Comparative Proteomic Analysis of tPVAT during Ang II Infusion
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Animals and Experimental Protocol
2.3. Adipose Tissue Transplantation
2.4. H&E Staining
2.5. Immunohistochemical Analysis
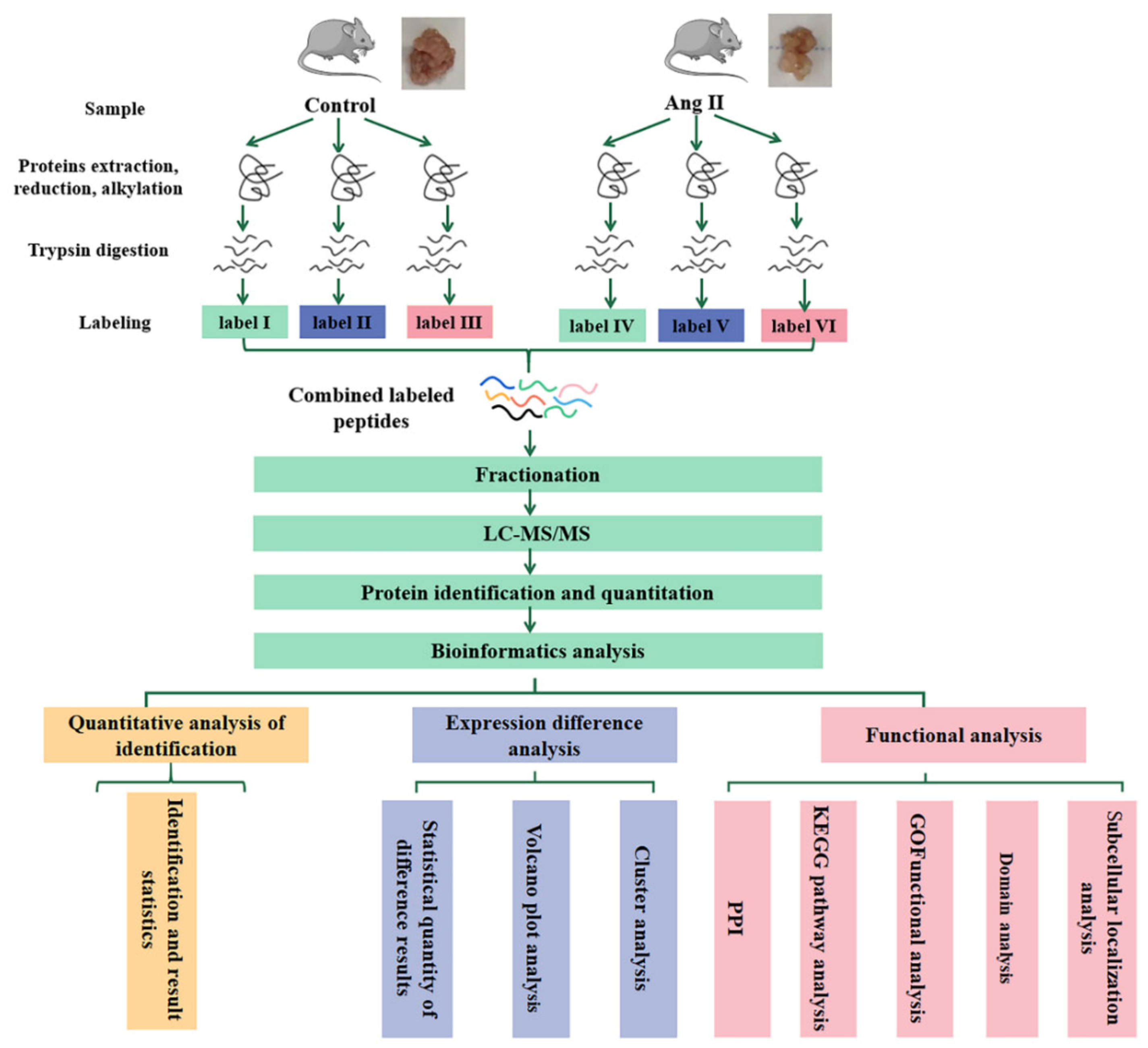
2.6. Quantitative Proteomic Profiling by Tandem Mass Tag (TMT) Technology
2.7. Identification and Quantitation of Proteins
2.8. Bioinformatic Analysis
2.8.1. Cluster Analysis
2.8.2. Subcellular Localization
2.8.3. GO Analysis
2.8.4. KEGG Pathway
2.8.5. PPI Analysis
2.9. Statistical Analysis
3. Results
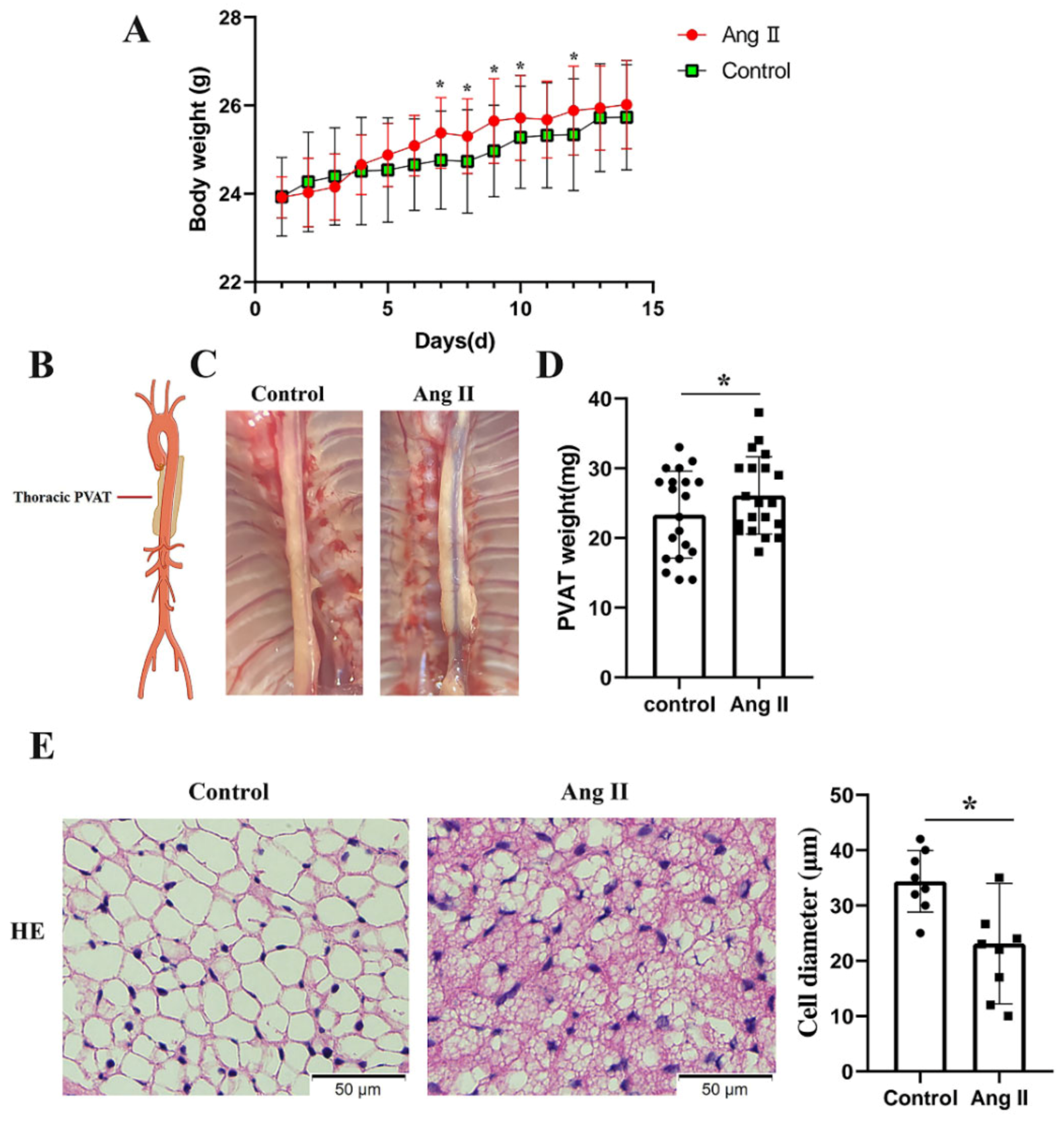
3.1. Pathological Conversion of tPVAT after Ang II Infusion
3.2. Molecular Changes during tPVAT Pathogenesis Analyzed by LC-MS/MS
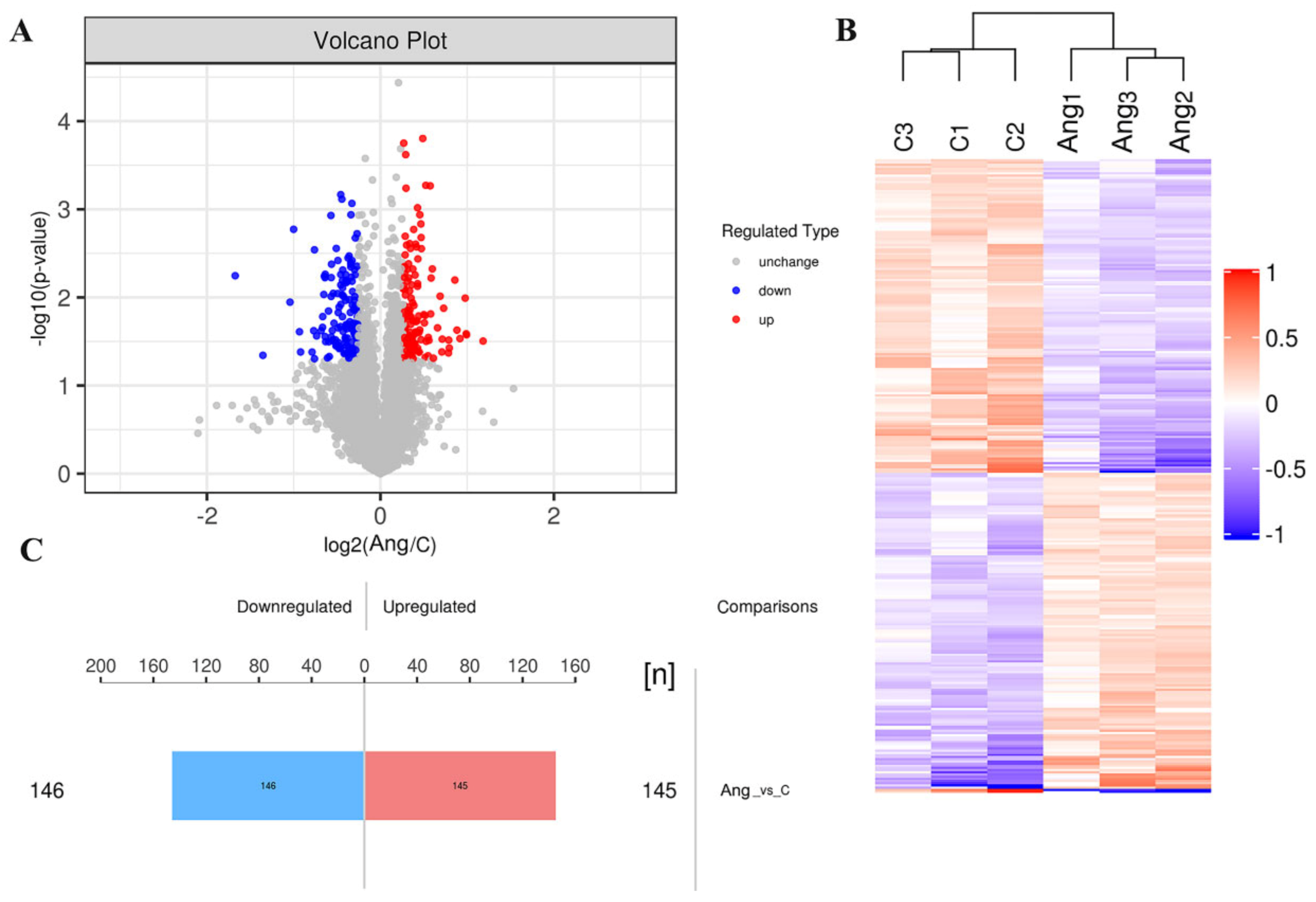
3.3. Identification of DEPs
3.4. Bioinformatics Analysis of the Altered Proteins during Ang II Infusion
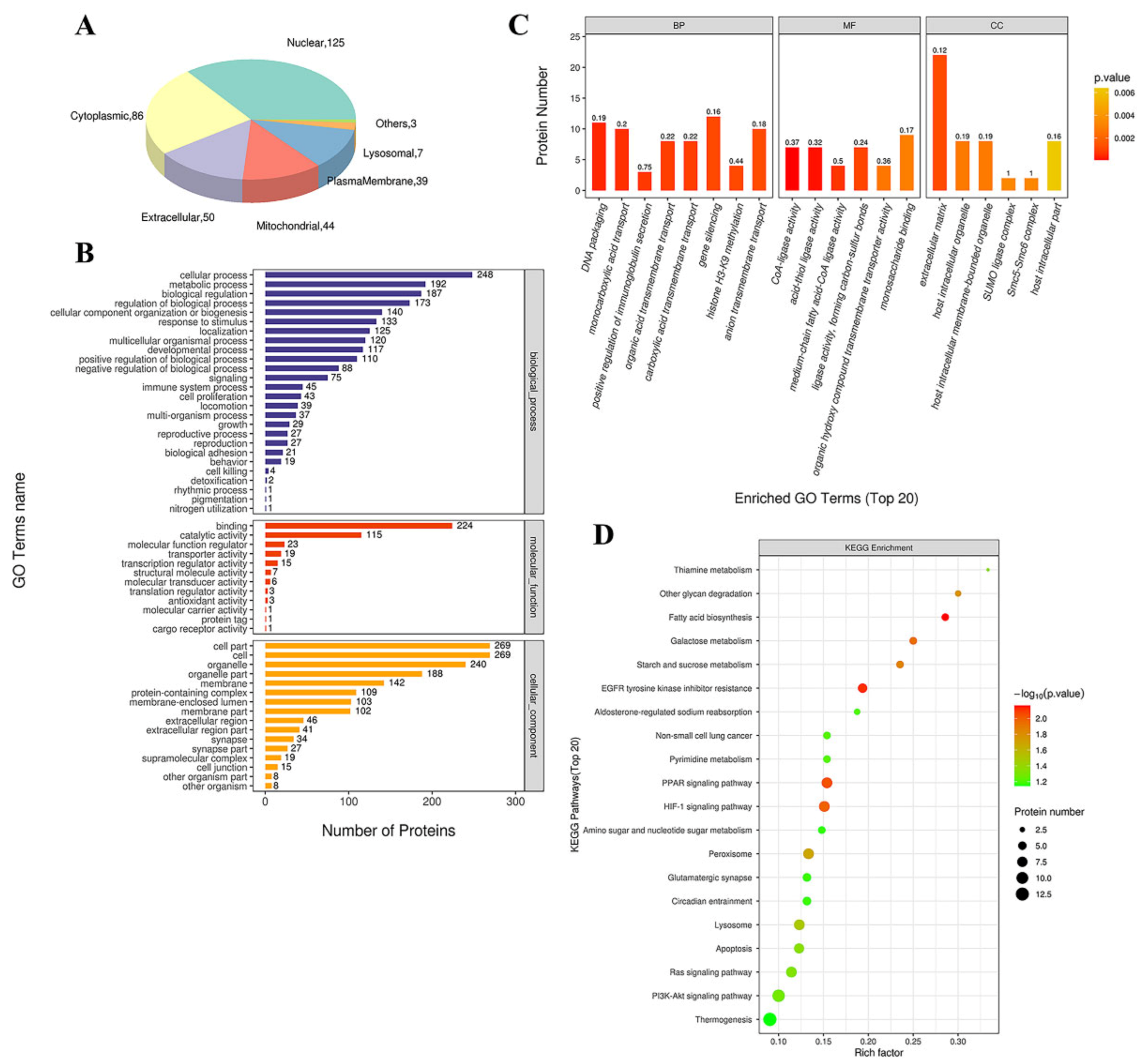
3.4.1. GO Functional Annotation and Analysis
3.4.2. KEGG Pathway Analysis
3.4.3. PPI Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bourmoum, M.; Charles, R.; Claing, A. The GTPase ARF6 controls ROS production to mediate angiotensin II-induced vascular smooth muscle cell proliferation. PLoS ONE 2016, 11, e0148097. [Google Scholar] [CrossRef] [PubMed]
- Nour-Eldine, W.; Ghantous, C.M.; Zibara, K.; Dib, L.; Issaa, H.; Itani, H.A.; El-Zein, N.; Zeidan, A. Adiponectin attenuates angiotensin II-induced vascular smooth muscle cell remodeling through nitric oxide and the RhoA/ROCK pathway. Front. Pharmacol. 2016, 7, 86. [Google Scholar] [CrossRef] [PubMed]
- Dai, F.; Qi, Y.; Guan, W.; Meng, G.; Liu, Z.; Zhang, T.; Yao, W. RhoGDI stability is regulated by SUMOylation and ubiquitination via the AT1 receptor and participates in Ang II-induced smooth muscle proliferation and vascular remodeling. Atherosclerosis 2019, 288, 124–136. [Google Scholar] [CrossRef] [PubMed]
- Siegel-Axel, D.I.; Häring, H.U. Perivascular adipose tissue: An unique fat compartment relevant for the cardiometabolic syndrome. Rev. Endocr. Metab. Disord. 2016, 17, 51–60. [Google Scholar] [CrossRef]
- Fernández-Alfonso, M.S.; Gil-Ortega, M.; Aranguez, I.; Souza, D.; Dreifaldt, M.; Somoza, B.; Dashwood, M.R. Role of PVAT in coronary atherosclerosis and vein graft patency: Friend or foe? Br. J. Pharmacol. 2017, 174, 3561–3572. [Google Scholar] [CrossRef]
- Qi, X.Y.; Qu, S.L.; Xiong, W.H.; Rom, O.; Chang, L.; Jiang, Z.S. Perivascular adipose tissue (PVAT) in atherosclerosis: A double-edged sword. Cardiovasc. Diabetol. 2018, 17, 134. [Google Scholar] [CrossRef]
- Langin, D.; Arner, P. Importance of TNF alpha and neutral lipases in human adipose tissue lipolysis. Trends. Endocrinol. Metab. 2006, 17, 314–320. [Google Scholar] [CrossRef]
- Cao, H. Adipocytokines in obesity and metabolic disease. J. Endocrinol. 2014, 220, T47–T59. [Google Scholar] [CrossRef]
- Cannon, B.; Nedergaard, J. Brown adipose tissue: Function and physiological significance. Physiol. Rev. 2004, 84, 277–359. [Google Scholar] [CrossRef]
- Kern, P.A.; Finlin, B.S.; Zhu, B.; Rasouli, N.; McGehee, R.E., Jr.; Westgate, P.M.; Dupont-Versteegden, E.E. The effects of temperature and seasons on subcutaneous white adipose tissue in humans: Evidence for thermogenic gene induction. J. Clin. Endocrinol. Metab. 2014, 99, E2772–E2779. [Google Scholar] [CrossRef]
- Lin, J.Z.; Martagón, A.J.; Cimini, S.L.; Gonzalez, D.D.; Tinkey, D.W.; Biter, A.; Baxter, J.D.; Webb, P.; Gustafsson, J.Å.; Hartig, S.M.; et al. Pharmacological activation of thyroid hormone receptors elicits a functional conversion of white to brown fat. Cell. Rep. 2015, 13, 1528–1537. [Google Scholar] [CrossRef]
- Wu, J.; Boström, P.; Sparks, L.M.; Ye, L.; Choi, J.H.; Giang, A.H.; Khandekar, M.; Virtanen, K.A.; Nuutila, P.; Schaart, G.; et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 2012, 150, 366–376. [Google Scholar] [CrossRef]
- Uysal, K.T.; Wiesbrock, S.M.; Marino, M.W.; Hotamisligil, G.S. Protection from obesity-induced insulin resistance in mice lacking TNF-alpha function. Nature 1997, 389, 610–614. [Google Scholar] [CrossRef]
- Janke, J.; Engeli, S.; Gorzelniak, K.; Luft, F.C.; Sharma, A.M. Mature adipocytes inhibit in vitro differentiation of human preadipocytes via angiotensin type 1 receptors. Diabetes 2002, 51, 1699–1707. [Google Scholar] [CrossRef]
