Differential Expression of the Demosponge (Suberites domuncula) Carotenoid Oxygenases in Response to Light: Protection Mechanism Against the Self-Produced Toxic Protein (Suberitine)
Abstract
:1. Introduction
2. Results
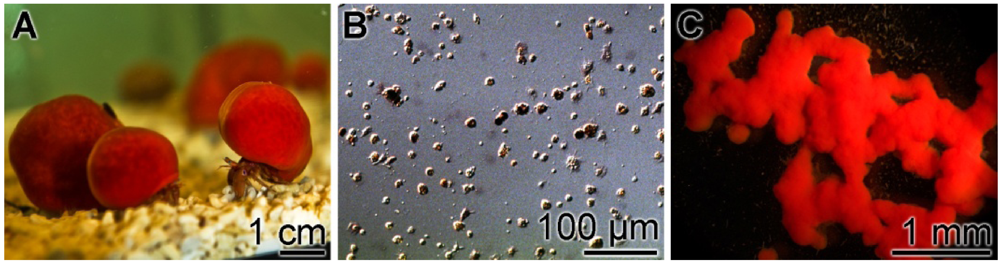
2.1. Suberitine: Purification
| Fraction | Total Protein (mg) | Total Hemagglutination (units) | Purification (fold) | Recovery (%) | Specific Activity (units/mg) |
|---|---|---|---|---|---|
| I: Crude fraction | 218.4 | 102,370 | 1 | 100 | 469.6 |
| II: (NH4)2SO4 precipitation | 13.02 | 47,940 | 7.8 | 47 | 3687.7 |
