Non-Coding RNAs and Wnt/β-Catenin Signaling Pathway in Gastric Cancer: From EMT to Drug Resistance
Abstract
Simple Summary
Abstract
1. Introduction
1.1. Gastric Cancer
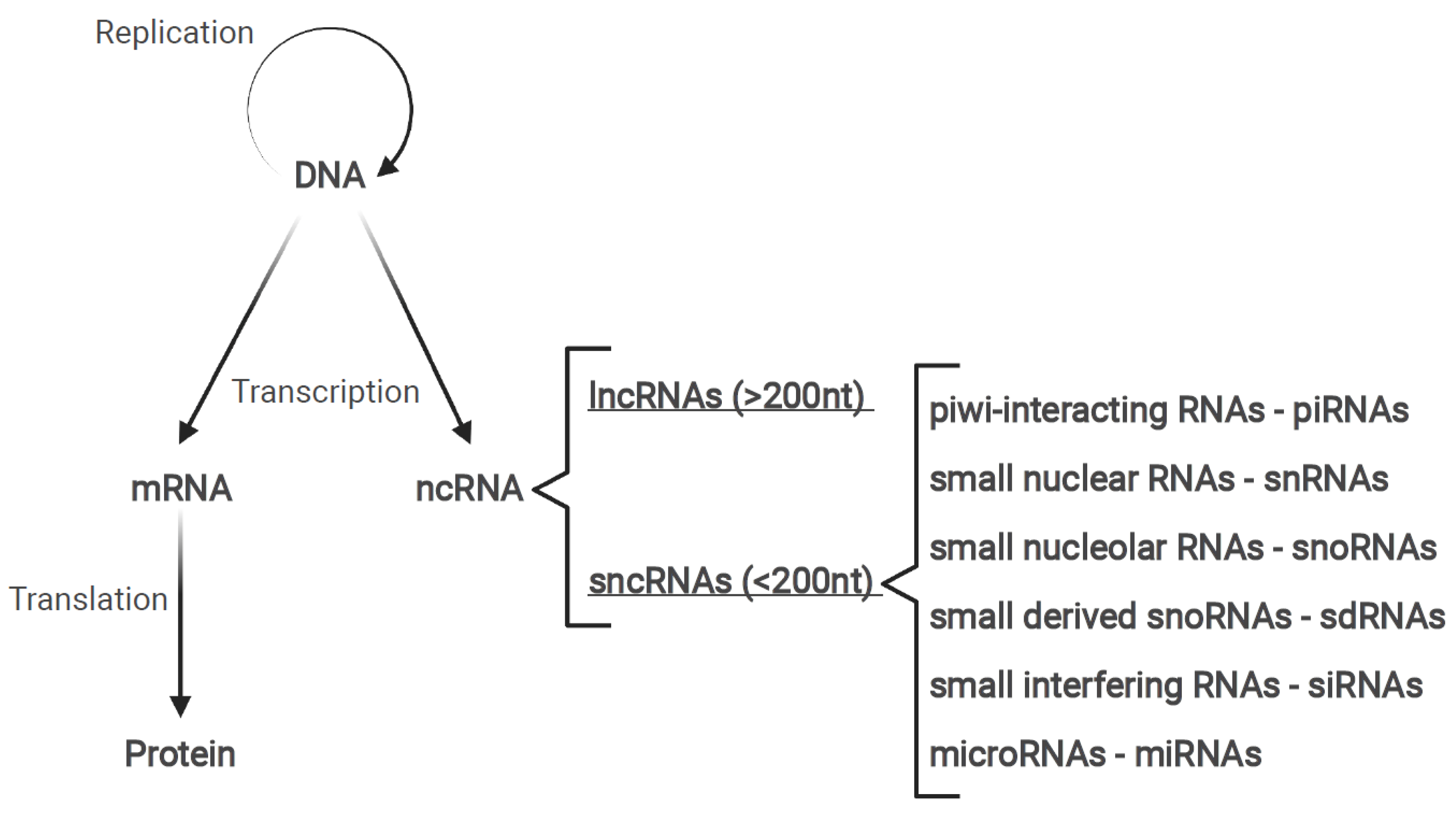
1.2. Non-Coding RNAs
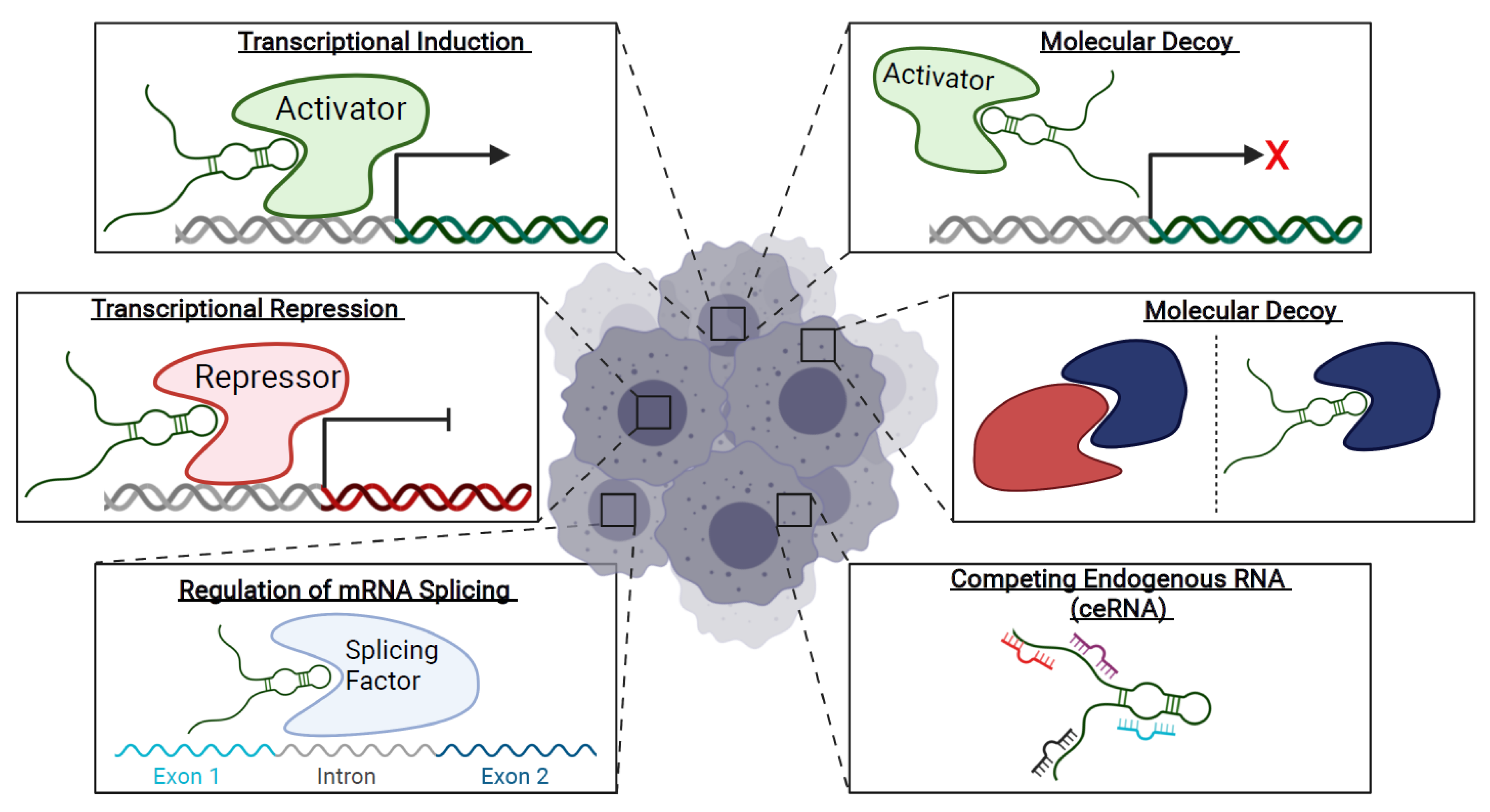
1.3. Long Non-Coding RNAs
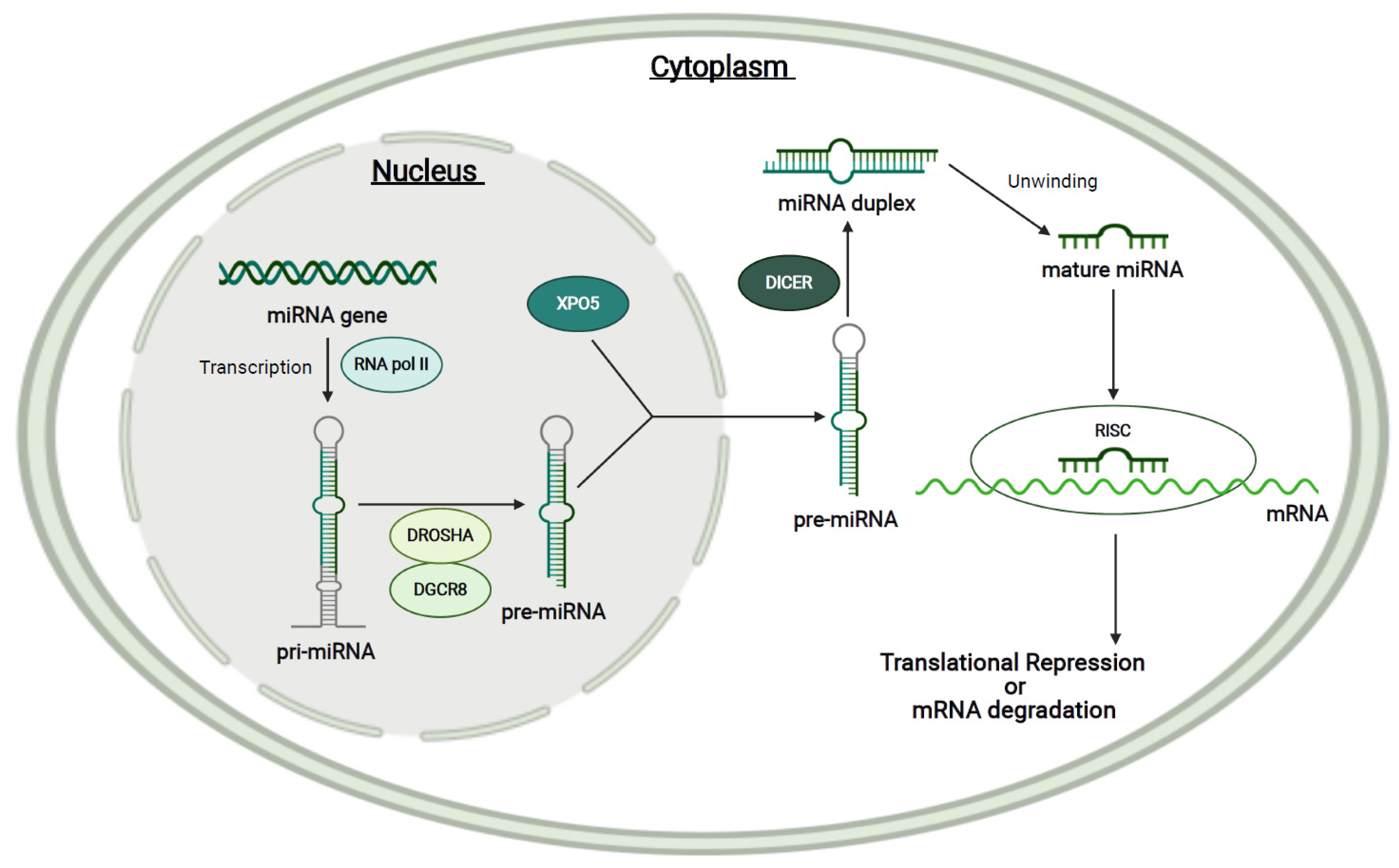
1.4. MicroRNAs
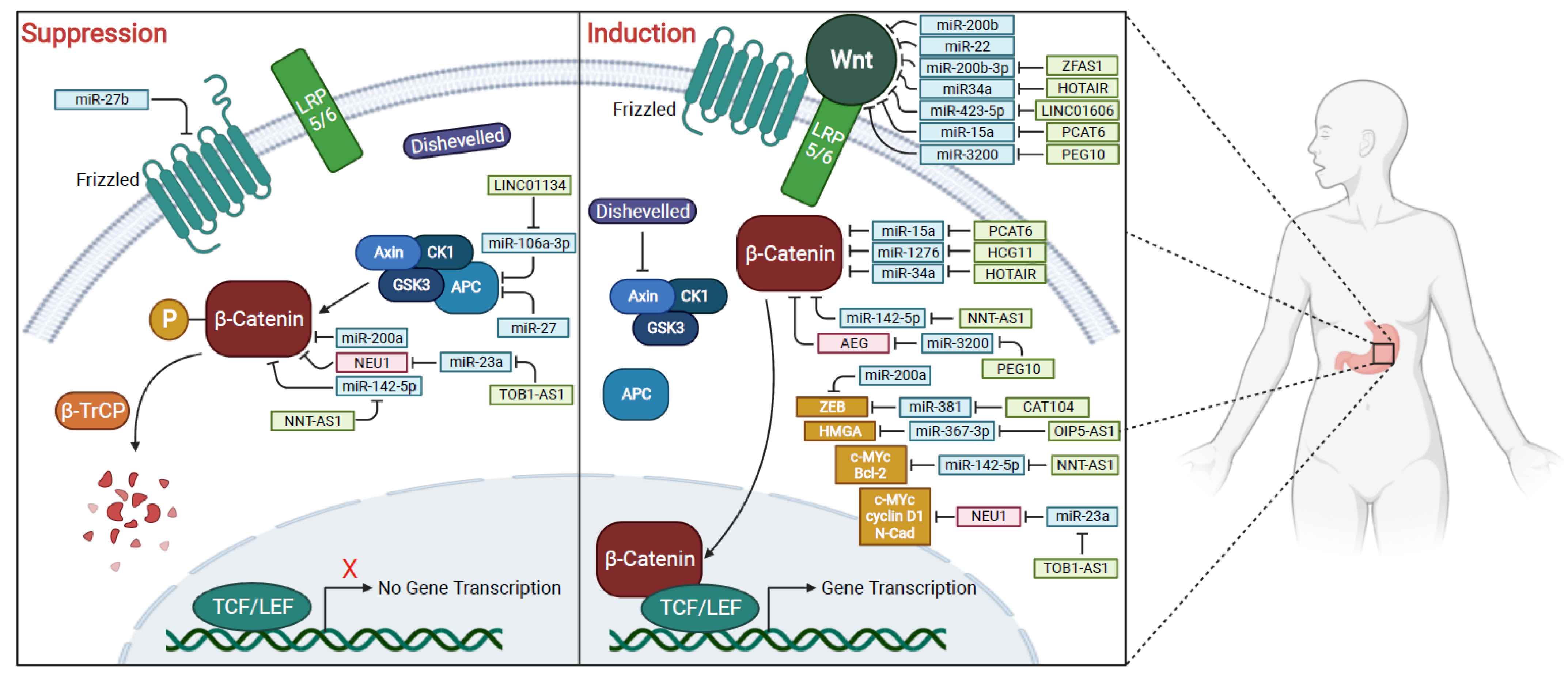
1.5. Wnt/β-Catenin Pathway
2. MicroRNAs Associated with Wnt/β-Catenin Signaling Pathway
2.1. Epithelial-to-Mesenchymal Transition (EMT) and Metastasis Increase in Gastric Cancer
| OncomiRNAs | Targets | Consequence on Wnt/β-Catenin Pathway | Consequence on EMT | Major Effects | Study Type | Ref. |
|---|---|---|---|---|---|---|
| miR-27 | APC | Induction | Induction | ↑ZEB1, ZEB2, Slug, Vimentin ↓E-cadherin | In vitro | [60] |
| miR-199a-5p | E-cadherin | Induction | Induction | ↓E-cadherin levels under SRF action | In vitro and in vivo | [61] |
| miR-194 | SUFU | Induction | Induction | ↑β-catenin | In vitro and in vivo | [62] |
| miR-192 | SMG-1 | Induction | Induction | ↓SMG↓E-cadherin, ↑N-cadherin | In vitro and in vivo | [63] |
| miR-215 | SMG-1 | Induction | Induction | ↓SMG↓E-cadherin ↑N-cadherin | In vitro and in vivo | [63] |
| miR-188-5p | PTEN | Induction | Induction | ↑Phospho-Ser9 of GSK3β; ↑Wnt | In vitro and in vivo | [64] |
| miR-675 | PITX1 | Induction | Induction | ↑Cell proliferation, migration and invasion | In vitro | [65] |
| Tumor Suppressor miRNAs | Targets | Consequence for Wnt/β-Catenin Pathway | Consequence for EMT | Major Effects | Study Type | Ref. |
|---|---|---|---|---|---|---|
| miR-200a | ZEB1, ZEB2 and β-catenin | Suppression | Suppression | ↓N-cadherin ↓β-catenin ↑E-cadherin | In vitro and in vivo | [66,67] |
| miR-29c-3p | KIAA1199 | Suppression | Suppression | ↓KIAA1199 ↓N-cadherin ↑E-cadherin ↑Axin2 | In vitro and in vivo | [68] |
| miR-381 | CUL4B | Suppression | Suppression | ↓Cell proliferation and invasion | In vitro and in vivo | [69] |
| miR-489 | CUL4B | Suppression | Suppression | ↓Cell proliferation and invasion | In vitro and in vivo | [69] |
| miR-338 | EphA2 | Suppression | Suppression | ↓EMT-related markers | In vitro | [70] |
| miR-375-3p | YWHAZ | Suppression | Suppression | ↓Cell proliferation and invasion | In vitro | [71] |
| miR-338-3p | SOX5 | Suppression | Suppression | ↓Cell proliferation | In vitro and tissues. | [72] |
| miR-873 | STRA6 | Suppression | Suppression | ↓Cell migration and invasion | In vitro and in vivo | [73] |
2.2. Cell Proliferation, Migration, Invasion, and Apoptosis
| miRNAs | Status in GC | Targets | Consequence on Wnt/β-Catenin Pathway | Cell Proliferation | Cell Migration and Invasion | Apoptosis | Tumor Growth | Study Type | Ref. |
|---|---|---|---|---|---|---|---|---|---|
| miR-200b | ↓ | Wnt-1 | Suppression | ↓ | - | - | ↓ | In vitro and in vivo | [75] |
| miR-22 | ↓ | Wnt-1 | Suppression | ↓ | - | - | ↓ | In vitro and in vivo | [75] |
| miR-19 | ↓ | MEF2D | Suppression | ↓ | - | - | ↓ | In vitro and in vivo | [76] |
| miR-27b | ↓ | FZD7 | Suppression | ↓ | - | - | - | In vitro | [77] |
| miR-511 | ↓ | TRIM24 | Suppression | ↓ | ↓ | - | - | In vitro | [78] |
| miR-185 | ↓ | TRIM29 | Suppression | ↓ | - | - | - | In vitro | [79] |
| miR-532 | ↑ | NKD1 | Induction | ↑ | ↑ | - | - | In vitro | [80] |
| miR-324-3p | ↑ | Smad4 | Induction | ↑ | ↑ | - | ↑ | In vivo, In vitro and organoids | [81] |
| miR-195-5p | ↓ | Not elucidated | Suppression | ↓ | - | ↑ | - | In vitro and samples | [82] |
| miR-18a | ↑ | SMAD2 | Induction | ↑ | ↑ | - | - | In vitro and in vivo | [83] |
| miR-19a | ↑ | SMAD2 | Induction | ↑ | ↑ | - | - | In vitro and in vivo | [83] |
| miR-876-5p | ↓ | WNT5A and MITF | Suppression | ↓ | ↓ | ↑ | ↓ | In vitro and in vivo | [84] |
2.3. Drug Resistance
| Gene or Protein That Targets miRNA | miRNAs | Status in GC | Targets | Consequence on Wnt/β-Catenin Pathway | Sphere Formation | Cell Lines Used | Study Type | Ref. |
|---|---|---|---|---|---|---|---|---|
| Not elucidated | miR-483-5p | ↑ | Not elucidated | Induction | ↑ | MKN-45 | In vitro | [85] |
| SLC34A2 | miR-25 | ↑ | GSK3β | Induction | ↑ | MKN-28 MKN-45 | In vitro and in vivo | [86] |
| Not elucidated | miR-501-5p | ↑ | DKK1 NKD1 GSK3β | Induction | ↑ | SGC-7901 HGC-27 MGC-803 MKN-28 BGC-823 | In vitro | [87] |
| BRD4 | miR-216a-3p | ↓ | Wnt3a | Suppression | ↓ | AGS, BGC-823 MKN-45 MGC-803 SGC-7901 GES-1 | In vitro | [88] |
3. Long Non-Coding RNAs Associated with Wnt/β-Catenin Signaling Pathway
| lncRNA | Proposed Mechanism | Expression in GC | Cancer Related Phenotype | Ref. |
|---|---|---|---|---|
| HOTAIR | ceRNA—miR34a | Upregulated | Drug Resistance | [90] |
