Diatom Derived Polyunsaturated Aldehydes Do Not Structure the Planktonic Microbial Community in a Mesocosm Study
Abstract
:1. Introduction
2. Results
2.1. Phytoplankton Development
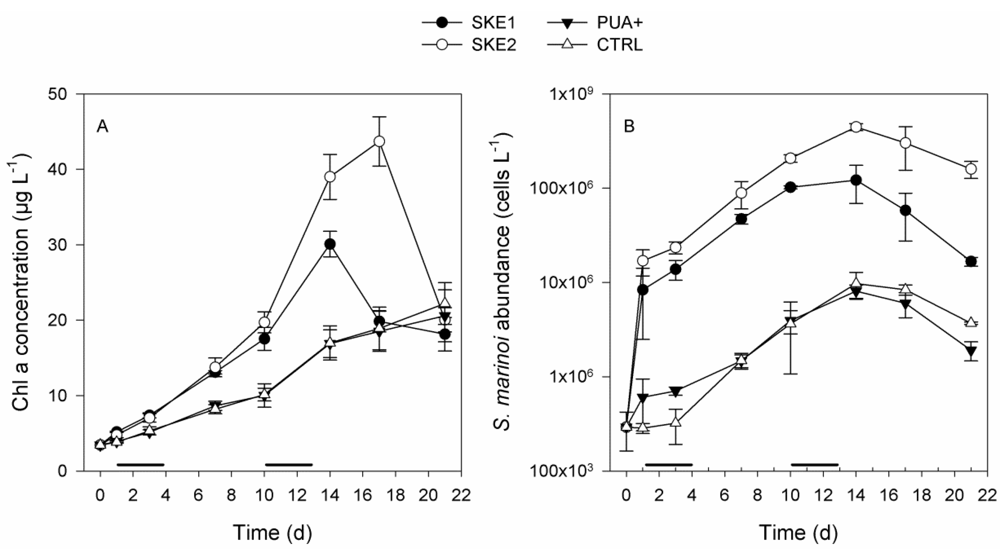
2.2. Bacterial and Viral Abundance
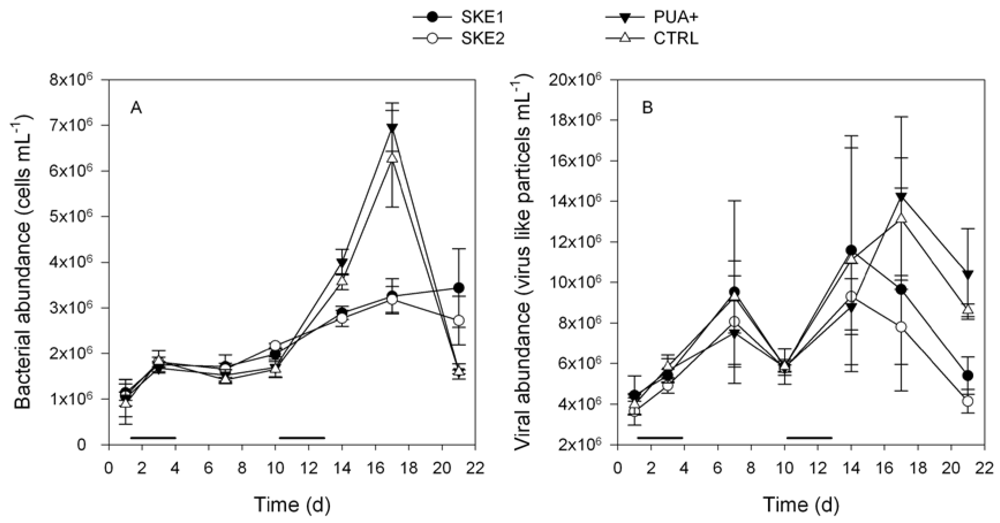
2.3. Dissolved PUA Concentration
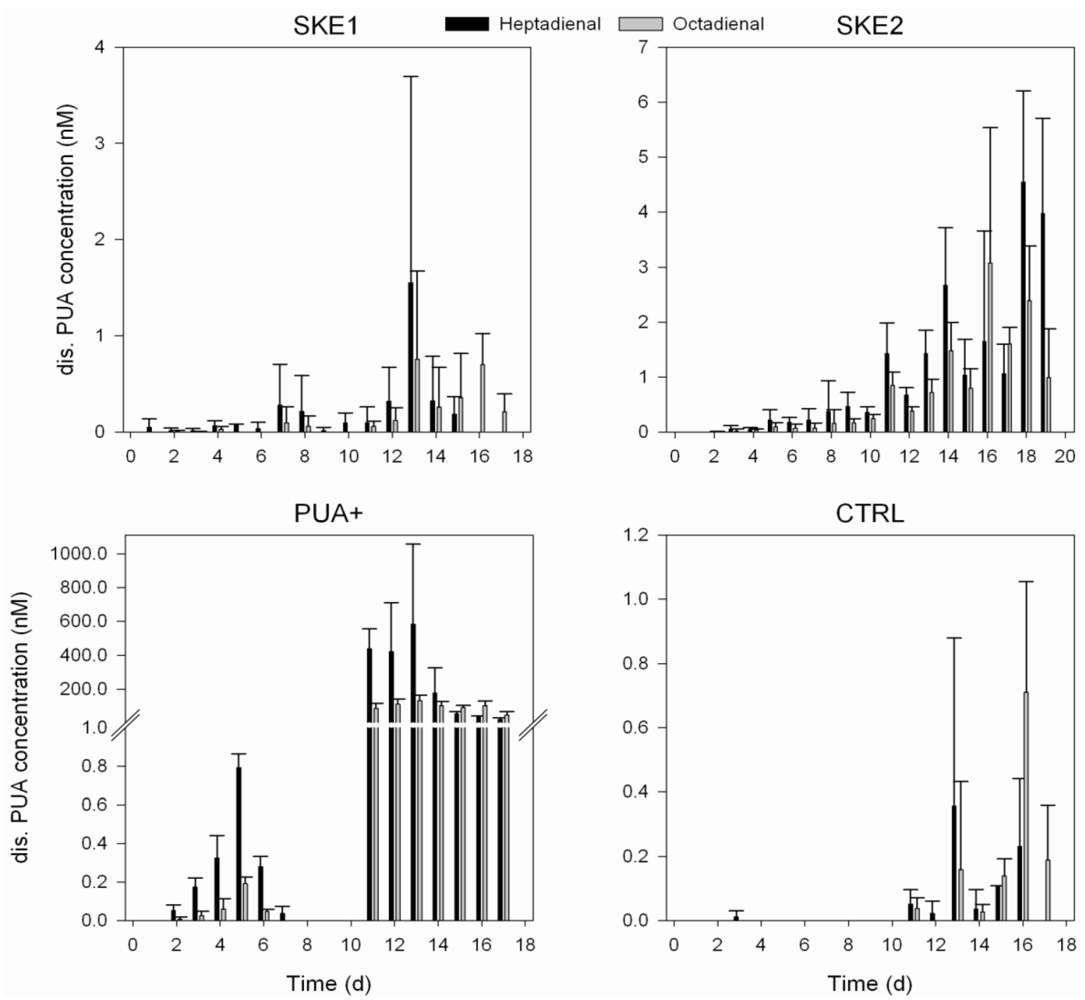
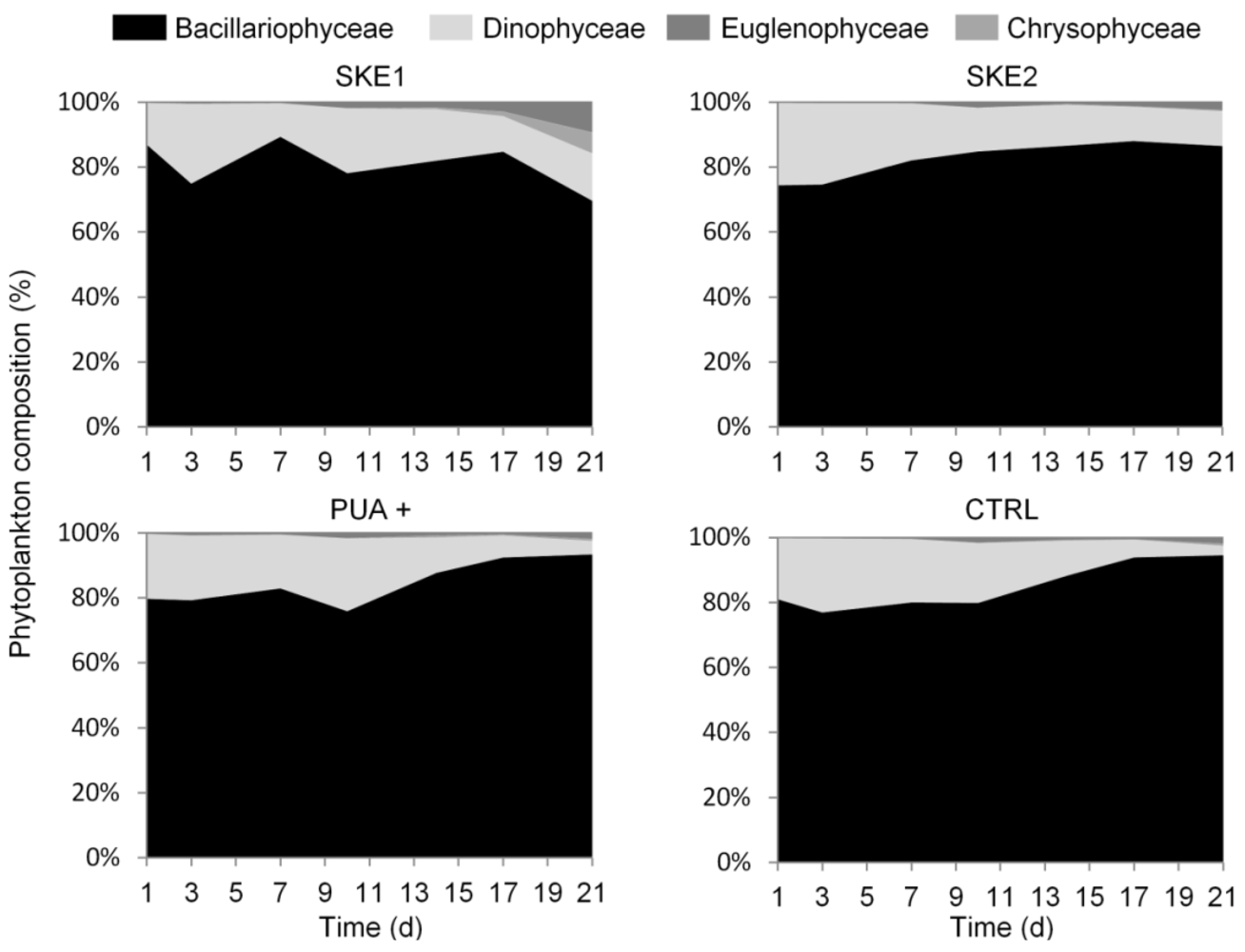
2.4. Bacterial Community Composition
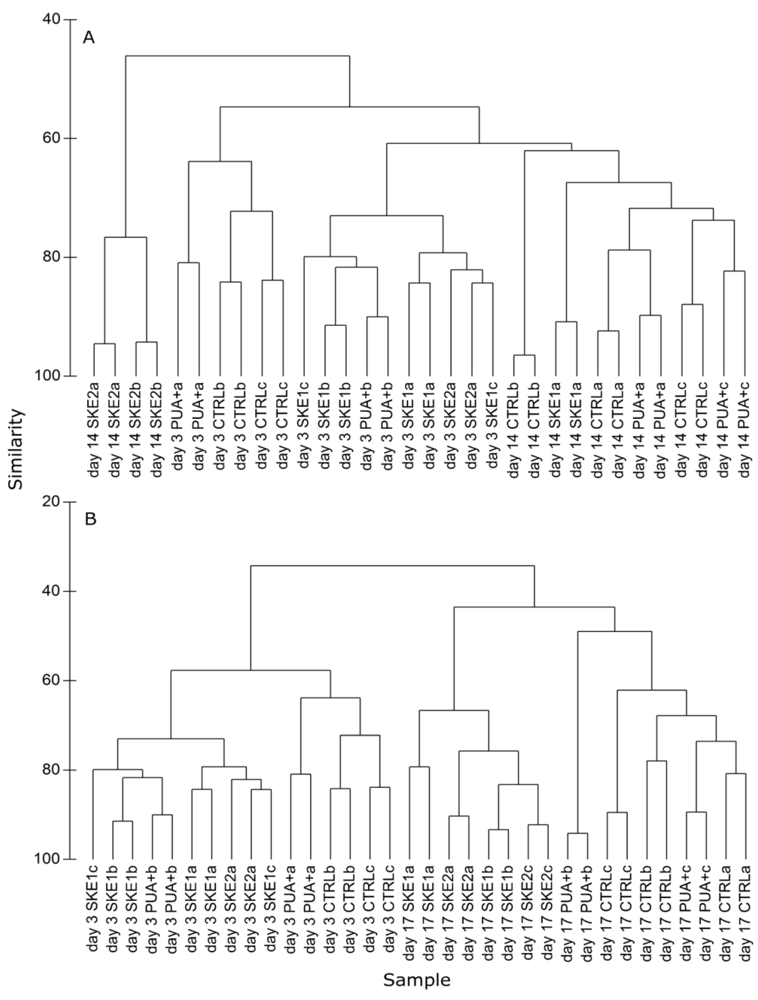
3. Discussion
4. Experimental Section
4.1. Experimental Design
4.2. General Sampling
4.3. Chlorophyll a Concentration
4.4. Phytoplankton Abundance
4.5. Bacterial Abundance
4.6. Viral Abundance
4.7. PUA Analysis
4.8. Bacterial Community Composition
4.9. Statistical Analysis
5. Conclusion
Acknowledgements
References
- Pohnert, G. Diatom/copepod interactions in plankton: The indirect chemical defense of unicellular algae. ChemBioChem 2005, 6, 946–959. [Google Scholar]
- Ianora, A.; Miralto, A.; Poulet, S.A.; Carotenuto, Y.; Buttino, I.; Romano, G.; Casotti, R.; Pohnert, G.; Wichard, T.; Colucci-D’Amato, L.; et al. Aldehyde suppression of copepod recruitment in blooms of a ubiquitous planktonic diatom. Nature 2004, 429, 403–407. [Google Scholar]
- Adolph, S.; Bach, S.; Blondel, M.; Cueff, A.; Moreau, M.; Pohnert, G.; Poulet, S.; Wichard, T.; Zuccaro, A. Cytotoxicity of diatom-derived oxylipins in organisms belonging to different phyla. J. Exp. Biol. 2004, 207, 2935–2946. [Google Scholar]