| III: Sephadex G-100 | 3.24 | 32,640 | 21.4 | 32 | 10,074.1 |
| IV: Sephadex G-50 | 1.05 | 27,530 | 55.8 | 27 | 26,219.1 |
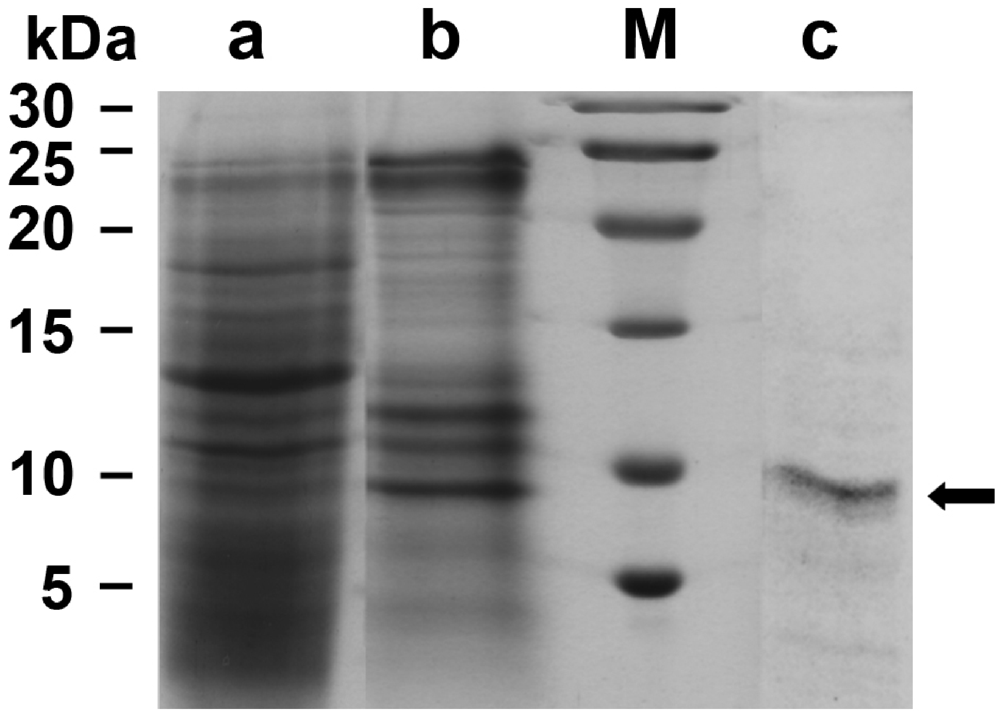
2.2. Suberitine: Biological Activity

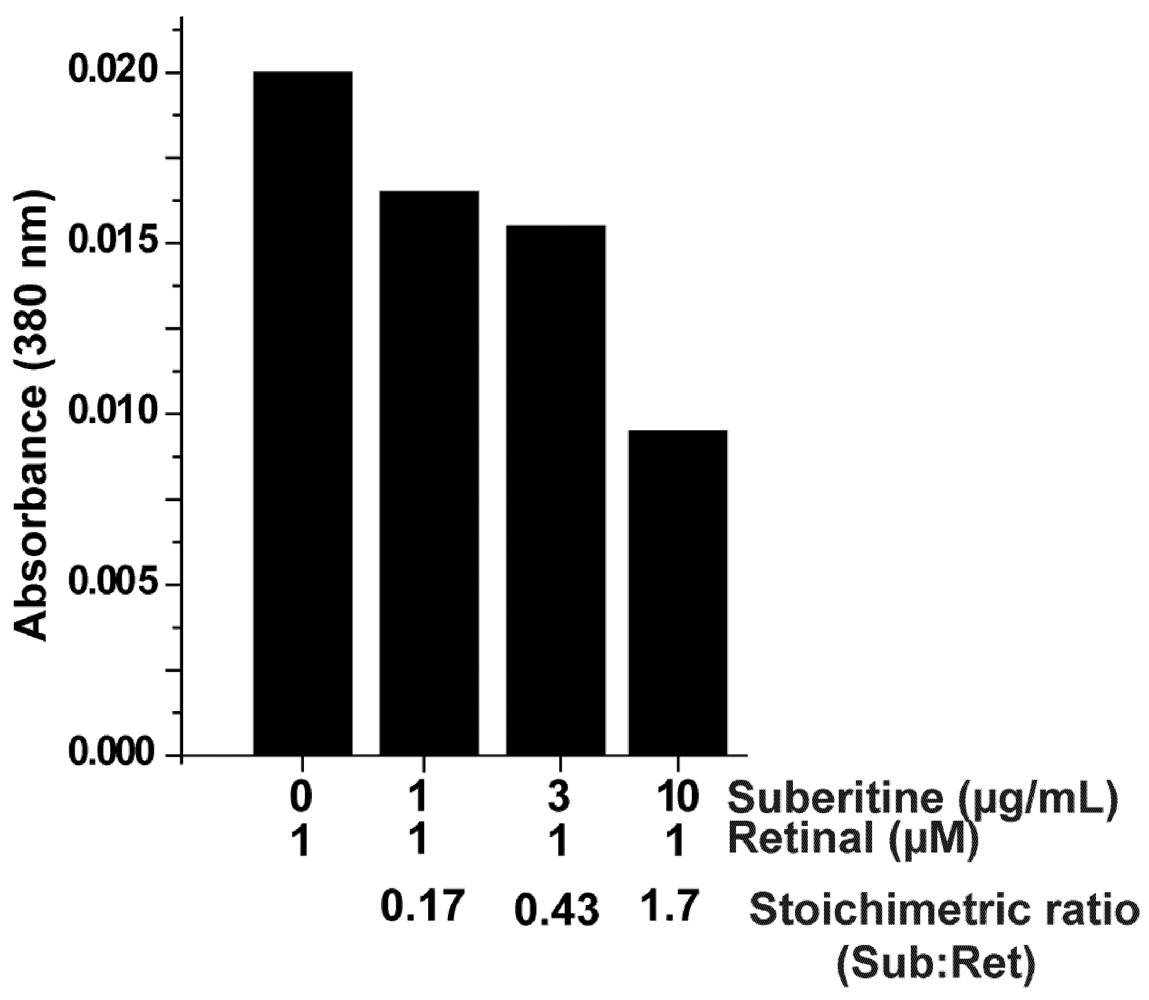
2.3. Interaction of Suberitine with Retinal
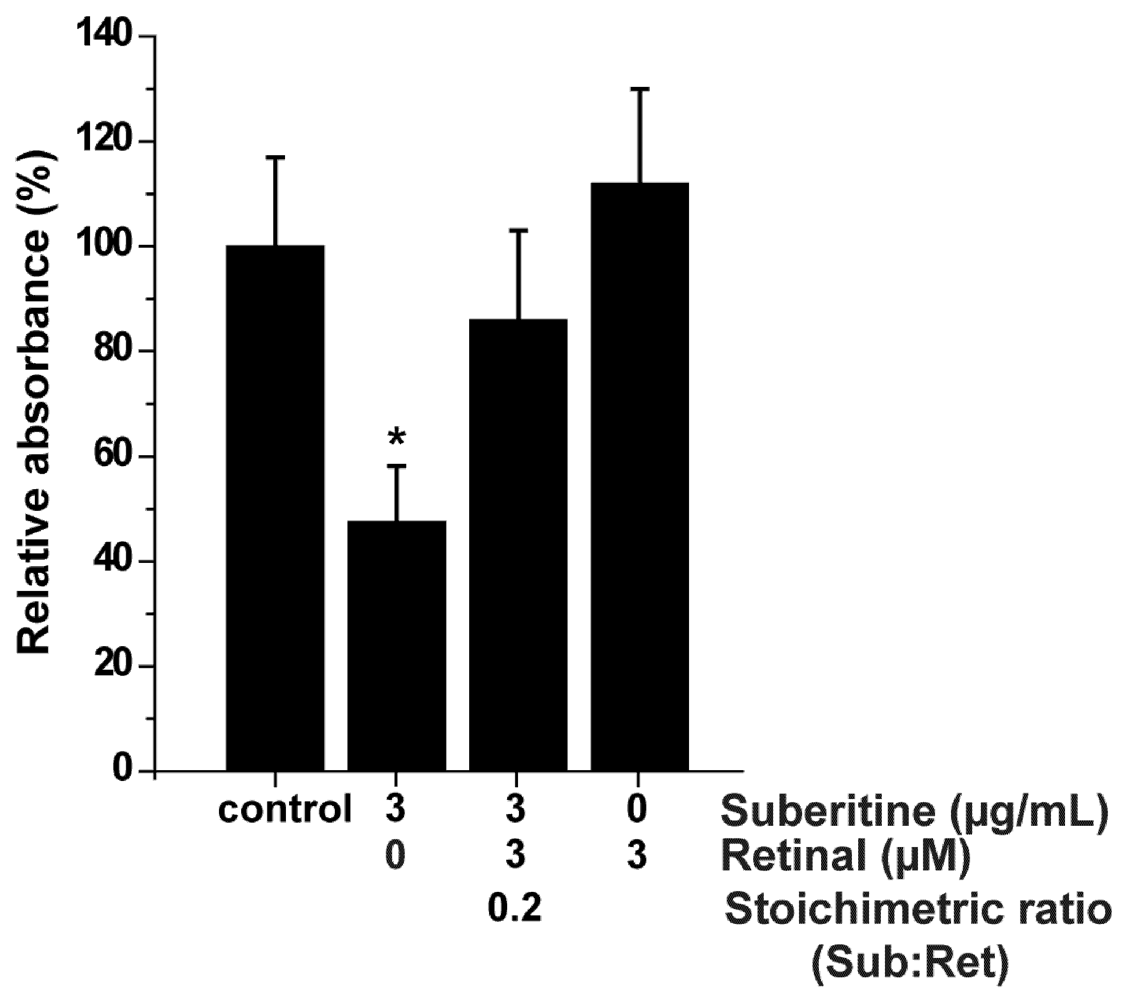
2.4. Effect of Suberitine on the Viability of Sponge Cells
2.5. Molecular Cloning of the Two (Related) Carotenoid Oxygenases
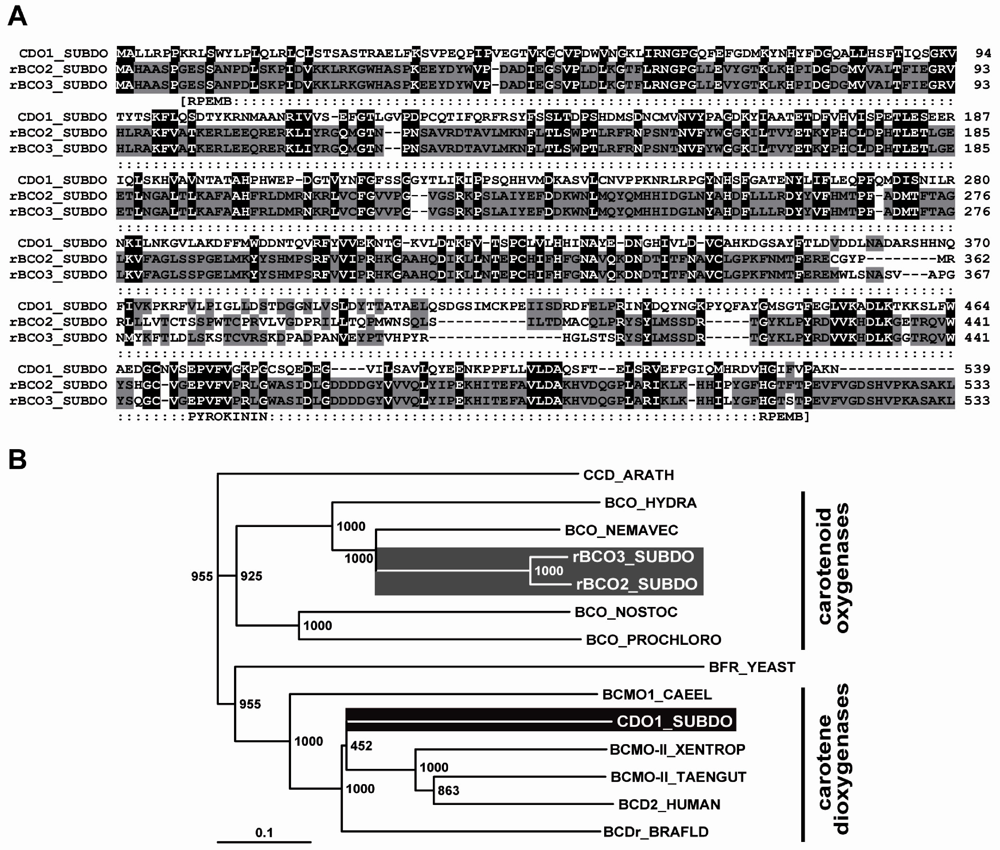
2.6. Determination of Oxygenase Activity in Bacterial System
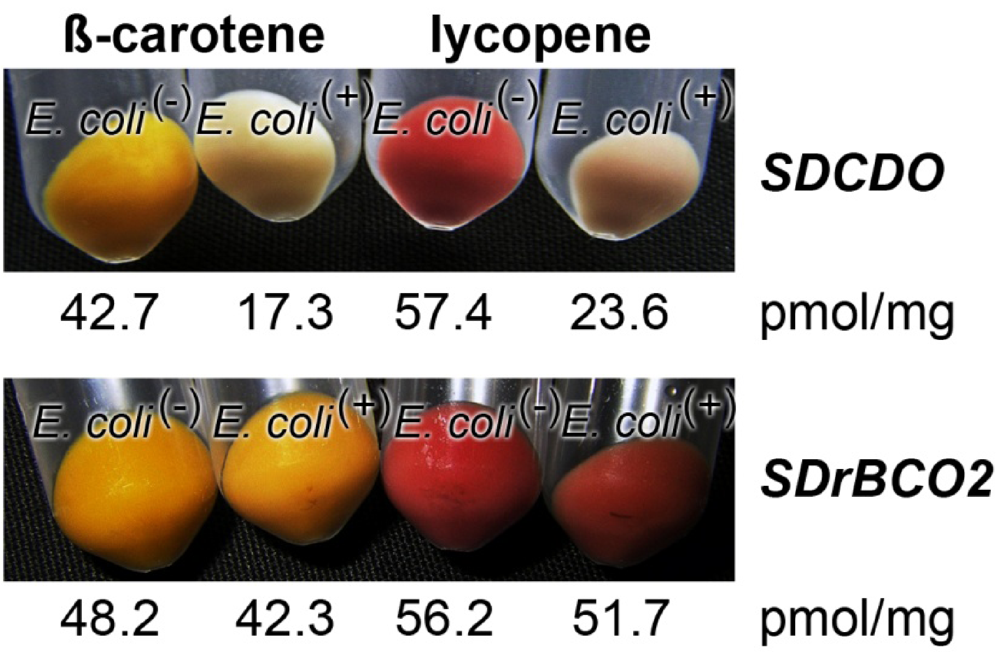