| CAT104 | ceRNA—miR-381 | Upregulated | Cell proliferation, migration, invasion, and apoptosis | [91] |
| LINC01133 | ceRNA—miR-106a-3p | Downregulated | Cell proliferation, migration, and invasion | [92] |
| LINC01606 | ceRNA—miR-423-5p | Upregulated | Cell migration and invasion | [93] |
| LINC00052 | interacts with β-catenin and SMYD2 | Upregulated | Cell proliferation, migration, and invasion | [94] |
| BCAR4 | Not elucidated | Upregulated | Drug resistance | [95] |
| FEZF1-AS1 | Not elucidated | Upregulated | Cell proliferation | [96] |
| MALAT1 | Not elucidated | Upregulated | Cell proliferation, migration, invasion, and apoptosis | [97] |
| ENST00000434223 | Not elucidated | Downregulated | Cell proliferation, migration, invasion, and apoptosis | [98] |
| lnc-GNAT1-1 | Not elucidated | Downregulated | Cell proliferation, migration, and invasion | [99] |
| PEG10 | ceRNA—miR-3200 | Upregulated | Cell proliferation, migration, invasion, and apoptosis | [100] |
| TP73-AS1 | Not elucidated | Upregulated | Cell proliferation and invasion | [101] |
| LINC01225 | Not elucidated | Upregulated | Cell proliferation, migration, and invasion | [102] |
| TOB1-AS1 | ceRNA—miR-23a | Upregulated | Cell proliferation, migration, invasion, and apoptosis | [103] |
| ZFAS1 | ceRNA—miRNA-200b-3p | Upregulated | Cell proliferation, migration, invasion, and drug resistance | [104] |
| MIR4435-2HG | Not elucidated | Upregulated | Cell proliferation, migration, invasion, and apoptosis | [105] |
| LINC01314 | Not elucidated | Downregulated | Cell migration, invasion, and angiogenesis | [106] |
| HCG11 | ceRNA—miR-1276 | Upregulated | Cell proliferation, migration, and apoptosis | [107] |
| GATA6-AS1 | interaction with EZH2 | Downregulated | Cell proliferation, migration, and invasion | [108] |
| GASL1 | Not elucidated | Downregulated | Cell proliferation | [109] |
| LINC01503 | Not elucidated | Upregulated | Cell proliferation and invasion | [110] |
| PCAT6 | ceRNA—miR-15a | Upregulated | Cell proliferation and apoptosis | [111] |
| OIP5-AS1 | ceRNA—miR-367-3p | Upregulated | Cell proliferation and apoptosis | [112] |
| NNT-AS1 | ceRNA—miR-142-5p | Upregulated | Cell proliferation, migration, invasion, and apoptosis | [113] |
| HOXC-AS1 | interaction with eIF4A3 | Upregulated | Cell proliferation and apoptosis | [114] |
| FAM83H-AS1 | Not elucidated | Upregulated | Drug resistance | [115] |
3.1. Cell Proliferation
3.2. Cell Migration and Invasion
3.3. Apoptosis
3.4. Drug Resistance
3.5. Angiogenesis
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Cancer statistics 2018: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Group, H.C.C. Gastric cancer and Helicobacter pylori: A combined analysis of 12 case control studies nested within prospective cohorts. Gut 2001, 49, 347–353. [Google Scholar] [CrossRef]
- Hino, R.; Uozaki, H.; Murakami, N.; Ushiku, T.; Shinozaki, A.; Ishikawa, S.; Morikawa, T.; Nakaya, T.; Sakatani, T.; Takada, K. Activation of DNA methyltransferase 1 by EBV latent membrane protein 2A leads to promoter hypermethylation of PTEN gene in gastric carcinoma. Cancer Res. 2009, 69, 2766–2774. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.L.; Abar, L.; Norat, T. WCRF-AICR continuous update project: Systematic literature review of prospective studies on circulating 25-hydroxyvitamin D and kidney cancer risk. J. Steroid Biochem. Mol. Biol. 2016, 164, 85–89. [Google Scholar] [CrossRef] [PubMed][Green Version]
- International Agency for Research on Cancer. A Review of Human Carcinogens: Personal Habits and Indoor Combustions; World Health Organization: Lyon, France, 2012; Volume 100. [Google Scholar]
- Karimi, P.; Islami, F.; Anandasabapathy, S.; Freedman, N.D.; Kamangar, F. Gastric cancer: Descriptive epidemiology, risk factors, screening, and prevention. Cancer Epidemiol. Prev. Biomark. 2014, 23, 700–713. [Google Scholar] [CrossRef]
- Mukaisho, K.-I.; Nakayama, T.; Hagiwara, T.; Hattori, T.; Sugihara, H. Two distinct etiologies of gastric cardia adenocarcinoma: Interactions among pH, Helicobacter pylori, and bile acids. Front. Microbiol. 2015, 6, 412. [Google Scholar] [CrossRef]
- Lauren, P. The two histological main types of gastric carcinoma: Diffuse and so-called intestinal-type carcinoma: An attempt at a histo-clinical classification. J. Acta Pathol. Microbiol. Scand. 1965, 64, 31–49. [Google Scholar] [CrossRef]
- Henson, D.E.; Dittus, C.; Younes, M.; Nguyen, H.; Albores-Saavedra, J. Differential trends in the intestinal and diffuse types of gastric carcinoma in the United States, 1973–2000: Increase in the signet ring cell type. Arch. Pathol. Lab. Med. 2004, 128, 765–770. [Google Scholar] [CrossRef] [PubMed]
