Disruption of Colorectal Cancer Network by Polyphyllins Reveals Pivotal Entities with Implications for Chemoimmunotherapy
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Culture and Reagents
2.2. Cytotoxicity Assays
2.3. Clonogenic Assay
2.4. Muse Cell Analysis
2.5. Proteomic Analysis
2.6. Statistical Analysis
3. Results
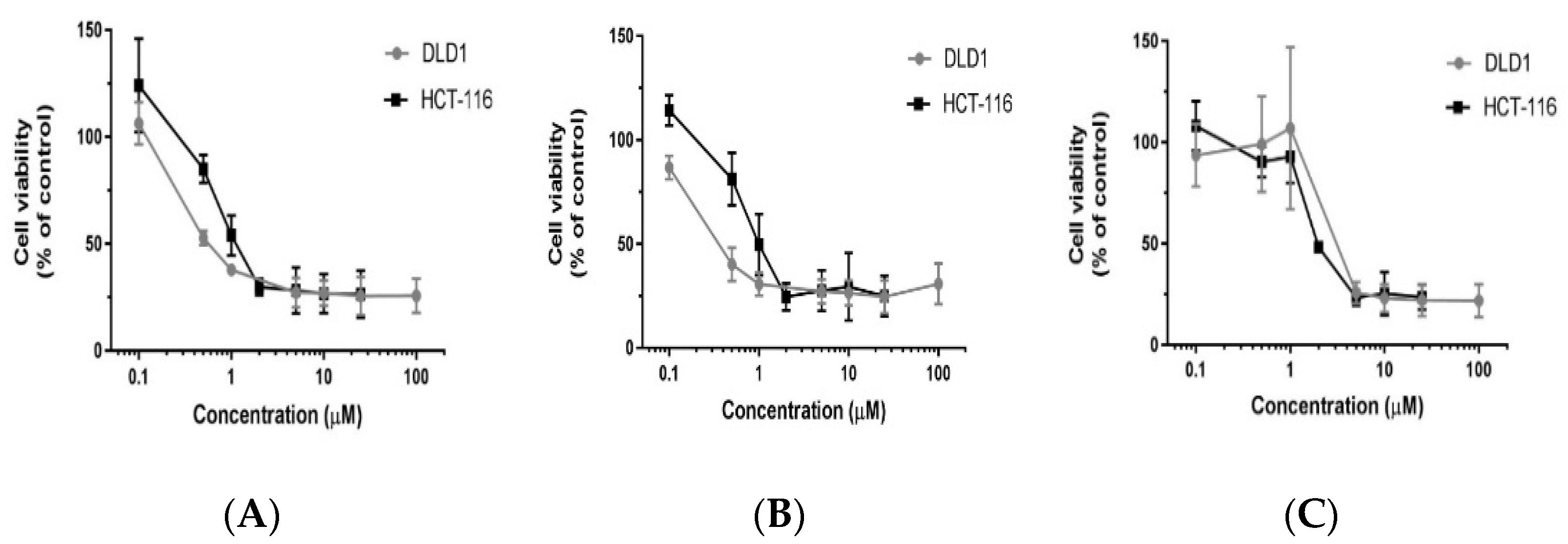
3.1. Polyphyllin D (PD) and Its Structural Analogs PII and PG Inhibit the Viability of Colorectal Cancer Cells
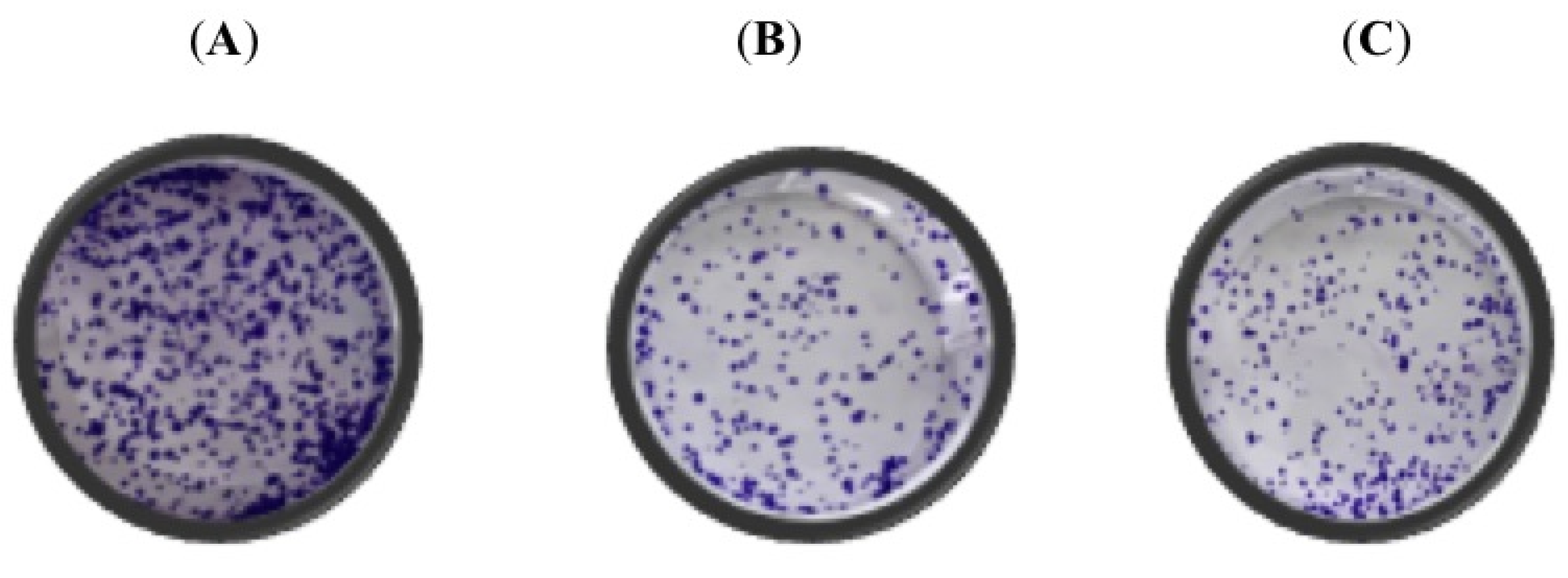
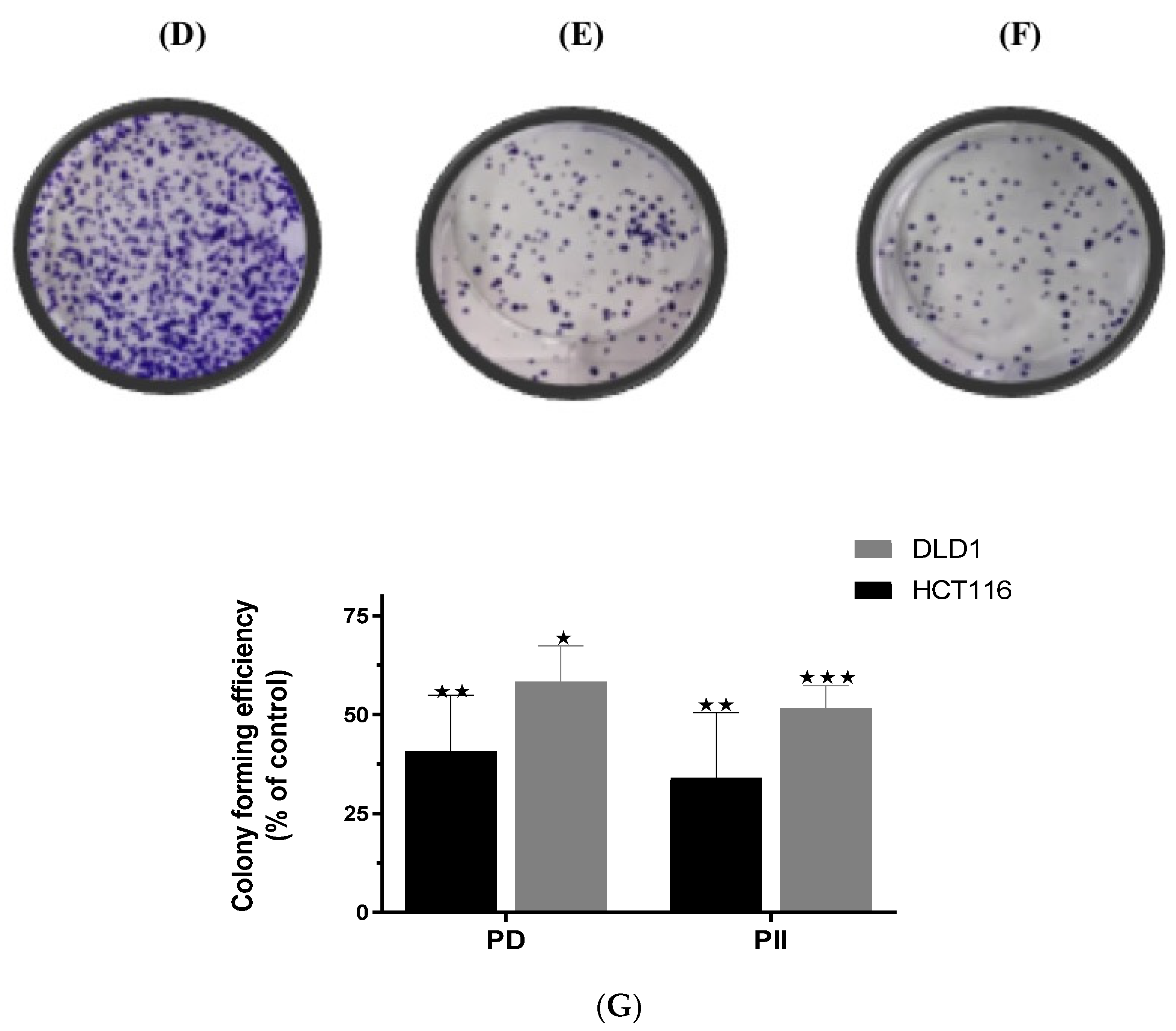
3.2. Polyphyllin D (PD) and PII Impact the Colony Forming Efficiency of DLD-1 and HCT116 Cells Leading to a Sustained Response
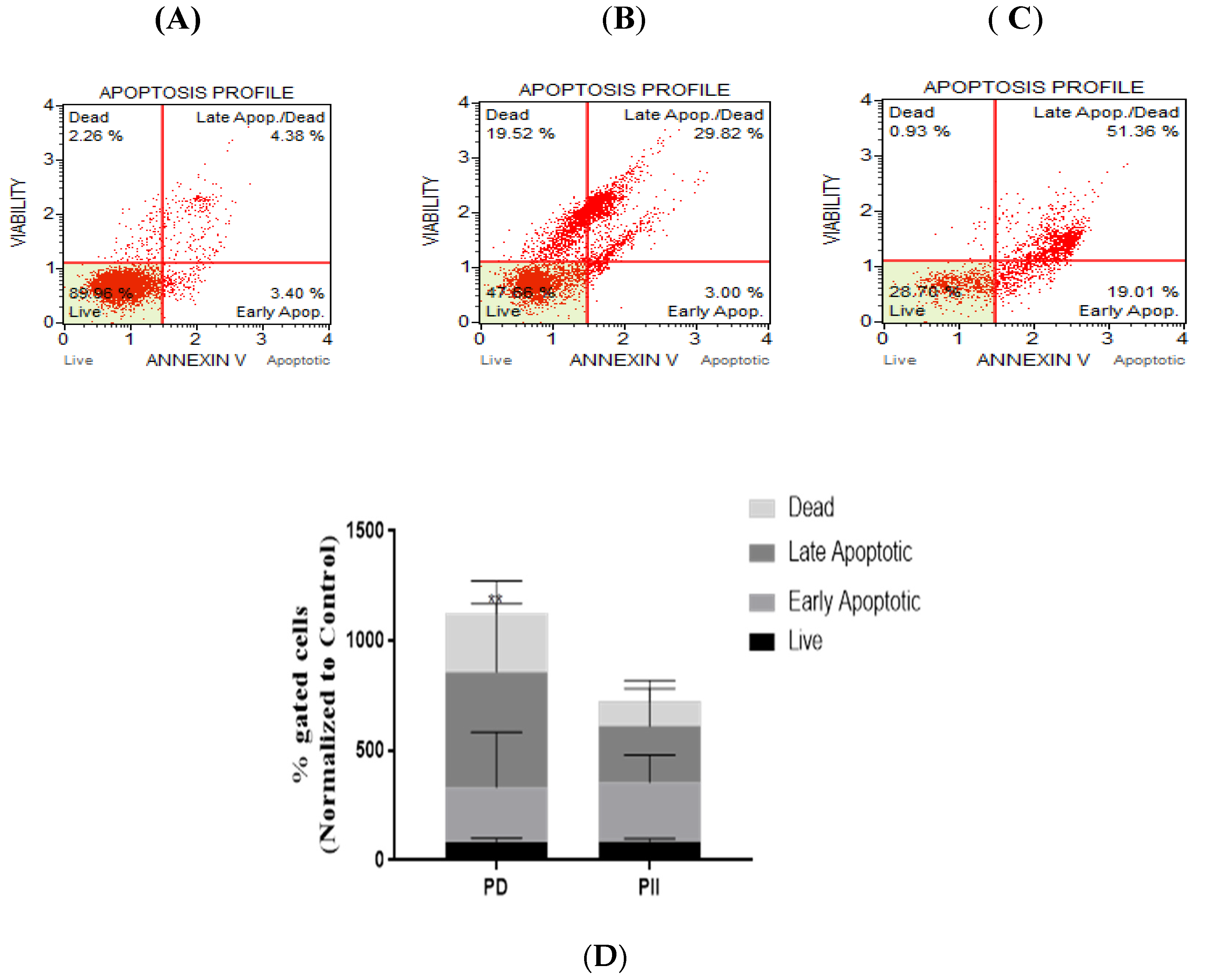
3.3. Elucidation of the Mechanism of Action of PD and PII Revealed Impact on Programmed Cell Death Pathway in DLD-1 Cells
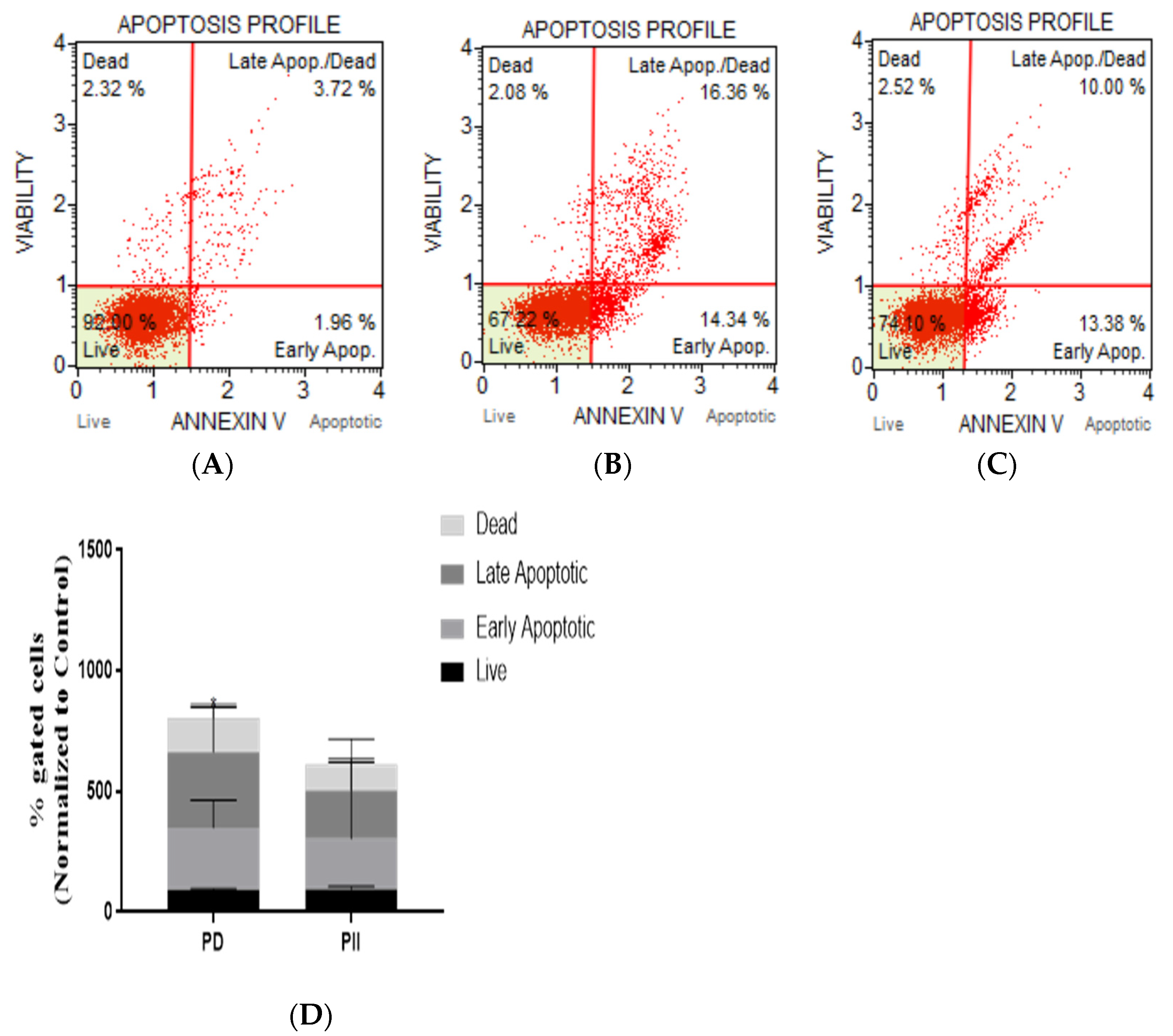
3.4. Studies on the Mechanism of Action of Polyphyllins PD and PII Revealed Activation of Programmed Cell Death Pathway in HCT116 Cells
3.5. Proteomic Analysis Revealed Key Entities of the Signaling Circuits in DLD-1 Cells That Were Differentially Regulated on Administration of PD and PII
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- United States Cancer Incidence, Centers for Disease Control and Prevention. Available online: https://gis.cdc.gov/Cancer/USCS/DataViz.html (accessed on 31 January 2022).
- Ayanian, J.Z.; Zaslavsky, A.M.; Fuchs, C.S.; Guadagnoli, E.; Creech, C.M.; Cress, R.D.; O’Connor, L.C.; West, D.W.; Allen, M.E.; Wolf, R.E.; et al. Use of adjuvant chemotherapy and radiation therapy for colorectal cancer in a population-based cohort. J. Clin. Oncol. 2003, 21, 1293–1300. [Google Scholar] [CrossRef]