- Than, A.; Leow, M.K.; Chen, P. Control of adipogenesis by the autocrine interplays between angiotensin 1-7/Mas receptor and angiotensin II/AT1 receptor signaling pathways. J. Biol. Chem. 2013, 288, 15520–15531. [Google Scholar] [CrossRef]
- Sysoeva, V.Y.; Ageeva, L.V.; Tyurin-Kuzmin, P.A.; Sharonov, G.V.; Dyikanov, D.T.; Kalinina, N.I.; Tkachuk, V.A. Local angiotensin II promotes adipogenic differentiation of human adipose tissue mesenchymal stem cells through type 2 angiotensin receptor. Stem. Cell. Res. 2017, 25, 115–122. [Google Scholar] [CrossRef]
- Wiśniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef]
- Francois, A.A.; Obasanjo-Blackshire, K.; Clark, J.E.; Boguslavskyi, A.; Holt, M.R.; Parker, P.J.; Marber, M.S.; Heads, R.J. Loss of Protein Kinase Novel 1 (PKN1) is associated with mild systolic and diastolic contractile dysfunction, increased phospholamban Thr17 phosphorylation, and exacerbated ischaemia-reperfusion injury. Cardiovasc. Res. 2018, 114, 138–157. [Google Scholar] [CrossRef]
- Fang, Y.; Liao, G.; Yu, B. LSD1/KDM1A inhibitors in clinical trials: Advances and prospects. J. Hematol. Oncol. 2019, 12, 129. [Google Scholar] [CrossRef]
- Fairbanks, A.J. Chemoenzymatic synthesis of glycoproteins. Curr. Opin. Chem. Biol. 2019, 53, 9–15. [Google Scholar] [CrossRef]
- Farmer, S.R. Molecular determinants of brown adipocyte formation and function. Genes. Dev. 2008, 22, 1269–1275. [Google Scholar] [CrossRef] [PubMed]
- Moretti, F.; Bergman, P.; Dodgson, S.; Marcellin, D.; Claerr, I.; Goodwin, J.M.; DeJesus, R.; Kang, Z.; Antczak, C.; Begue, D.; et al. TMEM41B is a novel regulator of autophagy and lipid mobilization. EMBO. Rep. 2018, 19, e45889. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.H.; Won, J.H.; Cheng, N.; Lau, L.F. The matricellular protein CCN1 in tissue injury repair. J. Cell. Commun. Signal. 2018, 12, 273–279. [Google Scholar] [CrossRef] [PubMed]
- Ren, B.; Liu, M.; Ni, J.; Tian, J. Role of selenoprotein f in protein folding and secretion: Potential involvement in human disease. Nutrients 2018, 10, 1619. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.H.; Yin, R.X.; Chen, W.X.; Cao, X.L.; Chen, Y.M. Association between the TIMD4-HAVCR1 variants and serum lipid levels, coronary heart disease and ischemic stroke risk and atorvastatin lipid-lowering efficacy. Biosci. Rep. 2018, 38, BSR20171058. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Shah, A.P.; Li, Z.; Li, M.; Tamashiro, K.L.; Baraban, J.M. Genetic inactivation of the translin/trax microRNA-degrading enzyme phenocopies the robust adiposity induced by Translin (Tsn) deletion. Mol. Metab. 2020, 40, 101013. [Google Scholar] [CrossRef] [PubMed]
- Hung, C.S.; Lin, J.C. Alternatively spliced MBNL1 isoforms exhibit differential influence on enhancing brown adipogenesis. Biochim. Biophys. Acta. Gene. Regul. Mech. 2020, 1863, 194437. [Google Scholar] [CrossRef]
- Zheng, X.; Ren, B.; Li, X.; Yan, H.; Xie, Q.; Liu, H.; Zhou, J.; Tian, J.; Huang, K. Selenoprotein F knockout leads to glucose and lipid metabolism disorders in mice. J. Biol. Inorg. Chem. 2020, 25, 1009–1022. [Google Scholar] [CrossRef]
- Cheng, C.K.; Bakar, H.A.; Gollasch, M.; Huang, Y. Perivascular adipose tissue: The sixth man of the cardiovascular system. Cardiovasc. Drugs. Ther. 2018, 32, 481–502. [Google Scholar] [CrossRef]
- Miao, C.Y.; Li, Z.Y. The role of perivascular adipose tissue in vascular smooth muscle cell growth. Br. J. Pharmacol. 2012, 165, 643–658. [Google Scholar] [CrossRef]
- Campbell, D.J.; Habener, J.F. Cellular localization of angiotensinogen gene expression in brown adipose tissue and mesentery: Quantification of messenger ribonucleic acid abundance using hybridization in situ. Endocrinology. 1987, 121, 1616–1626. [Google Scholar] [CrossRef]
- Cassis, L.A.; Police, S.B.; Yiannikouris, F.; Thatcher, S.E. Local adipose tissue renin-angiotensin system. Curr. Hypertens. Rep. 2008, 10, 93–98. [Google Scholar] [CrossRef]
- Hanzu, F.A.; Musri, M.M.; Sánchez-Herrero, A.; Claret, M.; Esteban, Y.; Kaliman, P.; Gomis, R.; Párrizas, M. Histone demethylase KDM1A represses inflammatory gene expression in preadipocytes. Obesity 2013, 21, E616–E625. [Google Scholar] [CrossRef]
