Involvement of Eukaryotic Small RNA Pathways in Host Defense and Viral Pathogenesis
Abstract
:1. Small RNA Biogenesis
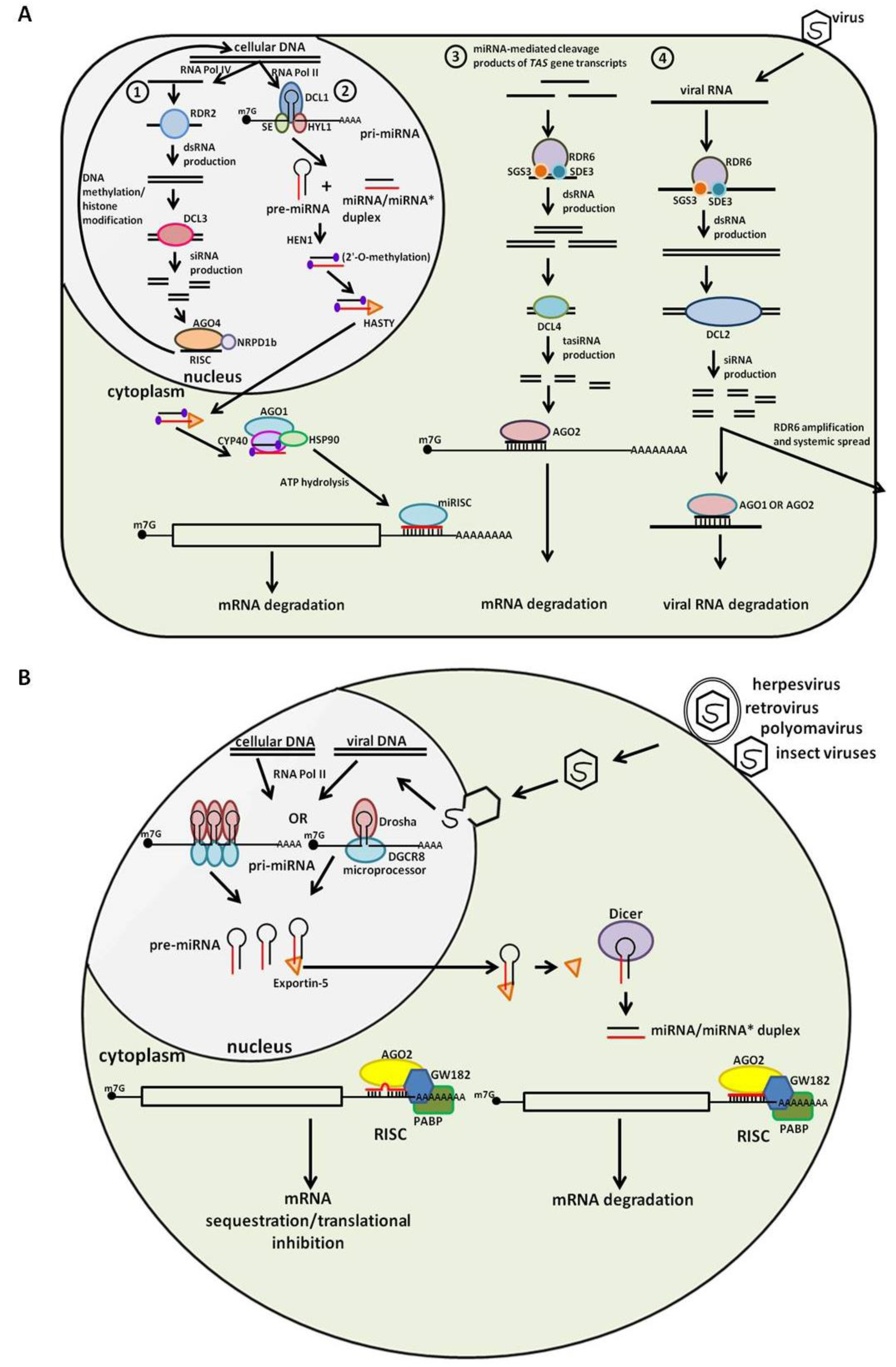
2. Plant Small RNA: An Overview
3. Animal Small RNA: An Overview
4. Plant Immunity and Small RNA
5. Animal Immunity and Small RNA
6. Pathogenic Manipulation of Small RNA Pathways
| Virus | microRNA | Effect | Ref. |
|---|---|---|---|
| HSV1 | miR-H2-3p | Targets viral ICP0; regulates latency | [29] |
| KSHV | miR-K12-4-5p | Targets RBL2; modulates epigenetic regulation | [30] |
| KSHV | miR-K12-7 | Targets viral RTA; regulates latency | [30,31,32, 33] |
| KSHV | miR-K9* | Targets viral RTA; regulates latency | [33] |
| HCMV | miR-UL112-1 | Targets host restriction factor BclAf1; enhances viral replication | [34,35] |
| KSHV | miR-K12-11 | Hosts miR-155 ortholog; regulates cell proliferation | [35] |
| MDV1 | miR-M4 | Hosts miR-155 ortholog; regulates cell proliferation | [36] |
| EBV | miR-BART1-3p, miR-BART5-5p, miR-BART22-3p | orthologs to host miR-29a/b/c, miR-18a/b andmiR-520d/miR-524-5p, respectively; regulate apoptosis and the cell cycle | [37] |
| BLV | blv-miR-B4 | Hosts miR-29 ortholog; contributes to viral-induced lymphoma | [38] |
| KSHV, EBV, HCMV | miR-K12-7, miR-BART2-5p, miR-UL112 | Target host MICB; virus immune evasion | [39] |
| Virus | Effector Molecule | Targets of interest | Effect on virus | Reference |
|---|---|---|---|---|
| EBV | miR-155 (induced) | Associated with viral transformation | [41,42,43,44] | |
| EBV | miR-34a (induced by EBV LMP1) | Enhanced viral growth | [50,51] | |
| EBV | miR-200b, miR-429 | ZEB1, ZEB2 | Regulation of viral latency | [52,53] |
| HCMV | miR-92a (downregulated) | GATA-2 | Increased IL-10, involved in viral latency | [54] |
| HSV1 | miR-101 (downregulated) | ATP5B | Enhances virus replication | [55] |
| MDV (MSB1 cell line) | miR-221, miR-222 | p27Kip1 | Associated with viral transformation | [57] |
| HIV | miR-28, miR-125b, miR-150, miR-223, miR-382 | multiple HIV genes | Maintenance of latency in resting T-cells | [62] |
| HCV | miR-122 | Interacts with the HCV genome | Enhances viral replication | [68,69,70] |
| Dengue | NS4B | Downregulates Dicer, Drosha, AGO1 and AGO2 | Enhances viral replication | [67] |
| TSWV | Viral NS proteins | Binds to dsRNA and blocks Dicer-mediated cleavage | Disrupts antiviral response | [72] |
| CMV | Viral protein 2b | Binds to and inhibits the function of AGO1; also directly binds to small RNAs to block RISC function | Disrupts antiviral response | [73] |
| TEV | Viral helper component protease | Interacts with doubled-stranded-siRNA and prevents strand separation | Disrupts antiviral response | [74] |
| TBSV | Viral protein p19 | Blocks RISC loading by binding to small dsRNA duplexes | Disrupts antiviral response | [75] |
| RGDV | pns11 | Upregulation of miR-160, miR-162, miR-167, miR-168 | Increased viral pathogenesis | [76] |