- Buttino, I.; de Rosa, G.; Carotenuto, Y.; Mazzella, M.; Ianora, A.; Esposito, F.; Vitiello, V.; Quaglia, F.; La Rotonda, M.I.; Miralto, A. Aldehyde-encapsulating liposomes impair marine grazer survivorship. J. Exp. Biol. 2008, 211, 1426–1433. [Google Scholar]
- Ianora, A.; Miralto, A. Toxigenic effects of diatoms on grazers, phytoplankton and other microbes: A review. Ecotoxicology 2010, 19, 493–511. [Google Scholar]
- Leflaive, J.; Ten-Hage, L. Chemical interactions in diatoms: Role of polyunsaturated aldehydes and precursors. New Phytol. 2009, 184, 794–805. [Google Scholar]
- Halsband-Lenk, C.; Pierson, J.J.; Leising, A.W. Reproduction of Pseudocalanus newmani (Copepoda: Calanoida) is deleteriously affected by diatom blooms—A field study. Prog. Oceanogr. 2005, 67, 332–348. [Google Scholar] [CrossRef]
- Irigoien, X.; Harris, R.P.; Head, R.N.; Harbour, D. The influence of diatom abundance on the egg production rate of Calanus helgolandicus in the English Channel. Limnol. Oceanogr. 2000, 45, 1433–1439. [Google Scholar] [CrossRef]
- Sommer, U. Copepod growth and diatoms: Insensitivity of Acartia tonsa to the composition of semi-natural plankton mixtures manipulated by silicon:nitrogen ratios in mesocosms. Oecologia 2009, 159, 207–215. [Google Scholar] [CrossRef]
- Poulet, S.A.; Cueff, A.; Wichard, T.; Marchetti, J.; Dancie, C.; Pohnert, G. Influence of diatoms on copepod reproduction. III. Consequences of abnormal oocyte maturation on reproductive factors in Calanus helgolandicus. Mar. Biol. 2007, 152, 415–428. [Google Scholar] [CrossRef]
