Cannabidiol Decreases Intestinal Inflammation in the Ovariectomized Murine Model of Postmenopause
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals and Materials
2.2. Animals and Treatment
2.3. Ribonucleic Acid Sequencing (RNA-Seq)
2.4. RNA Sequencing Data Analysis
2.5. LC-MS Analysis of Bile Acids
2.5.1. Preparation of Liver and Colon Samples
2.5.2. LC-MS Analysis
2.6. Ileal Organoid Experiments
2.7. MTT Analysis for Cell Viability
2.8. qPCR of Ileal Organoids and Liver Tissue
2.9. Statistics
3. Results
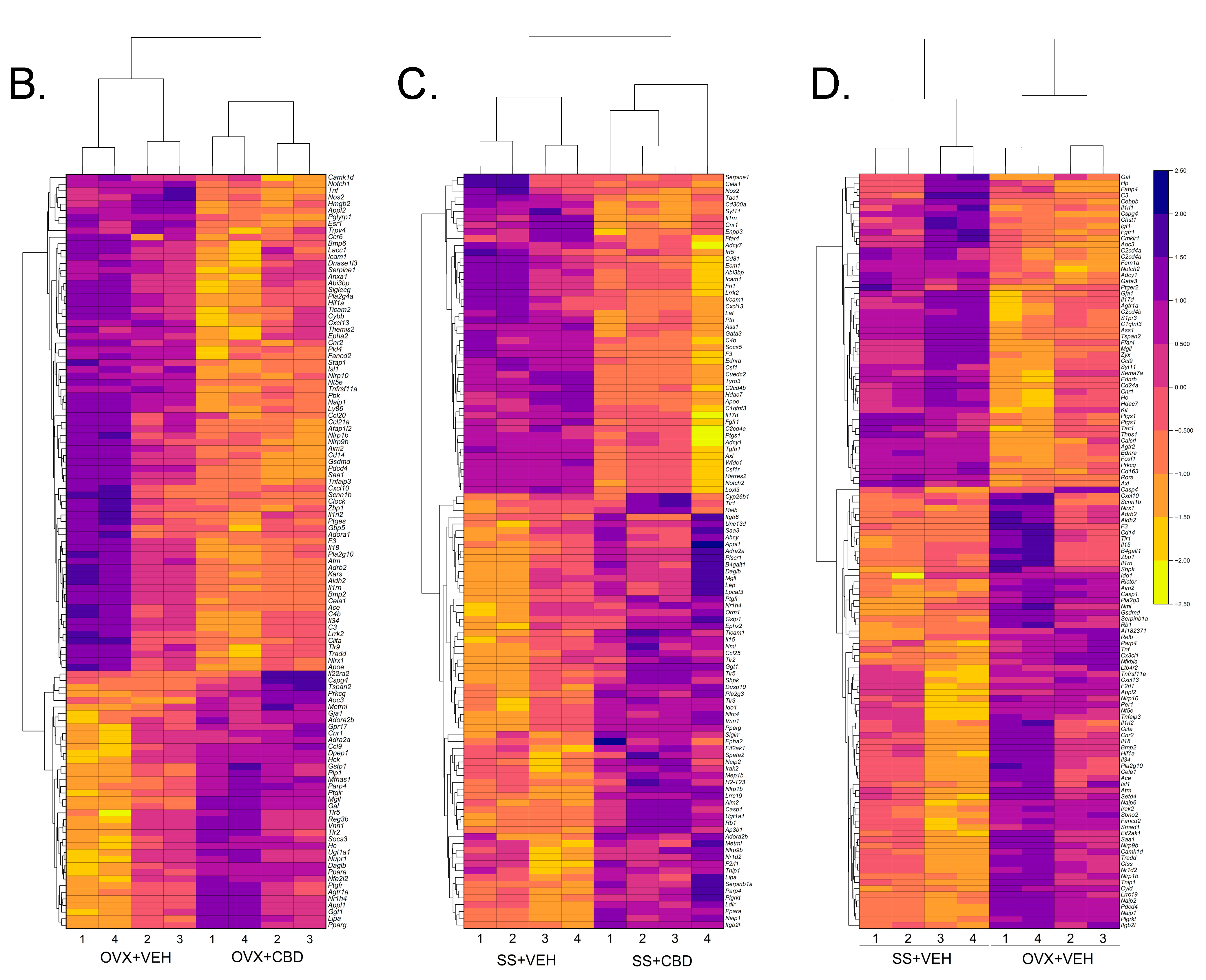
3.1. CBD-Induced Inflammatory Response Pathway Changes in E2-Deficient and -Sufficient Female Mice
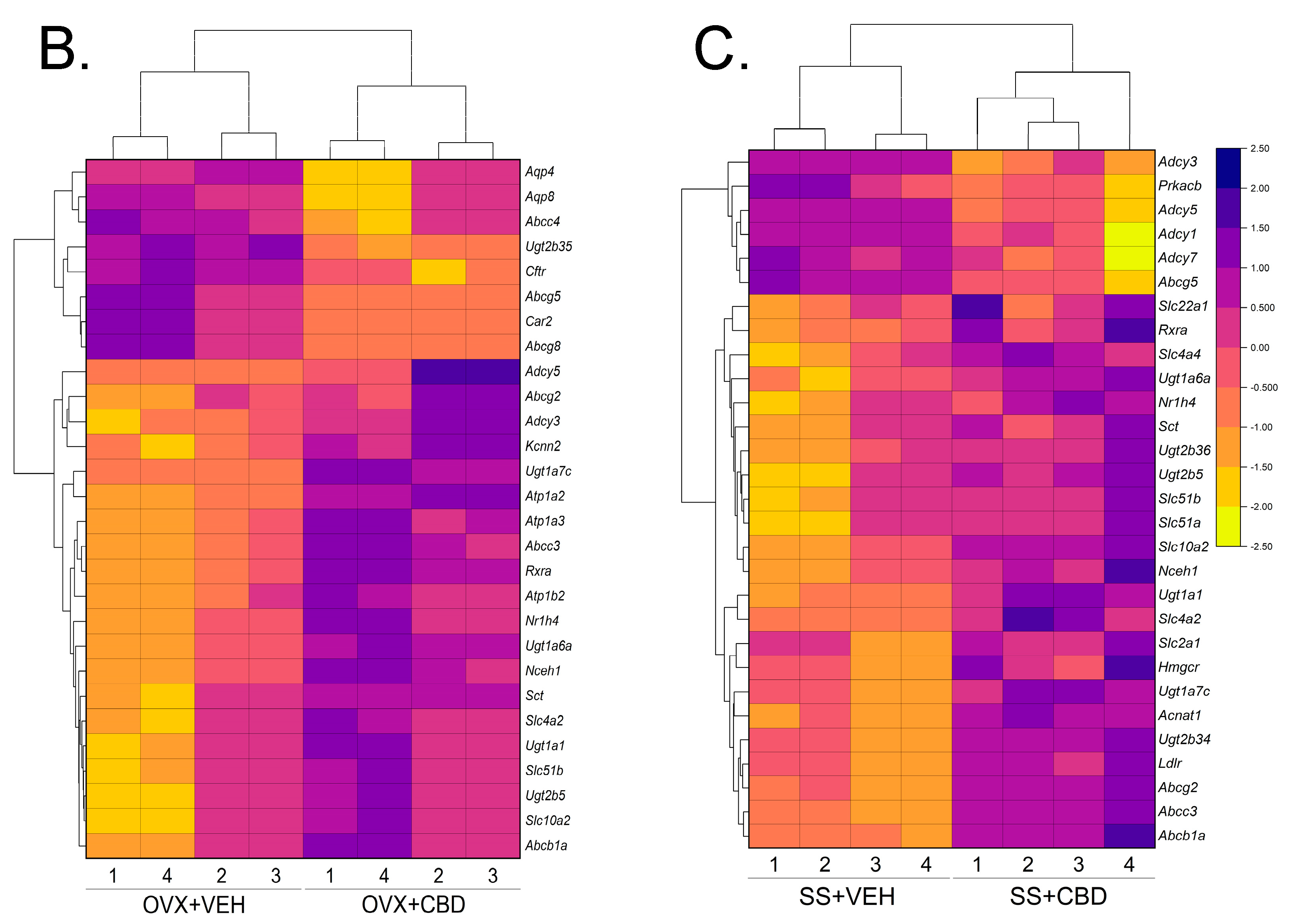
3.2. CBD-Induced Bile Secretion Pathway Changes in E2-Deficient and -Sufficient Female Mice
3.3. Colon Content and Hepatic BA Profiles
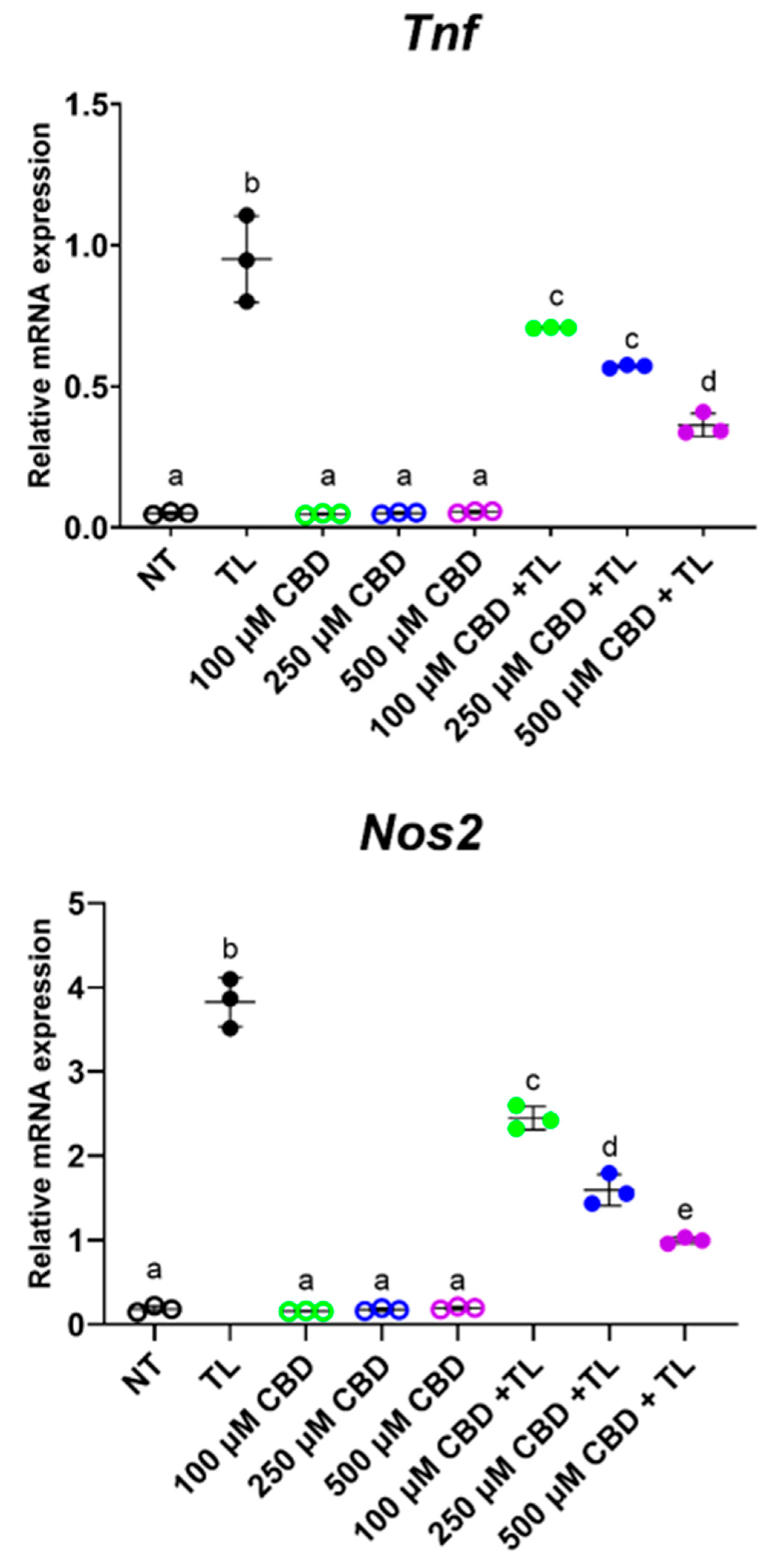
3.4. CBD Suppressed Inflammation in Ileal Organoids
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Peacock, K.; Ketvertis, K.M.; Doerr, C. Menopause (Nursing); StatPearls Publishing: Treasure Island, FL, USA, 2021. [Google Scholar]
- Col, N.F.; Guthrie, J.R.; Politi, M.; Dennerstein, L. Duration of vasomotor symptoms in middle-aged women: A longitudinal study. Menopause 2009, 16, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Freeman, E.W.; Sammel, M.D.; Lin, H.; Liu, Z.; Gracia, C.R. Duration of menopausal hot flushes and associated risk factors. Obstet. Gynecol. 2011, 117, 1095. [Google Scholar] [CrossRef] [PubMed]
- Freeman, E.W.; Sammel, M.D.; Sanders, R.J. Risk of long term hot flashes after natural menopause: Evidence from the Penn Ovarian Aging Cohort. Menopause 2014, 21, 924. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-Y.; Chassaing, B.; Tyagi, A.M.; Vaccaro, C.; Luo, T.; Adams, J.; Darby, T.M.; Weitzmann, M.N.; Mulle, J.G.; Gewirtz, A.T. Sex steroid deficiency–associated bone loss is microbiota dependent and prevented by probiotics. J. Clin. Investig. 2016, 126, 2049–2063. [Google Scholar] [CrossRef]
- Shieh, A.; Epeldegui, M.; Karlamangla, A.S.; Greendale, G.A. Gut permeability, inflammation, and bone density across the menopause transition. JCI Insight 2020, 5, e134092. [Google Scholar] [CrossRef]
- Woods, N.F.; Mitchell, E.S. Symptoms during the perimenopause: Prevalence, severity, trajectory, and significance in women’s lives. Am. J. Med. 2005, 118, 14–24. [Google Scholar] [CrossRef]
- Mauvais-Jarvis, F.; Clegg, D.J.; Hevener, A.L. The role of estrogens in control of energy balance and glucose homeostasis. Endocr. Rev. 2013, 34, 309–338. [Google Scholar] [CrossRef]