| PSTVd | miR-396, miR-319, miR-159, miR403 (downregulated) | Transcription factors associated with plant morphology and development | Increased viroid pathogenesis | [77] |
8. Small RNAs in Prokaryotic Immunity
9. Conclusions
Acknowledgements
Conflicts of Interest
References
- Okamura, K. Diversity of animal small RNA pathways and their biological utility. Wiley Interdiscip. Rev. RNA 2012, 3, 351–368. [Google Scholar] [CrossRef]
- Janga, S.C.; Vallabhaneni, S. MicroRNAs as post-transcriptional machines and their interplay with cellular networks. Adv. Exp. Med. Biol. 2011, 722, 59–74. [Google Scholar]
- Hohn, T.; Vazquez, F. RNA silencing pathways of plants: Silencing and its suppression by plant DNA viruses. Biochim. Biophys. Acta 2011, 1809, 588–600. [Google Scholar]
- Jaskiewicz, L.; Filipowicz, W. Role of Dicer in posttranscriptional RNA silencing. Curr. Top. Microbiol. Immunol. 2008, 320, 77–97. [Google Scholar] [CrossRef]
- Martínez de Alba, A.E.; Jauvion, V.; Mallory, A.C.; Bouteiller, N.; Vaucheret, H. The miRNA pathway limits AGO1 availability during siRNA-mediated PTGS defense against exogenous RNA. Nucleic Acids Res. 2011, 39, 9339–9344. [Google Scholar] [CrossRef]
- Earley, K.W.; Pontvianne, F.; Wierzbick, A.T.; Blevins, T.; Tucker, S.; Costa-Nunes, P.; Pontes, O.; Pikaard, C.S. Mechanisms of HDA6-mediated rRNA gene silencing: Suppression of intergenic Pol II transcription and differential effects on maintenance versus siRNA-directed cytosine methylation. Genes Dev. 2010, 24, 1119–1132. [Google Scholar] [CrossRef]
- Czech, B.; Hannon, G.J. Small RNA sorting: Matchmaking for Argonautes. Nat. Rev. Genet. 2011, 12, 19–31. [Google Scholar] [CrossRef]
- Llave, C. Virus-derived small interfering RNAs at the core of plant-virus interactions. Trends Plant Sci. 2010, 15, 701–707. [Google Scholar] [CrossRef]
- Coll, N.S.; Epple, P.; Dangl, J.L. Programmed cell death in the plant immune system. Cell Death Differ. 2011, 18, 1247–1256. [Google Scholar] [CrossRef]
- Bittel, P.; Robatzek, S. Microbe-associated molecular patterns (MAMPs) probe plant immunity. Curr. Opin. Plant Biol. 2007, 10, 335–341. [Google Scholar] [CrossRef]
- Xiao, S.; Wang, W.; Yang, X. Evolution of Resistance Genes in Plants. In Innate immunity of plants, animals and humans; Heine, H., Ed.; Springer-Verlag: Berlin, Germany, 2008; pp. 1–5. [Google Scholar]
- Jin, H. Endogenous small RNAs and antibacterial immunity in plants. FEBS Lett. 2008, 582, 2679–2684. [Google Scholar] [CrossRef]
- Fahlgren, N.; Howell, M.D.; Kasschau, K.D.; Chapman, E.J.; Sullivan, C.M.; Cumbie, J.S.; Givan, S.A.; Law, T.F.; Grant, S.R.; Dangl, J.L.; et al. High-throughput sequencing of Arabidopsis microRNAs: evidence for frequent birth and death of MIRNA genes. PLoS One 2007, 2, e219. [Google Scholar] [CrossRef]
- Shivaprasad, P.V.; Chen, H.M.; Patel, K.; Bond, D.M.; Santos, B.A.; Baulcombe, D.C. A microRNA superfamily regulates nucleotide binding site-leucine-rich repeats and other mRNAs. Plant Cell 2012, 24, 859–874. [Google Scholar] [CrossRef]