- Van der Woude, C.; Kleibeuker, J.; Tiebosch, A.; Homan, M.; Beuving, A.; Jansen, P.; Moshage, H. Diffuse and intestinal type gastric carcinomas differ in their expression of apoptosis related proteins. J. Pathol. 2003, 56, 699–702. [Google Scholar] [CrossRef]
- Gigek, C.O.; Calcagno, D.Q.; Rasmussen, L.T.; Santos, L.C.; Leal, M.F.; Wisnieski, F.; Burbano, R.R.; Lourenco, L.G.; Lopes-Filho, G.J.; Smith, M.A.C. Genetic variants in gastric cancer: Risks and clinical implications. J. Exp. Mol. Pathol. 2017, 103, 101–111. [Google Scholar] [CrossRef]
- Sánchez, Y.; Huarte, M. Long non-coding RNAs: Challenges for diagnosis and therapies. Nucleic Acid Ther. 2013, 23, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Bajic, V.B.; Zhang, Z. On the classification of long non-coding RNAs. RNA Biol. 2013, 10, 924–933. [Google Scholar] [CrossRef]
- Nature, E.P.C. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- Chakraborty, M.; Chatterjee, A.; Krithika, S.; Vasulu, T. A Statistical Analysis of MicroRNA: Classification, Identification and Conservation Based on Structure and Function. In Growth Curve and Structural Equation Modeling; Springer International Publishing: Cham, Switzerland, 2015; pp. 223–258. [Google Scholar]
- Mercer, T.R.; Dinger, M.E.; Sunkin, S.M.; Mehler, M.F.; Mattick, J.S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. USA 2008, 105, 716–721. [Google Scholar] [CrossRef]
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef]
- Choudhuri, S. Small noncoding RNAs: Biogenesis, function, and emerging significance in toxicology. J. Biochem. Mol. Toxicol 2010, 24, 195–216. [Google Scholar] [CrossRef]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef] [PubMed]
- Negishi, M.; Wongpalee, S.P.; Sarkar, S.; Park, J.; Lee, K.Y.; Shibata, Y.; Reon, B.J.; Abounader, R.; Suzuki, Y.; Sugano, S. A new lncRNA, APTR, associates with and represses the CDKN1A/p21 promoter by recruiting polycomb proteins. PLoS ONE 2014, 9, e95216. [Google Scholar] [CrossRef]
- Hung, T.; Wang, Y.; Lin, M.F.; Koegel, A.K.; Kotake, Y.; Grant, G.D.; Horlings, H.M.; Shah, N.; Umbricht, C.; Wang, P. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 2011, 43, 621–629. [Google Scholar] [CrossRef]
- Yin, G.; Tian, P.; BuHe, A.; Yan, W.; Li, T.; Sun, Z. LncRNA LINC00689 Promotes the Progression of Gastric Cancer Through Upregulation of ADAM9 by Sponging miR-526b-3p. Cancer Manag. Res. 2020, 12, 4227–4239. [Google Scholar] [CrossRef]
- Kretz, M.; Siprashvili, Z.; Chu, C.; Webster, D.E.; Zehnder, A.; Qu, K.; Lee, C.S.; Flockhart, R.J.; Groff, A.F.; Chow, J. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 2013, 493, 231–235. [Google Scholar] [CrossRef]
- Liu, B.; Sun, L.; Liu, Q.; Gong, C.; Yao, Y.; Lv, X.; Lin, L.; Yao, H.; Su, F.; Li, D. A cytoplasmic NF-κB interacting long noncoding RNA blocks IκB phosphorylation and suppresses breast cancer metastasis. Cancer Cell 2015, 27, 370–381. [Google Scholar] [CrossRef]
- Fang, S.; Zhang, L.; Guo, J.; Niu, Y.; Wu, Y.; Li, H.; Zhao, L.; Li, X.; Teng, X.; Sun, X. NONCODEV5: A comprehensive annotation database for long non-coding RNAs. Nucleic Acids Res. 2018, 46, D308–D314. [Google Scholar] [CrossRef] [PubMed]
- Frankish, A.; Diekhans, M.; Ferreira, A.-M.; Johnson, R.; Jungreis, I.; Loveland, J.; Mudge, J.M.; Sisu, C.; Wright, J.; Armstrong, J. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 2019, 47, D766–D773. [Google Scholar] [CrossRef]
- Tsagakis, I.; Douka, K.; Birds, I.; Aspden, J.L. Long non-coding RNAs in development and disease: Conservation to mechanisms. J. Pathol. 2020, 250, 480–495. [Google Scholar] [CrossRef]
- Bartel, D. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Williams, A.E. Functional aspects of animal microRNAs. Cell Mol. Life Sci. 2008, 65, 545–562. [Google Scholar] [CrossRef] [PubMed]
- Siomi, H.; Siomi, M.C. On the road to reading the RNA-interference code. Nature 2009, 457, 396–404. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, X. miRDB: An online database for prediction of functional microRNA targets. Nucleic Acids Res. 2020, 48, D127–D131. [Google Scholar] [CrossRef]
- Lee, Y.S.; Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 2007, 21, 1025–1030. [Google Scholar] [CrossRef]
- Pereira, J.; Santos, M.; Delabio, R.; Barbosa, M.; Smith, M.; Payão, S.; Rasmussen, L. Analysis of Gene Expression of miRNA-106b-5p and TRAIL in the Apoptosis Pathway in Gastric Cancer. Genes 2020, 11, 393. [Google Scholar] [CrossRef] [PubMed]
- Sharma, R.; Chopra, V. Effect of the Wingless (wg1) mutation on wing and haltere development in Drosophila melanogaster. Dev. Biol. 1976, 48, 461–465. [Google Scholar] [CrossRef]
- Nüsslein-Volhard, C.; Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 1980, 287, 795. [Google Scholar] [CrossRef] [PubMed]
