbee, R.; Grossman, A.R.; Kroth, P.G. In vivo characterization of diatom multipartite plastid targeting signals. J. Cell Sci. 2002, 115, 4061–4069. [Google Scholar] [CrossRef]
- Liang, Y.; Maeda, Y.; Sunaga, Y.; Muto, M.; Matsumoto, M.; Yoshino, T.; Tanaka, T. Biosynthesis of polyunsaturated fatty acids in the oleaginous marine diatom Fistulifera sp. strain JPCC DA0580. Mar. Drugs 2013, 11, 5008–5023. [Google Scholar] [CrossRef]
- Nemoto, M.; Maeda, Y.; Muto, M.; Tanaka, M.; Yoshino, T.; Mayama, S.; Tanaka, T. Identification of a frustule-associated protein of the marine pennate diatom Fistulifera sp. strain JPCC DA0580. Mar. Genomics 2014. [Google Scholar] [CrossRef]
- Bendtsen, J.D.; Nielsen, H.; von Heijne, G.; Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 2004, 340, 783–795. [Google Scholar]
- Abell, B.M.; Holbrook, L.A.; Abenes, M.; Murphy, D.J.; Hills, M.J.; Moloney, M.M. Role of the proline knot motif in oleosin endoplasmic reticulum topology and oil body targeting. Plant Cell 1997, 9, 1481–1493. [Google Scholar] [CrossRef]
- Huang, A.H. Oleosins and oil bodies in seeds and other organs. Plant Physiol. 1996, 110, 1055–1061. [Google Scholar]
- Kilian, O.; Kroth, P.G. Identification and characterization of a new conserved motif within the presequence of proteins targeted into complex diatom plastids. Plant J. 2005, 41, 175–183. [Google Scholar] [CrossRef]
- Stork, S.; Lau, J.; Moog, D.; Maier, U.G. Three old and one new: Protein import into red algal-derived plastids surrounded by four membranes. Protoplasma 2013, 250, 1013–1023. [Google Scholar] [CrossRef]
- Gibbs, S.P. The chloroplast endoplasmic reticulum: Structure, function, and evolutionary significance. Int. Rev. Cytol. 1981, 72, 49–99. [Google Scholar] [CrossRef]
- Gruber, A.; Vugrinec, S.; Hempel, F.; Gould, S.B.; Maier, U.G.; Kroth, P.G. Protein targeting into complex diatom plastids: Functional characterisation of a specific targeting motif. Plant Mol. Biol. 2007, 64, 519–530. [Google Scholar] [CrossRef]
- Peschke, M.; Moog, D.; Klingl, A.; Maier, U.G.; Hempel, F. Evidence for glycoprotein transport into complex plastids. Proc. Natl. Acad. Sci. USA 2013, 110, 10860–10865. [Google Scholar]
- Knothe, G. Dependence of biodiesel fuel properties on the structure of fatty acid alkyl esters. Fuel Process. Technol. 2005, 86, 1059–1070. [Google Scholar]
- Guillard, R.R.; Ryther, J.H. Studies of marine planktonic diatoms. I. Cyclotella nana Hustedt, and Detonula confervacea (cleve) Gran. Can. J. Microbiol. 1962, 8, 229–239. [Google Scholar] [CrossRef]
- Tanaka, T.; Maeda, Y.; Veluchamy, A.; Tanaka, M.; Bowler, C.; Muto, M.; Sunaga, Y.; Tanaka, M.; Yoshino, T.; Taniguchi, T.; et al. Genome and transcriptome analyses of theoleaginous diatom Fistulifera sp. reveals the oil accumulation mechanism. 2014; to be submitted for publication. [Google Scholar]
- Muto, M.; Fukuda, Y.; Nemoto, M.; Yoshino, T.; Matsunaga, T.; Tanaka, T. Establishment of a genetic transformation system for the marine pennate diatom Fistulifera sp. strain JPCC DA0580—A high triglyceride producer. Mar. Biotechnol. 2013, 15, 48–55. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Maeda, Y.; Sunaga, Y.; Yoshino, T.; Tanaka, T. Oleosome-Associated Protein of the Oleaginous Diatom Fistulifera solaris Contains an Endoplasmic Reticulum-Targeting Signal Sequence. Mar. Drugs 2014, 12, 3892-3903. https://doi.org/10.3390/md12073892
Maeda Y, Sunaga Y, Yoshino T, Tanaka T. Oleosome-Associated Protein of the Oleaginous Diatom Fistulifera solaris Contains an Endoplasmic Reticulum-Targeting Signal Sequence. Marine Drugs. 2014; 12(7):3892-3903. https://doi.org/10.3390/md12073892
Chicago/Turabian StyleMaeda, Yoshiaki, Yoshihiko Sunaga, Tomoko Yoshino, and Tsuyoshi Tanaka. 2014. "Oleosome-Associated Protein of the Oleaginous Diatom Fistulifera solaris Contains an Endoplasmic Reticulum-Targeting Signal Sequence" Marine Drugs 12, no. 7: 3892-3903. https://doi.org/10.3390/md12073892
APA StyleMaeda, Y., Sunaga, Y., Yoshino, T., & Tanaka, T. (2014). Oleosome-Associated Protein of the Oleaginous Diatom Fistulifera solaris Contains an Endoplasmic Reticulum-Targeting Signal Sequence. Marine Drugs, 12(7), 3892-3903. https://doi.org/10.3390/md12073892




