An Apple a Day Keeps the Doctor Away: Potential Role of miRNA 146 on Macrophages Treated with Exosomes Derived from Apples
Abstract
:1. Introduction
2. Materials and Methods
2.1. ADNVs Isolation and Purification
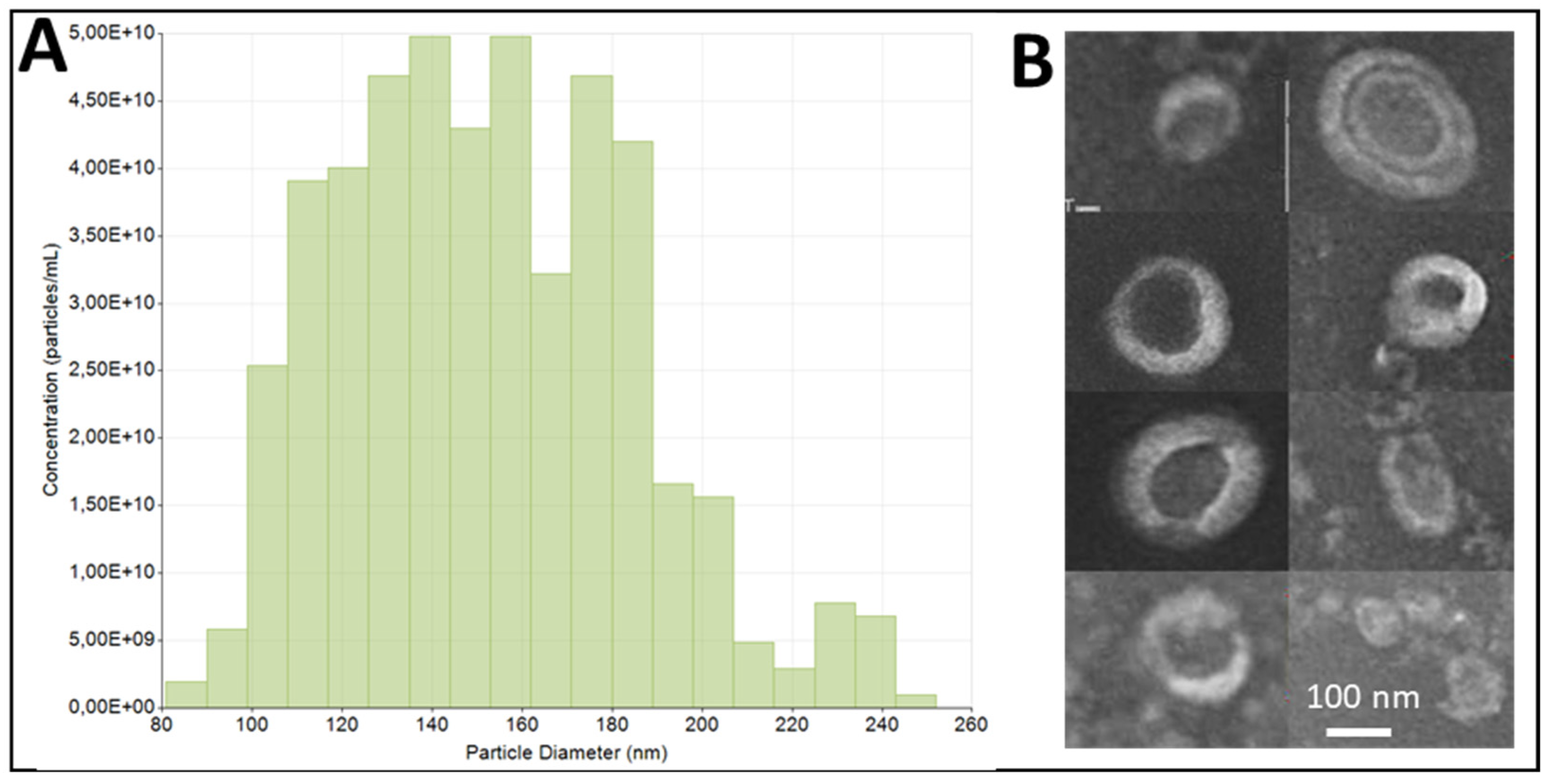
2.2. Size Distribution and Concentration Measurement of ADNVs
2.3. Cell Culture
2.4. Transmission Electron Microscopy
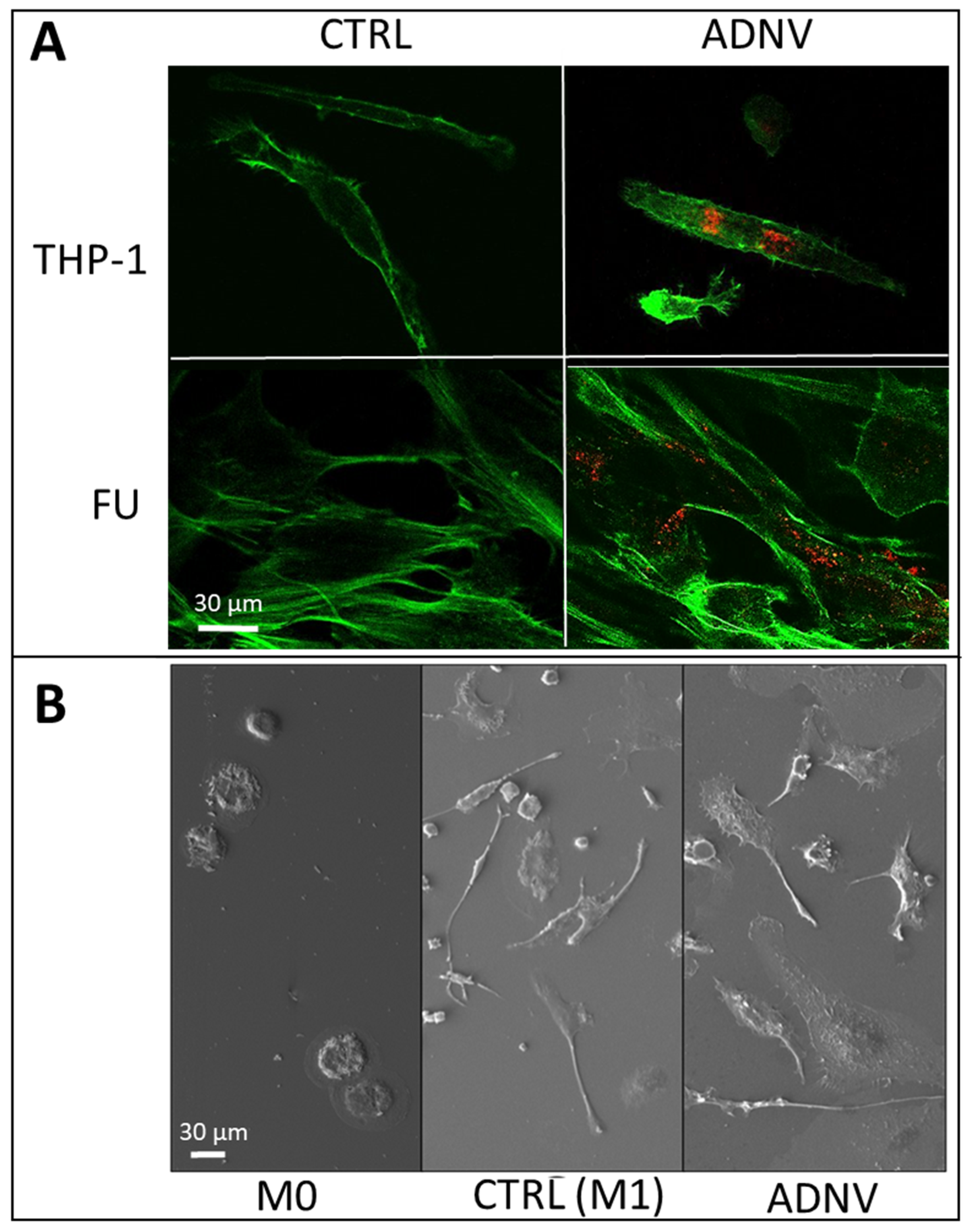
2.5. Scanning Electron Microscopy
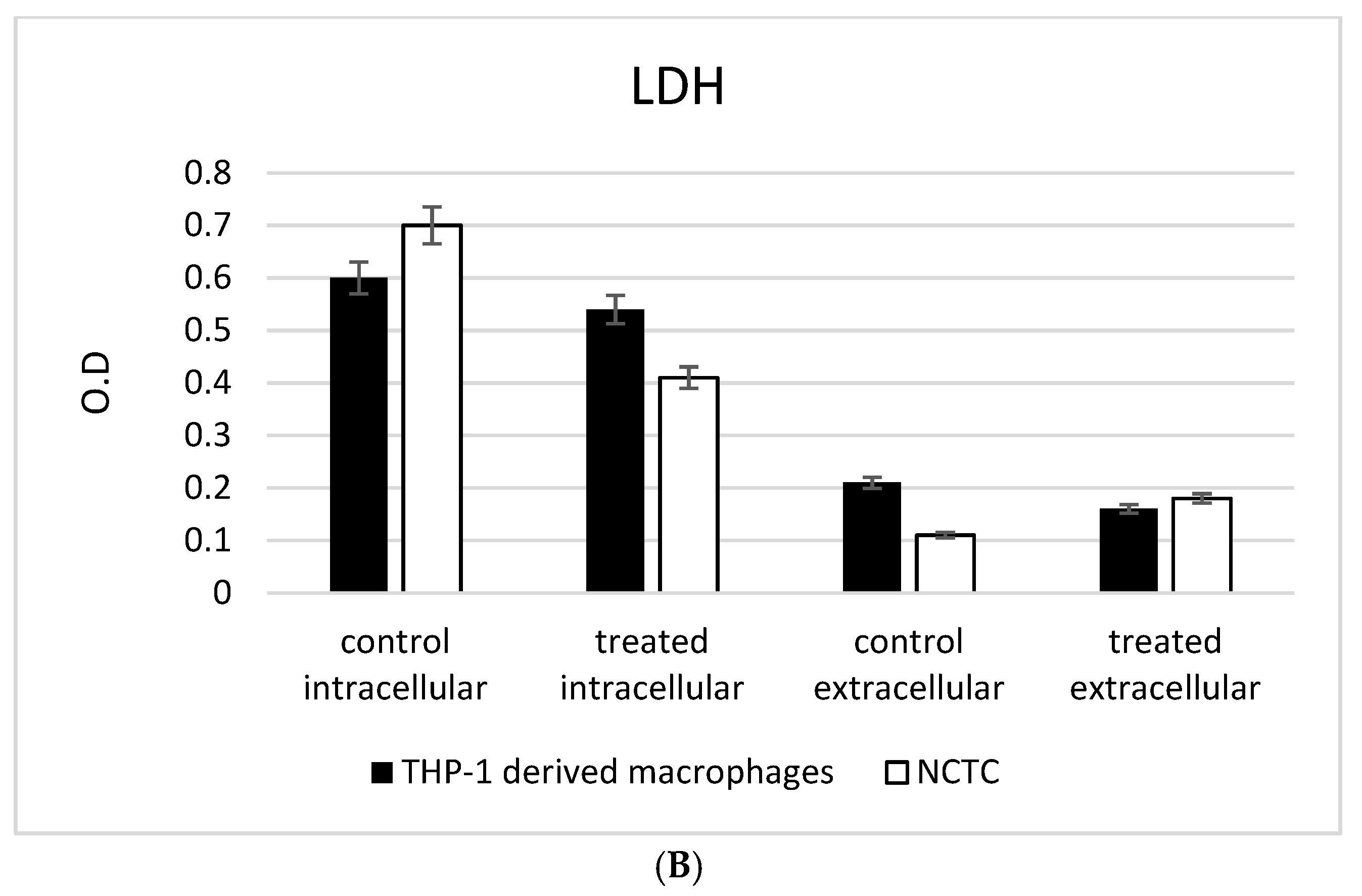
2.6. LDH and MTT Assays
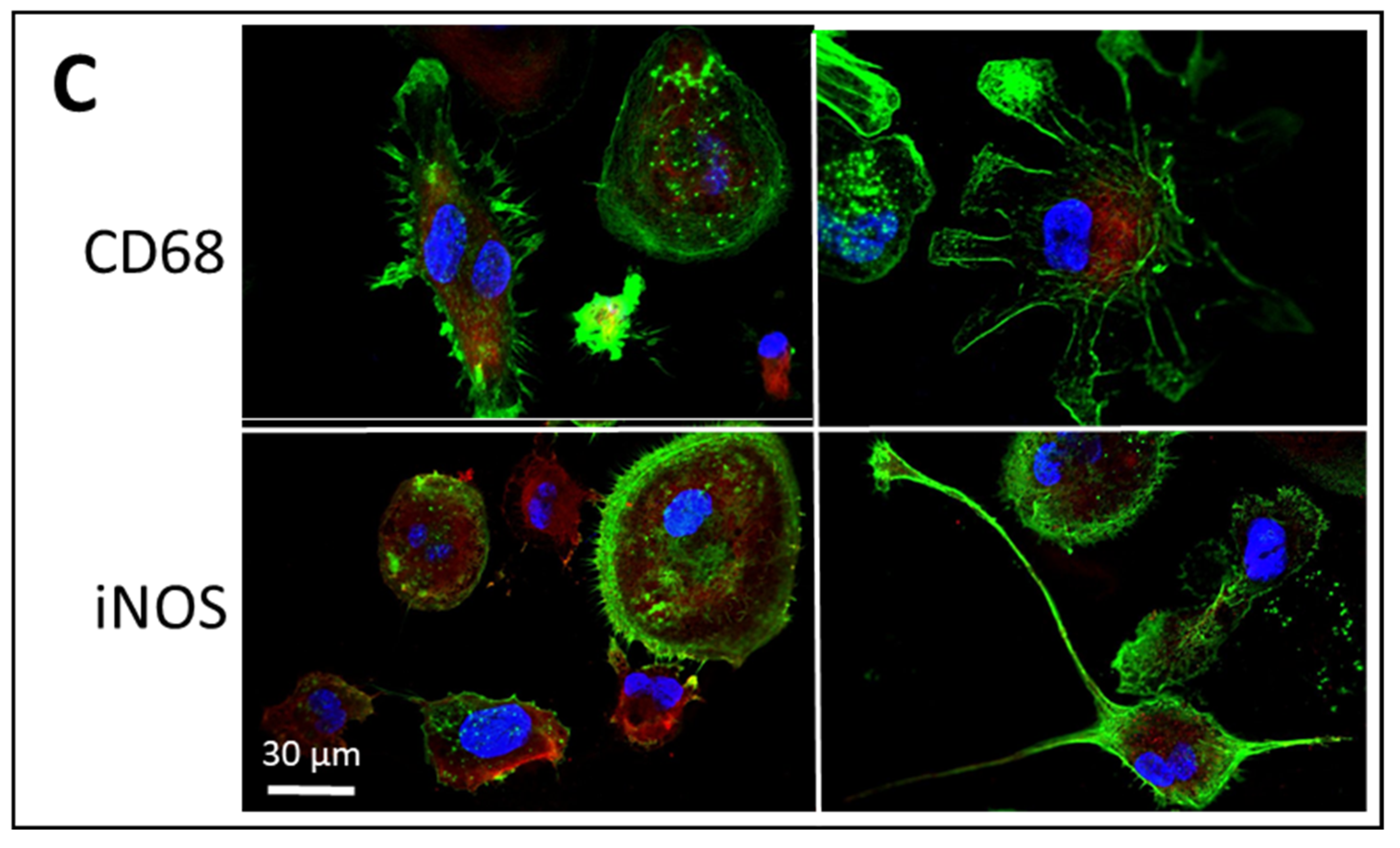
2.7. Fluorescence Labelling and Imaging
2.8. RNA Extraction, Sequencing, and RT-qPCR
2.9. Statistical Analysis
3. Results
3.1. ADNV Characterization
3.2. Uptake of ADNVs
3.3. ADNVs Effects on Cell Viability
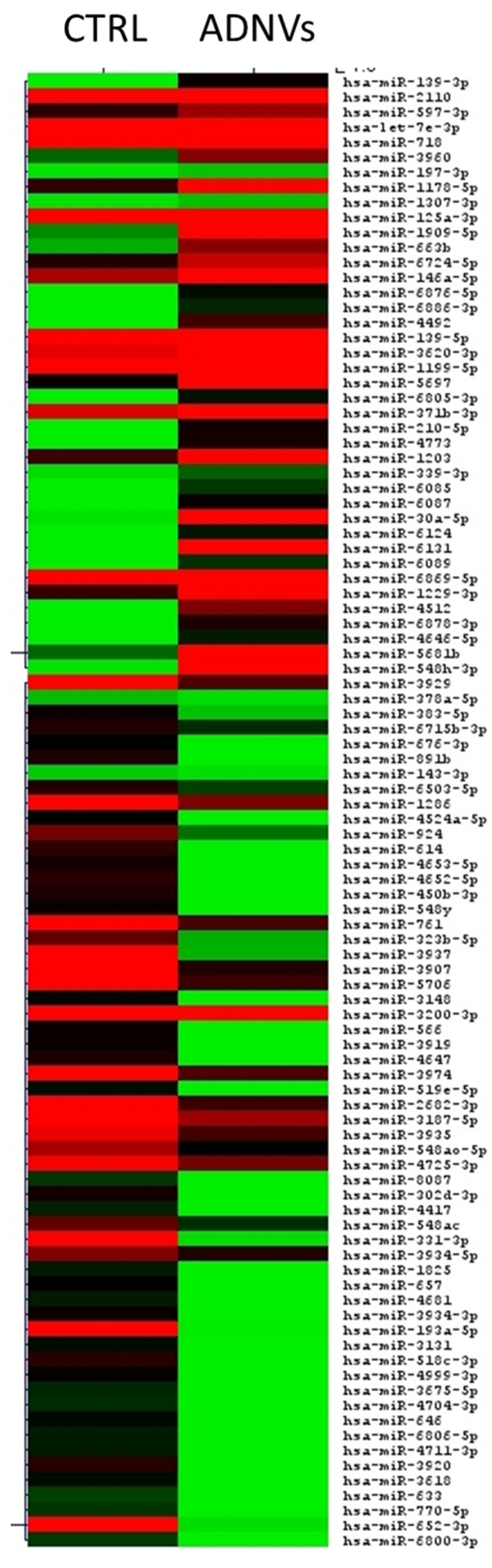
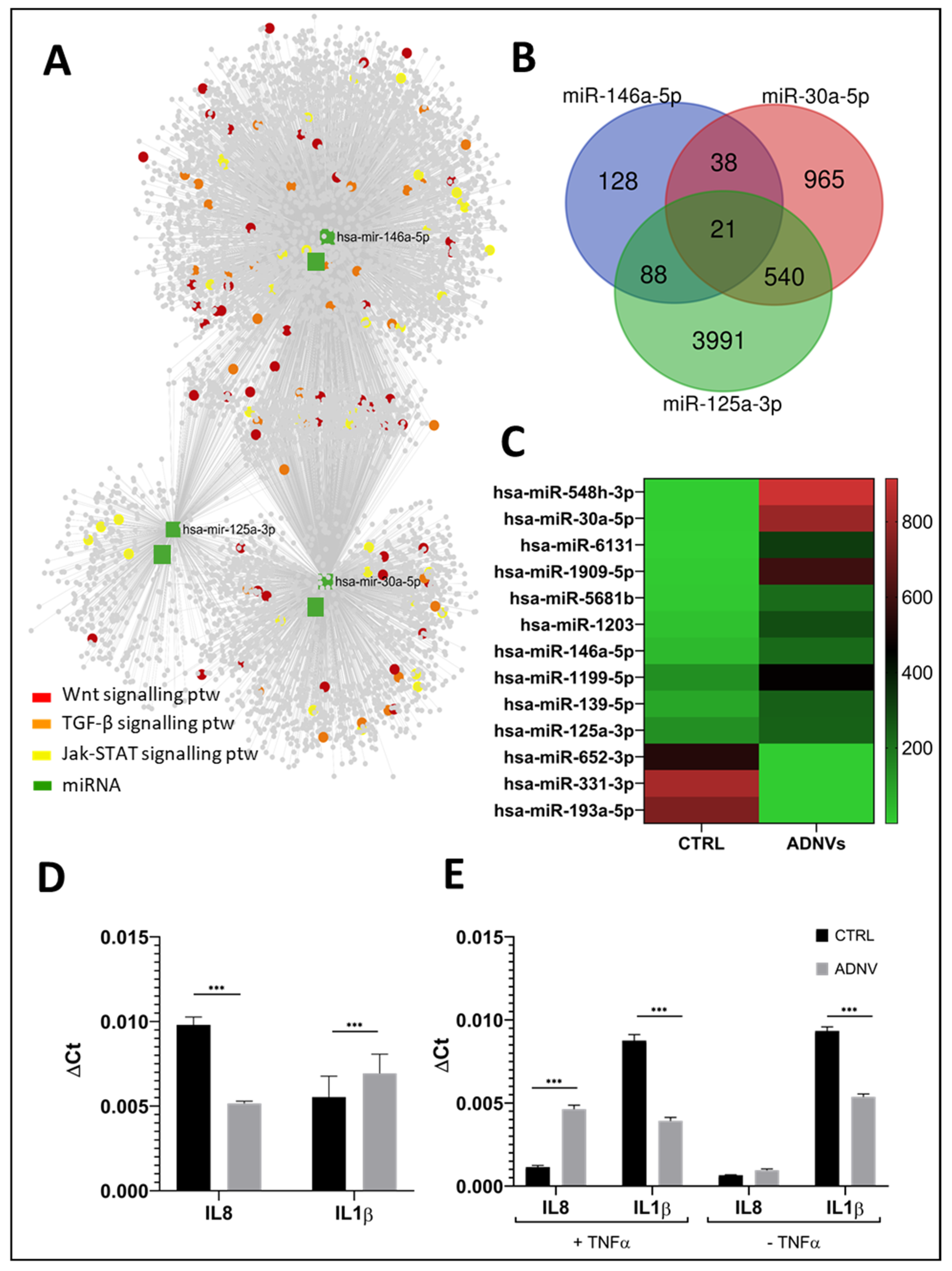
3.4. Differential miRNA Expression Analysis and Target Gene Prediction
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tsao, R. Chemistry and biochemistry of dietary polyphenols. Nutrients 2010, 2, 1231–1246. [Google Scholar] [CrossRef]
- Frankel, E.N. Nutritional Benefits of Flavonoids. Food Factors Cancer Prevention; Springer: Tokyo, Japan, 1997; pp. 613–616. [Google Scholar]
- Dias, J.S. Nutritional Quality and Health Benefits of Vegetables: A Review. Food Nutr. Sci. 2012, 3, 1354–1374. [Google Scholar] [CrossRef] [Green Version]
- Kaparapu, J.; Pragada, P.M.; Geddada, M.N.R. Fruits and Vegetables and its Nutritional Benefits. In Functional Foods and Nutraceuticals; Springer: Cham, Switzerland, 2020; pp. 241–260. [Google Scholar]
- Das, L.; Bhaumik, E.; Raychaudhuri, U.; Chakraborty, R. Role of nutraceuticals in human health. J. Food Sci. Technol. 2012, 49, 173–183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baldini, N.; Torreggiani, E.; Roncuzzi, L.; Perut, F.; Zini, N.; Avnet, S. Exosome-like Nanovesicles Isolated from Citrus limon L. Exert Antioxidative Effect. Curr. Pharm. Biotechnol. 2018, 19, 877–885. [Google Scholar] [CrossRef] [PubMed]
- Perut, F.; Roncuzzi, L.; Avnet, S.; Massa, A.; Zini, N.; Sabbadini, S.; Giampieri, F.; Mezzetti, B.; Baldini, N. Strawberry-Derived Exosome-Like Nanoparticles Prevent Oxidative Stress in Human Mesenchymal Stromal Cells. Biomolecules 2021, 11, 87. [Google Scholar] [CrossRef] [PubMed]