- Brina, D.; Miluzio, A.; Ricciardi, S.; Clarke, K.; Davidsen, P.K.; Viero, G.; Tebaldi, T.; Offenhäuser, N.; Rozman, J.; Rathkolb, B.; et al. eIF6 coordinates insulin sensitivity and lipid metabolism by coupling translation to transcription. Nat. Commun. 2015, 6, 8261. [Google Scholar] [CrossRef]
- Xiong, Y.; Wang, E.; Huang, Y.; Guo, X.; Yu, Y.; Du, Q.; Ding, X.; Sun, Y. Inhibition of lysine-specific demethylase-1 (LSD1/KDM1A) promotes the adipogenic differentiation of hESCs through H3K4 methylation. Stem. Cell. Rev. Rep. 2016, 12, 298–304. [Google Scholar] [CrossRef]
- Liu, C.; Wang, J.; Wei, Y.; Zhang, W.; Geng, M.; Yuan, Y.; Chen, Y.; Sun, Y.; Chen, H.; Zhang, Y.; et al. Fat-specific knockout of Mecp2 upregulates slpi to reduce obesity by enhancing browning. Diabetes 2020, 69, 35–47. [Google Scholar] [CrossRef]
- Reilly, S.M.; Hung, C.W.; Ahmadian, M.; Zhao, P.; Keinan, O.; Gomez, A.V.; DeLuca, J.H.; Dadpey, B.; Lu, D.; Zaid, J.; et al. Catecholamines suppress fatty acid re-esterification and increase oxidation in white adipocytes via STAT3. Nat. Metab. 2020, 2, 620–634. [Google Scholar] [CrossRef]
- Xu, Y.; Wang, N.; Tan, H.Y.; Li, S.; Zhang, C.; Zhang, Z.; Feng, Y. Panax notoginseng saponins modulate the gut microbiota to promote thermogenesis and beige adipocyte reconstruction via leptin-mediated AMPKα/STAT3 signaling in diet-induced obesity. Theranostics 2020, 10, 11302–11323. [Google Scholar] [CrossRef]
- Lakhani, H.V.; Zehra, M.; Pillai, S.S.; Puri, N.; Shapiro, J.I.; Abraham, N.G.; Sodhi, K. Beneficial role of HO-1-SIRT1 axis in attenuating angiotensin II-induced adipocyte dysfunction. Int. J. Mol. Sci. 2019, 20, 3205. [Google Scholar] [CrossRef]
- Goel, H.; Melot, J.; Krinock, M.D.; Kumar, A.; Nadar, S.K.; Lip, G.Y.H. Heart-type fatty acid-binding protein: An overlooked cardiac biomarker. Ann. Med. 2020, 52, 444–461. [Google Scholar] [CrossRef]
- Zhuang, L.; Mao, Y.; Liu, Z.; Li, C.; Jin, Q.; Lu, L.; Tao, R.; Yan, X.; Chen, K. FABP3 deficiency exacerbates metabolic derangement in cardiac hypertrophy and heart failure via PPARα pathway. Front. Cardiovasc. Med. 2021, 8, 722908. [Google Scholar] [CrossRef] [PubMed]
- Betz, M.J.; Enerbäck, S. Targeting thermogenesis in brown fat and muscle to treat obesity and metabolic disease. Nat. Rev. Endocrinol. 2018, 14, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Eugene Chen, Y. Editorial: The Yin and Yang of perivascular adipose tissue in vascular disease. Cardiovasc. Drugs. Ther. 2018, 32, 477–479. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Qi, Y.; Dai, F.; Gu, J.; Yao, W. PVAT: An important guardian of the cardiovascular system. Histol. Histopathol. 2020, 35, 779–787. [Google Scholar]
- Dalen, K.T.; Schoonjans, K.; Ulven, S.M.; Weedon-Fekjaer, M.S.; Bentzen, T.G.; Koutnikova, H.; Auwerx, J.; Nebb, H.I. Adipose tissue expression of the lipid droplet-associating proteins S3-12 and perilipin is controlled by peroxisome proliferator-activated receptor-gamma. Diabetes 2004, 53, 1243–1252. [Google Scholar] [CrossRef]
- Smith, C.E.; Ordovás, J.M. Update on perilipin polymorphisms and obesity. Nutr. Rev. 2012, 70, 611–621. [Google Scholar] [CrossRef][Green Version]
- Westhoff, C.C.; Mrozinski, J.; Riedel, I.; Heid, H.W.; Moll, R. Perilipin 1 is a highly specific marker for adipocytic differentiation in sarcomas with intermediate sensitivity. J. Cancer Res. Clin. Oncol. 2017, 143, 225–232. [Google Scholar] [CrossRef]
- Han, X.; Zhu, J.; Zhang, X.; Song, Q.; Ding, J.; Lu, M.; Sun, S.; Hu, G. Plin4-dependent lipid droplets hamper neuronal mitophagy in the MPTP/p-induced mouse model of Parkinson’s disease. Front. Neurosci. 2018, 12, 397. [Google Scholar] [CrossRef]
- Bernardi, R.E.; Olevska, A.; Morella, I.; Fasano, S.; Santos, E.; Brambilla, R.; Spanagel, R. The inhibition of RasGRF2, but not RasGRF1, alters cocaine reward in mice. J. Neurosci. 2019, 39, 6325–6338. [Google Scholar] [CrossRef]
- Huang, H.; Fan, Y.; Gao, Z.; Wang, W.; Shao, N.; Zhang, L.; Yang, Y.; Zhu, W.; Chen, Z.; Hu, J.; et al. HIF-1α contributes to Ang II-induced inflammatory cytokine production in podocytes. BMC. Pharmacol. Toxicol. 2019, 20, 59. [Google Scholar] [CrossRef]
- Yan, X.; Zhao, R.; Feng, X.; Mu, J.; Li, Y.; Chen, Y.; Li, C.; Yao, Q.; Cai, L.; Jin, L.; et al. Sialyltransferase7A promotes angiotensin II-induced cardiomyocyte hypertrophy via HIF-1α-TAK1 signalling pathway. Cardiovasc. Res. 2020, 116, 114–126. [Google Scholar] [CrossRef]