- Edwards, M.S.; Chadda, S.D.; Zhao, Z.; Barber, B.L.; Sykes, D.P. A systematic review of treatment guidelines for metastatic colorectal cancer. Colorectal Dis. 2012, 14, e31–e47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fakih, M.G. Metastatic colorectal cancer: Current state and future directions. J. Clin. Oncol. 2015, 33, 1809–1824. [Google Scholar] [CrossRef] [PubMed]
- Folprecht, G.; Grothey, A.; Alberts, S.; Raab, H.R.; Kohne, C.H. Neoadjuvant treatment of unresectable colorectal liver metastases: Correlation between tumour response and resection rates. Ann. Oncol. 2005, 16, 1311–1319. [Google Scholar] [CrossRef] [PubMed]
- Bukhari, S.N.A.; Alotaibi, N.H.; Ahmad, W.; Alharbi, K.S.; Abdelgawad, M.A.; Al-Sanea, M.M.; Ahmad, M.M.; Amjad, M.W.; Raja, M.A.G.; Hussain, M.A. Evaluation of Ligustrazine-Based Synthetic Compounds for their Antiproliferative Effects. Med. Chem. 2021, 17, 956–962. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Qin, H.L.; Leng, J.; Ameeduzzafar; Amjad, M.W.; Raja, M.A.G.; Hussain, M.A.; Bukhari, S.N.A. Synthesis and biological evaluation of new tetramethylpyrazine-based chalcone derivatives as potential anti-Alzheimer agents. Chem. Biol. Drug Des. 2018, 92, 1859–1866. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Suzuki, T.; Hara, F.; Yasui, T.; Uga, N.; Naoe, A. Polyphyllin D, a steroidal saponin in Paris polyphylla, induces apoptosis and necroptosis cell death of neuroblastoma cells. Pediatr. Surg. Int. 2017, 33, 713–719. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zheng, Q.; Chen, W.; Wu, M.; Pan, G.; Yang, K.; Li, X.; Man, S.; Teng, Y.; Yu, P.; et al. Chemosensitizing effect of Paris Saponin I on Camptothecin and 10-hydroxycamptothecin in lung cancer cells via p38 MAPK, ERK, and Akt signaling pathways. Eur. J. Med. Chem. 2017, 125, 760–769. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Wu, J.; Zheng, F.; Tang, Q.; Wu, W.; Hann, S.S. Inhibition of EZH2 via activation of SAPK/JNK and reduction of p65 and DNMT1 as a novel mechanism in inhibition of human lung cancer cells by polyphyllin I. J. Exp. Clin. Cancer Res. 2016, 35, 112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, H.; Zhao, P.J.; Su, D.; Feng, J.; Ma, S.L. Paris saponin I induces apoptosis via increasing the Bax/Bcl-2 ratio and caspase-3 expression in gefitinib-resistant non-small cell lung cancer in vitro and in vivo. Mol. Med. Rep. 2014, 9, 2265–2272. [Google Scholar] [CrossRef] [PubMed]
- Gu, L.; Feng, J.; Zheng, Z.; Xu, H.; Yu, W. Polyphyllin I inhibits the growth of ovarian cancer cells in nude mice. Oncol. Lett. 2016, 12, 4969–4974. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheung, J.Y.; Ong, R.C.; Suen, Y.K.; Ooi, V.; Wong, H.N.; Mak, T.C.; Fung, K.P.; Yu, B.; Kong, S.K. Polyphyllin D is a potent apoptosis inducer in drug-resistant HepG2 cells. Cancer Lett. 2005, 217, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.; Dai, X.; He, X.; Zhou, R.; Zhang, B.; Hu, G.; Huang, Z.; Fan, X. Paris saponin I induces G(2)/M cell cycle arrest and apoptosis in human gastric carcinoma SGC7901 cells. J. Huazhong Univ. Sci. Technol. Med. Sci. 2011, 31, 768–772. [Google Scholar] [CrossRef] [PubMed]
- Yue, G.; Wei, J.; Qian, X.; Yu, L.; Zou, Z.; Guan, W.; Wang, H.; Shen, J.; Liu, B. Synergistic anticancer effects of polyphyllin I and evodiamine on freshly-removed human gastric tumors. PLoS ONE 2013, 8, e65164. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Li, Q.; Lu, P.; Chen, Q. Polyphyllin D induces apoptosis in U87 human glioma cells through the c-Jun NH2-terminal kinase pathway. J. Med. Food 2014, 17, 1036–1042. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Li, X.; He, X.; Qiu, X.; Jin, X.L.; Zhao, X.Y.; Xu, R.Z. Polyphyllin I induces cell cycle arrest and apoptosis in human myeloma cells via modulating beta-catenin signaling pathway. Eur. J. Haematol. 2016, 97, 371–378. [Google Scholar] [CrossRef]