- Regente, M.; Corti-Monzón, G.; Maldonado, A.M.; Pinedo, M.; Jorrín, J.; de la Canal, L. Vesicular fractions of sunflower apoplastic fluids are associated with potential exosome marker proteins. FEBS Lett. 2009, 583, 3363–3366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chekanova, J.A.; Gregory, B.D.; Reverdatto, S.V.; Chen, H.; Kumar, R.; Hooker, T.; Yazaki, J.; Li, P.; Skiba, N.; Peng, Q.; et al. Genome-Wide High-Resolution Mapping of Exosome Substrates Reveals Hidden Features in the Arabidopsis Transcriptome. Cell 2007, 131, 1340–1353. [Google Scholar] [CrossRef] [Green Version]
- Sagini, K.; Urbanelli, L.; Buratta, S.; Leonardi, L.; Emiliani, C. Nanovesicles from Plants as Edible Carriers of Bioactive Compounds. AgroLife Sci. J. 2017, 6, 167–171. [Google Scholar]
- Kalarikkal, S.P.; Prasad, D.; Kasiappan, R.; Chaudhari, S.R.; Sundaram, G.M. A cost-effective polyethylene glycol-based method for the isolation of functional edible nanoparticles from ginger rhizomes. Sci. Rep. 2020, 10, 4456. [Google Scholar] [CrossRef] [PubMed]
- di Gioia, S.; Conese, M. Biological properties and therapeutic effects of plant—Derived nanovesicles. Open Med. 2020, 15, 1096–1122. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Zhang, M.; Merlin, D. Advances in Plant-derived Edible Nanoparticle-based lipid Nano-drug Delivery Systems as Therapeutic Nanomedicines. J. Mater. Chem. B 2018, 6, 1312–1321. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Qiao, L.; Wang, M.; He, B.; Lin, F.M.; Palmquist, J.; Huang, S.D.; Jin, H. Plants send small RNAs in extracellular vesicles to fungal pathogen to silence virulence genes. Science 2018, 360, 1126–1129. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, Q.; Hückelhoven, R.; Kogel, K.H.; Van Bel, A.J. Multivesicular bodies participate in a cell wall-associated defence response in barley leaves attacked by the pathogenic powdery mildew fungus. Cell. Microbiol. 2006, 8, 1009–1019. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Chen, X. Intercellular and systemic trafficking of RNAs in plants. Nat. Plants 2018, 4, 869–878. [Google Scholar] [CrossRef] [PubMed]