- Poulet, S.A.; Wichard, T.; Ledoux, J.B.; Lebreton, B.; Marchetti, J.; Dancie, C.; Bonnet, D.; Cueff, A.; Morin, P.; Pohnert, G. Influence of diatoms on copepod reproduction. I. Field and laboratory observations related to Calanus helgolandicus egg production. Mar. Ecol. Prog. Ser. 2006, 308, 129–142. [Google Scholar] [CrossRef]
- Wichard, T.; Poulet, S.A.; Boulesteix, A.L.; Ledoux, J.B.; Lebreton, B.; Marchetti, J.; Pohnert, G. Influence of diatoms on copepod reproduction. II. Uncorrelated effects of diatom-derived α, β, γ, δ-unsaturated aldehydes and polyunsaturated fatty acids on Calanus helgolandicus in the field. Prog. Oceanogr. 2008, 77, 30–44. [Google Scholar] [CrossRef]
- Jonasdottir, S.; Dutz, J.; Koski, M.; Yebra, L.; Jakobsen, H.H.; Vidoudez, C.; Pohnert, G.; Nejstgaard, J.C. Extensive cross-disciplinary analysis of biological and chemical control of Calanus finmarchicus reproduction during an aldehyde forming diatom bloom in mesocosms. Mar. Biol. 2011, 158, 1943–1963. [Google Scholar] [CrossRef]
- Flynn, K.J.; Irigoien, X. Aldehyde-induced insidious effects cannot be considered as a diatom defence mechanism against copepods. Mar. Ecol. Prog. Ser. 2009, 377, 79–89. [Google Scholar]
- Vardi, A.; Formiggini, F.; Casotti, R.; de Martino, A.; Ribalet, F.; Miralto, A.; Bowler, C. A stress surveillance system based on calcium and nitric oxide in marine diatoms. PLoS Biol. 2006, 4, 411–419. [Google Scholar]
- Vidoudez, C.; Pohnert, G. Growth phase-specific release of polyunsaturated aldehydes by the diatom Skeletonema marinoi. J. Plankton Res. 2008, 30, 1305–1313. [Google Scholar] [CrossRef]