- Sharma, N.M.; Haibara, A.S.; Katsurada, K.; Nandi, S.S.; Liu, X.; Zheng, H.; Patel, K.P. Central Ang II (Angiotensin II)-mediated sympathoexcitation: Role for HIF-1α (hypoxia-inducible factor-1α) facilitated glutamatergic tone in the paraventricular nucleus of the hypothalamus. Hypertension 2021, 77, 147–157. [Google Scholar] [CrossRef]
- Qi, D.; Wei, M.; Jiao, S.; Song, Y.; Wang, X.; Xie, G.; Taranto, J.; Liu, Y.; Duan, Y.; Yu, B.; et al. Hypoxia inducible factor 1α in vascular smooth muscle cells promotes angiotensin II-induced vascular remodeling via activation of CCL7-mediated macrophage recruitment. Cell Death Dis. 2019, 10, 544. [Google Scholar] [CrossRef]
- Yang, X.; Bao, M.; Fang, Y.; Yu, X.; Ji, J.; Ding, X. STAT3/HIF-1α signaling activation mediates peritoneal fibrosis induced by high glucose. J. Transl. Med. 2021, 19, 283. [Google Scholar] [CrossRef]


| Accession NO. | Gene Name | Protein Name | Accession NO. | Gene Name | Protein Name |
|---|---|---|---|---|---|
| Q6ZQ88 | KDM1A | Lysine-specific histone demethylase 1A | Q9CPV3 | MRPL42 | 39S ribosomal protein L42, mitochondrial |
| P70268 | PKN1 | Serine/threonine-protein kinase N1 | Q8VCW8 | ACSF2 | Medium-chain acyl-CoA ligase ACSF2, mitochondrial |
| Q8BX80 | ENGase | Cytosolic endo-beta-N-acetylglucosaminidase | Q9D0F9 | PGM1 | Phosphoglucomutase-1 |
| O88712 | CtBP1 | C-terminal-binding protein 1 | P53986 | SLC16A1 | Monocarboxylate transporter 1 |
| Q8K1A5 | Tmem41b | Transmembrane protein 41B | Q8BGQ1 | VIPAS39 | Spermatogenesis-defective protein 39 homolog |
| P70392 | RasGRF2 | Ras-specific guanine nucleotide-releasing factor 2 | Q9D2R0 | AACS | Acetoacetyl-CoA synthetase |
| Q61096 | PRTN3 | Myeloblastin | Q9CXR1 | DHRS7 | Dehydrogenase/reductase SDR family member 7 |
| Q8VD66 | ABHD4 | (Lyso)-N-acylphosphatidylethanolamine lipase | Q8K3K7 | AGPAT2 | 1-acyl-sn-glycerol-3-phosphate acyltransferase beta |
| Q6Y7W8 | GIGYF2 | GRB10-interacting GYF protein 2 | Q924N4 | SLC12A6 | Solute carrier family 12 member 6 |
| Q61490 | ALCAM | CD166 antigen | Q922Z0 | DDO | D-aspartate oxidase |
| Q8CHD8 | RAB11FIP3 | Rab11-family interacting protein 3 | Q14DH7 | ACSS3 | Acyl-CoA synthetase short-chain family member 3, mitochondrial |
| Q8R054 | SRPX2 | Sushi repeat-containing protein SRPX2 | Q00898 | SERPINA1E | Alpha-1-antitrypsin 1–5 |
| Q3UVG3 | FAM91A1 | Protein FAM91A1 | Q9JJF9 | SPPL2a | Signal peptide peptidase-like 2A |
| O89017 | LGMN | Legumain | Q9CS72 | FILIP1 | Filamin-A-interacting protein 1 |
| Q8R1R3 | StarD7 | StAR-related lipid transfer protein 7, mitochondrial | Q9QXG4 | ACSS2 | Acetyl-coenzyme A synthetase, cytoplasmic |
| Q924W5 | Smc6 | Structural maintenance of chromosomes protein 6 | P28063 | PSMB8 | Proteasome subunit beta type-8 |
| Q64516 | GK | Glycerol kinase | Q5RKZ7 | MOCS1 | Molybdenum cofactor biosynthesis protein 1 |
| Q9D023 | MPC2 | Mitochondrial pyruvate carrier 2 | P67984 | Rpl22 | 60S ribosomal protein L22 |
| E9Q1P8 | IRF2BP2 | Interferon regulatory factor 2-binding protein 2 | Q99KF1 | TMED9 | Transmembrane emp24 domain-containing protein 9 |
| P12242 | UPC1 | Mitochondrial brown fat uncoupling protein 1 | Q9CY73 | MRPL44 | 39S ribosomal protein L44, mitochondrial |
| P11404 | FABP3 | Fatty acid-binding protein, heart | Q8BHJ6 | SERINC5 | Serine incorporator 5 |
| P09470 | ACE | Angiotensin-converting enzyme | P70699 | GAA | Lysosomal alpha-glucosidase |
| Q8BMB3 | eIF4E2 | Eukaryotic translation initiation factor 4E type 2 | Q3U4G3 | XXYLT1 | Xyloside xylosyltransferase 1 |
| Q9D136 | OGFOD3 | 2-oxoglutarate and iron-dependent oxygenase domain-containing protein 3 | Q8C7H1 | MMAA | Methylmalonic aciduria type A homolog, mitochondrial |
| Q62009 | POSTN | Periostin | Q80VW7 | AKNA | Microtubule organization protein AKNA |
| Q78JN3 | Eci3 | Enoyl-CoA delta isomerase 3, peroxisomal | O54940 | BNIP2 | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2 |
| P68433 | H3C1 | Histone H3.1 | Q6PEE2 | CTIF | CBP80/20-dependent translation initiation factor |
| Q5SF07 | IGF2BP2 | Insulin-like growth factor 2 mRNA-binding protein 2 | Q9D4H1 | EXOC2 | Exocyst complex component 2 |
| Q9JM90 | STAP1 | Signal-transducing adaptor protein 1 | Q8CFV9 | RFK | Riboflavin kinase |