- Mu, J.; Zhuang, X.; Wang, Q.; Jiang, H.; Deng, Z.B.; Wang, B.; Zhang, L.; Kakar, S.; Jun, Y.; Miller, D.; et al. Interspecies communication between plant and mouse gut host cells through edible plant derived exosome-like nanoparticles. Mol. Nutr. Food Res. 2014, 58, 1561–1573. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Hou, D.; Chen, X.; Li, D.; Zhu, L.; Zhang, Y.; Li, J.; Bian, Z.; Liang, X.; Cai, X.; et al. Exogenous plant MIR168a specifically targets mammalian LDLRAP1: Evidence of cross-kingdom regulation by microRNA. Cell Res. 2012, 22, 107–126. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Viennois, E.; Prasad, M.; Zhang, Y.; Wang, L.; Zhang, Z.; Han, M.K.; Xiao, B.; Xu, C.; Srinivasan, S.; et al. Edible ginger-derived nanoparticles: A novel therapeutic approach for the prevention and treatment of in-flammatory bowel disease and colitis-associated cancer. Biomaterials 2016, 101, 321–340. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujita, D.; Arai, T.; Komori, H.; Shirasaki, Y.; Wakayama, T.; Nakanishi, T.; Tamai, I. Apple-Derived Nanoparticles Modulate Expression of Organic-Anion-Transporting Polypeptide (OATP) 2B1 in Caco-2 Cells. Mol. Pharm. 2018, 15, 5772–5780. [Google Scholar] [CrossRef] [PubMed]
- Deng, Z.; Rong, Y.; Teng, Y.; Mu, J.; Zhuang, X.; Tseng, M.; Samykutty, A.; Zhang, L.; Yan, J.; Miller, D.; et al. Broccoli-Derived Nanoparticle Inhibits Mouse Colitis by Activating Dendritic Cell AMP-Activated Protein Kinase. Mol. Ther. 2017, 25, 1641–1654. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, I.-M.; Rajakumar, G.; Venkidasamy, B.; Subramanian, U.; Thiruvengadam, M. Exosomes: Current use and future applications. Clin. Chim. Acta 2020, 500, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Kalinowska, M.; Bielawska, A.; Lewandowska-Siwkiewicz, H.; Priebe, W.; Lewandowski, W. Apples: Content of phenolic compounds vs. variety, part of apple and cultivation model, extraction of phenolic compounds, biological properties. Plant Physiol. Biochem. 2014, 84, 169–188. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.H. Health benefits of fruit and vegetables are from additive and synergistic combinations of phytochemicals. Am. J. Clin. Nutr. 2003, 78, 517S–520S. [Google Scholar] [CrossRef]