- Ribalet, F.; Intertaglia, L.; Lebaron, P.; Casotti, R. Differential effect of three polyunsaturated aldehydes on marine bacterial isolates. Aquat. Toxicol. 2008, 86, 249–255. [Google Scholar]
- Vidoudez, C.; Nejstgaard, J.C.; Jakobsen, H.H.; Pohnert, G. Dynamics of dissolved and particulate polyunsaturated aldehydes in mesocosms inoculated with different densities of the diatom Skeletonema marinoi. Mar. Drugs 2011, 9, 345–358. [Google Scholar] [CrossRef]
- Vidoudez, C.; Casotti, R.; Bastianini, M.; Pohnert, G. Quantification of dissolved and particulate polyunsaturated aldehydes in the Adriatic Sea. Mar. Drugs 2011, 9, 500–513. [Google Scholar]
- Balestra, C.; Alonso-Saez, L.; Gasol, J.M.; Casotti, R. Group-specific effects on coastal bacterioplankton of polyunsaturated aldehydes produced by diatoms. Aquat. Microb. Ecol. 2011, 63, 123–131. [Google Scholar]
- Romano, G.; Russo, G.L.; Buttino, I.; Ianora, A.; Miralto, A. A marine diatom-derived aldehyde induces apoptosis in copepod and sea urchin embryos. J. Exp. Biol. 2003, 206, 3487–3494. [Google Scholar]
- Ask, J.; Reinikainen, M.; Bamstedt, U. Variation in hatching success and egg production of Eurytemora affinis (Calanoida, Copepoda) from the Gulf of Bothnia, Baltic Sea, in relation to abundance and clonal differences of diatoms. J. Plankton Res. 2006, 28, 683–694. [Google Scholar] [CrossRef]
- Riemann, L.; Steward, G.F.; Azam, F. Dynamics of bacterial community composition and activity during a mesocosm diatom bloom. Appl. Environ. Microbiol. 2000, 66, 578–587. [Google Scholar]