2.7. Effect of Light on the Expression of the Carotene Dioxygenase and the Carotenoid Oxygenase 2
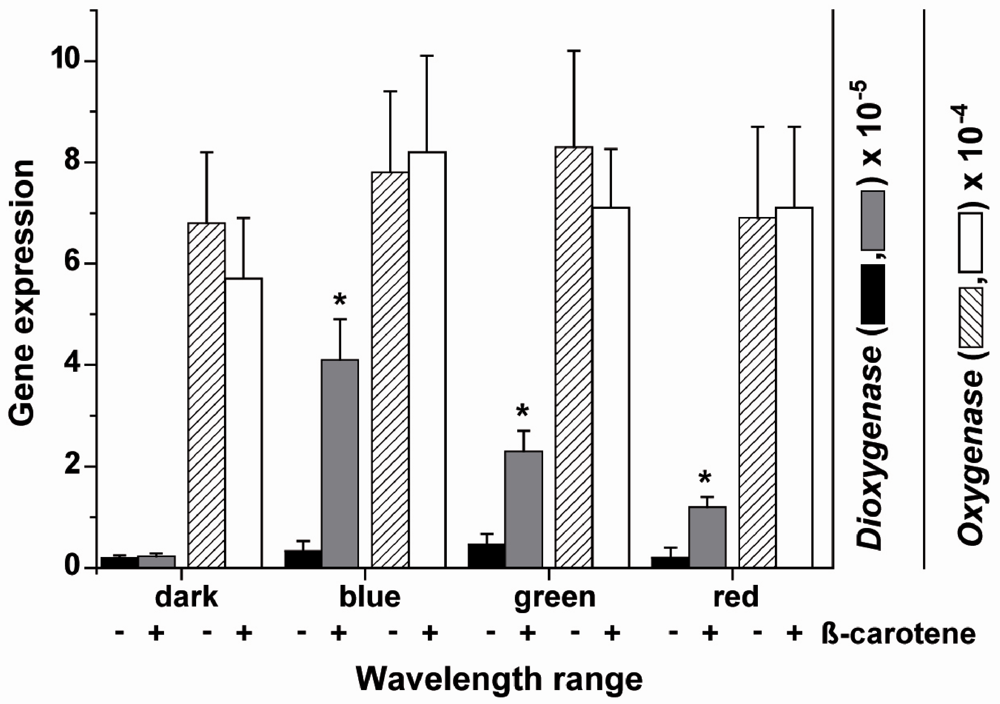
3. Discussion
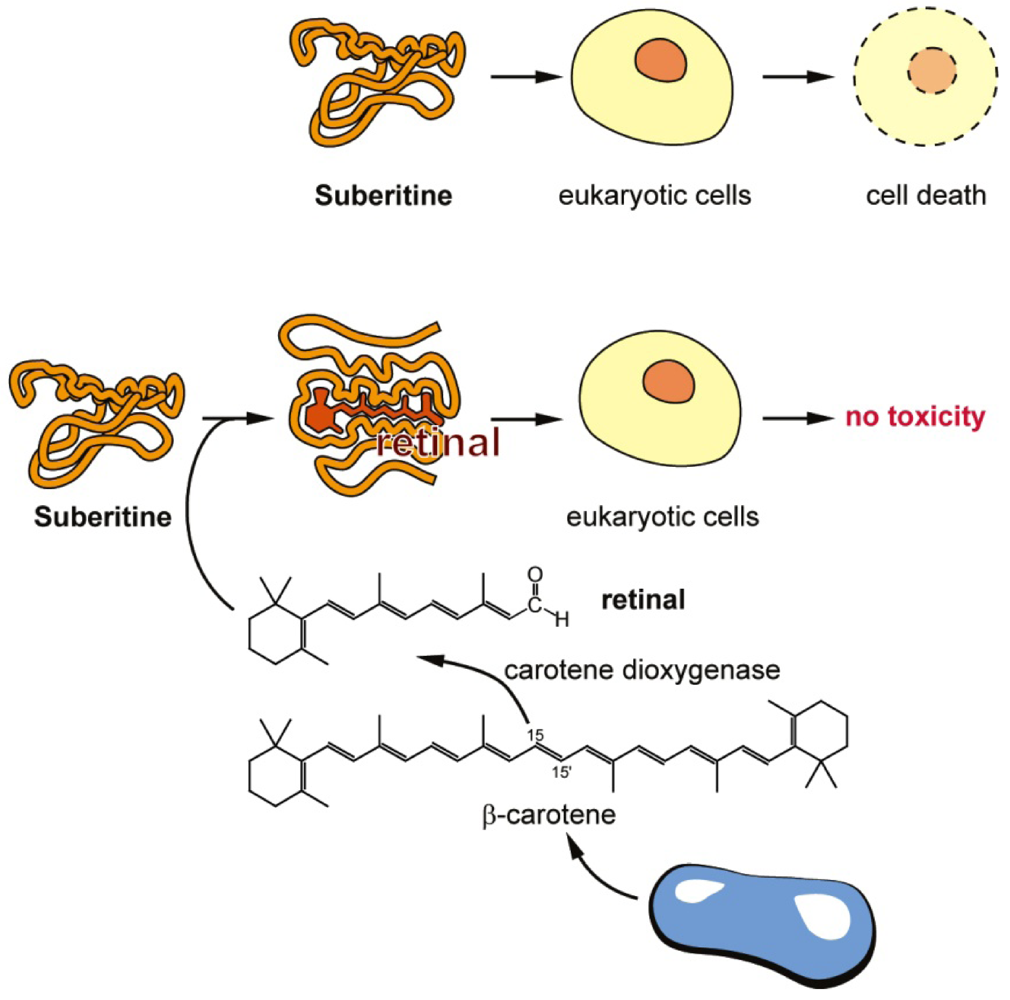
4. Experimental Section
4.1. Sponge and Primmorphs
4.2. Exposure of Primmorphs to Light
4.3. Purification of Suberitine
4.4. SDS-PAGE Analysis
4.5. Testing for Hemolytic Activity
4.6. Interaction of Suberitine with Retinal
4.7. Quenching of the Optical Absorption Spectrum of Retinal by Suberitine
4.8. MTT Sponge Cell Viability Assay
4.9. Molecular Cloning of S. domuncula (Related) Carotenoid Oxygenases
4.10. Sequence Analyses
