Gene Expression Profiling in Early Breast Cancer—Patient Stratification Based on Molecular and Tumor Microenvironment Features
Abstract
:1. Introduction
2. Clinical Evidence Supporting the Use of Canonical Gene Expression Signatures
3. Use of Gene Expression Signatures Tackling Additional Clinical Endpoints
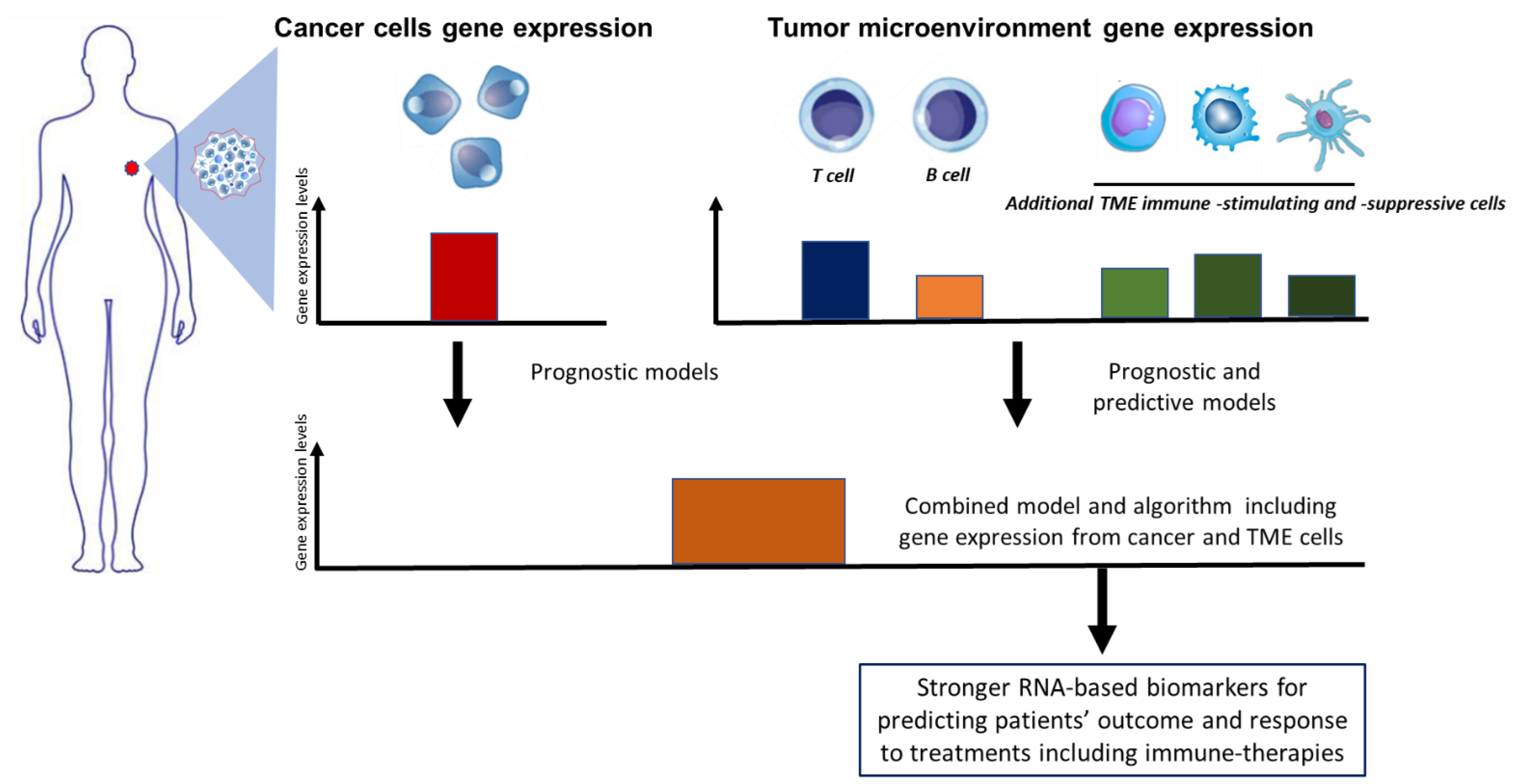
4. Immune-Related Gene Signatures Complementing Established Molecular Assays
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Loibl, S.; Poortmans, P.; Morrow, M.; Denkert, C.; Curigliano, G. Breast cancer. Lancet 2021, 397, 1750–1769. [Google Scholar] [CrossRef]
- Early Breast Cancer Trialists’ Collaborative Group (EBCTCG). Effects of Chemotherapy and Hormonal Therapy for Early Breast Cancer on Recurrence and 15-Year Survival: An Overview of the Randomised Trials. Lancet 2005, 365, 1687–1717. [Google Scholar] [CrossRef]
- Geffen, D.B. Should decisions on adding adjuvant chemotherapy in early-stage ER-positive breast cancer be based on gene expression testing or clinicopathologic factors or both? Ann. Oncol. 2018, 29, 1096–1098. [Google Scholar] [CrossRef] [PubMed]
- Kwa, M.; Makris, A.; Esteva, F.J. Clinical utility of gene-expression signatures in early stage breast cancer. Nat. Rev. Clin. Oncol 2017, 14, 595–610. [Google Scholar] [CrossRef]
- Győrffy, B.; Hatzis, C.; Sanft, T.; Hofstatter, E.; Aktas, B.; Pusztai, L. Multigene Prognostic Tests in Breast Cancer: Past, Present, Future. Breast Cancer Res. 2015, 17, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reis-Filho, J.S.; Pusztai, L. Gene expression profiling in breast cancer: Classification, prognostication, and prediction. Lancet 2011, 378, 1812–1823. [Google Scholar] [CrossRef]
- Russnes, H.G.; Lingjærde, O.C.; Børresen-Dale, A.L.; Caldas, C. Breast Cancer Molecular Stratification: From Intrinsic Subtypes to Integrative Clusters. Am. J. Pathol. 2017, 187, 2152–2162. [Google Scholar] [CrossRef]
- Vieira, A.F.; Schmitt, F. An Update on Breast Cancer Multigene Prognostic Tests—Emergent Clinical Biomarkers. Front. Med. 2018, 5, 248. [Google Scholar] [CrossRef] [Green Version]
- Cognetti, F.; Biganzoli, L.; De Placido, S.; del Mastro, L.; Masetti, R.; Naso, G.; Pruneri, G.; Santini, D.; Tondini, C.A.; Tinterri, C.; et al. Multigene Tests for Breast Cancer: The Physician’s Perspective. Oncotarget 2021, 12, 936–947. [Google Scholar] [CrossRef]