- Collins, F.L.; Rios-Arce, N.D.; Atkinson, S.; Bierhalter, H.; Schoenherr, D.; Bazil, J.N.; McCabe, L.R.; Parameswaran, N. Temporal and regional intestinal changes in permeability, tight junction, and cytokine gene expression following ovariectomy-induced estrogen deficiency. Physiol. Rep. 2017, 5, e13263. [Google Scholar] [CrossRef]
- Yu, M.; Pal, S.; Paterson, C.W.; Li, J.-Y.; Tyagi, A.M.; Adams, J.; Coopersmith, C.M.; Weitzmann, M.N.; Pacifici, R. Ovariectomy induces bone loss via microbial-dependent trafficking of intestinal TNF+ T cells and Th17 cells. J. Clin. Investig. 2021, 131, e143137. [Google Scholar] [CrossRef]
- Harper-Harrison, G.; Shanahan, M.M. Hormone Replacement Therapy. In StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Grossman, D.C.; Curry, S.J.; Owens, D.K.; Barry, M.J.; Davidson, K.W.; Doubeni, C.A.; Epling, J.W.; Kemper, A.R.; Krist, A.H.; Kurth, A.E. Hormone therapy for the primary prevention of chronic conditions in postmenopausal women: US Preventive Services Task Force recommendation statement. JAMA 2017, 318, 2224–2233. [Google Scholar]
- Shoupe, D. HRT dosing regimens: Continuous versus cyclic-pros and cons. Int. J. Fertil. Women’s Med. 2001, 46, 7–15. [Google Scholar]
- Atalay, S.; Jarocka-Karpowicz, I.; Skrzydlewska, E. Antioxidative and anti-inflammatory properties of cannabidiol. Antioxidants 2019, 9, 21. [Google Scholar] [CrossRef] [PubMed]
- De Filippis, D.; Esposito, G.; Cirillo, C.; Cipriano, M.; De Winter, B.Y.; Scuderi, C.; Sarnelli, G.; Cuomo, R.; Steardo, L.; De Man, J.G. Cannabidiol reduces intestinal inflammation through the control of neuroimmune axis. PLoS ONE 2011, 6, e28159. [Google Scholar] [CrossRef] [PubMed]
- Nichols, J.M.; Kaplan, B.L. Immune responses regulated by cannabidiol. Cannabis Cannabinoid Res. 2020, 5, 12–31. [Google Scholar] [CrossRef] [PubMed]
- Cocetta, V.; Governa, P.; Borgonetti, V.; Tinazzi, M.; Peron, G.; Catanzaro, D.; Berretta, M.; Biagi, M.; Manetti, F.; Dall’Acqua, S. Cannabidiol isolated from Cannabis sativa L. protects intestinal barrier from in vitro inflammation and oxidative stress. Front. Pharmacol. 2021, 12, 641210. [Google Scholar] [CrossRef]
- Li, D.; Lin, Z.; Meng, Q.; Wang, K.; Wu, J.; Yan, H. Cannabidiol administration reduces sublesional cancellous bone loss in rats with severe spinal cord injury. Eur. J. Pharmacol. 2017, 809, 13–19. [Google Scholar] [CrossRef]