- Rhee, G.Y. Competition between an alga and an aquatic bacterium for phophorous. Limnol. Oceanogr. 1972, 17, 505–514. [Google Scholar]
- Sarno, D.; Kooistra, W.; Medlin, L.K.; Percopo, I.; Zingone, A. Diversity in the genus Skeletonema (Bacillariophyceae). II. An assessment of the taxonomy of S. costatum-like species with the description of four new species. J. Phycol. 2005, 41, 151–176. [Google Scholar] [CrossRef]
- Terekhova, V.E.; Aizdaicher, N.A.; Buzoleva, L.S.; Somov, G.P. Influence of extrametabolites of marine microalgae on the reproduction of the bacterium Listeria monocytogenes. Russ. J. Mar. Biol. 2009, 35, 355–358. [Google Scholar] [CrossRef]
- Bratbak, G.; Egge, J.K.; Heldal, M. Viral mortality of the marine alga Emiliania huxleyi (Haptophyceae) and termination of algal blooms. Mar. Ecol. Prog. Ser. 1993, 93, 39–48. [Google Scholar] [CrossRef]
- Jacquet, S.; Heldal, M.; Iglesias-Rodriguez, D.; Larsen, A.; Wilson, W.; Bratbak, G. Flow cytometric analysis of an Emiliana huxleyi bloom terminated by viral infection. Aquat. Microb. Ecol. 2002, 27, 111–124. [Google Scholar]
- Vardi, A.; van Mooy, B.A.S.; Fredricks, H.F.; Popendorf, K.J.; Ossolinski, J.E.; Haramaty, L.; Bidle, K.D. Viral glycosphingolipids induce lytic infection and cell death in marine phytoplankton. Science 2009, 326, 861–865. [Google Scholar]