- Richman, J.; Ring, A.; Dowsett, M.; Sestak, I. Clinical Validity of Clinical Treatment Score 5 (CTS5) for Estimating Risk of Late Recurrence in Unselected, Non-Trial Patients with Early Oestrogen Receptor-Positive Breast Cancer. Breast Cancer Res. Treat. 2021, 186, 115–123. [Google Scholar] [CrossRef]
- Lakhanpal, R.; Sestak, I.; Shadbolt, B.; Bennett, G.M.; Brown, M.; Phillips, T.; Zhang, Y.; Bullman, A.; Rezo, A. IHC4 Score plus Clinical Treatment Score Predicts Locoregional Recurrence in Early Breast Cancer. Breast 2016, 29, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Piccart, M.; van’t Veer, L.J.; Poncet, C.; Cardozo, J.M.N.L.; Delaloge, S.; Pierga, J.-Y.; Vuylsteke, P.; Brain, E.; Vrijaldenhoven, S.; Neijenhuis, P.A.; et al. 70-Gene Signature as an Aid for Treatment Decisions in Early Breast Cancer: Updated Results of the Phase 3 Randomised MINDACT Trial with an Exploratory Analysis by Age. Lancet Oncol. 2021, 22, 476–488. [Google Scholar] [CrossRef]
- Sparano, J.A.; Gray, R.J.; Ravdin, P.M.; Makower, D.F.; Pritchard, K.I.; Albain, K.S.; Hayes, D.F.; Geyer, C.E.; Dees, E.C.; Goetz, M.P.; et al. Clinical and Genomic Risk to Guide the Use of Adjuvant Therapy for Breast Cancer. N. Engl. J. Med. 2019, 380, 2395–2405. [Google Scholar] [CrossRef] [PubMed]
- Kalinsky, K.; Barlow, W.E.; Gralow, J.R.; Meric-Bernstam, F.; Albain, K.S.; Hayes, D.; Lin, N.; Perez, E.A.; Goldstein, L.J.; Chia, S.K.L.; et al. 21-Gene Assay to Inform Chemotherapy Benefit in Node-Positive Breast Cancer. N. Engl. J. Med. 2021, 385, 2336–2347. [Google Scholar] [CrossRef] [PubMed]
- OPTIMA: A Prospective Randomised Trial to Validate the Predictive Utility and Cost-Effectiveness of Gene Expression Test-Directed Chemotherapy Decisions—European Journal of Surgical Oncology. Available online: https://www.ejso.com/article/S0748-7983(16)30727-2/fulltext (accessed on 19 November 2021).
- Piccart, M.J.; Kalinsky, K.; Gray, R.; Barlow, W.E.; Poncet, C.; Cardoso, F.; Winer, E.; Sparano, J. Gene Expression Signatures for Tailoring Adjuvant Chemotherapy of Luminal Breast Cancer: Stronger Evidence, Greater Trust. Ann. Oncol. 2021, 32, 1077–1082. [Google Scholar] [CrossRef]
- Lopes Cardozo, J.; Drukker, C.; Schmidt, M.; van’t Veer, L.; Glas, A.; Witteveen, A.; Cardoso, F.; Piccart-Gebhart, M.J.; Poncet, C.; Rutgers, E.J. Outcome of Patients with an Ultralow Risk 70-Gene Signature in the MINDACT Trial. J. Clin. Oncol. 2021, 39, 500. [Google Scholar] [CrossRef]