| Q80YX1 | TNC | Tenascin | P29595 | NEDD8 | NEDD8 |
| Q9DCE5 | PAK1IP1 | p21-activated protein kinase-interacting protein 1 | Q9Z0P5 | TWF2 | Twinfilin-2 |
| Q8JZR0 | ACSL5 | Long-chain-fatty-acid--CoA ligase 5 | Q9CXJ1 | EARS2 | Probable glutamate--tRNA ligase, mitochondrial |
| Q9CXD9 | LRRc17 | Leucine-rich repeat-containing protein 17 | Q8VHQ9 | ACOT11 | Acyl-coenzyme A thioesterase 11 |
| Q8BWJ3 | PHKA2 | Phosphorylase b kinase regulatory subunit alpha, liver isoform | Q920B9 | SUPT16H | FACT complex subunit SPT16 |
| Q811L6 | MAST4 | Microtubule-associated serine/threonine-protein kinase 4 | Q9DCV4 | RMDN1 | Regulator of microtubule dynamics protein 1 |
| P61202 | COPS2 | COP9 signalosome complex subunit 2 | P06745 | GPI | Glucose-6-phosphate isomerase |
| Q1W617 | SHROOM4 | Protein SHROOM4 | O08600 | EndoG | Endonuclease G, mitochondrial |
| P63158 | HMGB1 | High mobility group protein B1 | O08746 | MATN2 | Matrilin-2 |
| P36536 | SAR1A | GTP-binding protein SAR1A | Q78J03 | MsrB2 | Methionine-R-sulfoxide reductase B2, mitochondrial |
| Q8BJ03 | COX15 | Cytochrome c oxidase assembly protein COX15 homolog | Q99L45 | EIF2S2 | Eukaryotic translation initiation factor 2 subunit 2 |
| Q8BHC0 | LYVE-1 | Lymphatic vessel endothelial hyaluronic acid receptor 1 | P97452 | BOP1 | Ribosome biogenesis protein BOP1 |
| Q8VE11 | MTMR6 | Myotubularin-related protein 6 | O08528 | HK2 | Hexokinase-2 |
| Q91V76 | C11orf54 | Ester hydrolase C11orf54 homolog | Q9Z1T1 | Ap3B1 | AP-3 complex subunit beta-1 |
| Q64008 | Rab34 | Ras-related protein Rab-34 | Q920A5 | SCPEP1 | Retinoid-inducible serine carboxypeptidase |
| P30993 | C5aR1 | C5a anaphylatoxin chemotactic receptor 1 | Q91VU0 | FAM3C | Protein FAM3C |
| Q91WC3 | ACSL6 | Long-chain-fatty-acid--CoA ligase 6 | Q9CQE7 | ERGIC3 | Endoplasmic reticulum-Golgi intermediate compartment protein 3 |
| Q7TPM3 | TRIM17 | E3 ubiquitin-protein ligase TRIM17 | P50580 | PA2G4 | Proliferation-associated protein 2G4 |
| P70122 | SBDS | Ribosome maturation protein SBDS | Q9CZD5 | MTIF3 | Translation initiation factor IF-3, mitochondrial |
| Q8CHG3 | GCC2 | GRIP and coiled-coil domain-containing protein 2 | P01631 | Ig kappa chain V-II region 26–10 | |
| Q80X95 | RRAGA | Ras-related GTP-binding protein A | Q9D125 | MRPS25 | 28S ribosomal protein S25, mitochondrial |
| Q8R3F5 | MCAT | Malonyl-CoA-acyl carrier protein transacylase, mitochondrial | Q5SW19 | CLUH | Clustered mitochondria protein homolog |
| Q99LJ6 | GPX7 | Glutathione peroxidase 7 | Q7TPE5 | SLC7A6OS | Probable RNA polymerase II nuclear localization protein SLC7A6OS |
| O70503 | HSD17B12 | Very-long-chain 3-oxoacyl-CoA reductase | P42227 | STAT3 | Signal transducer and activator of transcription 3 |
| P12382 | PFKL | ATP-dependent 6-phosphofructokinase, liver type | Q9Z0S1 | BPNT-1 | 3′(2′),5′-bisphosphate nucleotidase 1 |
| Q61704 | ITIH3 | Inter-alpha-trypsin inhibitor heavy chain H3 | P09242 | ALPL | Alkaline phosphatase, tissue-nonspecific isozyme |
| P06797 | CTSL | Cathepsin L1 | P12265 | GUSB | Beta-glucuronidase |
| Q80WG5 | LRRC8A | Volume-regulated anion channel subunit LRRC8A | Q61391 | MME | Neprilysin |
| Q9CQV4 | RETREG3 | Reticulophagy regulator 3 | P82198 | TGFBI | Transforming growth factor-beta-induced protein ig-h3 |
| Q8BGR6 | ARL15 | ADP-ribosylation factor-like protein 15 | Q9WUP7 | UCHL5 | Ubiquitin carboxyl-terminal hydrolase isozyme L5 |
| P11276 | FN1 | Fibronectin | P52196 | TST | Thiosulfate sulfurtransferase |
| P70460 | VASP | Vasodilator-stimulated phosphoprotein | Q9QZA0 | CA5B | Carbonic anhydrase 5B, mitochondrial |
| P06339 | H2-T23 | H-2 class I histocompatibility antigen, D-37 alpha chain | Q8QZY9 | SF3B4 | Splicing factor 3B subunit 4 |
| Q65Z40 | Wapl | Wings apart-like protein homolog | P34914 | EPHX2 | Bifunctional epoxide hydrolase 2 |
| P10126 | Eef1a1 | Elongation factor 1-alpha 1 | O54946 | DNAJB6 | DnaJ homolog subfamily B member 6 |
| P54823 | Ddx6 | Probable ATP-dependent RNA helicase DDX6 | Q9CQ86 | MIEN1 | Migration and invasion enhancer 1 |
| Q8BUE4 | Aifm2 | Apoptosis-inducing factor 2 | Q8CG46 | SMC5 | Structural maintenance of chromosomes protein 5 |
| O35143 | ATP5IF1 | ATPase inhibitor, mitochondrial | Q60715 | P4HA1 | Prolyl 4-hydroxylase subunit alpha-1 |