- Charde, M.S.; Ahmed, A.; Chakole, R.D. Apple phytochemicals for human benefits. Int. J. Pharmacol. Res. 2012, 1, 40. [Google Scholar] [CrossRef]
- Migicovsky, Z.; Gardner, K.M.; Richards, C.; Chao, C.T.; Schwaninger, H.R.; Fazio, G.; Zhong, G.-Y.; Myles, S. Genomic consequences of apple improvement. Hortic. Res. 2021, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Boyer, J.; Liu, R.H. Apple phytochemicals and their health benefits. Nutr. J. 2004, 3, 5. [Google Scholar] [CrossRef] [Green Version]
- Baxter, E.; Graham, A.; Re, N.; Carr, I.; Robinson, J.; Mackie, S.; Morgan, A. Standardized protocols for differentiation of THP-1 cells to macrophages with distinct M(IFNγ+LPS), M(IL-4) and M(IL-10) phenotypes. J. Immunol. Methods 2020, 478, 112721. [Google Scholar] [CrossRef]
- Breslin, S.; O’Driscoll, L. The relevance of using 3D cell cultures, in addition to 2D monolayer cultures, when evaluating breast cancer drug sensitivity and resistance. Oncotarget 2016, 7, 45745–45756. [Google Scholar] [CrossRef] [Green Version]
- Rostam, H.M.; Reynolds, P.M.; Alexander, M.R.; Gadegaard, N.; Ghaemmaghami, A.M. Image based Machine Learn-ing for identification of macrophage subsets OPEN. Sci. Rep. 2017, 7, 3521. [Google Scholar] [CrossRef]
- Zhu, X.; Liu, Q.; Wang, M.; Liang, M.; Yang, X.; Xu, X.; Zou, H.; Qiu, J. Activation of Sirt1 by Resveratrol Inhibits TNF-α Induced Inflammation in Fibroblasts. PLoS ONE 2011, 6, e27081. [Google Scholar] [CrossRef]
- Ju, S.; Mu, J.; Dokland, T.; Zhuang, X.; Wang, Q.; Jiang, H.; Xiang, X.; Deng, Z.-B.; Wang, B.; Zhang, L.; et al. Grape Exosome-like Nanoparticles Induce Intestinal Stem Cells and Protect Mice From DSS-Induced Colitis. Mol. Ther. 2013, 21, 1345–1357. [Google Scholar] [CrossRef] [Green Version]
- Murray, P.J.; Allen, J.E.; Biswas, S.K.; Fisher, E.A.; Gilroy, D.W.; Goerdt, S.; Gordon, S.; Hamilton, J.A.; Ivashkiv, L.B.; Lawrence, T.; et al. Macrophage Activation and Polarization: Nomenclature and Experimental Guidelines. Immunity 2014, 41, 14–20. [Google Scholar] [CrossRef] [Green Version]