- Cardoso, F.; van’t Veer, L.J.; Bogaerts, J.; Slaets, L.; Viale, G.; Delaloge, S.; Pierga, J.-Y.; Brain, E.; Causeret, S.; DeLorenzi, M.; et al. 70-Gene Signature as an Aid to Treatment Decisions in Early-Stage Breast Cancer. N. Engl. J. Med. 2016, 375, 717–729. [Google Scholar] [CrossRef] [Green Version]
- Sparano, J.A.; Gray, R.J.; Makower, D.F.; Pritchard, K.I.; Albain, K.S.; Hayes, D.F.; Geyer, C.E.; Dees, E.C.; Goetz, M.P.; Olson, J.A.; et al. Adjuvant Chemotherapy Guided by a 21-Gene Expression Assay in Breast Cancer. N. Engl. J. Med. 2018, 379, 111–121. [Google Scholar] [CrossRef] [Green Version]
- Nitz, U.; Gluz, O.; Christgen, M.; Kates, R.E.; Clemens, M.; Malter, W.; Nuding, B.; Aktas, B.; Kuemmel, S.; Reimer, T.; et al. Reducing Chemotherapy Use in Clinically High-Risk, Genomically Low-Risk PN0 and PN1 Early Breast Cancer Patients: Five-Year Data from the Prospective, Randomised Phase 3 West German Study Group (WSG) PlanB Trial. Breast Cancer Res. Treat. 2017, 165, 573–583. [Google Scholar] [CrossRef] [Green Version]
- Sparano, J.A.; Crager, M.R.; Tang, G.; Gray, R.J.; Stemmer, S.M.; Shak, S. Development and Validation of a Tool Integrating the 21-Gene Recurrence Score and Clinical-Pathological Features to Individualize Prognosis and Prediction of Chemotherapy Benefit in Early Breast Cancer. J. Clin. Oncol. 2021, 39, 557–564. [Google Scholar] [CrossRef]
- Stein, R.C.; Makris, A.; MacPherson, I.R.; Hughes-Davies, L.; Marshall, A.; Dotchin, G.; Cameron, D.A.; Kiely, B.E.; Tsang, J.; Naume, B.; et al. Optima: Optimal Personalised Treatment of Early Breast Cancer Using Multi-Parameter Analysis, an International Randomized Trial of Tumor Gene Expression Test-Directed Chemotherapy Treatment in a Largely Node-Positive Population. J. Clin. Oncol. 2021, 39, TPS599. [Google Scholar] [CrossRef]
- Brain, E.; Girre, V.; Rollot, F.; Bonnetain, F.; Debled, M.; Lacroix, M.; Baffert, S.; Latouche, A.; Falandry, C.; Peyro Saint Paul, H.P.; et al. ASTER 70s: Benefit of Adjuvant Chemotherapy for Estrogen Receptor-Positive HER2-Negative Breast Cancer in Women over 70 According to Genomic Grade—A French GERICO/UCBG UNICANCER Multicenter Phase III Trial. J. Clin. Oncol. 2012, 30, TPS667. [Google Scholar] [CrossRef]
- ClinicalTrials.Gov Beta. EndoPredict® Extended Endocrine—EXET TRIAL. Available online: https://clinicaltrials.gov/ct2/show/NCT04016935 (accessed on 19 November 2021).