| Q9Z0V8 | Timm17a | Mitochondrial import inner membrane translocase subunit Tim17-A | P15105 | GLUL | Glutamine synthetase |
| Q9CWX9 | Ddx47 | Probable ATP-dependent RNA helicase DDX47 | P08228 | SOD1 | Superoxide dismutase [Cu-Zn] |
| O35344 | Kpna3 | Importin subunit alpha-4 | Q5SWU9 | ACACA | Acetyl-CoA carboxylase 1 |
| O54950 | Prkag1 | 5’-AMP-activated protein kinase subunit gamma-1 | Q91Z96 | BMP2K | BMP-2-inducible protein kinase |
| Q99P65 | Slc29a3 | Equilibrative nucleoside transporter 3 | Q9CYK1 | WARS2 | Tryptophan--tRNA ligase, mitochondrial |
| O09111 | Ndufb11 | NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 11, mitochondrial |
| Accession NO. | Gene Name | Protein Name | Accession NO. | Gene Name | Protein Name |
|---|---|---|---|---|---|
| P18406 | CCN1 | CCN family member 1 | A2RTL5 | RSRC2 | Arginine/serine-rich coiled-coil protein 2 |
| Q09098 | Pate4 | Prostate and testis expressed protein 4 | Q8BL86 | MBLAC2 | Metallo-beta-lactamase domain-containing protein 2 |
| Q9ERR7 | Selenof | Selenoprotein F | Q6ZWY8 | TMSB10 | Thymosin beta-10 |
| O54916 | Reps1 | RalBP1-associated Eps domain-containing protein 1 | Q9D5V6 | Syap1 | Synapse-associated protein 1 |
| Q9CRB2 | NHP2 | H/ACA ribonucleoprotein complex subunit 2 | Q8BK08 | TMEM11 | Transmembrane protein 11, mitochondrial |
| Q6U7R4 | TIMD4 | T-cell immunoglobulin and mucin domain-containing protein 4 | Q80X80 | C2CD2L | Phospholipid transfer protein C2CD2L |
| Q9JKP5 | MBNL1 | Muscleblind-like protein 1 | Q9WV85 | NME3 | Nucleoside diphosphate kinase 3 |
| Q62348 | TSN | Translin | Q9DCL2 | CIAO2A | Cytosolic iron-sulfur assembly component 2A |
| Q8VCH7 | RDH10 | Retinol dehydrogenase 10 | Q61285 | ABCD2 | ATP-binding cassette sub-family D member 2 |
| Q3THF9 | COQ10B | Coenzyme Q-binding protein COQ10 homolog B, mitochondrial | P70333 | HNRNPH2 | Heterogeneous nuclear ribonucleoprotein H2 |
| Q9JK38 | GNPNAT1 | Glucosamine 6-phosphate N-acetyltransferase | Q69ZP3 | PNKD | Probable hydrolase PNKD |
| Q05144 | Rac2 | Ras-related C3 botulinum toxin substrate 2 | O35864 | COPS5 | COP9 signalosome complex subunit 5 |
| P60710 | ACTB | Actin, cytoplasmic 1 | Q61464 | ZNF638 | Zinc finger protein 638 |
| Q9Z140 | CPNE6 | Copine-6 | Q9JM14 | NT5C | 5’(3’)-deoxyribonucleotidase, cytosolic type |
| P47955 | RPLP1 | 60S acidic ribosomal protein P1 | Q9D1L0 | CHCHD2 | Coiled-coil-helix-coiled-coil-helix domain-containing protein 2 |
| P63073 | eIF4E | Eukaryotic translation initiation factor 4E | Q3TMW1 | CCDC102A | Coiled-coil domain-containing protein 102A |
| Q8CH72 | TRIM32 | E3 ubiquitin-protein ligase TRIM32 | Q9Z0V7 | TIMM17B | Mitochondrial import inner membrane translocase subunit Tim17-B |
| P83870 | PHF5A | PHD finger-like domain-containing protein 5A | O88895 | HDAC3 | Histone deacetylase 3 |
| P41317 | MBL2 | Mannose-binding protein C | P97370 | ATP1B3 | Sodium/potassium-transporting ATPase subunit beta-3 |
| P15864 | H1-2 | Histone H1.2 | P26645 | MARCKS | Myristoylated alanine-rich C-kinase substrate |
| Q91WU5 | AS3MT | Arsenite methyltransferase | Q3TEA8 | HP1BP3 | Heterochromatin protein 1-binding protein 3 |
| O54962 | BANF1 | Barrier-to-autointegration factor | Q9D1I2 | CARD19 | Caspase recruitment domain-containing protein 19 |
| O70451 | SLC16A7 | Monocarboxylate transporter 2 | Q3UEZ8 | SLC10A4 | Sodium/bile acid cotransporter 4 |
| P26011 | ITGB7 | Integrin beta-7 | Q99ME9 | GTPBP4 | Nucleolar GTP-binding protein 1 |
| Q9JHH9 | COPZ2 | Coatomer subunit zeta-2 | Q0VGM9 | RTEL1 | Regulator of telomere elongation helicase 1 |
| O54788 | DFFB | DNA fragmentation factor subunit beta | Q9JLQ0 | CD2AP | CD2-associated protein |
| P82348 | SGCG | Gamma-sarcoglycan | O70370 | CTSS | Cathepsin S |
| B1AVZ0 | UPRT | Uracil phosphoribosyltransferase homolog | Q69ZF3 | GBA2 | Non-lysosomal glucosylceramidase |
| Q9JHI7 | EXOSC9 | Exosome complex component RRP45 | Q8K124 | PLEKHO2 | Pleckstrin homology domain-containing family O member 2 |
| Q61207 | PSAP | Prosaposin | P19973 | LSP1 | Lymphocyte-specific protein 1 |
| O35638 | STAG2 | Cohesin subunit SA-2 | P62874 | GNB1 | Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
| Q91YE5 | BAZ2A | Bromodomain adjacent to zinc finger domain protein 2A | Q80VL1 | TDRKH | Tudor and KH domain-containing protein |