- Oates, N.A.; van Vliet, J.; Duffy, D.L.; Kroes, H.Y.; Martin, N.G.; Boomsma, D.I.; Campbell, M.; Coulthard, M.G.; Whitelaw, E.; Chong, S. Increased DNA methylation at the AXIN1 gene in a monozygotic twin from a pair discordant for a caudal duplication anomaly. Am. J. Hum. Genet. 2006, 79, 155–162. [Google Scholar] [CrossRef]
- Kanazawa, A.; Tsukada, S.; Sekine, A.; Tsunoda, T.; Takahashi, A.; Kashiwagi, A.; Tanaka, Y.; Babazono, T.; Matsuda, M.; Kaku, K. Association of the gene encoding wingless-type mammary tumor virus integration-site family member 5B (WNT5B) with type 2 diabetes. Am. J. Hum. Genet. 2004, 75, 832–843. [Google Scholar] [CrossRef] [PubMed]
- Clevers, H. Wnt/β-catenin signaling in development and disease. Cell 2006, 127, 469–480. [Google Scholar] [CrossRef]
- Veeman, M.T.; Axelrod, J.D.; Moon, R.T. A second canon: Functions and mechanisms of β-catenin-independent Wnt signaling. Dev. Cell 2003, 5, 367–377. [Google Scholar] [CrossRef]
- Savagner, P. Leaving the neighborhood: Molecular mechanisms involved during epithelial-mesenchymal transition. Bioessays 2001, 23, 912–923. [Google Scholar] [CrossRef]
- García-Castro, M.n.I.; Marcelle, C.; Bronner-Fraser, M. Ectodermal Wnt function as a neural crest inducer. Science 2002, 297, 848–851. [Google Scholar] [CrossRef] [PubMed]
- Arwert, E.N.; Hoste, E.; Watt, F.M. Epithelial stem cells, wound healing and cancer. Nat. Rev. Cancer 2012, 12, 170–180. [Google Scholar] [CrossRef]
- Gonzalez, D.M.; Medici, D. Signaling mechanisms of the epithelial-mesenchymal transition. Sci. Signal. 2014, 7, re8. [Google Scholar] [CrossRef]
- Tammela, T.; Sanchez-Rivera, F.J.; Cetinbas, N.M.; Wu, K.; Joshi, N.S.; Helenius, K.; Park, Y.; Azimi, R.; Kerper, N.R.; Wesselhoeft, R.A. A Wnt-producing niche drives proliferative potential and progression in lung adenocarcinoma. Nature 2017, 545, 355–359. [Google Scholar] [CrossRef] [PubMed]
- de Sousa e Melo, F.; Kurtova, A.V.; Harnoss, J.M.; Kljavin, N.; Hoeck, J.D.; Hung, J.; Anderson, J.E.; Storm, E.E.; Modrusan, Z.; Koeppen, H. A distinct role for Lgr5+ stem cells in primary and metastatic colon cancer. Nature 2017, 543, 676–680. [Google Scholar] [CrossRef] [PubMed]
- Batlle, E.; Clevers, H. Cancer stem cells revisited. Nat. Med. 2017, 23, 1124. [Google Scholar] [CrossRef] [PubMed]
- Koushyar, S.; Powell, A.G.; Vincan, E.; Phesse, T.J. Targeting Wnt Signaling for the Treatment of Gastric Cancer. Int. J. Mol. Sci. 2020, 21, 3927. [Google Scholar] [CrossRef] [PubMed]
- Pan, K.-F.; Liu, W.-G.; Zhang, L.; You, W.-C.; Lu, Y.-Y. Mutations in components of the Wnt signaling pathway in gastric cancer. World J. Gastroenterol. 2008, 14, 1570. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.P.A.; Fei, G.; Kahmann, S.; Müller, O.; Yu, J.; Sung, J.J.Y.; Malfertheiner, P. Increased beta-catenin mRNA levels and mutational alterations of the APC and beta-catenin gene are present in intestinal-type gastric cancer. Carcinogenesis 2002, 23, 87–91. [Google Scholar] [CrossRef]
- Clements, W.M.; Wang, J.; Sarnaik, A.; Kim, O.J.; MacDonald, J.; Fenoglio-Preiser, C.; Groden, J.; Lowy, A.M. beta-Catenin mutation is a frequent cause of Wnt pathway activation in gastric cancer. Cancer Res. 2002, 62, 3503–3506. [Google Scholar]
- Min, B.-H.; Hwang, J.; Kim, N.K.; Park, G.; Kang, S.Y.; Ahn, S.; Ahn, S.; Ha, S.Y.; Lee, Y.K.; Kushima, R.; et al. Dysregulated Wnt signalling and recurrent mutations of the tumour suppressor RNF43 in early gastric carcinogenesis. J. Pathol. 2016, 240, 304–314. [Google Scholar] [CrossRef]
- Radulescu, S.; Ridgway, R.A.; Cordero, J.; Athineos, D.; Salgueiro, P.; Poulsom, R.; Neumann, J.; Jung, A.; Patel, S.; Woodgett, J.; et al. Acute WNT signalling activation perturbs differentiation within the adult stomach and rapidly leads to tumour formation. Oncogene 2013, 32, 2048–2057. [Google Scholar] [CrossRef] [PubMed]
- Oshima, H.; Oshima, M. Mouse models of gastric tumors: Wnt activation and PGE2 induction. Pathol. Int. 2010, 60, 599–607. [Google Scholar] [CrossRef]
- Czyzewska, J.; Guzińska-Ustymowicz, K.; Ustymowicz, M.; Pryczynicz, A.; Kemona, A. The expression of E-cadherin-catenin complex in patients with advanced gastric cancer: Role in formation of metastasis. Folia Histochem. Cytobiol. 2010, 48, 37–45. [Google Scholar] [CrossRef]
- Sereno, M.; Castro, J.D.; Cejas, P.; García-Cabezas, M.A.; Belda, C.; Casado, E.; Feliu, J.; Gómez, C.; López, M.; Barón, M.G. Expression profile as predictor of relapse after adjuvant treatment in gastric cancer. J. Gastrointest. Cancer 2012, 43, 181–189. [Google Scholar] [CrossRef] [PubMed]
- Dongre, A.; Weinberg, R.A. New insights into the mechanisms of epithelial–mesenchymal transition and implications for cancer. Nat. Rev. Mol. Cell Biol. 2019, 20, 69–84. [Google Scholar] [CrossRef]