- Tomaru, Y.; Takao, Y.; Suzuki, H.; Nagumo, T.; Nagasaki, K. Isolation and characterization of a single-stranded RNA virus infecting the bloom-forming diatom Chaetoceros socialis. Appl. Environ. Microbiol. 2009, 75, 2375–2381. [Google Scholar] [CrossRef]
- Ribalet, F.; Berges, J.A.; Ianora, A.; Casotti, R. Growth inhibition of cultured marine phytoplankton by toxic algal-derived polyunsaturated aldehydes. Aquat. Toxicol. 2007, 85, 219–227. [Google Scholar]
- Grossart, H.P.; Levold, F.; Allgaier, M.; Simon, M.; Brinkhoff, T. Marine diatom species harbour distinct bacterial communities. Environ. Microbiol. 2005, 7, 860–873. [Google Scholar]
- Riemann, L.; Middelboe, M. Stability of bacterial and viral community compositions in Danish coastal waters as depicted by DNA fingerprinting techniques. Aquat. Microb. Ecol. 2002, 27, 219–232. [Google Scholar]
- Barofsky, A.; Vidoudez, C.; Pohnert, G. Metabolic profiling reveals growth stage variability in diatom exudates. Limnol. Oceanogr. Methods 2009, 7, 382–390. [Google Scholar]
- Paul, C.; Barofsky, A.; Vidoudez, C.; Pohnert, G. Diatom exudates influence metabolism and cell growth of co-cultured diatom species. Mar. Ecol. Prog. Ser. 2009, 389, 61–70. [Google Scholar]
- Available online: http://www.umf.umu.se/miljoovervakning/databasen-dbotnia/.
- Utermöhl, H. Zur Vervollkommnung der quantitativen Phytoplankton-Methodik. Mitt. Int. Ver. Theor. Angew. Limnol. 1958, 9, 1–38. [Google Scholar]
- Helcom, Baltic Sea Environment Proceedings No. 106., Biovolumes and Size-Classes of Phytoplankton in the Baltic Sea. Helsinki Commission Baltic Marine Environment Protection Commission: Helsinki, Finland, 2006.