| Q9CY57 | Chtop | Chromatin target of PRMT1 protein | Q9Z2D6 | MECP2 | Methyl-CpG-binding protein 2 |
| O55135 | eIF6 | Eukaryotic translation initiation factor 6 | Q61503 | NT5E | 5’-nucleotidase |
| Q8R3B7 | BRD8 | Bromodomain-containing protein 8 | Q8K2Q5 | CHCHD7 | Coiled-coil-helix-coiled-coil-helix domain-containing protein 7 |
| Q8R143 | PTTG1IP | Pituitary tumor-transforming gene 1 protein-interacting protein | Q8BP48 | MetAP1 | Methionine aminopeptidase 1 |
| Q9D2V5 | AAR2 | Protein AAR2 homolog | Q64237 | DBH | Dopamine beta-hydroxylase |
| P28798 | GRN | Progranulin | O35226 | PSMD4 | 26S proteasome non-ATPase regulatory subunit 4 |
| Q9R0Y5 | AK-1 | Adenylate kinase isoenzyme 1 | E9PZM4 | CHD2 | Chromodomain-helicase-DNA-binding protein 2 |
| P03899 | MTND3 | NADH-ubiquinone oxidoreductase chain 3 | P26369 | U2AF2 | Splicing factor U2AF 65 kDa subunit |
| Q8JZS0 | LIN7A | Protein lin-7 homolog A | P56379 | ATP5MPL | ATP synthase subunit ATP5MPL, mitochondrial |
| Q91V12 | ACOT7 | Cytosolic acyl coenzyme A thioester hydrolase | E9Q5C9 | NOLC1 | Nucleolar and coiled-body phosphoprotein 1 |
| Q00724 | RBP4 | Retinol-binding protein 4 | P59279 | RAB2B | Ras-related protein Rab-2B |
| Q78HU7 | GYPC | Glycophorin-C | Q8R332 | NUP58 | Nucleoporin p58/p45 |
| Q9QZB0 | RGS17 | Regulator of G-protein signaling 17 | P97384 | ANXA11 | Annexin A11 |
| Q8BGX2 | TIMM29 | Mitochondrial import inner membrane translocase subunit Tim29 | Q5XG73 | ACBD5 | Acyl-CoA-binding domain-containing protein 5 |
| P97429 | ANXA4 | Annexin A4 | Q60754 | MARCO | Macrophage receptor MARCO |
| O55186 | CD59a | CD59a glycoprotein | A2AJB2 | TMEM141 | Transmembrane protein 141 |
| Q8BLV3 | SLC9A7 | Sodium/hydrogen exchanger 7 | Q8K2I4 | MANBA | Beta-mannosidase |
| Q0VBL1 | TIGD2 | Tigger transposable element-derived protein 2 | P20444 | PRKCA | Protein kinase C alpha type |
| O09117 | SYPL1 | Synaptophysin-like protein 1 | P43276 | H1.5 | Histone H1.5 |
| Q9WTR5 | CDH13 | Cadherin-13 | Q8CGN5 | PLIN1 | Perilipin-1 |
| Q8VCF0 | MAVS | Mitochondrial antiviral-signaling protein | Q63844 | MAPK3 | Mitogen-activated protein kinase 3 |
| O88492 | PLIN4 | Perilipin-4 | Q3UMY5 | EML4 | Echinoderm microtubule-associated protein-like 4 |
| Q80US4 | ACTR5 | Actin-related protein 5 | Q6DIC0 | SNF2L2 | Probable global transcription activator SNF2L2 |
| Q8C3X8 | LMF2 | Lipase maturation factor 2 | P07356 | ANXA2 | Annexin A2 |
| P99028 | UQCRH | Cytochrome b-c1 complex subunit 6, mitochondrial | Q9WVH9 | FBLN5 | Fibulin-5 |
| P58742 | AAAS | Aladin | P43135 | NR2F2 | COUP transcription factor 2 |
| Q6PCP5 | Mff | Mitochondrial fission factor | Q8R4R6 | NUP35 | Nucleoporin NUP35 |
| O35488 | SLC27A2 | Very long-chain acyl-CoA synthetase | P11798 | CAMK2A | Calcium/calmodulin-dependent protein kinase type II subunit alpha |
| Q9DAS9 | GNG12 | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-12 | Q8BHL3 | TBC1D10B | TBC1 domain family member 10B |
| Q9D1G5 | LRRC57 | Leucine-rich repeat-containing protein 57 | Q8BTI8 | SRRNM2 | Serine/arginine repetitive matrix protein 2 |
| Q8BGD8 | COA6 | Cytochrome c oxidase assembly factor 6 homolog | P97952 | SCN1B | Sodium channel subunit beta-1 |
| Q9CRB8 | MTFP1 | Mitochondrial fission process protein 1 | P51912 | SLC1A5 | Neutral amino acid transporter B (0) |
| Q99MS7 | EHBP1L1 | EH domain-binding protein 1-like protein 1 | Q8BPB5 | EFEMP1 | EGF-containing fibulin-like extracellular matrix protein 1 |
| Q61029 | TMPO | Lamina-associated polypeptide 2, isoforms beta/delta/epsilon/gamma | Q64373 | BCL2L1 | Bcl-2-like protein 1 |
| Q91YU6 | LZTS2 | Leucine zipper putative tumor suppressor 2 | Q6P9Q4 | FHOD1 | FH1/FH2 domain-containing protein 1 |
| Q8K212 | PACS1 | Phosphofurin acidic cluster sorting protein 1 | Q99L88 | SNTB1 | Beta-1-syntrophin |
| Q3U3R4 | LMF1 | Lipase maturation factor 1 | Q80UZ0 | FGD5 | FYVE, RhoGEF and PH domain-containing protein 5 |
| Q9JI46 | NUDT3 | Diphosphoinositol polyphosphate phosphohydrolase 1 | O35704 | SPTLC1 | Serine palmitoyltransferase 1 |
| Q8K4G1 | LTBP-4 | Latent-transforming growth factor beta-binding protein 4 | A2ASQ1 | AGRN | Agrin |
| P70429 | EVL | Ena/VASP-like protein | Q9JIG8 | PRAF2 | PRA1 family protein 2 |
| C0HKD8 | MFAP1a | Microfibrillar-associated protein 1A | Q8R323 | RFC3 | Replication factor C subunit 3 |