- Wu, C.; Zhuang, Y.; Jiang, S.; Liu, S.; Zhou, J.; Wu, J.; Teng, Y.; Xia, B.; Wang, R.; Zou, X. Interaction between Wnt/β-catenin pathway and microRNAs regulates epithelial-mesenchymal transition in gastric cancer. Int. J. Oncol. 2016, 48, 2236–2246. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Liu, S.; Shi, R.; Zhao, G. miR-27 promotes human gastric cancer cell metastasis by inducing epithelial-to-mesenchymal transition. Cancer Genet. 2011, 204, 486–491. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; He, L.; Li, T.; Lu, Y.; Miao, Y.; Liang, S.; Guo, H.; Bai, M.; Xie, H.; Luo, G. SRF expedites metastasis and modulates the epithelial to mesenchymal transition by regulating miR-199a-5p expression in human gastric cancer. Cell Death Differ. 2014, 21, 1900–1913. [Google Scholar] [CrossRef]
- Peng, Y.; Zhang, X.; Ma, Q.; Yan, R.; Qin, Y.; Zhao, Y.; Cheng, Y.; Yang, M.; Wang, Q.; Feng, X. MiRNA-194 activates the Wnt/β-catenin signaling pathway in gastric cancer by targeting the negative Wnt regulator, SUFU. Cancer Lett. 2017, 385, 117–127. [Google Scholar] [CrossRef]
- Zhang, X.; Peng, Y.; Huang, Y.; Yang, M.; Yan, R.; Zhao, Y.; Cheng, Y.; Liu, X.; Deng, S.; Feng, X. SMG-1 inhibition by miR-192/-215 causes epithelial-mesenchymal transition in gastric carcinogenesis via activation of Wnt signaling. Cancer Med. 2018, 7, 146–156. [Google Scholar] [CrossRef]
- Li, Y.; Yan, X.; Shi, J.; He, Y.; Xu, J.; Lin, L.; Chen, W.; Lin, X.; Lin, X. Aberrantly expressed miR-188-5p promotes gastric cancer metastasis by activating Wnt/β-catenin signaling. BMC Cancer 2019, 19, 505. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Tian, Y.-C.; Mao, G.; Zhang, Y.-G.; Han, L. MiR-675 is frequently overexpressed in gastric cancer and enhances cell proliferation and invasion via targeting a potent anti-tumor gene PITX1. J. Cell. Signal. 2019, 62, 109352. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; Zhang, A.; Shi, Z.; Ma, F.; Pu, P.; Wang, T.; Zhang, J.; Kang, C.; Zhang, Q. MicroRNA-200a suppresses the Wnt/β-catenin signaling pathway by interacting with β-catenin. Int. J. Oncol. 2012, 40, 1162–1170. [Google Scholar] [CrossRef]
- Cong, N.; Du, P.; Zhang, A.; Shen, F.; Su, J.; Pu, P.; Wang, T.; Zjang, J.; Kang, C.; Zhang, Q. Downregulated microRNA-200a promotes EMT and tumor growth through the wnt/β-catenin pathway by targeting the E-cadherin repressors ZEB1/ZEB2 in gastric adenocarcinoma. Oncol. Rep. 2013, 29, 1579–1587. [Google Scholar] [CrossRef]
- Wang, L.; Yu, T.; Li, W.; Li, M.; Zuo, Q.; Zou, Q.; Xiao, B. The miR-29c-KIAA1199 axis regulates gastric cancer migration by binding with WBP11 and PTP4A3. Oncogene 2019, 38, 3134–3150. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z.; Zhong, M.; Wang, Y.; Yuan, X.; Guo, H.; Yao, Y.; Feng, M.; Chen, J.; Xiong, J.; Xiang, X. miR-381 and miR-489 suppress cell proliferation and invasion by targeting CUL4B via the Wnt/β-catenin pathway in gastric cancer. Int. J. Oncol. 2019, 54, 733–743. [Google Scholar] [CrossRef]
- Song, B.; Lin, H.; Dong, L.; Ma, J.; Jiang, Z. MicroRNA-338 inhibits proliferation, migration, and invasion of gastric cancer cells by the Wnt/β-catenin signaling pathway. Eur. Rev. Med. Pharm. Sci. 2018, 22, 1290–1296. [Google Scholar] [CrossRef]
- Guo, F.; Gao, Y.; Sui, G.; Jiao, D.; Sun, L.; Fu, Q.; Jin, C. miR-375-3p/YWHAZ/β-catenin axis regulates migration, invasion, EMT in gastric cancer cells. J. Clin. Exp. Pharmacol. Physiol. 2019, 46, 144–152. [Google Scholar] [CrossRef]
- Zheng, J.-J.; Que, Q.-Y.; Xu, H.-T.; Luo, D.-S.; Sun, Z.; Ni, J.-S.; Que, H.-F.; Ma, J.; Wu, D.; Shi, H. Hypoxia activates SOX5/Wnt/β-catenin signaling by suppressing MiR-338-3p in gastric cancer. J. Technol. Cancer Res. Treat. 2020, 19, 1533033820905825. [Google Scholar] [CrossRef]
- Lin, L.; Xiao, J.; Shi, L.; Chen, W.; Ge, Y.; Jiang, M.; Li, Z.; Fan, H.; Yang, L.; Xu, Z. STRA6 exerts oncogenic role in gastric tumorigenesis by acting as a crucial target of miR-873. J. Exp. Clin. Cancer Res. 2019, 38, 452. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Huang, Y.-X.; Zhang, R.; Hou, L.-D.; Liu, H.; Chen, X.-Y.; Zhu, J.-S.; Zhang, J. Toosendanin suppresses oncogenic phenotypes of human gastric carcinoma SGC-7901 cells partly via miR-200a-mediated downregulation of β-catenin pathway. Int. J. Oncol. 2017, 51, 1563–1573. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Kong, Y.; Guo, J.; Tang, Y.; Xie, X.; Yang, L.; Su, Q.; Xie, X. Diallyl disulfide suppresses proliferation and induces apoptosis in human gastric cancer through Wnt-1 signaling pathway by up-regulation of miR-200b and miR-22. Cancer Lett. 2013, 340, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Zhao, Y.-C. MEF2D/Wnt/β-catenin pathway regulates the proliferation of gastric cancer cells and is regulated by microRNA-19. Tumor Biol. 2016, 37, 9059–9069. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Lu, X.; Wu, X.; Xue, L.; Wang, X.; Xu, J. MicroRNA-27b suppresses Helicobacter pylori-induced gastric tumorigenesis through negatively regulating Frizzled7. Oncol. Rep. 2016, 35, 2441–2450. [Google Scholar] [CrossRef]