4.11. Determination of the Potential Carotenoid Oxygenase Activity in the Bacterial System
4.12. Quantitative Analysis of β-carotene and Lycopene
4.13. Quantitative Real-Time RT-PCR (qRT-PCR)
4.14. Additional Analytical Method
4.15. Statistical Analysis
4. Conclusion
Acknowledgements
- Samples Availability: Available from the authors.
References
- Bergmann, W.; Feeney, R.J. The isolation of a new thymine pentoside from sponges. J. Am. Chem. Soc. 1950, 72, 2809–2810. [Google Scholar]
- Sarma, A.S.; Daum, T.; Müller, W.E.G. Secondary metabolites from marine sponges: Part I: Origin and chemistry of new metabolites, and synthetic studies; Part II: Biological properties of new metabolites and physiological activities of avarol and related compounds isolated from Dysidea sp. In Akademie gemeinnütziger Wissenschaften zu Erfurt; Ullstein-Mosby Verlag: Berlin, Germany, 1993; pp. 1–168. [Google Scholar]
- Pomponi, S.A. The potential for the marine biotechnology industry. Trends Future Chall. US Natl. Ocean Coast. Policy 1998, 101–104. [Google Scholar]
- Dictionary of Marine Natural Products with CD-ROM; Blunt, J.W.; Munro, M.H.G. (Eds.) Chapman and Hall/CRC: Boca Raton, FL, USA, 2008; p. 2534.