- Ettl, J.; Blohmer, J.-U.; Denkert, C.; Keller, M.; Klein, E.; Kronenwett, R.; Neuser, P.; Paepke, S.; Schade-Brittinger, C.; Schnuppe, K.; et al. Abstract OT1-12-03: RESCUE: Reaching for Evidence-BaSed Chemotherapy Use in Endocrine Sensitive Breast Cancer —A Prospective Health Care Study on Risk Assessment by the Clinicomolecular Test EndoPredict® and Long-Term Patient Outcome in Early Luminal Breast Cancer. In Proceedings of the Ongoing Clinical Trials, American Association for Cancer Research, San Antonio, TX, USA, 15 February 2019; p. OT1-12-03. [Google Scholar]
- Cussac, A.L.; Pérez-García, J.; Guerrero, Á.; Bermejo, B.; Gil, M.; Carañana, V.; Morales, S.; de la Haba, J.; Fernández, M.; Alba, E.; et al. Abstract CT219: Neoadjuvant Letrozole and Palbociclib in Stage II-IIIB HR[+]/HER2[-] Breast Cancer with Oncotype DX Recurrence Score® (RS) 18-25 or 26-100. Analysis of RS Changes at Surgery (DxCARTES Trial). Cancer Res. 2019, 79, CT219. [Google Scholar] [CrossRef]
- Jung, J.G.; Kim, H.K.; Kim, Y.; Lee, H.-B.; Moon, H.-G.; Noh, D.-Y.; Han, W. Personalized Neoadjuvant Strategy in Luminal A Breast Cancer to Increase Breast Conserving Surgery (BCS) Rate [PLATO Study]. J. Clin. Oncol. 2020, 38, TPS603. [Google Scholar] [CrossRef]
- Puhalla, S.; Yothers, G.; Sing, A.P.; Julian, T.B.; Wolmark, N.; Jacobs, S.A. Abstract OT2-02-03: NSABP FB-13: An Assessment of the Biological and Clinical Effects of Palbociclib with Ovarian Suppression and Letrozole in the Neoadjuvant Treatment of Pts (Pts) with Premenopausal (PreM) Estrogen-Receptor Positive/HER2-Negative Primary Breast Cancer. In Proceedings of the Ongoing Clinical Trials; American Association for Cancer Research: San Antonio, TX, USA, 15 February 2020; p. OT2-02-03. [Google Scholar]
- Smith, I.; Robertson, J.; Kilburn, L.; Wilcox, M.; Evans, A.; Holcombe, C.; Horgan, K.; Kirwan, C.; Mallon, E.; Sibbering, M.; et al. Long-term outcome and prognostic value of Ki67 after perioperative endocrine therapy in postmenopausal women with hormone-sensitive early breast cancer (POETIC): An open-label, multicentre, parallel-group, randomised, phase 3 trial. Lancet Oncol. 2020, 21, 1443–1454. [Google Scholar] [CrossRef]
- ClinicalTrials.Gov Beta. POETIC-A TRIAL. Available online: https://clinicaltrials.gov/ct2/show/NCT04584853 (accessed on 19 November 2021).