- Blackburn, N.; Hagström, Å.; Wikner, J.; Cuadros-Hansson, R.; Bjørnsen, P.K. Rapid determination of bacterial abundance, biovolume, morphology, and growth by neural network-based image analysis. Appl. Environ. Microbiol. 1998, 64, 3246–3255. [Google Scholar]
- Marie, D.; Brussaard, C.P.D.; Thyrhaug, R.; Bratbak, G.; Vaulot, D. Enumeration of marine viruses in culture and natural samples by flow cytometry. Appl. Environ. Microbiol. 1999, 65, 45–52. [Google Scholar]
- Brussaard, C.P.D. Optimization of procedures for counting viruses by flow cytometry. Appl. Environ. Microbiol. 2004, 70, 1506–1513. [Google Scholar]
- Ghiglione, J.F.; Larcher, M.; Lebaron, P. Spatial and temporal scales of variation in bacterioplankton community structure in the NW Mediterranean Sea. Aquat. Microb. Ecol. 2005, 40, 229–240. [Google Scholar]
- Pandey, J.; Ganesan, K.; Jain, R.K. Variations in T-RFLP profiles with differing chemistries of fluorescent dyes used for labeling the PCR primers. J. Microbiol. Methods 2007, 68, 633–638. [Google Scholar]
- Smith, C.J.; Danilowicz, B.S.; Clear, A.K.; Costello, F.J.; Wilson, B.; Meijer, W.G. T-Align, a web-based tool for comparison of multiple terminal restriction fragment length polymorphism profiles. FEMS Microbiol. Ecol. 2005, 54, 375–380. [Google Scholar]
- Culman, S.W.; Bukowski, R.; Gauch, H.G.; Cadillo-Quiroz, H.; Buckley, D.H. T-REX: Software for the processing and analysis of T-RFLP data. BMC Bioinforma. 2009, 10. [Google Scholar]
- Abdo, Z.; Schuette, U.M.E.; Bent, S.J.; Williams, C.J.; Forney, L.J.; Joyce, P. Statistical methods for characterizing diversity of microbial communities by analysis of terminal restriction fragment length polymorphisms of 16S rRNA genes. Environ. Microbiol. 2006, 8, 929–938. [Google Scholar]
- Samples Availability: Available from the authors.
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Paul, C.; Reunamo, A.; Lindehoff, E.; Bergkvist, J.; Mausz, M.A.; Larsson, H.; Richter, H.; Wängberg, S.-Å.; Leskinen, P.; Båmstedt, U.; et al. Diatom Derived Polyunsaturated Aldehydes Do Not Structure the Planktonic Microbial Community in a Mesocosm Study. Mar. Drugs 2012, 10, 775-792. https://doi.org/10.3390/md10040775
Paul C, Reunamo A, Lindehoff E, Bergkvist J, Mausz MA, Larsson H, Richter H, Wängberg S-Å, Leskinen P, Båmstedt U, et al. Diatom Derived Polyunsaturated Aldehydes Do Not Structure the Planktonic Microbial Community in a Mesocosm Study. Marine Drugs. 2012; 10(4):775-792. https://doi.org/10.3390/md10040775
Chicago/Turabian StylePaul, Carsten, Anna Reunamo, Elin Lindehoff, Johanna Bergkvist, Michaela A. Mausz, Henrik Larsson, Hannes Richter, Sten-Åke Wängberg, Piia Leskinen, Ulf Båmstedt, and et al. 2012. "Diatom Derived Polyunsaturated Aldehydes Do Not Structure the Planktonic Microbial Community in a Mesocosm Study" Marine Drugs 10, no. 4: 775-792. https://doi.org/10.3390/md10040775
APA StylePaul, C., Reunamo, A., Lindehoff, E., Bergkvist, J., Mausz, M. A., Larsson, H., Richter, H., Wängberg, S.-Å., Leskinen, P., Båmstedt, U., & Pohnert, G. (2012). Diatom Derived Polyunsaturated Aldehydes Do Not Structure the Planktonic Microbial Community in a Mesocosm Study. Marine Drugs, 10(4), 775-792. https://doi.org/10.3390/md10040775