- Cai, X.; Guo, L.; Pei, F.; Chang, X.; Zhang, R. Polyphyllin G exhibits antimicrobial activity and exerts anticancer effects on human oral cancer OECM-1 cells by triggering G2/M cell cycle arrest by inactivating cdc25C-cdc2. Arch. Biochem. Biophys. 2018, 644, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, E.; Li, P. Chemotaxonomic studies of nine Paris species from China based on ultra-high performance liquid chromatography tandem mass spectrometry and Fourier transform infrared spectroscopy. J. Pharm. Biomed. Anal. 2017, 140, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Wong, E.L.; Shum, T.Y.; Che, C.M.; Hui, Y. Fluorophore-appended steroidal saponin (dioscin and polyphyllin D) derivatives. Org. Lett. 2005, 7, 669–672. [Google Scholar] [CrossRef] [PubMed]
- Man, S.; Gao, W.; Zhang, Y.; Huang, L.; Liu, C. Chemical study and medical application of saponins as anti-cancer agents. Fitoterapia 2010, 81, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.C.; Hsieh, M.J.; Chen, C.J.; Lin, J.T.; Lo, Y.S.; Chuang, Y.C.; Chien, S.Y.; Chen, M.K. Polyphyllin G induce apoptosis and autophagy in human nasopharyngeal cancer cells by modulation of AKT and mitogen-activated protein kinase pathways in vitro and in vivo. Oncotarget 2016, 7, 70276–70289. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hsieh, M.J.; Chien, S.Y.; Lin, J.T.; Yang, S.F.; Chen, M.K. Polyphyllin G induces apoptosis and autophagy cell death in human oral cancer cells. Phytomedicine 2016, 23, 1545–1554, Erratum in Phytomedicine 2018, 45, 120. [Google Scholar] [CrossRef] [PubMed]