- Jenkins, S.; Wesolowski, R.; Gatti-Mays, M.E. Improving Breast Cancer Responses to Immunotherapy—A Search for the Achilles Heel of the Tumor Microenvironment. Curr. Oncol. Rep. 2021, 23, 55. [Google Scholar] [CrossRef]
- Finak, G.; Bertos, N.; Pepin, F.; Sadekova, S.; Souleimanova, M.; Zhao, H.; Chen, H.; Omeroglu, G.; Meterissian, S.; Omeroglu, A.; et al. Stromal Gene Expression Predicts Clinical Outcome in Breast Cancer. Nat. Med. 2008, 14, 518–527. [Google Scholar] [CrossRef]
- Farmer, P.; Bonnefoi, H.; Anderle, P.; Cameron, D.; Wirapati, P.; Wirapati, P.; Becette, V.; André, S.; Piccart, M.; Campone, M.; et al. A Stroma-Related Gene Signature Predicts Resistance to Neoadjuvant Chemotherapy in Breast Cancer. Nat. Med. 2009, 15, 68–74. [Google Scholar] [CrossRef]
- Zheng, S.; Zou, Y.; Xie, X.; Liang, J.-Y.; Yang, A.; Yu, K.; Wang, J.; Tang, H.; Xie, X. Development and Validation of a Stromal Immune Phenotype Classifier for Predicting Immune Activity and Prognosis in Triple-Negative Breast Cancer. Int. J. Cancer 2020, 147, 542–553. [Google Scholar] [CrossRef]
- Winslow, S.; Lindquist, K.E.; Edsjö, A.; Larsson, C. The Expression Pattern of Matrix-Producing Tumor Stroma Is of Prognostic Importance in Breast Cancer. BMC Cancer 2016, 16, 841. [Google Scholar] [CrossRef] [Green Version]
- Desmedt, C.; Haibe-Kains, B.; Wirapati, P.; Buyse, M.; Larsimont, D.; Bontempi, G.; Delorenzi, M.; Piccart, M.; Sotiriou, C. Biological Processes Associated with Breast Cancer Clinical Outcome Depend on the Molecular Subtypes. Clin. Cancer Res. 2008, 14, 5158–5165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teschendorff, A.E.; Miremadi, A.; Pinder, S.E.; Ellis, I.O.; Caldas, C. An Immune Response Gene Expression Module Identifies a Good Prognosis Subtype in Estrogen Receptor Negative Breast Cancer. Genome Biol. 2007, 8, R157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teschendorff, A.E.; Caldas, C. A Robust Classifier of High Predictive Value to Identify Good Prognosis Patients in ER-Negative Breast Cancer. Breast Cancer Res. 2008, 10, R73. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rody, A.; Holtrich, U.; Pusztai, L.; Liedtke, C.; Gaetje, R.; Ruckhaeberle, E.; Solbach, C.; Hanker, L.; Ahr, A.; Metzler, D.; et al. T-Cell Metagene Predicts a Favorable Prognosis in Estrogen Receptor-Negative and HER2-Positive Breast Cancers. Breast Cancer Res. 2009, 11, R15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pilipow, K.; Darwich, A.; Losurdo, A. T-Cell-Based Breast Cancer Immunotherapy. Semin. Cancer Biol. 2021, 72, 90–101. [Google Scholar] [CrossRef]
- Yau, C.; Esserman, L.; Moore, D.H.; Waldman, F.; Sninsky, J.; Benz, C.C. A Multigene Predictor of Metastatic Outcome in Early Stage Hormone Receptor-Negative and Triple-Negative Breast Cancer. Breast Cancer Res. 2010, 12, R85. [Google Scholar] [CrossRef] [Green Version]
- Yau, C.; Sninsky, J.; Kwok, S.; Wang, A.; Degnim, A.; Ingle, J.N.; Gillett, C.; Tutt, A.; Waldman, F.; Moore, D.; et al. An Optimized Five-Gene Multi-Platform Predictor of Hormone Receptor Negative and Triple Negative Breast Cancer Metastatic Risk. Breast Cancer Res. 2013, 15, R103. [Google Scholar] [CrossRef] [Green Version]