- Napimoga, M.H.; Benatti, B.B.; Lima, F.O.; Alves, P.M.; Campos, A.C.; Pena-dos-Santos, D.R.; Severino, F.P.; Cunha, F.Q.; Guimarães, F.S. Cannabidiol decreases bone resorption by inhibiting RANK/RANKL expression and pro-inflammatory cytokines during experimental periodontitis in rats. Int. Immunopharmacol. 2009, 9, 216–222. [Google Scholar] [CrossRef]
- Raphael-Mizrahi, B.; Gabet, Y. The cannabinoids effect on bone formation and bone healing. Curr. Osteoporos. Rep. 2020, 18, 433–438. [Google Scholar] [CrossRef]
- Gaston, T.E.; Ampah, S.B.; Bebin, E.M.; Grayson, L.P.; Cutter, G.R.; Hernando, K.; Szaflarski, J.P. Long-term safety and efficacy of highly purified cannabidiol for treatment refractory epilepsy. Epilepsy Behav. 2021, 117, 107862. [Google Scholar] [CrossRef]
- Iffland, K.; Grotenhermen, F. An update on safety and side effects of cannabidiol: A review of clinical data and relevant animal studies. Cannabis Cannabinoid Res. 2017, 2, 139–154. [Google Scholar] [CrossRef]
- Larsen, C.; Shahinas, J. Dosage, efficacy and safety of cannabidiol administration in adults: A systematic review of human trials. J. Clin. Med. Res. 2020, 12, 129. [Google Scholar] [CrossRef]
- Machado Bergamaschi, M.; Helena Costa Queiroz, R.; Waldo Zuardi, A.; Crippa, A.S. Safety and side effects of cannabidiol, a Cannabis sativa constituent. Curr. Drug Saf. 2011, 6, 237–249. [Google Scholar] [CrossRef]
- Dahlgren, M.K.; El-Abboud, C.; Lambros, A.M.; Sagar, K.A.; Smith, R.T.; Gruber, S.A. A survey of medical cannabis use during perimenopause and postmenopause. Menopause 2022, 29, 1028–1036. [Google Scholar] [CrossRef]
- Sui, K.; Tveter, K.M.; Bawagan, F.G.; Buckendahl, P.; Martinez, S.A.; Jaffri, Z.H.; MacDonell, A.T.; Wu, Y.; Duran, R.M.; Shapses, S.A. Cannabidiol-Treated Ovariectomized Mice Show Improved Glucose, Energy, and Bone Metabolism with a Bloom in Lactobacillus. Front. Pharmacol. 2022, 2282. [Google Scholar] [CrossRef]
- Jansson, P.-A.; Curiac, D.; Ahrén, I.L.; Hansson, F.; Niskanen, T.M.; Sjögren, K.; Ohlsson, C. Probiotic treatment using a mix of three Lactobacillus strains for lumbar spine bone loss in postmenopausal women: A randomised, double-blind, placebo-controlled, multicentre trial. Lancet Rheumatol. 2019, 1, e154–e162. [Google Scholar] [CrossRef]
- Nilsson, A.; Sundh, D.; Bäckhed, F.; Lorentzon, M. Lactobacillus reuteri reduces bone loss in older women with low bone mineral density: A randomized, placebo-controlled, double-blind, clinical trial. J. Intern. Med. 2018, 284, 307–317. [Google Scholar] [CrossRef]
- Rizzoli, R.; Biver, E. Are probiotics the new calcium and vitamin D for bone health? Curr. Osteoporos. Rep. 2020, 18, 273–284. [Google Scholar] [CrossRef]