- Müller, W.; Yamazaki, Z.; Sögtrop, H.H.; Zahn, R.K. Action of 1-β-D-arabinofuranosylcytosine on mammalian tumor cells. 2. Inhibition of mammalian and oncogenic viral polymerases. Eur. J. Cancer 1972, 8, 421–428. [Google Scholar] [PubMed]
- Müller, W.E.G.; Zahn, R.K.; Beyer, R.; Falke, D. 9-β-D-arabinofuranosyladenine as a tool to study Herpes simplex virus DNA replication in vitro. Virology 1977, 76, 787–796. [Google Scholar]
- Mebs, D.; Weiler, I.; Heinke, H.F. Bioactive proteins from marine sponges: Screening of sponge extracts for hemagglutinating, hemolytic, ichthyotoxic and lethal properties and isolation and characterization of hemagglutinins. Toxicon 1985, 23, 955–962. [Google Scholar]
- Pfeifer, K.; Haasemann, M.; Gamulin, V.; Bretting, H.; Fahrenholz, F.; Müller, W.E.G. S-type lectins occur also in invertebrates: Unusual subunit composition and high conservation of the carbohydrate recognition domain in the lectin genes from the marine sponge Geodia cydonium. Glycobiology 1993, 3, 179–184. [Google Scholar]
- Ghosh, D.; Saha, M.; Sana, B.; Mukherjee, J. Marine enzymes. Adv. Biochem. Engin/Biotechnol. 2005, 96, 189–218. [Google Scholar] [CrossRef]
- Wiens, M.; Schröder, H.C.; Korzhev, M.; Wang, X.H.; Batel, R.; Müller, W.E.G. Inducible ASABF-type antimicrobial peptide from the sponge Suberites domuncula: microbicidal and hemolytic activity in vitro and toxic effect on molluscs in vivo. Mar. Drugs 2011, 9, 1969–1994. [Google Scholar]
- Wang, G. Antimicrobial Peptides: Discovery, Design and Novel Therapeutic Strategies; CABI: Oxfordshire, UK, 2010. [Google Scholar]
- Müller, W.E.G. Sponges (Porifera); Springer Press: Berlin, Germany, 2003. [Google Scholar]
- Müller, W.E.G.; Diehl-Seifert, B.; Sobel, C.; Bechtold, A.; Kljajic, Z.; Dorn, A. Sponge secondary metabolites: biochemical and ultrastructural localization of the antimitotic agent avarol in Dysidea avara. J. Histochem. Cytochem. 1986, 34, 1687–1690. [Google Scholar]
- Ebel, R.; Brenzinger, M.; Kunze, A.; Gross, H.J.; Proksch, P. Wound activation of protoxins in marine sponge Aplysina aerophoba. J. Chem. Ecol. 1997, 23, 1451–1462. [Google Scholar]
- Biesalski, H.K.; Doepner, G.; Tzimas, G.; Gamulin, V.; Schröder, H.C.; Batel, R.; Nau, H.; Müller, W.E.G. Modulation of myb gene expression in sponges by retinoic acid. Oncogene 1992, 7, 1765–1774. [Google Scholar]
- Müller, W.E.G.; Binder, M.; von Lintig, J.; Guo, Y.W.; Wang, X.H.; Kaandorp, J.A.; Wiens, M.; Schröder, H.C. Interaction of the retinoic acid signaling pathway with spicule formation in the marine sponge Suberites domuncula through activation of bone morphogenetic protein-1. Biochim. Biophys. Acta 2011, 1810, 1178–1194. [Google Scholar]
- Wiens, M.; Batel, R.; Korzhev, M.; Müller, W.E.G. Retinoid X receptor and retinoic acid response in the marine sponge Suberites domuncula. J. Exp. Biol. 2003, 206, 3261–3271. [Google Scholar]
- Richet, C. De l'action toxique de la subéritine (extrait aqueux de Suberites domuncula). C. R. Séanc. Soc. Biol. 1906, 61, 598–600. [Google Scholar]
- Cariello, L.; Zanetti, L. Suberitine, the toxic protein from the marine sponge, Suberites domuncula. Comp. Biochem. Physiol. 1979, 64C, 15–19. [Google Scholar]
- Arndt, W. Die Spongien als kryptotoxische Tiere. Zool. Jb. Abt. Allg. Zool. 1928, 45, 343–360. [Google Scholar]
- Cariello, L.; Salvato, B.; Jori, G. Partial characterization of suberitine, the neurotoxic protein purified from Suberites domuncula. Comp. Biochem. Physiol. 1980, 67B, 337–344. [Google Scholar]
- Cariello, L.; Tosti, E.; Zanetti, L. The hemolytic activity of suberitine. Comp. Biochem. Physiol. 1982, 73C, 91–93. [Google Scholar]
- Cariello, L.; Salvato, B.; Jori, G. The role of the cysteinyl and one of the tryptophyl residues in the neurotoxic action of suberitine. Experientia 1981, 37, 801–803. [Google Scholar]
- Cariello, L.; Zanetti, L. A blue carotenoprotein from the marine sponge Suberites domuncula: purification and properties. Mar. Biol. 1981, 62, 151–155. [Google Scholar]
- Giuliano, G.; Al-Babili, S.; von Lintig, J. Carotenoid oxygenases: Cleave it or leave it. Trends Plant Sci. 2003, 8, 145–149. [Google Scholar]