- Bareche, Y.; Buisseret, L.; Gruosso, T.; Girard, E.; Venet, D.; Dupont, F.; Desmedt, C.; Larsimont, D.; Park, M.; Rothé, F.; et al. Unraveling Triple-Negative Breast Cancer Tumor Microenvironment Heterogeneity: Towards an Optimized Treatment Approach. JNCI J. Natl. Cancer Inst. 2020, 112, 708–719. [Google Scholar] [CrossRef] [Green Version]
- Rody, A.; Karn, T.; Liedtke, C.; Pusztai, L.; Ruckhaeberle, E.; Hanker, L.; Gaetje, R.; Solbach, C.; Ahr, A.; Metzler, D.; et al. A Clinically Relevant Gene Signature in Triple Negative and Basal-like Breast Cancer. Breast Cancer Res. 2011, 13, R97. [Google Scholar] [CrossRef] [Green Version]
- Han, J.; Choi, Y.-L.; Kim, H.; Choi, J.Y.; Lee, S.K.; Lee, J.E.; Choi, J.-S.; Park, S.; Choi, J.-S.; Kim, Y.D.; et al. MMP11 and CD2 as Novel Prognostic Factors in Hormone Receptor-Negative, HER2-Positive Breast Cancer. Breast Cancer Res. Treat. 2017, 164, 41–56. [Google Scholar] [CrossRef] [Green Version]
- Schmidt, M.; Böhm, D.; von Törne, C.; Steiner, E.; Puhl, A.; Pilch, H.; Lehr, H.-A.; Hengstler, J.G.; Kölbl, H.; Gehrmann, M. The Humoral Immune System Has a Key Prognostic Impact in Node-Negative Breast Cancer. Cancer Res. 2008, 68, 5405–5413. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bianchini, G.; Qi, Y.; Alvarez, R.H.; Iwamoto, T.; Coutant, C.; Ibrahim, N.K.; Valero, V.; Cristofanilli, M.; Green, M.C.; Radvanyi, L.; et al. Molecular Anatomy of Breast Cancer Stroma and Its Prognostic Value in Estrogen Receptor–Positive and –Negative Cancers. J. Clin. Oncol. 2010, 28, 4316–4323. [Google Scholar] [CrossRef] [PubMed]
- Oh, E.; Choi, Y.-L.; Park, T.; Lee, S.; Nam, S.J.; Shin, Y.K. A Prognostic Model for Lymph Node-Negative Breast Cancer Patients Based on the Integration of Proliferation and Immunity. Breast Cancer Res. Treat. 2012, 132, 499–509. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Chou, J.; Tao, Y.; Wu, D.; Wu, X.; Li, X.; Li, Y.; Chu, Y.; Tang, F.; Shi, Y.; et al. An Assessment of Prognostic Immunity Markers in Breast Cancer. NPJ Breast Cancer 2018, 4, 1–9. [Google Scholar] [CrossRef]
- Lee, H.J.; Lee, J.-J.; Song, I.H.; Park, I.A.; Kang, J.; Yu, J.H.; Ahn, J.-H.; Gong, G. Prognostic and Predictive Value of NanoString-Based Immune-Related Gene Signatures in a Neoadjuvant Setting of Triple-Negative Breast Cancer: Relationship to Tumor-Infiltrating Lymphocytes. Breast Cancer Res. Treat. 2015, 151, 619–627. [Google Scholar] [CrossRef]
- Sinn, B.V.; Loibl, S.; Hanusch, C.A.; Zahm, D.M.; Sinn, H.P.; Untch, M.; Weber, K.; Karn, T.; Becker, C.; Marmé, F.; et al. Immune-related Gene Expression Predicts Response to Neoadjuvant Chemotherapy but not Additional Benefit from PD-L1 Inhibition in Women with Early Triple-negative Breast Cancer. Clin. Cancer Res. 2021, 27, 2584–2591. [Google Scholar] [CrossRef]
- Alcaraz-Sanabria, A.; Baliu-Piqué, M.; Saiz-Ladera, C.; Rojas, K.; Manzano, A.; Marquina, G.; Casado, A.; Cimas, F.J.; Pérez-Segura, P.; Pandiella, A.; et al. Genomic Signatures of Immune Activation Predict Outcome in Advanced Stages of Ovarian Cancer and Basal-Like Breast Tumors. Front. Oncol. 2020, 9, 1486. [Google Scholar] [CrossRef] [Green Version]