| Terms | Count | p Value | FDR | richFactor | Accession NO. |
|---|---|---|---|---|---|
| GO (gene ontology) | |||||
| Gene silencing (BP) | 12 | 0.0011 | 0.5010 | 0.1600 | Q91YE5, Q9Z2D6, O88712, Q6ZQ88, Q6Y7W8, Q9CY73, Q80VL1, P15864, P43276, O55135, P42227, P68433 |
| Monosaccharide binding (MF) | 9 | 0.0029 | 0.5010 | 0.1698 | P41317, Q8K2I4, O08528, Q60715, Q64237, Q9D136, P12382, P06745, Q9JK38 |
| Extracellular matrix (CC) | 22 | 0.0012 | 0.5010 | 0.1164 | P07356, P11276, Q9WVH9, P97429, Q8K4G1, Q62009, P97384, A2ASQ1, C0HKD8, P06797, P41317, Q8BPB5, Q80YX1, O08746, P82198, P18406, Q61704, Q8R054, Q60754, O70503, P09242, Q9CXD9 |
| KEGG (kyoto encyclopedia of genes and genomes) pathways | |||||
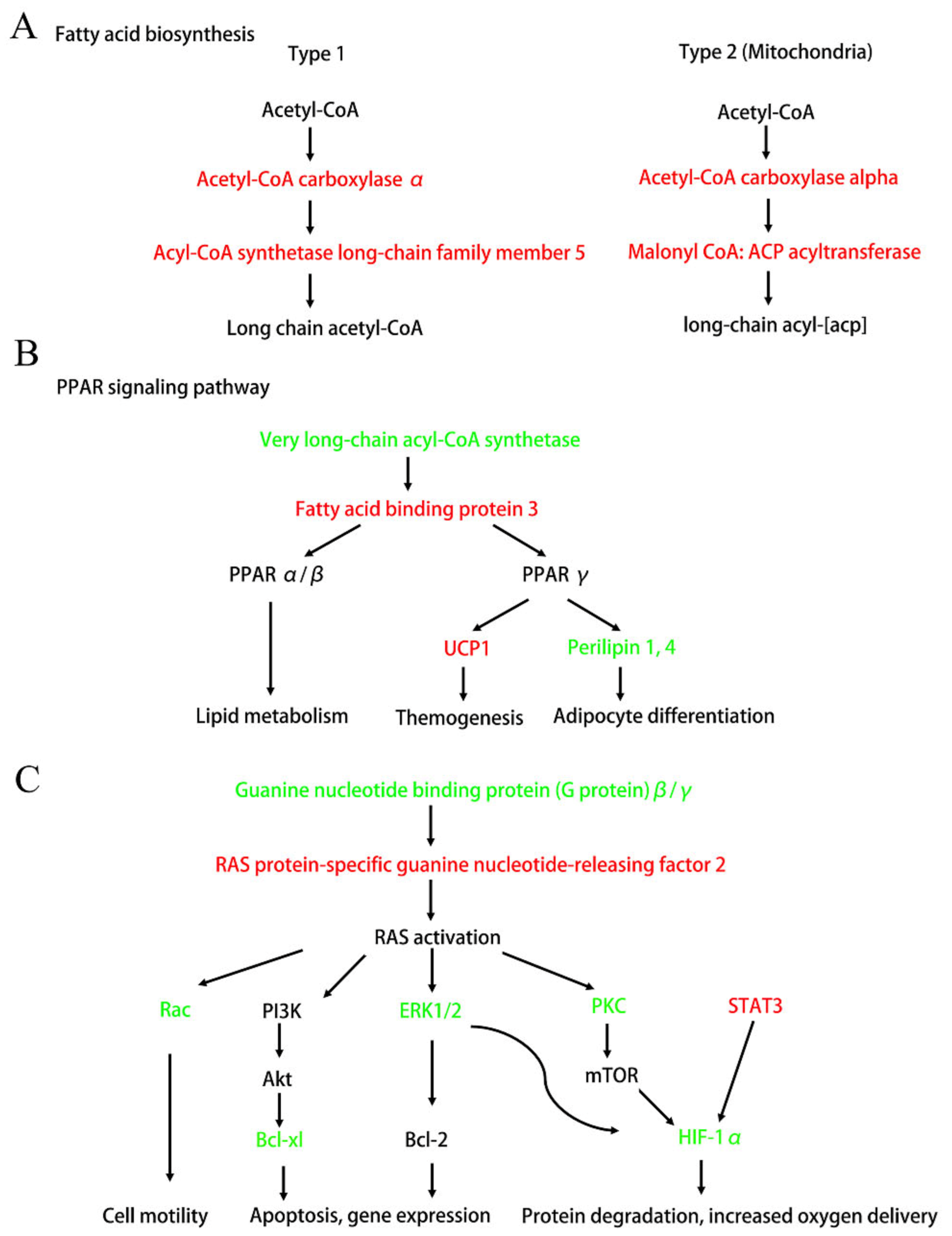
| Fatty acid biosynthesis | 4 | 0.0069 | 0.5894 | 0.2857 | Q5SWU9, Q8JZR0, Q8R3F5, Q91WC3 |
| EGFR tyrosine kinase inhibitor resistance | 6 | 0.0076 | 0.5894 | 0.1935 | Q63844, Q8BMB3, P63073, Q64373, P20444, P42227 |
| PPAR signaling pathway | 8 | 0.0092 | 0.5894 | 0.1538 | P12242, Q8CGN5, O88492, Q8JZR0, P11404, Q64516, O35488, Q91WC3 |
| HIF-1 signaling pathway | 8 | 0.0103 | 0.5894 | 0.1509 | P12382, O08528, P11798, Q63844, Q8BMB3, P63073, P20444, P42227 |
| Galactose metabolism | 4 | 0.0114 | 0.5894 | 0.2500 | P12382, Q9D0F9, O08528, P70699 |
| Starch and sucrose metabolism | 4 | 0.0143 | 0.6137 | 0.2353 | Q9D0F9, O08528, P06745, P70699 |
| Other glycan degradation | 3 | 0.0169 | 0.6230 | 0.3000 | Q8BX80, Q69ZF3, Q8K2I4 |
| Peroxisome | 8 | 0.0210 | 0.6757 | 0.1333 | P08228, Q8JZR0, P34914, Q78JN3, Q61285, O35488, Q91WC3, Q922Z0 |
| Lysosome | 8 | 0.0323 | 0.7758 | 0.1231 | Q61207, Q9Z1T1, P06797, O89017, P12265, P70699, O70370, Q8K2I4 |
| Thiamine metabolism | 2 | 0.0427 | 0.7758 | 0.3333 | P09242, Q9R0Y5 |
| Apoptosis | 7 | 0.0443 | 0.7758 | 0.1228 | P60710, O08600, P06797, Q63844, Q64373, O70370, O54788 |
| Ras signaling pathway | 8 | 0.0472 | 0.7758 | 0.1143 | P62874, Q05144, Q9DAS9, Q63844, Q64373, P20444, P70392, Q9D4H1 |
| Accession NO. | Gene Name | Protein Name | Degree | Up/Down Regulation |
|---|---|---|---|---|
| P11276 | FN1 | Fibronectin | 28 | up |
| P60710 | ACTB | Actin, cytoplasmic 1 | 25 | down |
| Q63844 | MAPK3 | Mitogen-activated protein kinase 3 | 17 | down |
| Q9CRB2 | NHP2 | H/ACA ribonucleoprotein complex subunit 2 | 17 | down |
| P42227 | STAT3 | Signal transducer and activator of transcription 3 | 16 | up |
| P50580 | PA2G4 | Proliferation-associated protein 2G4 | 15 | up |
| Q99ME9 | GTPBP4 | Nucleolar GTP-binding protein 1 | 14 | down |
| O55135 | eIF6 | Eukaryotic translation initiation factor 6 | 14 | down |
| P61202 | COPS2 | COP9 signalosome complex subunit 2 | 12 | up |
| Q64373 | BCL2L1 | Bcl-2-like protein 1 | 11 | down |
| O35488 | SLC27A2 | Very long-chain acyl-CoA synthetase | 11 | down |
| Q5SWU9 | ACACA | Acetyl-CoA carboxylase 1 | 11 | up |
| P67984 | Rpl22 | 60S ribosomal protein L22 | 11 | up |
| O08528 | HK2 | Hexokinase-2 | 10 | up |
| P08228 | SOD1 | Superoxide dismutase [Cu-Zn] | 10 | up |
| P63073 | eIF4E | Eukaryotic translation initiation factor 4E | 10 | down |
| P63158 | HMGB1 | High mobility group protein B1 | 10 | up |
| Q99L45 | EIF2S2 | Eukaryotic translation initiation factor 2 subunit 2 | 10 | up |
| Q6ZQ88 | KDM1A | Lysine-specific histone demethylase 1A | 10 | up |
| Q9WUP7 | UCHL5 | Ubiquitin carboxyl-terminal hydrolase isozyme L5 | 10 | up |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liang, X.; Guan, H.; Sun, J.; Qi, Y.; Yao, W. Comparative Proteomic Analysis of tPVAT during Ang II Infusion. Biomedicines 2021, 9, 1820. https://doi.org/10.3390/biomedicines9121820
Liang X, Guan H, Sun J, Qi Y, Yao W. Comparative Proteomic Analysis of tPVAT during Ang II Infusion. Biomedicines. 2021; 9(12):1820. https://doi.org/10.3390/biomedicines9121820
Chicago/Turabian StyleLiang, Xiuying, Haijing Guan, Jingwen Sun, Yan Qi, and Wenjuan Yao. 2021. "Comparative Proteomic Analysis of tPVAT during Ang II Infusion" Biomedicines 9, no. 12: 1820. https://doi.org/10.3390/biomedicines9121820
APA StyleLiang, X., Guan, H., Sun, J., Qi, Y., & Yao, W. (2021). Comparative Proteomic Analysis of tPVAT during Ang II Infusion. Biomedicines, 9(12), 1820. https://doi.org/10.3390/biomedicines9121820