- Rehorek, M.; Heyn, M.P. Binding of all-trans-retinal to the purple membrane. Evidence for cooperativity and determination of the extinction coefficient. Biochemistry 1979, 18, 4977–4983. [Google Scholar]
- Edwards, A.J.; Nguyen, C.H.; You, C.S.; Swanson, J.E.; Emenhiser, C.; Parker, R.S. α- and β-Carotene from a commercial carrot puree are more bioavailable to humans than from boiled-mashed carrots, as determined using an extrinsic stable isotope reference method. J. Nutr. 2002, 132, 159–167. [Google Scholar] [PubMed]
- Farrens, D.L.; Khorana, H.G. Structure and function in rhodopsin. Measurement of the rate of metarhodopsin II decay by fluorescence spectroscopy. J. Biol. Chem. 1995, 270, 5073–5076. [Google Scholar]
- Ottolenghi, M.; Sheves, M. The photophysics and photochemistry of retinal proteins. Isr. J. Chem. 1995, 35, 193–513. [Google Scholar]
- Jang, D.J.; el-Sayed, M.A. Tryptophan fluorescence quenching as a monitor for the protein conformation changes occurring during the photocycle of bacteriorhodopsin under different perturbations. Proc. Natl. Acad. Sci. USA 1989, 86, 5815–5819. [Google Scholar]
- Wiens, M.; Perović-Ottstadt, S.; Müller, I.M.; Müller, W.E.G. Allograft rejection in the mixed cell reaction system of the demosponge Suberites domuncula is controlled by differential expression of apoptotic genes. Immunogenetics 2004, 56, 597–610. [Google Scholar]
- Kiefer, C.; Hessel, S.; Lampert, J.M.; Vogt, K.; Lederer, M.O.; Breithaupt, D.E.; von Lintig, J. Identification and characterization of a mammalian enzyme catalyzing the asymmetric oxidative cleavage of provitamin A. J. Biol. Chem. 2001, 276, 14110–14116. [Google Scholar]
- Sponge sequence database found under SpongeBase. Available online: https://octavia.vk.medizin.uni-mainz.de/login.cgi (accessed on 15 August 2010).
- Amir, H.; Karas, M.; Giat, J.; Danilenko, M.; Levy, R.; Yermiahu, T.; Levy, J.; Sharoni, Y. Lycopene and 1,25-dihydroxyvitamin D3 cooperate in the inhibition of cell cycle progression and induction of differentiation in HL-60 leukemic cells. Nutr. Cancer 1999, 33, 105–112. [Google Scholar]
- von Lintig, J.; Vogt, K. Filling the gap in vitamin A research. Molecular identification of an enzyme cleaving β-carotene to retinal. J. Biol. Chem. 2000, 275, 11915–11920. [Google Scholar]
- Liaaen-Jensen, S.; Renstrøm, B.; Ramdahl, T.; Hallenstvet, M.; Bergquist, P. Carotenoids of marine sponges. Biochem. Syst. Ecol. 1982, 10, 167–174. [Google Scholar]
- Müller, W.E.G.; Kasueske, M.; Wang, X.H.; Schröder, H.C.; Wang, Y.; Pisignano, D.; Wiens, M. Luciferase a light source for the silica-based optical waveguides (spicules) in the demosponge Suberites domuncula. Cell Mol. Life. Sci. 2009, 66, 537–552. [Google Scholar]
- Müller, W.E.G.; Wang, X.H.; Schröder, H.C.; Korzhev, M.; Grebenjuk, V.A.; Markl, J.S.; Jochum, K.P.; Pisignano, D.; Wiens, M. A cryptochrome-based photosensory system in the siliceous sponge Suberites domuncula (Demospongiae). FEBS J. 2010, 277, 1182–1201. [Google Scholar]
- Rangarajan, K. Light penetration in the inshore waters of Porto Novo. Proc. Indian Acad. Sci 1959, 49B, 271–279. [Google Scholar]
- Simkin, A.J.; Underwood, B.A.; Auldridge, M.; Loucas, H.M.; Shibuya, K.; Schmelz, E.; Clark, D.G.; Klee, H.J. Circadian regulation of the PhCCD1 carotenoid cleavage dioxygenase controls emission of β-ionone, a fragrance volatile of Petunia flowers. Plant Physiol. 2004, 136, 3504–3514. [Google Scholar]
- LePennec, G.; Perović, S.; Ammar, M.S.A.; Grebenjuk, V.A.; Steffen, R.; Müller, W.E.G. Cultivation of primmorphs from the marine sponge Suberites domuncula: morphogenetic potential of silicon and iron. J. Biotechnol. 2003, 100, 93–108. [Google Scholar]
- Müller, W.E.G.; Wiens, M.; Batel, R.; Steffen, R.; Schröder, H.C.; Borojevic, R.; Custodio, M.R. Establishment of a primary cell culture from a sponge: primmorphs from Suberites domuncula. Mar. Ecol. Progr. Ser. 1999, 178, 205–219. [Google Scholar]
- Wiens, M.; Wang, X.; Unger, A.; Schröder, H.C.; Grebenjuk, V.A.; Pisignano, D.; Jochum, K.P.; Müller, W.E.G. Flashing light signaling circuit in sponges: Endogenous light generation after tissue ablation in Suberites domuncula. J. Cell Biochem. 2010, 111, 1377–1389. [Google Scholar]