- Pérez-Pena, J.; Tibor Fekete, J.; Páez, R.; Baliu-Piqué, M.; García-Saenz, J.Á.; García-Barberán, V.; Manzano, A.; Pérez-Segura, P.; Esparis-Ogando, A.; Pandiella, A.; et al. A Transcriptomic Immunologic Signature Predicts Favorable Outcome in Neoadjuvant Chemotherapy Treated Triple Negative Breast Tumors. Front. Immunol. 2019, 10, 2802. [Google Scholar] [CrossRef]
- Cui, Y.; Li, B.; Pollom, E.L.; Horst, K.C.; Li, R. Integrating Radiosensitivity and Immune Gene Signatures for Predicting Benefit of Radiotherapy in Breast Cancer. Clin. Cancer Res. 2018, 24, 4754–4762. [Google Scholar] [CrossRef] [Green Version]
- Győrffy, B.; Pongor, L.; Bottai, G.; Li, X.; Budczies, J.; Szabó, A.; Hatzis, C.; Pusztai, L.; Santarpia, L. An Integrative Bioinformatics Approach Reveals Coding and Non-Coding Gene Variants Associated with Gene Expression Profiles and Outcome in Breast Cancer Molecular Subtypes. Br. J. Cancer 2018, 118, 1107–1114. [Google Scholar] [CrossRef] [Green Version]
| Clinical Trials (NCT#) and Study Phase | GEP Assay | Breast Cancer Population | Clinical Endpoints | Study Description |
|---|---|---|---|---|
| OPTIMA (ISRCTN 42400492) Phase III | Prosigna/PAM50 | HR+/HER2−, pN1–2 or pN1mi with pT ≥ 20 mm or pN0 with pT ≥ 30 mm | iDFS | A de-escalation clinical trial to tailor adjuvant therapy for HR+/HER2− high clinical risk breast cancer |
| RxPONDER (NCT01272037) Phase III | Oncotype DX | HR+/HER2−, 1–3 N+, recurrence score ≤ 25 | iDFS | Role of recurrent score in predicting benefit of adjuvant chemotherapy in women with LN+ disease |
| GERICO 11/ASTER 70s (NCT01564056) Phase III | Genomic Grade test | HR+/HER2−, N0 or N+, age ≥ 70 years, PS ≤ 2 | OS | Adjuvant systemic treatment for ER+/HER2− breast cancers in patients over 70 years of age according to genomic grade index: chemotherapy + endocrine therapy vs. endocrine therapy |
| EXET (NCT04016935) Observational | EndoPredict | HR+/HER2−, stage I–III, 0–3 N+, potential adjuvant chemotherapy, no neoadjuvant chemotherapy | DRFS | Extended Endocrine Trial: A prospective registry study testing the impact of EndoPredict on extended endocrine therapy decisions and patients’ outcomes |
| RESCUE (NCT03503799) Observational | EndoPredict | HR+/HER2−, stage I–III, 0–3 N+ | DRFS | Prospective assessment of disease progression in breast cancer patients undergoing EndoPredict test—a care research study |
| DxCARTES (NCT03819010) Phase II | Oncotype DX | HR+/HER2−, Ki67 ≥ 20, stage II-IIIb T2–T4, N0–N2, age ≥ 18, PS ≤ 1 | Molecular differences on recurrence score between pre- and post-treatment | Neoadjuvant letrozole + palbociclib in HR+/HER2− patients with stage II–IIIb breast cancer, and pretreatment recurrence scores: 18–25 or 26–100 by Oncotype DX testing. Analysis of recurrent score and pathological feature changes at time of surgery |