- Sattar, S.; Song, Y.; Anstead, J.A.; Sunkar, R.; Thompson, G.A. Cucumis melo microRNA expression profile during aphid herbivory in a resistant and susceptible interaction. Mol. Plant Microbe Interact. 2012, 25, 839–848. [Google Scholar] [CrossRef]
- Katiyar-Agarwal, S.; Gao, S.; Vivian-Smith, A.; Jin, H. A novel class of bacteria-induced small RNAs in Arabidopsis. Genes Dev. 2007, 21, 3123–3134. [Google Scholar] [CrossRef]
- Calarco, J.P.; Martienssen, R.A. Genome reprogramming and small interfering RNA in the Arabidopsis germline. Curr. Opin. Genet. Dev. 2011, 21, 134–139. [Google Scholar] [CrossRef]
- Kuchen, S.; Resch, W.; Yamane, A.; Kuo, N.; Li, Z.; Chakraborty, T.; Wei, L.; Laurence, A.; Yasuda, T.; Peng, S.; et al. Regulation of microRNA expression and abundance during lymphopoiesis. Immunity 2010, 32, 828–839. [Google Scholar] [CrossRef]
- Luers, A.J.; Loudig, O.D.; Berman, J.W. MicroRNAs are expressed and processed by human primary macrophages. Cell Immunol. 2010, 263, 1–8. [Google Scholar] [CrossRef]
- Forres, A.R.; Kanamori-Katayama, M.; Tomaru, Y.; Lassmann, T.; Ninomiya, N.; Takahashi, Y.; de Hoon, M.J.; Kubosaki, A.; Kaiho, A.; Suzuki, M.; et al. Induction of microRNAs, mir-155, mir-222, mir-424 and mir-503, promotes monocytic differentiation through combinatorial regulation. Leukemia 2010, 24, 460–466. [Google Scholar] [CrossRef]
- Dudda, J.C.; Salaun, B.; Ji, Y.; Palmer, D.C.; Monnot, G.C.; Merck, E.; Boudousquie, C.; Utzschneider, D.T.; Escobar, T.M.; Perret, R.; et al. MicroRNA-155 is required for effector CD8+ T cell responses to virus infection and cancer. Immunity 2013, 38, 742–753. [Google Scholar] [CrossRef]
- O’Connell, R.M.; Taganov, K.D.; Boldin, M.P.; Cheng, G.; Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 2007, 104, 1604–1609. [Google Scholar]
- Wang, P.; Hou, J.; Lin, L.; Wang, C.; Liu, X.; Li, D.; Ma, F.; Wang, Z.; Cao, X. Inducible microRNA-155 feedback promotes type I IFN signaling in antiviral innate immunity by targeting suppressor of cytokine signaling 1. J. Immunol. 2010, 185, 6226–6233. [Google Scholar] [CrossRef]
- Quinn, S.R.; O’Neill, L.A. A trio of microRNAs that control Toll-like receptor signalling. Int. Immunol. 2011, 23, 421–425. [Google Scholar] [CrossRef]
- O’Neill, L.A.; Sheedy, F.J.; McCoy, C.E. MicroRNAs: The fine-tuners of Toll-like receptor signalling. Nat. Rev. Immunol. 2011, 11, 163–175. [Google Scholar]
- Li, Y.; Li, J.; Belisle, S.; Baskin, C.R.; Tumpey, T.M.; Katze, M.G. Differential microRNA expression and virulence of avian, 1918 reassortant, and reconstructed 1918 influenza A viruses. Virology 2011, 421, 105–113. [Google Scholar] [CrossRef]
- Vaz, C.; Mer, A.S.; Bhattacharya, A.; Ramaswamy, R. MicroRNAs modulate the dynamics of the NF-κB signaling pathway. PLoS One 2011, 6, e27774. [Google Scholar]
- miRBase. Available online: http://www.mirbase.org/ (accessed on 26 October 2013).