- Müller, W.E.G.; Müller, I.; Pondeljak, V.; Kurelec, B.; Zahn, R.K. Species-specific aggregation factor in sponges. Isolation, purification and characterization of the aggregation factor from Suberites domuncula. Differentiation 1978, 10, 45–53. [Google Scholar] [CrossRef]
- Determann, H. Gel Chromatography-Gel Filtration, 2nd ed; Springer Press: Berlin, Germany, 1969. [Google Scholar]
- Kljajic, Z.; Schröder, H.C.; Rottmann, M.; Cuperlovic, M.; Movesian, M.; Uhlenbruck, G.; Gasic, M.; Zahn, R.K.; Müller, W.E.G. A D-mannose specific lectin from Gerardia savaglia that inhibits nucleocytoplasmic transport of mRNA. Eur. J. Biochem. 1987, 169, 97–104. [Google Scholar]
- Horwitz, J.; Heller, J. Modification of tryptophan residues in retinol-binding protein and prealbumin with 2-hydroxy-5-nitrobenzyl bromide. Effects of the modification of the protein-retinol and protein-protein interaction. J. Biol. Chem. 1974, 249, 7181–7185. [Google Scholar] [PubMed]
- Gonzalez-Fernandez, F.; Bevilacqua, T.; Lee, K.I.; Chandrashekar, R.; Hsu, L.; Garlipp, M.A.; Griswold, J.B.; Crouch, R.K.; Ghosh, D. Retinol-binding site in interphotoreceptor retinoid-binding protein (IRBP): A novel hydrophobic cavity. Invest. Ophthalmol. Vis. Sci. 2009, 50, 5577–5586. [Google Scholar]
- Zhang, X.; Le Pennec, G.; Steffen, R.; Müller, W.E.G.; Zhang, W. Application of a MTT assay for screening nutritional factors in growth media of primary sponge cell culture. Biotechnol. Progr. 2004, 20, 151–155. [Google Scholar]
- Eckert, C.; Schröder, H.C.; Brandt, D.; Perovic-Ottstadt, S.; Müller, W.E.G. A histochemical and electron microscopic analysis of the spiculogenesis in the demosponge Suberites domuncula. J. Histochem. Cytochem. 2006, 54, 1031–1040. [Google Scholar]
- BLAST Assembled RefSeq Genomes. Available online: http://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 31 January 2011).
- FASTA Assembled RefSeq Genomes. Available online: http://www.ebi.ac.uk/fasta33/ (accessed on 31 January 2011).
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar]
- Motif scan analysis using the internet database. Available online: http://myhits.isbsib.ch/cgi-bin/motif_scan (accessed on 31 January 2011).
- Nicholas, K.B.; Nicholas, H.B.; Deerfield, D.W. GeneDoc: Analysis and visualization of genetic variation. EMBnet News 1997, 4, 1–4. [Google Scholar]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, 2002–2007. [Google Scholar]
- Compton, S.; Jones, C.G. Mechanism of dye response and interference in the Bradford protein assay. Anal. Biochem. 1985, 151, 369–374. [Google Scholar]
- Sachs, L. Angewandte Statistik; Springer: Berlin, Germany, 1984; p. 242. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Müller, W.E.G.; Wang, X.; Binder, M.; Lintig, J.v.; Wiens, M.; Schröder, H.C. Differential Expression of the Demosponge (Suberites domuncula) Carotenoid Oxygenases in Response to Light: Protection Mechanism Against the Self-Produced Toxic Protein (Suberitine). Mar. Drugs 2012, 10, 177-199. https://doi.org/10.3390/md10010177
Müller WEG, Wang X, Binder M, Lintig Jv, Wiens M, Schröder HC. Differential Expression of the Demosponge (Suberites domuncula) Carotenoid Oxygenases in Response to Light: Protection Mechanism Against the Self-Produced Toxic Protein (Suberitine). Marine Drugs. 2012; 10(1):177-199. https://doi.org/10.3390/md10010177
Chicago/Turabian StyleMüller, Werner E. G., Xiaohong Wang, Michael Binder, Johannes von Lintig, Matthias Wiens, and Heinz C. Schröder. 2012. "Differential Expression of the Demosponge (Suberites domuncula) Carotenoid Oxygenases in Response to Light: Protection Mechanism Against the Self-Produced Toxic Protein (Suberitine)" Marine Drugs 10, no. 1: 177-199. https://doi.org/10.3390/md10010177
APA StyleMüller, W. E. G., Wang, X., Binder, M., Lintig, J. v., Wiens, M., & Schröder, H. C. (2012). Differential Expression of the Demosponge (Suberites domuncula) Carotenoid Oxygenases in Response to Light: Protection Mechanism Against the Self-Produced Toxic Protein (Suberitine). Marine Drugs, 10(1), 177-199. https://doi.org/10.3390/md10010177