- Sapra, L.; Dar, H.Y.; Bhardwaj, A.; Pandey, A.; Kumari, S.; Azam, Z.; Upmanyu, V.; Anwar, A.; Shukla, P.; Mishra, P.K. Lactobacillus rhamnosus attenuates bone loss and maintains bone health by skewing Treg-Th17 cell balance in Ovx mice. Sci. Rep. 2021, 11, 1807. [Google Scholar] [CrossRef]
- Yang, L.-C.; Lin, S.-W.; Li, I.-C.; Chen, Y.-P.; Tzu, S.-Y.; Chou, W.; Chen, C.-C.; Lin, W.-C.; Chen, Y.-L.; Lin, W.-H. Lactobacillus plantarum GKM3 and Lactobacillus paracasei GKS6 supplementation ameliorates bone loss in ovariectomized mice by promoting osteoblast differentiation and inhibiting osteoclast formation. Nutrients 2020, 12, 1914. [Google Scholar] [CrossRef]
- Yu, J.; Cao, G.; Yuan, S.; Luo, C.; Yu, J.; Cai, M. Probiotic supplements and bone health in postmenopausal women: A meta-analysis of randomised controlled trials. BMJ Open 2021, 11, e041393. [Google Scholar] [CrossRef]
- Ridlon, J.M.; Kang, D.J.; Hylemon, P.B.; Bajaj, J.S. Bile acids and the gut microbiome. Curr. Opin. Gastroenterol. 2014, 30, 332. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.L.; Takeda, K.; Sundrud, M.S. Emerging roles of bile acids in mucosal immunity and inflammation. Mucosal Immunol. 2019, 12, 851–861. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, T.R.; Haeusler, R.A. Bile acids in glucose metabolism and insulin signalling—Mechanisms and research needs. Nat. Rev. Endocrinol. 2019, 15, 701–712. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.W.; An, J.H.; Park, H.; Yang, J.Y.; Choi, H.J.; Kim, S.W.; Park, Y.J.; Kim, S.Y.; Yim, M.; Baek, W.Y. Positive regulation of osteogenesis by bile acid through FXR. J. Bone Miner. Res. 2013, 28, 2109–2121. [Google Scholar] [CrossRef] [PubMed]
- Perucca, E.; Bialer, M. Critical aspects affecting cannabidiol oral bioavailability and metabolic elimination, and related clinical implications. CNS Drugs 2020, 34, 795–800. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Ge, S.X.; Jung, D.; Yao, R. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 2020, 36, 2628–2629. [Google Scholar] [CrossRef]
- Pang, Z.; Zhou, G.; Ewald, J.; Chang, L.; Hacariz, O.; Basu, N.; Xia, J. Using MetaboAnalyst 5.0 for LC–HRMS spectra processing, multi-omics integration and covariate adjustment of global metabolomics data. Nat. Protoc. 2022, 17, 1735–1761. [Google Scholar] [CrossRef]
- Mahe, M.M.; Aihara, E.; Schumacher, M.A.; Zavros, Y.; Montrose, M.H.; Helmrath, M.A.; Sato, T.; Shroyer, N.F. Establishment of gastrointestinal epithelial organoids. Curr. Protoc. Mouse Biol. 2013, 3, 217–240. [Google Scholar] [CrossRef]