- Umbach, J.L.; Kramer, M.F.; Jurak, I.; Karnowski, H.W.; Coen, D.M.; Cullen, B.R. MicroRNAs expressed by herpes simplex virus 1 during latent infection regulate viral mRNAs. Nature. 2008, 454, 780–783. [Google Scholar]
- Malterer, G.; Dölken, L.; Haas, J. The miRNA-targetome of KSHV and EBV in human B-cells. RNA Biol. 2011, 8, 30–34. [Google Scholar]
- Lin, X.; Liang, D.; He, Z.; Deng, Q.; Robertson, E.S.; Lan, K. miR-K12–7-5p encoded by Kaposi’s sarcoma-associated herpesvirus stabilizes the latent state by targeting viral ORF50/RTA. PLoS One 2011, 6, e16224. [Google Scholar]
- Lu, F.; Stedman, W.; Yousef, M.; Renne, R.; Lieberman, P.M. Epigenetic regulation of Kaposi’s sarcoma-associated herpesvirus latency by virus-encoded microRNAs that target Rta and the cellular Rbl2-DNMT pathway. J. Virol. 2010, 84, 2697–2706. [Google Scholar]
- Bellare, P.; Ganem, D. Regulation of KSHV lytic switch protein expression by a virus-encoded microRNA: An evolutionary adaptation that fine-tunes lytic reactivation. Cell Host Microbe 2009, 6, 570–575. [Google Scholar] [CrossRef]
- Lee, S.H.; Kalejta, R.F.; Kerry, J.; Semmes, O.J.; O’Connor, C.M.; Khan, Z.; Garcia, B.A.; Shenk, T.; Murphy, E. BclAF1 restriction factor is neutralized by proteasomal degradation and microRNA repression during human cytomegalovirus infection. Proc. Natl. Acad. Sci. USA 2012, 109, 9575–9580. [Google Scholar] [CrossRef]
- Saffert, R.T.; Kalejta, R.F. Inactivating a cellular intrinsic immune defense mediated by DAXX is the mechanism through which the human cytomegalovirus pp71 protein stimulates viral immediate-early gene expression. J. Virol. 2006, 80, 3863–3871. [Google Scholar] [CrossRef]
- Morgan, R.; Anderson, A.; Bernberg, E.; Kamboj, S.; Huang, E.; Lagasse, G.; Isaacs, G.; Parcells, M.; Meyers, B.C; Green, P.J.; et al. Sequence conservation and differential expression of Marek’s disease virus microRNAs. J. Virol. 2008, 82, 12213–12220. [Google Scholar] [CrossRef]
- Kincaid, R.P.; Burke, J.M.; Sullivan, C.S. RNA virus microRNA that mimics a B-cell oncomiR. Proc. Natl. Acad. Sci. USA 2012, 109, 3077–3082. [Google Scholar] [CrossRef]
- Nachmani, D.; Stern-Ginossar, N.; Sarid, R.; Mandelboim, O. Diverse herpesvirus microRNAs target the stress-induced immune ligand MICB to escape recognition by natural killer cells. Cell Host Microbe 2009, 5, 376–385. [Google Scholar] [CrossRef]
- Forte, E.; Salinas, R.E.; Chang, C.; Zhou, T.; Linnstaedt, S.D.; Gottwein, E.; Jacobs, C.; Jima, D.; Li, Q.J.; Dave, S.S.; et al. The Epstein-Barr virus (EBV)-induced tumor suppressor microRNA MiR-34a is growth promoting in EBV-infected B cells. J. Virol. 2012, 86, 6889–6898. [Google Scholar] [CrossRef]
- Qin, Z.; Jakymiw, A.; Findlay, V.; Parsons, C. KSHV-Encoded MicroRNAs: Lessons for viral cancer pathogenesis and emerging concepts. Int. J. Cell Biol. 2012, 2012. [Google Scholar] [CrossRef]
- Lu, F.; Weidmer, A.; Liu, C.G.; Volinia, S.; Croce, C.M.; Lieberman, P.M. Epstein-Barr virus-induced miR-155 attenuates NF-kappaB signaling and stabilizes latent virus persistence. J. Virol. 2008, 82, 10436–10443. [Google Scholar] [CrossRef]
- Gatto, G.; Rossi, A.; Rossi, D.; Kroening, S.; Bonatti, S.; Mallardo, M. Epstein-Barr virus latent membrane protein 1 trans-activates miR-155 transcription through the NF-κB pathway. Nucleic Acids Res. 2008, 36, 6608–6619. [Google Scholar] [CrossRef]