- Shiratori, H.; Feinweber, C.; Luckhardt, S.; Linke, B.; Resch, E.; Geisslinger, G.; Weigert, A.; Parnham, M.J. THP-1 and human peripheral blood mononuclear cell-derived macrophages differ in their capacity to polarize in vitro. Mol. Immunol. 2017, 88, 58–68. [Google Scholar] [CrossRef] [PubMed]
- DMcGonagle; Sharif, K.; O’Regan, A.; Bridgewood, C. The Role of Cytokines including Interleukin-6 in COVID-19 in-duced Pneumonia and Macrophage Activation Syndrome-Like Disease. Autoimmun. Rev. 2020, 19, 102537. [Google Scholar] [CrossRef] [PubMed]
- Takaoka, Y.; Niwa, S.; Nagai, H. Interleukin-1B Induces Interleukin-6 Production through the Production of Prostaglan-din E2 in Human Osteoblasts, MG-63 Cells. J. Biochem. 1999, 126, 553–558. [Google Scholar] [CrossRef]
- Dorrington, M.G.; Fraser, I.D.C. NF-κB signaling in macrophages: Dynamics, crosstalk, and signal integration. Front. Immunol. 2019, 10, 705. [Google Scholar] [CrossRef] [PubMed]
- Oeckinghaus, A.; Ghosh, S. The NF-κB Family of Transcription Factors and Its Regulation. Cold Spring Harb. Perspect. Biol. 2009, 1, a000034. [Google Scholar] [CrossRef]
- Hsieh, J.Y.; Huang, T.S.; Cheng, S.M.; Lin, W.S.; Tsai, T.N.; Lee, O.K.; Wang, H.W. miR-146a-5p circuitry uncouples cell proliferation and migration, but not differentiation, in human mesenchy-mal stem cells. Nucleic Acids Res. 2013, 41, 9753–9763. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saba, R.; Sorensen, D.L.; Booth, S.A. MicroRNA-146a: A Dominant, Negative Regulator of the Innate Immune Response. Front. Immunol. 2014, 5, 578. [Google Scholar] [CrossRef]
- Lee, H.M.; Kim, T.S.; Jo, E.K. MiR-146 and miR-125 in the regulation of innate immunity andinflammation. BMB Rep. 2016, 49, 311. [Google Scholar] [CrossRef] [Green Version]
- Boldin, M.P.; Taganov, K.D.; Rao, D.; Yang, L.; Zhao, J.L.; Kalwani, M.; Garcia-Flores, Y.; Luong, M.; Devrekanli, A.; Xu, J.; et al. miR-146a is a significant brake on autoimmunity, myeloproliferation, and cancer in mice. J. Exp. Med. 2011, 208, 1189–1201. [Google Scholar] [CrossRef]
- Taganov, K.D.; Boldin, M.; Chang, K.-J.; Baltimore, D. NF-B-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 2006, 103, 12481–12486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meisgen, F.; Landén, N.X.; Wang, A.; Réthi, B.; Bouez, C.; Zuccolo, M.; Gueniche, A.; Ståhle, M.; Sonkoly, E.; Breton, L.; et al. MiR-146a Negatively Regulates TLR2-Induced Inflammatory Responses in Keratinocytes. J. Investig. Dermatol. 2014, 134, 1931–1940. [Google Scholar] [CrossRef] [Green Version]