- Tveter, K.M.; Villa-Rodriguez, J.A.; Cabales, A.J.; Zhang, L.; Bawagan, F.G.; Duran, R.M.; Roopchand, D.E. Polyphenol-induced improvements in glucose metabolism are associated with bile acid signaling to intestinal farnesoid X receptor. BMJ Open Diabetes Res Care 2020, 8, e001386. [Google Scholar] [CrossRef]
- Aziz, A.I.; Nguyen, L.C.; Oumeslakht, L.; Bensussan, A.; Ben Mkaddem, S. Cannabinoids as Immune System Modulators: Cannabidiol Potential Therapeutic Approaches and Limitations. Cannabis Cannabinoid Res. 2022. [Google Scholar] [CrossRef]
- Della Torre, S.; Benedusi, V.; Fontana, R.; Maggi, A. Energy metabolism and fertility: A balance preserved for female health. Nat. Rev. Endocrinol. 2014, 10, 13–23. [Google Scholar] [CrossRef]
- Zenone, M.A.; Snyder, J.; Crooks, V.A. What are the informational pathways that shape people’s use of cannabidiol for medical purposes? J. Cannabis Res. 2021, 3. [Google Scholar] [CrossRef]
- Soleymanpour, M.; Saderholm, S.; Kavuluru, R. Therapeutic Claims in Cannabidiol (CBD) Marketing Messages on Twitter. In Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Houston, TX, USA, 9–12 December 2021; pp. 3083–3088. [Google Scholar] [CrossRef]
- Benedusi, V.; Martini, E.; Kallikourdis, M.; Villa, A.; Meda, C.; Maggi, A. Ovariectomy shortens the life span of female mice. Oncotarget 2015, 6, 10801–10811. [Google Scholar] [CrossRef]
- Miquel, J.; Ramírez-Boscá, A.; Ramírez-Bosca, J.V.; Alperi, J.D. Menopause: A review on the role of oxygen stress and favorable effects of dietary antioxidants. Arch. Gerontol. Geriatr. 2006, 42, 289–306. [Google Scholar] [CrossRef]
- Pereira, S.R.; Hackett, B.; O’Driscoll, D.N.; Sun, M.C.; Downer, E.J. Cannabidiol modulation of oxidative stress and signalling. Neuronal Signal. 2021, 5, NS20200080. [Google Scholar] [CrossRef]
- Pérez, L.M.; Hooshmand, B.; Mangialasche, F.; Mecocci, P.; Smith, A.D.; Refsum, H.; Inzitari, M.; Fratiglioni, L.; Rizzuto, D.; Calderón-Larrañaga, A. Glutathione serum levels and rate of multimorbidity development in older adults. J. Gerontol. Ser. A 2020, 75, 1089–1094. [Google Scholar] [CrossRef]
- McGuire, P.; Robson, P.; Cubala, W.J.; Vasile, D.; Morrison, P.D.; Barron, R.; Taylor, A.; Wright, S. Cannabidiol (CBD) as an adjunctive therapy in schizophrenia: A multicenter randomized controlled trial. Am. J. Psychiatry 2018, 175, 225–231. [Google Scholar] [CrossRef]
- Saft, C.; von Hein, S.M.; Lücke, T.; Thiels, C.; Peball, M.; Djamshidian, A.; Heim, B.; Seppi, K. Cannabinoids for treatment of dystonia in Huntington’s disease. J. Huntingt. Dis. 2018, 7, 167–173. [Google Scholar] [CrossRef]