- Linnstaedt, S.D.; Gottwein, E.; Skalsky, R.L.; Luftig, M.A.; Cullen, B.R. Virally induced cellular microRNA miR-155 plays a key role in B-cell immortalization by Epstein-Barr virus. J. Virol. 2010, 84, 11670–11678. [Google Scholar] [CrossRef]
- Skalsky, R.L.; Samols, M.A.; Plaisance, K.B.; Boss, I.W.; Riva, A.; Lopez, M.C.; Baker, H.V.; Renne, R. Kaposi’s sarcoma-associated herpesvirus encodes an ortholog of miR-155. J. Virol. 2007, 81, 12836–12845. [Google Scholar] [CrossRef]
- Dahlke, C.; Maul, K.; Christalla, T.; Walz, N.; Schult, P.; Stocking, C.; Grundhoff, A. A microRNA encoded by Kaposi sarcoma-associated herpesvirus promotes B-cell expansion in vivo. PLoS One. 2012, 7, e49435. [Google Scholar]
- Sin, S.H.; Kim, Y.B.; Dittmer, D.P. Latency locus complements MicroRNA 155 deficiency in vivo. J. Virol. 2013, 87, 11908–11911. [Google Scholar] [CrossRef]
- Zhao, Y.; Xu, H.; Yao, Y.; Smith, L.P.; Kgosana, L.; Green, J.; Petherbridge, L.; Baigent, S.J.; Nair, V. Critical role of the virus-encoded microRNA-155 ortholog in the induction of Marek’s disease lymphomas. PLoS Pathog. 2011, 7, e1001305. [Google Scholar] [CrossRef]
- Burnside, J.; Morgan, R. Emerging roles of chicken and viral microRNAs in avian disease. BMC Proc. 2011, 5. [Google Scholar] [CrossRef]
- Riley, K.J.; Rabinowitz, G.S.; Yario, T.A.; Luna, J.M.; Darnell, R.B.; Steitz, J.A. EBV and human microRNAs co-target oncogenic and apoptotic viral and human genes during latency. EMBO J. 2012, 31, 2207–2221. [Google Scholar] [CrossRef]
- Chang, T.C.; Wentzel, E.A.; Kent, O.A.; Ramachandran, K.; Mullendore, M.; Lee, K.H.; Feldmann, G.; Yamakuchi, M.; Ferlito, M.; Lowenstein, C.J.; et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol. Cell 2007, 26, 745–752. [Google Scholar] [CrossRef]
- Ellis-Connell, A.L.; Iempridee, T.; Xu, I.; Mertz, J.E. Cellular microRNAs 200b and 429 regulate the Epstein-Barr virus switch between latency and lytic replication. J. Virol. 2010, 84, 10329–10343. [Google Scholar] [CrossRef]
- Ellis, A.L.; Wang, Z.; Yu, X.; Mertz, J.E. Either ZEB1 or ZEB2/SIP1 can play a central role in regulating the Epstein-Barr virus latent-lytic switch in a cell type-specific manner. J. Virol. 2010, 84, 6139–6152. [Google Scholar] [CrossRef]
- Poole, E.; McGregor-Dallas, S.R.; Colston, J.; Joseph, R.S.; Sinclair, J. Virally induced changes in cellular microRNAs maintain latency of human cytomegalovirus in CD34+ progenitors. J. Gen. Virol. 2011, 92, 1539–1549. [Google Scholar] [CrossRef]
- Zheng, S.Q.; Li, Y.X.; Zhang, Y.; Li, X.; Tang, H. MiR-101 regulates HSV-1 replication by targeting ATP5B. Antiviral Res. 2011, 89, 219–226. [Google Scholar]
- Anselmo, A.; Flori, L.; Jaffrezic, F.; Rutigliano, T.; Cecere, M.; Cortes-Perez, N.; Lefèvre, F.; Rogel-Gaillard, C.; Giuffra, E. Co-expression of host and viral microRNAs in porcine dendritic cells infected by the pseudorabies virus. PLoS One 2011, 6, e17374. [Google Scholar] [CrossRef]
- Lambeth, L.S.; Yao, Y.; Smith, L.P.; Zhao, Y.; Nair, V. MicroRNAs 221 and 222 target p27Kip1 in Marek’s disease virus-transformed tumour cell line MSB-1. J. Gen. Virol. 2009, 90, 1164–1171. [Google Scholar] [CrossRef]