| PLATO (NCT03900637) Phase II | MammaPrint | HR+/HER2−, stage I–IIIA, ineligible for BCS, age ≥ 19, PS ≤ 2 | Conversion rate | Multicenter study to increase BCS rate with personalized neoadjuvant strategy in HR+/HER2− breast cancer for whom BCS is not feasible. Neoadjuvant chemotherapy and endocrine therapy are conducted on genomic high and low risk patients, respectively |
| NSABP FB-13 (NCT03628066) Phase II | Oncotype DX | HR+/HER2−, T2–T4, candidate for neoadjuvant HT, premenopausal status, PS ≤ 1, recurrence score < 26 | Complete cell cycle arrest (percentage of patients with a Ki67 < 2.7%) | Neoadjuvant treatment for premenopausal HR+/HER2− breast cancer patients and evaluation of the clinical and biological effects of palbociclib with ovarian suppression and letrozole |
| POETIC-A (NCT04584853) Phase III | AIR-CIS | HR+/HER2−, postmenopausal status, ≥1.5 cm, grade 3 and/or Ki67 level ≥ 20% | Time to tumor recurrence (local or distant disease) | Preoperative endocrine therapy for individualized therapy with abemaciclib |
| Gene Arrays | RNA-seq | |
|---|---|---|
| Number of genes | Whole transcriptome | Whole transcriptome |
| Section of genes | Targeted gene sequences | Entire genes |
| Bioinformatics complexity | Established tools | Multiple pipelines available |
| Alternative splicing | Can be included | Feasible but large read counts needed |
| Strength | Low cost, established pipelines, quality control straightforward | Higher specificity Higher sensitivity for genes expressed either at low or very high level Higher levels of reproducibility |
| Weakness | Gene sequence must be set upfront cross-hybridization bias | Complex processing, potential GC content bias (mapping ambiguity for paralogous sequences), longer handling time, currently higher cost |
| Opportunity | High throughput, can be transformed to PCR | Can be used to identify novel transcribed regions, gene variations (e.g., mutations), assess allele-specific expression |
| Commercially available assays for breast cancer | MammaPrint, Oncotype Dx, Prosigna, etc. | None available yet |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Munkácsy, G.; Santarpia, L.; Győrffy, B. Gene Expression Profiling in Early Breast Cancer—Patient Stratification Based on Molecular and Tumor Microenvironment Features. Biomedicines 2022, 10, 248. https://doi.org/10.3390/biomedicines10020248
Munkácsy G, Santarpia L, Győrffy B. Gene Expression Profiling in Early Breast Cancer—Patient Stratification Based on Molecular and Tumor Microenvironment Features. Biomedicines. 2022; 10(2):248. https://doi.org/10.3390/biomedicines10020248
Chicago/Turabian StyleMunkácsy, Gyöngyi, Libero Santarpia, and Balázs Győrffy. 2022. "Gene Expression Profiling in Early Breast Cancer—Patient Stratification Based on Molecular and Tumor Microenvironment Features" Biomedicines 10, no. 2: 248. https://doi.org/10.3390/biomedicines10020248
APA StyleMunkácsy, G., Santarpia, L., & Győrffy, B. (2022). Gene Expression Profiling in Early Breast Cancer—Patient Stratification Based on Molecular and Tumor Microenvironment Features. Biomedicines, 10(2), 248. https://doi.org/10.3390/biomedicines10020248