- Singer, E.; Judkins, J.; Salomonis, N.; Matlaf, L.; Soteropoulos, P.; McAllister, S.; Soroceanu, L. Reactive oxygen species-mediated therapeutic response and resistance in glioblastoma. Cell Death Dis. 2015, 6, e1601. [Google Scholar] [CrossRef] [PubMed]
- Vomhof-DeKrey, E.E.; Picklo Sr, M.J. The Nrf2-antioxidant response element pathway: A target for regulating energy metabolism. J. Nutr. Biochem. 2012, 23, 1201–1206. [Google Scholar] [CrossRef] [PubMed]
- Vargas, M.R.; Johnson, J.A. The Nrf2–ARE cytoprotective pathway in astrocytes. Expert Rev. Mol. Med. 2009, 11, e17. [Google Scholar] [CrossRef] [PubMed]
- Janbandhu, V.; Tallapragada, V.; Patrick, R.; Li, Y.; Abeygunawardena, D.; Humphreys, D.T.; Martin, E.M.; Ward, A.O.; Contreras, O.; Farbehi, N. Hif-1a suppresses ROS-induced proliferation of cardiac fibroblasts following myocardial infarction. Cell Stem Cell 2022, 29, 281–297.e12. [Google Scholar] [CrossRef] [PubMed]
- Miyauchi, Y.; Sato, Y.; Kobayashi, T.; Yoshida, S.; Mori, T.; Kanagawa, H.; Katsuyama, E.; Fujie, A.; Hao, W.; Miyamoto, K. HIF1α is required for osteoclast activation by estrogen deficiency in postmenopausal osteoporosis. Proc. Natl. Acad. Sci. USA 2013, 110, 16568–16573. [Google Scholar] [CrossRef] [PubMed]
- Shao, J.; Zhang, Y.; Yang, T.; Qi, J.; Zhang, L.; Deng, L. HIF-1α disturbs osteoblasts and osteoclasts coupling in bone remodeling by up-regulating OPG expression. In Vitro Cell. Dev. Biol.-Anim. 2015, 51, 808–814. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Lin, Z.; Cao, S.; Janowska, I.; Sonomoto, K.; Andreev, D.; Katharina, K.; Wen, J.; Knaup, K.X.; Wiesener, M.S. Estrogen-mediated downregulation of HIF-1α signaling in B lymphocytes influences postmenopausal bone loss. Bone Res. 2022, 10, 15. [Google Scholar] [CrossRef]
- Ray, P.D.; Huang, B.-W.; Tsuji, Y. Reactive oxygen species (ROS) homeostasis and redox regulation in cellular signaling. Cell. Signal. 2012, 24, 981–990. [Google Scholar] [CrossRef]
- Yang, S.; Lian, G. ROS and diseases: Role in metabolism and energy supply. Mol. Cell. Biochem. 2020, 467, 1–12. [Google Scholar] [CrossRef]
- Silva Sofrás, F.M.; Desimone, M.F. Entourage Effect And Analytical Chemistry: Chromatography As A Tool In The Analysis Of The Secondary Metabolism of Cannabis sativa L. Curr. Pharm. Des. 2022. online ahead of print. [Google Scholar] [CrossRef]
- Bautista, J.L.; Yu, S.; Tian, L. Flavonoids in Cannabis sativa: Biosynthesis, Bioactivities, and Biotechnology. ACS Omega 2021, 6, 5119–5123. [Google Scholar] [CrossRef]