- Bauman, Y.; Nachmani, D.; Vitenshtein, A.; Tsukerman, P.; Drayman, N.; Stern-Ginossar, N.; Lankry, D.; Gruda, R.; Mandelboim, O. An identical miRNA of the human JC and BK polyoma viruses targets the stress-induced ligand ULBP3 to escape immune elimination. Cell Host Microbe 2011, 9, 93–102. [Google Scholar] [CrossRef]
- Sullivan, C.S.; Sung, C.K.; Pack, C.D.; Grundhoff, A.; Lukacher, A.E.; Benjamin, T.L.; Ganem, D. Murine Polyomavirus encodes a microRNA that cleaves early RNA transcripts but is not essential for experimental infection. Virology 2009, 387, 157–167. [Google Scholar] [CrossRef]
- Sullivan, C.S.; Grundhoff, A.T.; Tevethia, S.; Pipas, J.M.; Ganem, D. SV40-encoded microRNAs regulate viral gene expression and reduce susceptibility to cytotoxic T cells. Nature 2005, 435, 682–686. [Google Scholar] [CrossRef]
- Chen, C.J.; Kincaid, R.P.; Seo, G.J.; Bennett, M.D.; Sullivan, C.S. Insights into Polyomaviridae microRNA function derived from study of the bandicoot papillomatosis carcinomatosis viruses. J. Virol. 2011, 85, 4487–4500. [Google Scholar] [CrossRef]
- Huang, J.; Wang, F.; Argyris, E.; Chen, K.; Liang, Z.; Tian, H.; Huang, W.; Squires, K.; Verlinghieri, G.; Zhang, H. Cellular microRNAs contribute to HIV-1 latency in resting primary CD4+ T lymphocytes. Nat. Med. 2007, 13, 1241–1247. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, S.; Huang, C.; Zhang, W.; Hu, Y.; Wei, B. MicroRNAome of splenic macrophages in hypersplenism due to portal hypertension in hepatitis B virus-related cirrhosis. Exp. Biol. Med. 2008, 233, 1454–1461. [Google Scholar] [CrossRef]
- Li, Y.; Chan, E.Y.; Li, J.; Ni, C.; Peng, X.; Rosenzweig, E.; Tumpey, T.M.; Katze, M.G. MicroRNA expression and virulence in pandemic influenza virus-infected mice. J. Virol. 2010, 84, 3023–3032. [Google Scholar]
- Loveday, E.K.; Svinti, V.; Diederich, S.; Pasick, J.; Jean, F. Temporal- and strain-specific host microRNA molecular signatures associated with swine-origin H1N1 and avian-origin H7N7 influenza A virus infection. J. Virol. 2012, 86, 6109–6122. [Google Scholar]
- Grinberg, M.; Gilad, S.; Meiri, E.; Levy, A.; Isakov, O.; Ronen, R.; Shomron, N.; Bentwich, Z.; Shemer-Avni, Y. Vaccinia virus infection suppresses the cell microRNA machinery. Arch. Virol. 2012, 157, 1719–1727. [Google Scholar] [CrossRef]
- Kakumani, P.K.; Ponia, S.S.; Rajgokul, K.S.; Sood, V.; Chinnappan, M.; Banerjea, A.C.; Medigeshi, G.R.; Malhotra, P.; Mukherjee, S.K.; Bhatnagar, R.K. Role of RNA Interference (RNAi) in dengue virus replication and identification of NS4B as an RNAi suppressor. J. Virol. 2013, 87, 8870–8883. [Google Scholar] [CrossRef]
- Jopling, C.L.; Yi, M.; Lancaster, A.M.; Lemon, S.M.; Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science 2005, 309, 1577–1581. [Google Scholar] [CrossRef]
- Lewis, A.P.; Jopling, C.L. Regulation and biological function of the liver-specific miR-122. Biochem. Soc. Trans. 2010, 38, 1553–1557. [Google Scholar] [CrossRef]
- Fukuhara, T.; Kambara, H.; Shiokawa, M.; Ono, C.; Katoh, H.; Morita, E.; Okuzaki, D.; Maehara, Y.; Koike, K.; Matsuura, Y. Expression of MicroRNA miR-122 Facilitates an Efficient Replication in Nonhepatic Cells upon Infection with Hepatitis C Virus. J. Virol. 2012, 86, 7918–7933. [Google Scholar] [CrossRef]