- Fang, Z.; Zhang, L.; Liao, Q.; Wang, Y.; Yu, F.; Feng, M.; Xiang, X.; Xiong, J. Regulation of TRIM24 by miR-511 modulates cell proliferation in gastric cancer. J. Exp. Clin. Cancer Res. 2017, 36, 17. [Google Scholar] [CrossRef] [PubMed]
- Qiu, F.; Xiong, J.-P.; Deng, J.; Xiang, X.-J. TRIM29 functions as an oncogene in gastric cancer and is regulated by miR-185. Int. J. Clin. Exp. Pathol. 2015, 8, 5053. [Google Scholar] [PubMed]
- Hu, S.; Zheng, Q.; Wu, H.; Wang, C.; Liu, T.; Zhou, W. miR-532 promoted gastric cancer migration and invasion by targeting NKD1. Life Sci. 2017, 177, 15–19. [Google Scholar] [CrossRef]
- Sun, G.-L.; Li, Z.; Wang, W.-Z.; Chen, Z.; Zhang, L.; Li, Q.; Wei, S.; Li, B.-W.; Xu, J.-H.; Chen, L. miR-324-3p promotes gastric cancer development by activating Smad4-mediated Wnt/beta-catenin signaling pathway. J. Gastroenterol. 2018, 53, 725–739. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Wu, Q. Effect of inhibition to Yes-related proteins-mediated Wnt/β-catenin signaling pathway through miR-195-5p on apoptosis of gastric cancer cells. J. Eur. Rev. Med. Pharm. Sci. 2019, 23, 6486–6496. [Google Scholar] [CrossRef]
- Yuan, J.; Tan, L.; Yin, Z.; Zhu, W.; Tao, K.; Wang, G.; Shi, W.; Gao, J. MIR17HG-miR-18a/19a axis, regulated by interferon regulatory factor-1, promotes gastric cancer metastasis via Wnt/β-catenin signalling. Cell Death Dis. 2019, 10, 454. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Yu, Z.; Tan, Q.; Wei, C.; Tang, Q.; Wang, L.; Hong, Y. MiR-876-5p regulates gastric cancer cell proliferation, apoptosis and migration through targeting WNT5A and MITF. Biosci. Rep. 2019, 39, BSR20190066. [Google Scholar] [CrossRef]
- Wu, K.; Ma, L.; Zhu, J. miR-483-5p promotes growth, invasion and self-renewal of gastric cancer stem cells by Wnt/β-catenin signaling. Mol. Med. Rep. 2016, 14, 3421–3428. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Guo, X.; Zhang, L.; Yang, F.; Qin, L.; Zhang, D.; Qin, Y. SLC34A2 regulates miR-25-Gsk3β signaling pathway to affect tumor progression in gastric cancer stem cell-like cells. Mol. Carcinog. 2018, 57, 440–450. [Google Scholar] [CrossRef] [PubMed]
- Fan, D.; Ren, B.; Yang, X.; Liu, J.; Zhang, Z. Upregulation of miR-501-5p activates the wnt/β-catenin signaling pathway and enhances stem cell-like phenotype in gastric cancer. J. Exp. Clin. Cancer Res. 2016, 35, 177. [Google Scholar] [CrossRef]
- Song, H.; Shi, L.; Xu, Y.; Xu, T.; Fan, R.; Cao, M.; Xu, W.; Song, J. BRD4 promotes the stemness of gastric cancer cells via attenuating miR-216a-3p-mediated inhibition of Wnt/β-catenin signaling. Eur. J. Pharmacol. 2019, 852, 189–197. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Cheng, C.; Qin, Y.; Zhi, Q.; Wang, J.; Qin, C. Knockdown of long non-coding RNA HOTAIR inhibits cisplatin resistance of gastric cancer cells through inhibiting the PI3K/Akt and Wnt/β-catenin signaling pathways by up-regulating miR-34a. Int. J. Biol. Macromol. 2018, 107, 2620–2629. [Google Scholar] [CrossRef]
- Yuan, G.; Quan, J.; Dong, D.; Wang, Q. Long noncoding RNA CAT104 promotes cell viability, migration, and invasion in gastric carcinoma cells through activation of microrna-381-inhibiting zinc finger E-box-binding homeobox 1 (ZEB1) expression. Oncol. Res. 2018, 26, 1037–1046. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.-Z.; Cheng, T.-T.; He, Q.-J.; Lei, Z.-Y.; Chi, J.; Tang, Z.; Liao, Q.-X.; Zhang, H.; Zeng, L.-S.; Cui, S.-Z. LINC01133 as ceRNA inhibits gastric cancer progression by sponging miR-106a-3p to regulate APC expression and the Wnt/β-catenin pathway. Mol. Cancer 2018, 17, 126. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Tan, W.; Jia, W.; Liu, Z.; Ye, P.; Fu, Z.; Lu, F.; Xiang, W.; Tang, L.; Yao, L. The long non-coding RNA LINC01606 contributes to the metastasis and invasion of human gastric cancer and is associated with Wnt/β-catenin signaling. J. Int. J. Biochem. Cell Biol. 2018, 103, 125–134. [Google Scholar] [CrossRef]
- Shan, Y.; Ying, R.; Jia, Z.; Kong, W.; Wu, Y.; Zheng, S.; Jin, H. LINC00052 promotes gastric cancer cell proliferation and metastasis via activating the Wnt/β-catenin signaling pathway. Oncol. Res. 2017, 25, 1589–1599. [Google Scholar] [CrossRef]