- LaVigne, J.E.; Hecksel, R.; Keresztes, A.; Streicher, J.M. Cannabis sativa terpenes are cannabimimetic and selectively enhance cannabinoid activity. Sci. Rep. 2021, 11, 8232. [Google Scholar] [CrossRef]
- Anderson, L.L.; Etchart, M.G.; Bahceci, D.; Golembiewski, T.A.; Arnold, J.C. Cannabis constituents interact at the drug efflux pump BCRP to markedly increase plasma cannabidiolic acid concentrations. Sci. Rep. 2021, 11, 14948. [Google Scholar] [CrossRef]
- Pamplona, F.A.; da Silva, L.R.; Coan, A.C. Potential Clinical Benefits of CBD-Rich Cannabis Extracts Over Purified CBD in Treatment-Resistant Epilepsy: Observational Data Meta-analysis. Front. Neurol. 2018, 9, 759. [Google Scholar] [CrossRef]
- Skinner, C.M.; Nookaew, I.; Ewing, L.E.; Wongsurawat, T.; Jenjaroenpun, P.; Quick, C.M.; Yee, E.U.; Piccolo, B.D.; ElSohly, M.; Walker, L.A.; et al. Potential Probiotic or Trigger of Gut Inflammation—The Janus-Faced Nature of Cannabidiol-Rich Cannabis Extract. J. Diet. Suppl. 2020, 17, 543–560. [Google Scholar] [CrossRef]
- Santoro, A.; Mele, E.; Marino, M.; Viggiano, A.; Nori, S.L.; Meccariello, R. The Complex Interplay between Endocannabinoid System and the Estrogen System in Central Nervous System and Periphery. Int. J. Mol. Sci. 2021, 22, 972. [Google Scholar] [CrossRef]
- Di Marzo, V.; Piscitelli, F. The Endocannabinoid System and its Modulation by Phytocannabinoids. Neurotherapeutics 2015, 12, 692–698. [Google Scholar] [CrossRef]
- Joshi, N.; Onaivi, E.S. Endocannabinoid System Components: Overview and Tissue Distribution. In Recent Advances in Cannabinoid Physiology and Pathology; Advances in Experimental Medicine and Biology; Springer: Cham, Switzerland, 2019; Volume 1162, pp. 1–12. [Google Scholar] [CrossRef]
- Britch, S.C.; Babalonis, S.; Walsh, S.L. Cannabidiol: Pharmacology and therapeutic targets. Psychopharmacology 2021, 238, 9–28. [Google Scholar] [CrossRef]
- Ibeas Bih, C.; Chen, T.; Nunn, A.V.; Bazelot, M.; Dallas, M.; Whalley, B.J. Molecular Targets of Cannabidiol in Neurological Disorders. Neurotherapeutics 2015, 12, 699–730. [Google Scholar] [CrossRef]
- Bab, I.; Zimmer, A. Cannabinoid receptors and the regulation of bone mass. Br. J. Pharmacol. 2008, 153, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Feng, J.; Sun, L.; Xuan, Y.-W.; Wen, L.; Li, Y.-x.; Yang, S.; Zhu, B.; Tian, X.-Y.; Li, S. Cannabidiol Promotes Osteogenic Differentiation of Bone Marrow Mesenchymal Stem Cells in the Inflammatory Microenvironment via the CB2-dependent p38 MAPK Signaling Pathway. Int. J. Stem Cells 2022, 15, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Benamar, K.; Yondorf, M.; Meissler, J.J.; Geller, E.B.; Tallarida, R.J.; Eisenstein, T.K.; Adler, M.W. A novel role of cannabinoids: Implication in the fever induced by bacterial lipopolysaccharide. J. Pharmacol. Exp. Ther. 2007, 320, 1127–1133. [Google Scholar] [CrossRef] [PubMed]