- Lodise, O.; Patil, K.; Karshenboym, I.; Prombo, S.; Chukwueke, C.; Pai, S.B. Inhibition of Prostate Cancer Cells by 4,5-Dicaffeoylquinic Acid through Cell Cycle Arrest. Prostate Cancer 2019, 2019, 4520645. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, H.; Patil, K.; Vashi, A.; Wang, Y.; Strickland, E.; Pai, S.B. Cellular molecular and proteomic profiling deciphers the SIRT1 controlled cell death pathways in esophageal adenocarcinoma cells. Cancer Treat. Res. Commun. 2021, 26, 100271. [Google Scholar] [CrossRef] [PubMed]
- Dhir, H.; Choudhury, M.; Patil, K.; Cheung, C.; Bodlak, A.; Pardo, D.; Adams, A.; Travaglino, S.; Rojas, J.A.; Pai, S.B. Interception of Signaling Circuits of Esophageal Adenocarcinoma Cells by Resveratrol Reveals Molecular and Immunomodulatory Signatures. Cancers 2021, 13, 5811. [Google Scholar] [CrossRef] [PubMed]
- Jayasena, T.; Poljak, A.; Braidy, N.; Zhong, L.; Rowlands, B.; Muenchhoff, J.; Grant, R.; Smythe, G.; Teo, C.; Raftery, M.; et al. Application of Targeted Mass Spectrometry for the Quantification of Sirtuins in the Central Nervous System. Sci. Rep. 2016, 6, 35391. [Google Scholar] [CrossRef] [Green Version]
- Shimbara, N.; Orino, E.; Sone, S.; Ogura, T.; Takashina, M.; Shono, M.; Tamura, T.; Yasuda, H.; Tanaka, K.; Ichihara, A. Regulation of gene expression of proteasomes (multi-protease complexes) during growth and differentiation of human hematopoietic cells. J. Biol. Chem. 1992, 267, 18100–18109. [Google Scholar] [CrossRef]
- Teicher, B.A.; Tomaszewski, J.E. Proteasome inhibitors. Biochem. Pharmacol. 2015, 96, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Ahmad, S.S.; Glatzle, J.; Bajaeifer, K.; Buhler, S.; Lehmann, T.; Konigsrainer, I.; Vollmer, J.P.; Sipos, B.; Ahmad, S.S.; Northoff, H.; et al. Phosphoglycerate kinase 1 as a promoter of metastasis in colon cancer. Int. J. Oncol. 2013, 43, 586–590. [Google Scholar] [CrossRef] [Green Version]
- Li, H.X.; Sun, X.Y.; Yang, S.M.; Wang, Q.; Wang, Z.Y. Peroxiredoxin 1 promoted tumor metastasis and angiogenesis in colorectal cancer. Pathol. Res. Pract. 2018, 214, 655–660. [Google Scholar] [CrossRef]