- Wang, L.; Chunyan, Q.; Zhou, Y.; He, Q.; Ma, Y.; Ga, Y.; Wang, X. BCAR4 increase cisplatin resistance and predicted poor survival in gastric cancer patients. Eur. Rev. Med. Pharm. Sci. 2017, 21, 4064–4070. [Google Scholar]
- Wu, X.; Zhang, P.; Zhu, H.; Li, S.; Chen, X.; Shi, L. Long noncoding RNA FEZF1-AS1 indicates a poor prognosis of gastric cancer and promotes tumorigenesis via activation of Wnt signaling pathway. Biomed. Pharm. 2017, 96, 1103–1108. [Google Scholar] [CrossRef]
- Lee, N.K.; Lee, J.H.; Ivan, C.; Ling, H.; Zhang, X.; Park, C.H.; Calin, G.A.; Lee, S.K. MALAT1 promoted invasiveness of gastric adenocarcinoma. BMC Cancer 2017, 17, 46. [Google Scholar] [CrossRef]
- Zhao, Y.-X.; Liu, J.-F.; Sun, W.-J.; Zeng, R.-F.; Li, T.; Ma, R.-M. Long non-coding RNA-ENST00000434223 suppresses tumor progression in gastric cancer cells through the Wnt/β-catenin signaling pathway. Int. J. Biol. Macromol. 2018, 120, 491–501. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Shuai, T.; Li, B.; Zhu, L.; Li, X. Long non-coding RNA lnc-GNAT1-1 inhibits gastric cancer cell proliferation and invasion through the Wnt/β-catenin pathway in Helicobacter pylori infection. Mol. Med. Rep. 2018, 18, 4009–4015. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chu, X.; Zhang, D.; Kong, D. Knockdown of long non-coding RNA PEG10 inhibits growth, migration and invasion of gastric carcinoma cells by up-regulating miR-3200. Neoplasma 2018, 65, 769–778. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xiao, S.; Wang, B.; Li, Y.; Chen, Q. Knockdown of lncRNA TP73-AS1 inhibits gastric cancer cell proliferation and invasion via the WNT/β-catenin signaling pathway. Oncol. Lett. 2018, 16, 3248–3254. [Google Scholar] [CrossRef]
- Xu, Y.; Zhang, G.; Zou, C.; Qi, W.; Gong, Z.; Zhang, G.; Ma, G.; Zhang, W.; Jiang, P. Long non-coding RNA LINC01225 promotes proliferation, invasion and migration of gastric cancer via Wnt/β-catenin signalling pathway. J. Cell. Mol. Med. 2019, 23, 7581–7591. [Google Scholar] [CrossRef]
- Jiang, K.; Zhi, X.; Ma, Y.; Zhou, L. Long non-coding RNA TOB1-AS1 modulates cell proliferation, apoptosis, migration and invasion through miR-23a/NEU1 axis via Wnt/b-catenin pathway in gastric cancer. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 9890–9899. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Li, Y.; Xu, W.; He, L.; Tan, Y.; Xu, H. Long non-coding RNA ZFAS1 regulates the malignant progression of gastric cancer via the microRNA-200b-3p/Wnt1 axis. Biosci. Biotechnol. Biochem. 2019, 83, 1289–1299. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wu, M.; Lu, Y.; He, K.; Cai, X.; Yu, X.; Lu, J.; Teng, L. LncRNA MIR4435-2HG targets desmoplakin and promotes growth and metastasis of gastric cancer by activating Wnt/β-catenin signaling. Aging 2019, 11, 6657. [Google Scholar] [CrossRef]
- Tang, L.; Wen, J.-B.; Wen, P.; Li, X.; Gong, M.; Li, Q. Long non-coding RNA LINC01314 represses cell migration, invasion, and angiogenesis in gastric cancer via the Wnt/β-catenin signaling pathway by down-regulating KLK4. Cancer Cell Int. 2019, 19, 94. [Google Scholar] [CrossRef]
- Zhang, H.; Huang, H.; Xu, X.; Wang, H.; Wang, J.; Yao, Z.; Xu, X.; Wu, Q.; Xu, F. LncRNA HCG11 promotes proliferation and migration in gastric cancer via targeting miR-1276/CTNNB1 and activating Wnt signaling pathway. Cancer Cell Int. 2019, 19, 350. [Google Scholar] [CrossRef]
- Li, Z.-T.; Zhang, X.; Wang, D.-W.; Xu, J.; Kou, K.-J.; Wang, Z.-W.; Yong, G.; Liang, D.-S.; Sun, X.-Y. Overexpressed lncRNA GATA6-AS1 Inhibits LNM and EMT via FZD4 through the Wnt/β-Catenin Signaling Pathway in GC. Mol. Ther. Nucleic Acids 2020, 19, 827–840. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.; Li, X.; Yu, Y.; Chen, J. LncRNA GASL1 inhibits tumor growth in gastric carcinoma by inactivating the Wnt/β-catenin signaling pathway. Exp. Ther. Med. 2019, 17, 4039–4045. [Google Scholar] [CrossRef]
- Ding, J.; Shi, F.; Xie, G.; Zhu, Y. Long Non-coding RNA LINC01503 Promotes Gastric Cancer Cell Proliferation and Invasion by Regulating Wnt Signaling. Dig. Dis. Sci. 2020, 66, 452–459. [Google Scholar] [CrossRef]
- Dong, D.; Lun, Y.; Sun, B.; Sun, H.; Wang, Q.; Yuan, G.; Quan, J. Silencing of long non-coding RNA PCAT6 restrains gastric cancer cell proliferation and epithelial-mesenchymal transition by targeting microRNA-15a. Gen. Physiol. Biophys. 2020, 39, 1–12. [Google Scholar] [CrossRef]
- Tao, Y.; Wan, X.; Fan, Q.; Wang, Y.; Sun, H.; Ma, L.; Sun, C.; Wu, Y. Long non-coding RNA OIP5-AS1 promotes the growth of gastric cancer through the miR-367-3p/HMGA2 axis. Dig. Liver Dis. 2020, 52, 773–779. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, K.; Hou, Y. Long non-coding RNA NNT-AS1 knockdown represses the progression of gastric cancer via modulating the miR-142-5p/SOX4/Wnt/β-catenin signaling pathway. Mol. Med. Rep. 2020, 22, 687–696. [Google Scholar] [CrossRef]