- Marin, J.J.G. Bile Acids in Physiology, Pathology and Pharmacology. Curr. Drug Metab. 2015, 17, 4–29. [Google Scholar] [CrossRef]
- Lázár, B.; László, S.B.; Hutka, B.; Tóth, A.S.; Mohammadzadeh, A.; Berekméri, E.; Ágg, B.; Balogh, M.; Sajtos, V.; Király, K. A comprehensive time course and correlation analysis of indomethacin-induced inflammation, bile acid alterations and dysbiosis in the rat small intestine. Biochem. Pharmacol. 2021, 190, 114590. [Google Scholar] [CrossRef] [PubMed]
- Nakano, Y.; Tajima, M.; Sugiyama, E.; Sato, V.H.; Sato, H. Development of a Novel Nano-emulsion Formulation to Improve Intestinal Absorption of Cannabidiol. Med. Cannabis Cannabinoids 2019, 2, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Bertolini, A. Bile acids and their receptors: Modulators and therapeutic targets in liver inflammation. Semin. Immunopathol. 2022, 44, 547–564. [Google Scholar] [CrossRef]
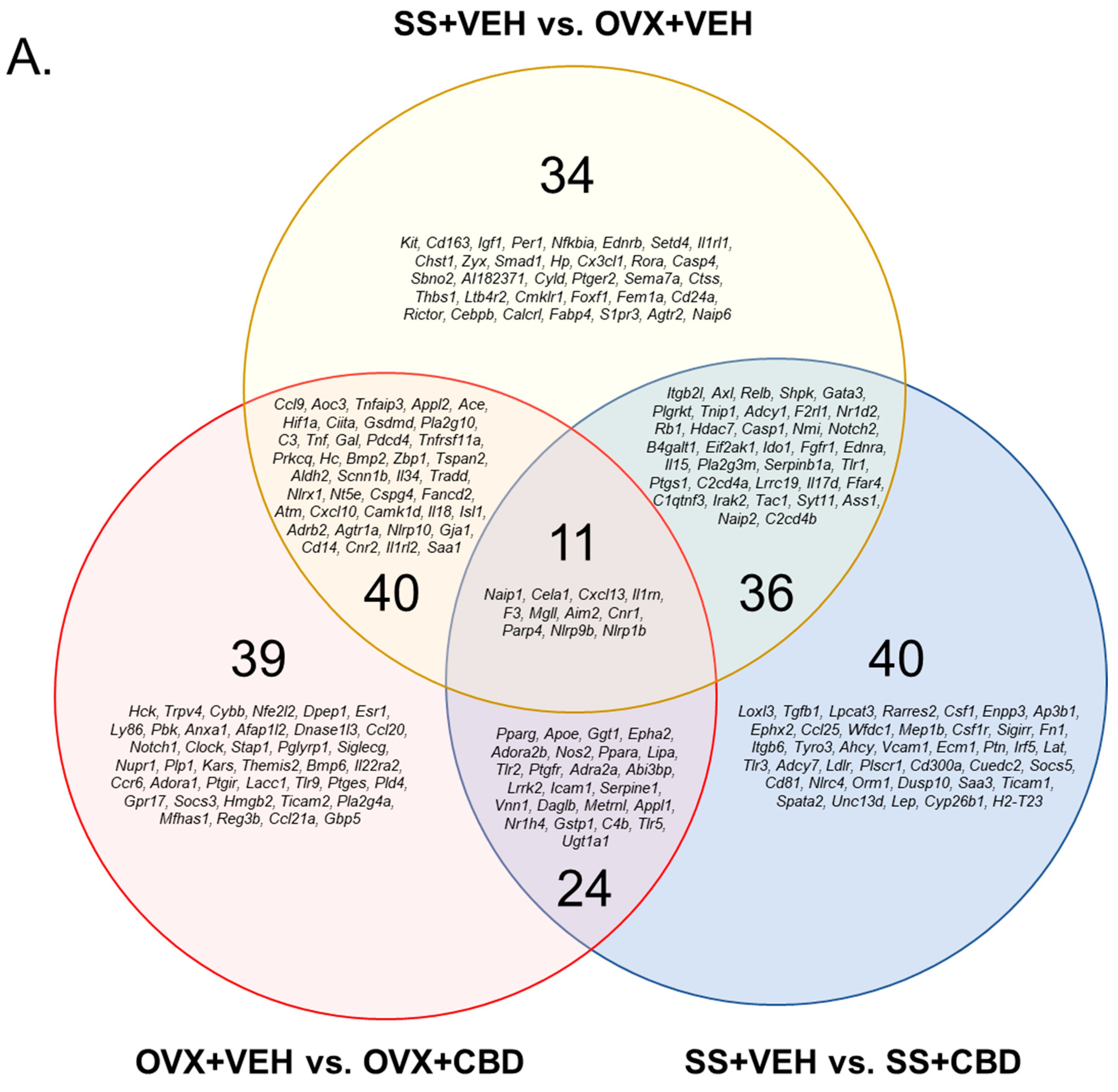

| Upregulated | Downregulated | Total Changed | |
|---|---|---|---|
| OVX+VEH vs. OVX+CBD | 1334 | 1255 | 2589 |
| SS+VEH vs. SS+CBD | 964 | 13,544 | 14,508 |
| SS+VEH vs. OVX+VEH | 1552 | 1610 | 3162 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bacalia, K.M.A.; Tveter, K.M.; Palmer, H.; Douyere, J.; Martinez, S.; Sui, K.; Roopchand, D.E. Cannabidiol Decreases Intestinal Inflammation in the Ovariectomized Murine Model of Postmenopause. Biomedicines 2023, 11, 74. https://doi.org/10.3390/biomedicines11010074
Bacalia KMA, Tveter KM, Palmer H, Douyere J, Martinez S, Sui K, Roopchand DE. Cannabidiol Decreases Intestinal Inflammation in the Ovariectomized Murine Model of Postmenopause. Biomedicines. 2023; 11(1):74. https://doi.org/10.3390/biomedicines11010074
Chicago/Turabian StyleBacalia, Karen Mae A., Kevin M. Tveter, Hayley Palmer, Jeffrey Douyere, Savannah Martinez, Ke Sui, and Diana E. Roopchand. 2023. "Cannabidiol Decreases Intestinal Inflammation in the Ovariectomized Murine Model of Postmenopause" Biomedicines 11, no. 1: 74. https://doi.org/10.3390/biomedicines11010074
APA StyleBacalia, K. M. A., Tveter, K. M., Palmer, H., Douyere, J., Martinez, S., Sui, K., & Roopchand, D. E. (2023). Cannabidiol Decreases Intestinal Inflammation in the Ovariectomized Murine Model of Postmenopause. Biomedicines, 11(1), 74. https://doi.org/10.3390/biomedicines11010074