- Fischer, J.; Eglinton, T.W.; Frizelle, F.A.; Hampton, M.B. Peroxiredoxins in Colorectal Cancer: Predictive Biomarkers of Radiation Response and Therapeutic Targets to Increase Radiation Sensitivity? Antioxidants 2018, 7, 136. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, S.; Ma, Y.; Tong, Q.; Yang, J.; Liu, J.; Wang, Y.; Li, G.; Zeng, J.; Fang, S.; Li, F.; et al. Cullin-5 neddylation-mediated NOXA degradation is enhanced by PRDX1 oligomers in colorectal cancer. Cell Death Dis. 2021, 12, 265. [Google Scholar] [CrossRef]
- Hu, X.; Du, S.; Yu, J.; Yang, X.; Yang, C.; Zhou, D.; Wang, Q.; Qin, S.; Yan, X.; He, L.; et al. Common housekeeping proteins are upregulated in colorectal adenocarcinoma and hepatocellular carcinoma, making the total protein a better “housekeeper”. Oncotarget 2016, 7, 66679–66688. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pan, X.; Hobbs, R.P.; Coulombe, P.A. The expanding significance of keratin intermediate filaments in normal and diseased epithelia. Curr. Opin. Cell Biol. 2013, 25, 47–56. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferraro, A.; Boni, T.; Pintzas, A. EZH2 regulates cofilin activity and colon cancer cell migration by targeting ITGA2 gene. PLoS ONE 2014, 9, e115276. [Google Scholar] [CrossRef]
- Mouneimne, G.; Hansen, S.D.; Selfors, L.M.; Petrak, L.; Hickey, M.M.; Gallegos, L.L.; Simpson, K.J.; Lim, J.; Gertler, F.B.; Hartwig, J.H.; et al. Differential remodeling of actin cytoskeleton architecture by profilin isoforms leads to distinct effects on cell migration and invasion. Cancer Cell 2012, 22, 615–630. [Google Scholar] [CrossRef] [Green Version]
- Karamouzis, M.V.; Gorgoulis, V.G.; Papavassiliou, A.G. Transcription factors and neoplasia: Vistas in novel drug design. Clin. Cancer Res. 2002, 8, 949–961. [Google Scholar]