- Zhou, C.; An, N.; Cao, C.; Wang, G. lncRNA HOXC-AS1 promotes gastric cancer via binding eIF4AIII by activating Wnt/β-catenin signaling. J. Gene Med. 2020, 22, e3202. [Google Scholar] [CrossRef]
- Wang, B.; Guan, G.; Zhao, D. Silence of FAM83H-AS1 promotes chemosensitivity of gastric cancer through Wnt/β-catenin signaling pathway. Biomed. Pharmacother. 2020, 125, 109961. [Google Scholar] [CrossRef]
- Guo, W.; Wang, Q.; Zhan, Y.; Chen, X.; Yu, Q.; Zhang, J.; Wang, Y.; Xu, X.-j.; Zhu, L. Transcriptome sequencing uncovers a three–long noncoding RNA signature in predicting breast cancer survival. Sci. Rep. 2016, 6, 27931. [Google Scholar] [CrossRef] [PubMed]
- Lamouille, S.; Xu, J.; Derynck, R. Molecular mechanisms of epithelial–mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2014, 15, 178–196. [Google Scholar] [CrossRef]
- Zeng, H.-F.; Qiu, H.-Y.; Feng, F.-B. Long noncoding RNA LINC01133 functions as an miR-422a sponge to aggravate the tumorigenesis of human osteosarcoma. Oncol. Res. 2018, 26, 335–343. [Google Scholar] [CrossRef]
- Zhang, J.; Zhu, N.; Chen, X. A novel long noncoding RNA LINC01133 is upregulated in lung squamous cell cancer and predicts survival. Tumor Biol. 2015, 36, 7465–7471. [Google Scholar] [CrossRef]
- Xu, W.; He, L.; Li, Y.; Tan, Y.; Zhang, F.; Xu, H. Silencing of lncRNA ZFAS1 inhibits malignancies by blocking Wnt/β-catenin signaling in gastric cancer cells. Biosci. Biotechnol. Biochem. 2018, 82, 456–465. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Xie, S.; Li, Q.; Ma, J.; Wang, G. Large intervening non-coding RNA HOTAIR is associated with hepatocellular carcinoma progression. J. Int. Med. Res. 2011, 39, 2119–2128. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.-H.; Sun, M.; Nie, F.-Q.; Ge, Y.-B.; Zhang, E.-B.; Yin, D.-D.; Kong, R.; Xia, R.; Lu, K.-H.; Li, J.-H. Lnc RNA HOTAIR functions as a competing endogenous RNA to regulate HER2 expression by sponging miR-331-3p in gastric cancer. Mol. Cancer 2014, 13, 92. [Google Scholar] [CrossRef]
- Zhao, W.; An, Y.; Liang, Y.; Xie, X. Role of HOTAIR long noncoding RNA in metastatic progression of lung cancer. Eur. Rev. Med. Pharm. Sci. 2014, 18, 1930–1936. [Google Scholar]
- Leal, M.F.; do Nascimento, J.L.M.; da Silva, C.E.A.; Lamarão, M.F.V.; Calcagno, D.Q.; Khayat, A.S.; Assumpção, P.P.; Cabral, I.R.; Smith, M.d.A.C.; Burbano, R.R. Establishment and conventional cytogenetic characterization of three gastric cancer cell lines. Cancer Genet. Cytogenet. 2009, 195, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Lima, E.M.; Rissino, J.D.; Harada, M.L.; Assumpção, P.P.; Demachki, S.; Guimarães, A.C.; Casartelli, C.; Smith, M.d.A.C.; Burbano, R.R. Conventional cytogenetic characterization of a new cell line, ACP01, established from a primary human gastric tumor. J. Braz. J. Med. Biol. Res. 2004, 37, 1831–1838. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, H.F.; Alcântara, D.; Matos, L.A.; Sousa, J.M.D.C.; Leal, M.F.; Smith, M.D.A.C.; Burbano, R.R.; Bahia, M.D.O. Cytogenetic characterization and evaluation of c-MYC gene amplification in PG100, a new Brazilian gastric cancer cell line. Braz. J. Med. Biol. Res. 2010, 43, 717–721. [Google Scholar] [CrossRef] [PubMed]
- Anauate, A.C.; Leal, M.F.; Wisnieski, F.; Santos, L.C.; Gigek, C.O.; Chen, E.S.; Geraldis, J.C.; Calcagno, D.Q.; Assumpcao, P.P.; Demachki, S.; et al. Identification of suitable reference genes for miRNA expression normalization in gastric cancer. Gene 2017, 621, 59–68. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Takao Real Karia, B.; Albuquerque Pinto, C.; Gigek, C.O.; Wisnieski, F.; Arruda Cardoso Smith, M. Non-Coding RNAs and Wnt/β-Catenin Signaling Pathway in Gastric Cancer: From EMT to Drug Resistance. Onco 2021, 1, 140-157. https://doi.org/10.3390/onco1020012
Takao Real Karia B, Albuquerque Pinto C, Gigek CO, Wisnieski F, Arruda Cardoso Smith M. Non-Coding RNAs and Wnt/β-Catenin Signaling Pathway in Gastric Cancer: From EMT to Drug Resistance. Onco. 2021; 1(2):140-157. https://doi.org/10.3390/onco1020012
Chicago/Turabian StyleTakao Real Karia, Bruno, Camila Albuquerque Pinto, Carolina Oliveira Gigek, Fernanda Wisnieski, and Marilia Arruda Cardoso Smith. 2021. "Non-Coding RNAs and Wnt/β-Catenin Signaling Pathway in Gastric Cancer: From EMT to Drug Resistance" Onco 1, no. 2: 140-157. https://doi.org/10.3390/onco1020012
APA StyleTakao Real Karia, B., Albuquerque Pinto, C., Gigek, C. O., Wisnieski, F., & Arruda Cardoso Smith, M. (2021). Non-Coding RNAs and Wnt/β-Catenin Signaling Pathway in Gastric Cancer: From EMT to Drug Resistance. Onco, 1(2), 140-157. https://doi.org/10.3390/onco1020012