- Paik, J.H.; Jang, J.-Y.; Jeon, Y.K.; Kim, W.Y.; Kim, T.M.; Heo, D.S.; Kim, C.-W. MicroRNA-146a Downregulates NFκB Activity via Targeting TRAF6 and Functions as a Tumor Suppressor Having Strong Prognostic Implications in NK/T Cell Lymphoma. Clin. Cancer Res. 2011, 17, 4761–4771. [Google Scholar] [CrossRef] [Green Version]
- Curtale, G.; Rubino, M.; Locati, M. MicroRNAs as Molecular Switches in Macrophage Activation. Front. Immunol. 2019, 10, 799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, G.; Abraham, E. MicroRNAs in immune response and macrophage polarization. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 170–177. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Zhang, M.; Zhong, M.; Suo, Q.; Lv, K. Expression profiles of miRNAs in polarized macrophages. Int. J. Mol. Med. 2013, 31, 797–802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Busch, S.; Auth, E.; Scholl, F.; Huenecke, S.; Koehl, U.; Suess, B.; Steinhilber, D. 5-Lipoxygenase Is a Direct Target of miR-19a-3p and miR-125b-5p. J. Immunol. 2015, 194, 1646–1653. [Google Scholar] [CrossRef] [Green Version]
- Shinde, A.V.; Frangogiannis, N.G. Fibroblasts in myocardial infarction: A role in inflammation and repair. J. Mol. Cell. Cardiol. 2014, 70, 74–82. [Google Scholar] [CrossRef] [Green Version]
- Tung, K.H.; Ernstoff, M.S.; Allen, C.; La Shu, S. A Review of Exosomes and their Role in The Tumor Microenvironment and Host-Tumor “Macroenvironment”. J. Immunol. Sci. 2019, 3, 4–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, K. Signaling Cross Talk between TGF-β/Smad and Other Signaling Pathways. Cold Spring Harb. Perspect. Biol. 2017, 9, a022137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, S.; Bae, E.; Kim, H.-S.; Kim, T.-A.; Byun, K.; Kim, B.; Hong, S.; Im, J.P.; Yun, C.; Lee, B.; et al. TRAF6 Mediates IL-1β/LPS-Induced Suppression of TGF-β Signaling through Its Interaction with the Type III TGF-β Receptor. PLoS ONE 2012, 7, e32705. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamashita, M.; Fatyol, K.; Jin, C.; Wang, X.; Liu, Z.; Zhang, Y.E. TRAF6 Mediates Smad-Independent Activation of JNK and p38 by TGF-β. Mol. Cell 2008, 31, 918–924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gkouveris, I.; Nikitakis, N.G. Role of JNK signaling in oral cancer: A mini review. Tumor Biol. 2017, 39, 1010428317711659. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.; Cai, L.; Liu, J.; Wang, G.; Li, H.; Wang, X.; Xu, W.; Ren, M.; Feng, L.; Liu, P.; et al. MicroRNA-30a-5p Inhibits the Growth of Renal Cell Carcinoma by Modulating GRP78 Expression. Cell. Physiol. Biochem. 2017, 43, 2405–2419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ye, Y.-Y.; Mei, J.-W.; Xiang, S.-S.; Li, H.-F.; Ma, Q.; Song, X.-L.; Wang, Z.; Zhang, Y.-J.; Liu, Y.-C.; Jin, Y.-P.; et al. MicroRNA-30a-5p inhibits gallbladder cancer cell proliferation, migration and metastasis by targeting E2F7. Cell Death Dis. 2018, 9, 410. [Google Scholar] [CrossRef] [PubMed]
- Li, W.F.; Dai, H.; Ou, Q.; Zuo, G.Q.; Liu, C.A. Overexpression of microRNA-30a-5p inhibits liver cancer cell prolif-eration and induces apoptosis by targeting MTDH/PTEN/AKT pathway. Tumor Biol. 2016, 37, 5885–5895. [Google Scholar] [CrossRef]
- Di Stefano, L.; Jensen, M.R.; Helin, K. E2F7, a novel E2F featuring DP-independent repression of a subset of E2F-regulated genes. EMBO J. 2003, 22, 6289–6298. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, W.; Zhou, C.; Luo, M.; Shi, X.; Li, Y.; Sun, Z.; Zhou, F.; Chen, Z.; He, J. MiR-652-3p is upregulated in non-small cell lung cancer and promotes proliferation and metastasis by di-rectly targeting Lgl1. Oncotarget 2016, 7, 16703. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; Dongol, S.; Qiu, C.; Xu, Y.; Sun, C.; Zhang, Z.; Yang, X.; Zhang, Q.; Kong, B. miR-652 Promotes Tumor Proliferation and Metastasis by Targeting RORA in Endometrial Cancer. Mol. Cancer Res. 2018, 16, 1927–1939. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, X.; Luo, H.; Li, X.; Tian, X.; Peng, B.; Liu, S.; Zhan, T.; Wan, Y.; Chen, W.; Li, Y.; et al. miR-331-3p functions as an oncogene by targeting ST7L in pancreatic cancer. Carcinogenesis 2018, 39, 1006–1015. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Chu, F.; Cao, Y.; Shao, J.; Wang, F. Serum miR-182 and miR-331-3p as diagnostic and prognostic markers in pa-tients with hepatocellular carcinoma. Tumor Biol. 2015, 36, 7439–7447. [Google Scholar] [CrossRef] [PubMed]