- Xu, T.; Zou, Q.; Wu, J.; Yu, B.; Xu, Z.; Cai, H.; Zhang, W. Heterogeneous nuclear ribonucleoprotein U-like 1 and Poly (ADP-ribose) polymerase 1 are downregulated in renal cell carcinoma and connected with the prognosis. Cancer Biomark. 2013, 13, 411–415. [Google Scholar] [CrossRef] [PubMed]
- Oji, Y.; Tatsumi, N.; Fukuda, M.; Nakatsuka, S.; Aoyagi, S.; Hirata, E.; Nanchi, I.; Fujiki, F.; Nakajima, H.; Yamamoto, Y.; et al. The translation elongation factor eEF2 is a novel tumorassociated antigen overexpressed in various types of cancers. Int. J. Oncol. 2014, 44, 1461–1469. [Google Scholar] [CrossRef] [Green Version]
- Sun, W.; Wei, X.; Niu, A.; Ma, X.; Li, J.J.; Gao, D. Enhanced anti-colon cancer immune responses with modified eEF2-derived peptides. Cancer Lett. 2015, 369, 112–123. [Google Scholar] [CrossRef] [Green Version]
- Sharma, R.; Palanisamy, A.; Dhama, K.; Mal, G.; Singh, B.; Singh, K.P. Exploring the possible use of saponin adjuvants in COVID-19 vaccine. Hum. Vaccines Immunother. 2020, 16, 2944–2953. [Google Scholar] [CrossRef] [PubMed]
- Skene, C.D.; Sutton, P. Saponin-adjuvanted particulate vaccines for clinical use. Methods 2006, 40, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Fraga, M.F.; Ballestar, E.; Villar-Garea, A.; Boix-Chornet, M.; Espada, J.; Schotta, G.; Bonaldi, T.; Haydon, C.; Ropero, S.; Petrie, K.; et al. Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat. Genet. 2005, 37, 391–400. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, B.; Gamallat, Y.; Khan, M.F.; Din, S.R.; Israr, M.; Ahmad, M.; Tahir, N.; Azam, N.; Rahman, K.U.; Xin, W.; et al. Natural Polyphyllins (I, II, D, VI, VII) Reverses Cancer Through Apoptosis, Autophagy, Mitophagy, Inflammation, and Necroptosis. Onco Targets Ther. 2021, 14, 1821–1841. [Google Scholar] [CrossRef] [PubMed]