- Norman, K.L.; Sarnow, P. Modulation of hepatitis C virus RNA abundance and the isoprenoid biosynthesis pathway by microRNA miR-122 involves distinct mechanisms. J. Virol. 2010, 84, 666–670. [Google Scholar] [CrossRef]
- Shimakami, T.; Yamane, D.; Welsch, C.; Hensley, L.; Jangra, R.K.; Lemon, S.M. Base pairing between hepatitis C virus RNA and microRNA 122 3' of its seed sequence is essential for genome stabilization and production of infectious virus. J. Virol. 2012, 86, 7372–7383. [Google Scholar]
- González, I.; Martínez, L.; Rakitina, D.V.; Lewsey, M.G.; Atencio, F.A.; Llave, C.; Kalinina, N.O.; Carr, J.P.; Palukaitis, P.; Canto, T. Cucumber mosaic virus 2b protein subcellular targets and interactions: Their significance to RNA silencing suppressor activity. Mol. Plant Microbe Interact. 2010, 23, 294–303. [Google Scholar]
- Kataya, A.R.; Suliman, M.N.; Kalantidis, K.; Livieratos, I.C. Cucurbit yellow stunting disorder virus p25 is a suppressor of post-transcriptional gene silencing. Virus Res. 2009, 145, 48–53. [Google Scholar] [CrossRef]
- Ahn, J.W.; Lee, J.S.; Davarpanah, S.J.; Jeon, J.H.; Park, Y.I.; Liu, J.R.; Jeong, W.J. Host-dependent suppression of RNA silencing mediated by the viral suppressor p19 in potato. Planta 2011, 234, 1065–1072. [Google Scholar] [CrossRef]
- Sansregret, R.; Dufour, V.; Langlois, M.; Daayf, F.; Dunoyer, P.; Voinnet, O.; Bouarab, K. Extreme resistance as a host counter-counter defense against viral suppression of RNA silencing. PLoS Pathog. 2013, 9, e1003435. [Google Scholar] [CrossRef]
- Pacheco, R.; García-Marcos, A.; Barajas, D.; Martiáñez, J.; Tenllado, F. PVX-potyvirus synergistic infections differentially alter microRNA accumulation in Nicotiana benthamiana. Virus Res. 2012, 165, 231–235. [Google Scholar] [CrossRef]
- Maier, L.K.; Fischer, S.; Stoll, B.; Brendel, J.; Pfeiffer, F.; Dyall-Smith, M.; Marchfelder, A. The immune system of halophilic archaea. Mob. Genet. Elements 2012, 2, 228–232. [Google Scholar] [CrossRef]
- Rajaram, M.V.; Ni, B.; Morris, J.D.; Brooks, M.N.; Carlson, T.K.; Bakthavachalu, B.; Schoenberg, D.R.; Torrelles, J.B.; Schlesinger, L.S. Mycobacterium tuberculosis lipomannan blocks TNF biosynthesis by regulating macrophage MAPK-activated protein kinase 2 (MK2) and microRNA miR-125b. Proc. Natl. Acad. Sci. USA 2011, 108, 17408–17413. [Google Scholar] [CrossRef]
- Izar, B.; Mannala, G.K.; Mraheil, M.A.; Chakraborty, T.; Hain, T. microRNA Response to Listeria monocytogenes Infection in Epithelial Cells. Int. J. Mol. Sci. 2012, 13, 1173–1185. [Google Scholar] [CrossRef]
- Burgyán, J.; Havelda, Z. Viral suppressors of RNA silencing. Trends Plant Sci. 2011, 16, 265–272. [Google Scholar] [CrossRef]
- Pantaleo, V.; Saldarelli, P.; Miozzi, L.; Giampetruzzi, A.; Gisel, A.; Moxon, S.; Dalmay, T.; Bisztray, G.; Burgyan, J. Deep sequencing analysis of viral short RNAs from an infected Pinot Noir grapevine. Virology 2010, 408, 49–56. [Google Scholar] [CrossRef]
- Shen, W.J.; Ruan, X.L.; Li, X.S.; Zhao, Q.; Li, H.P. RNA silencing suppressor Pns11 of rice gall dwarf virus induces virus-like symptoms in transgenic rice. Arch. Virol. 2012, 157, 1531–1539. [Google Scholar] [CrossRef]
- Ding, B. Viroids: Self-replicating, mobile, and fast-evolving noncoding regulatory RNAs. Wiley Interdiscip. Rev. RNA 2010, 1, 362–375. [Google Scholar] [CrossRef]