| Gene | FOR | REV |
|---|---|---|
| IL-1β | CCTGTCCTGCGTGTTGAAAGA | GGGAACTGGGCAGACTCAAA |
| IL-8 | ACTCCAAACCTTTCCACCCC | TTCTCAGCCCTCTTCAAAAACT |
| ACTB1 1 | GCATCCACGAAACTACCTTCAACTC | CTTGATCTTCATTGTGCTGGGTG |
| miRNAs | Function (miRNet) |
|---|---|
| miR-125a-3p | Inflammation; Apoptosis; Tumor-suppressing miRNAs |
| miR-30a-5p | Innate immunity; Immune response |
| miR-146a-5p | Regulation of NF-kb; Innate immunity; Immune response; Cell proliferation; Cell death |
| miR-331-3p | Regulation of AKT pathway; Glucose metabolism; Toxicity |
| miR-139-5p | T cell differentiation |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trentini, M.; Zanotti, F.; Tiengo, E.; Camponogara, F.; Degasperi, M.; Licastro, D.; Lovatti, L.; Zavan, B. An Apple a Day Keeps the Doctor Away: Potential Role of miRNA 146 on Macrophages Treated with Exosomes Derived from Apples. Biomedicines 2022, 10, 415. https://doi.org/10.3390/biomedicines10020415
Trentini M, Zanotti F, Tiengo E, Camponogara F, Degasperi M, Licastro D, Lovatti L, Zavan B. An Apple a Day Keeps the Doctor Away: Potential Role of miRNA 146 on Macrophages Treated with Exosomes Derived from Apples. Biomedicines. 2022; 10(2):415. https://doi.org/10.3390/biomedicines10020415
Chicago/Turabian StyleTrentini, Martina, Federica Zanotti, Elena Tiengo, Francesca Camponogara, Margherita Degasperi, Danilo Licastro, Luca Lovatti, and Barbara Zavan. 2022. "An Apple a Day Keeps the Doctor Away: Potential Role of miRNA 146 on Macrophages Treated with Exosomes Derived from Apples" Biomedicines 10, no. 2: 415. https://doi.org/10.3390/biomedicines10020415
APA StyleTrentini, M., Zanotti, F., Tiengo, E., Camponogara, F., Degasperi, M., Licastro, D., Lovatti, L., & Zavan, B. (2022). An Apple a Day Keeps the Doctor Away: Potential Role of miRNA 146 on Macrophages Treated with Exosomes Derived from Apples. Biomedicines, 10(2), 415. https://doi.org/10.3390/biomedicines10020415