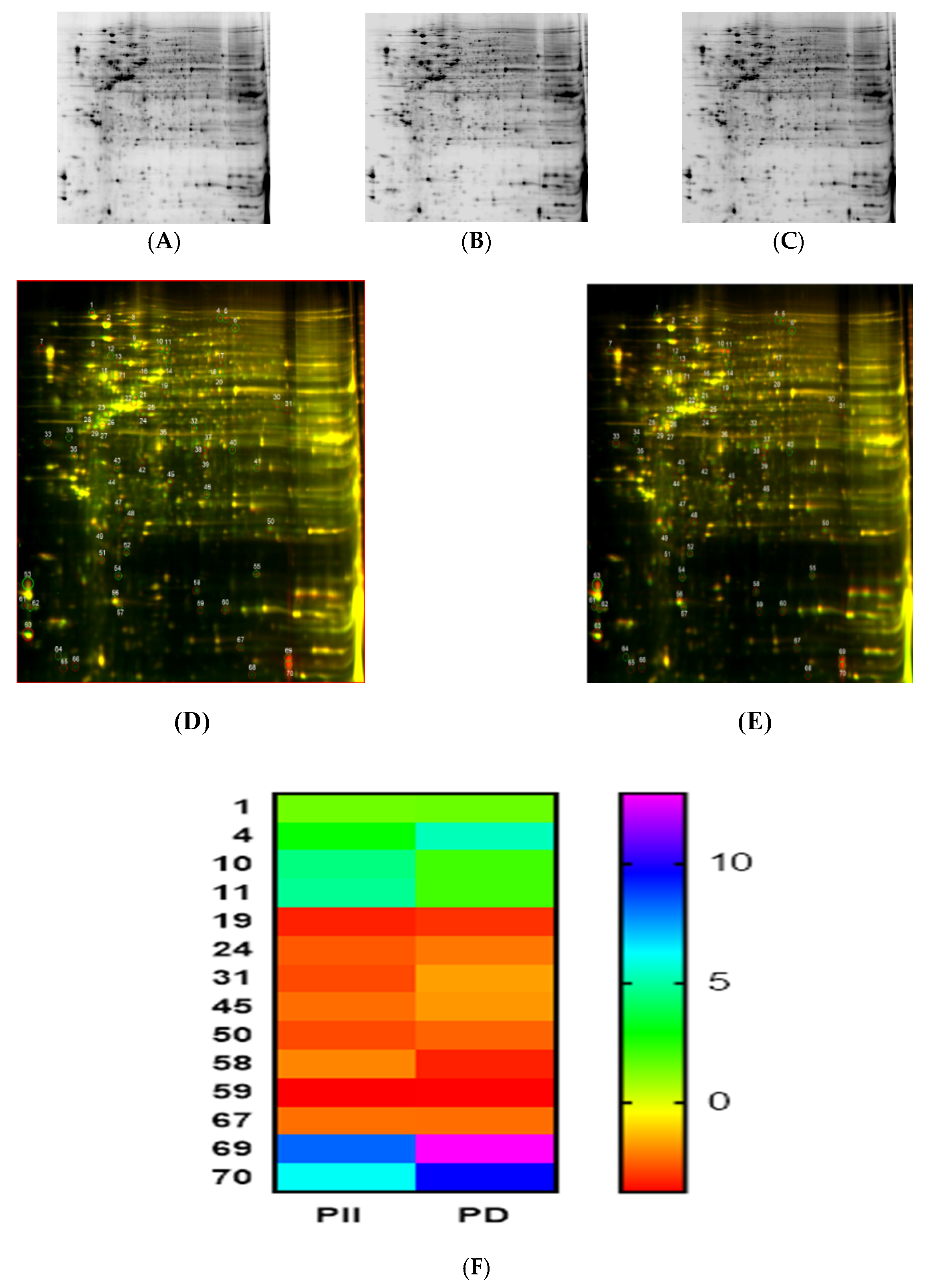
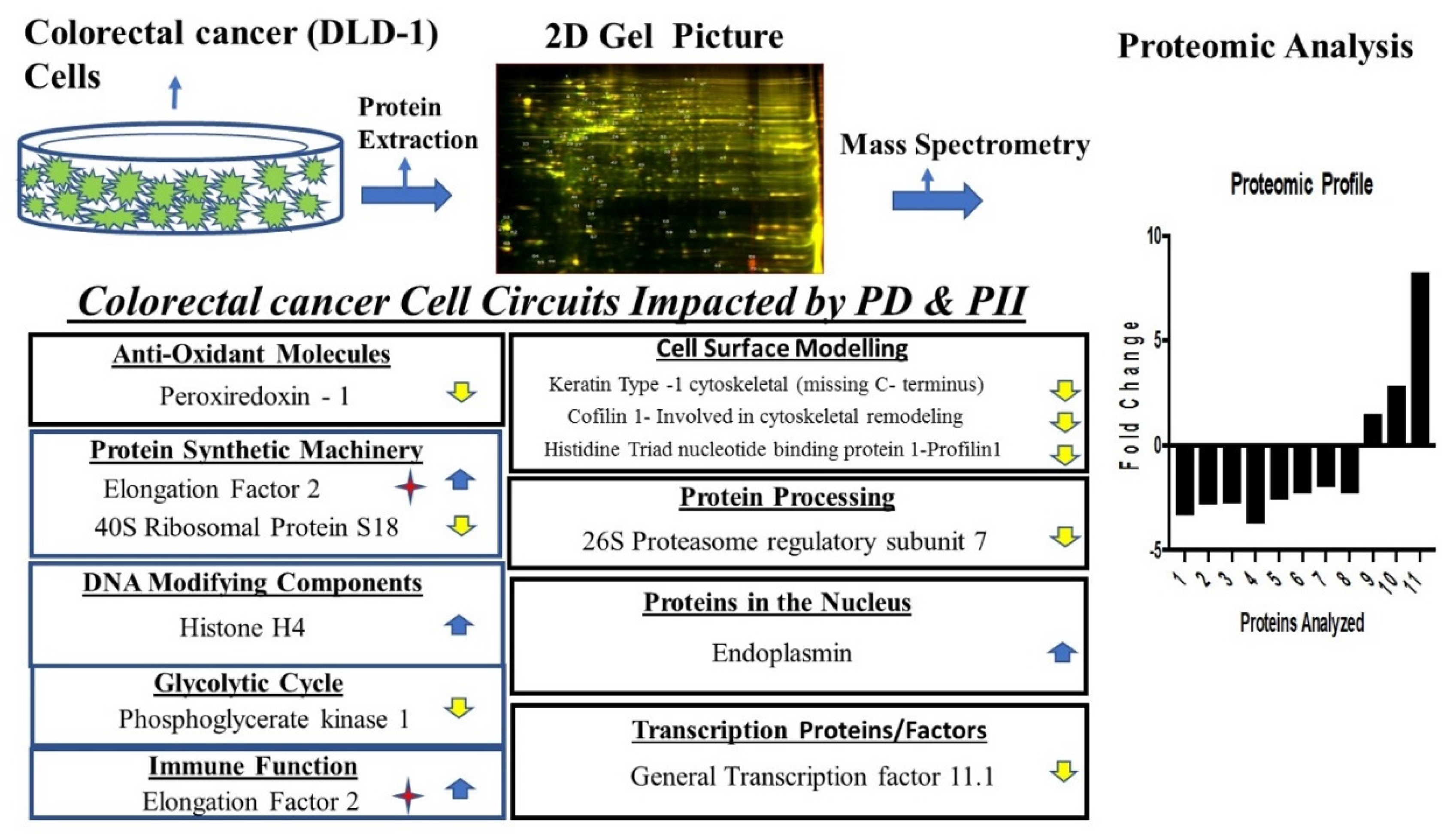
| SI # | Fold Change | Proteins Identified |
|---|---|---|
| PII PD | ||
| 1 | −3.33 −3.10 | 26S Proteasome regulatory subunit 7 |
| 2 | −2.81 −1.67 | Phosphoglycerate Kinase 1 |
| 3 | −2.80 −2.46 | Peroxiredoxin-1 |
| 4 | −3.74 −3.73 | 40S Ribosomal Protein S18 |
| 5 | −2.60 −2.20 | Keratin type 1 cytoskeletal |
| 6 | −2.32 −1.77 | General Transcription factor 11.1 |
| 7 | −2.01 −3.33 | Cofilin 1 |
| 8 | −2.30 −2.33 | Histidine triad nucleotide-binding protein 1-Profilin1 |
| 9 | +1.48 +1.55 | Endoplasmin |
| 10 | +2.88 +5.33 | Elongation Factor 2 |
| 11 | +8.28 +12.94 | Histone H4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Siripuram, R.; Bartolek, Z.; Patil, K.; Gill, S.S.; Pai, S.B. Disruption of Colorectal Cancer Network by Polyphyllins Reveals Pivotal Entities with Implications for Chemoimmunotherapy. Biomedicines 2022, 10, 583. https://doi.org/10.3390/biomedicines10030583
Siripuram R, Bartolek Z, Patil K, Gill SS, Pai SB. Disruption of Colorectal Cancer Network by Polyphyllins Reveals Pivotal Entities with Implications for Chemoimmunotherapy. Biomedicines. 2022; 10(3):583. https://doi.org/10.3390/biomedicines10030583
Chicago/Turabian StyleSiripuram, Ram, Zinka Bartolek, Ketki Patil, Saj S. Gill, and S. Balakrishna Pai. 2022. "Disruption of Colorectal Cancer Network by Polyphyllins Reveals Pivotal Entities with Implications for Chemoimmunotherapy" Biomedicines 10, no. 3: 583. https://doi.org/10.3390/biomedicines10030583
APA StyleSiripuram, R., Bartolek, Z., Patil, K., Gill, S. S., & Pai, S. B. (2022). Disruption of Colorectal Cancer Network by Polyphyllins Reveals Pivotal Entities with Implications for Chemoimmunotherapy. Biomedicines, 10(3), 583. https://doi.org/10.3390/biomedicines10030583