- Navarro, B.; Pantaleo, V.; Gisel, A.; Moxon, S.; Dalmay, T.; Bisztray, G.; di Serio, F.; Burgyán, J. Deep sequencing of viroid-derived small RNAs from grapevine provides new insights on the role of RNA silencing in plant-viroid interaction. PLoS One 2009, 4, e7686. [Google Scholar] [CrossRef]
- St-Pierre, P.; Hassen, I.F.; Thompson, D.; Perreault, J.P. Characterization of the siRNAs associated with peach latent mosaic viroid infection. Virology 2009, 383, 178–182. [Google Scholar] [CrossRef]
- Diermann, N.; Matoušek, J.; Junge, M.; Riesner, D.; Steger, G. Characterization of plant miRNAs and small RNAs derived from potato spindle tuber viroid (PSTVd) in infected tomato. Biol. Chem. 2010, 391, 1379–1390. [Google Scholar]
- Makarova, K.S.; Wolf, Y.I.; Koonin, E.V. Comparative genomics of defense systems in archaea and bacteria. Nucleic Acids Res. 2013, 41, 4360–4377. [Google Scholar] [CrossRef]
- Szczepankowska, A. Role of CRISPR/cas system in the development of bacteriophage resistance. Adv. Virus Res. 2012, 82, 289–338. [Google Scholar] [CrossRef]
- Westra, E.R.; Staals, R.H.; Gort, G.; Høgh, S.; Neumann, S.; de la Cruz, F.; Fineran, P.C.; Brouns, S.J. CRISPR-Cas systems preferentially target the leading regions of MOBF conjugative plasmids. RNA Biol. 2013, 10, 749–761. [Google Scholar] [CrossRef]
- Savitskaya, E.; Semenova, E.; Dedkov, V.; Metlitskaya, A.; Severinov, K. High-throughput analysis of type I-E CRISPR/Cas spacer acquisition in E. coli. RNA Biol. 2013, 10, 716–725. [Google Scholar]
- Han, P.; Niestemski, L.R.; Barrick, J.E.; Deem, M.W. Physical model of the immune response of bacteria against bacteriophage through the adaptive CRISPR-Cas immune system. Phys. Biol. 2013, 10. [Google Scholar] [CrossRef]
- Lange, S.J.; Alkhnbashi, O.S.; Rose, D.; Will, S.; Backofen, R. CRISPRmap: An automated classification of repeat conservation in prokaryotic adaptive immune systems. Nucleic Acids Res. 2013, 41, 8034–8044. [Google Scholar] [CrossRef]
- Cai, F.; Axen, S.D.; Kerfeld, C.A. Evidence for the widespread distribution of CRISPR-Cas system in the Phylum Cyanobacteria. RNA Biol. 2013, 10, 687–693. [Google Scholar] [CrossRef]
- Díez-Villaseñor, C.; Guzmán, N.M.; Almendros, C.; García-Martínez, J.; Mojica, F.J. CRISPR-spacer integration reporter plasmids reveal distinct genuine acquisition specificities among CRISPR-Cas I-E variants of Escherichia coli. RNA Biol. 2013, 10, 792–802. [Google Scholar] [CrossRef] [Green Version]
- Jaubert, C.; Danioux, C.; Oberto, J.; Cortez, D.; Bize, A.; Krupovic, M.; She, Q.; Forterre, P.; Prangishvili, D.; Sezonov, G. Genomics and genetics of Sulfolobus islandicus LAL14/1, a model hyperthermophilic archaeon. Open Biol. 2013, 3. [Google Scholar] [CrossRef]
- Bondy-Denomy, J.; Pawluk, A.; Maxwell, K.L.; Davidson, A.R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 2013, 493, 429–432. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hicks, J.; Liu, H.-C. Involvement of Eukaryotic Small RNA Pathways in Host Defense and Viral Pathogenesis. Viruses 2013, 5, 2659-2678. https://doi.org/10.3390/v5112659
Hicks J, Liu H-C. Involvement of Eukaryotic Small RNA Pathways in Host Defense and Viral Pathogenesis. Viruses. 2013; 5(11):2659-2678. https://doi.org/10.3390/v5112659
Chicago/Turabian StyleHicks, Julie, and Hsiao-Ching Liu. 2013. "Involvement of Eukaryotic Small RNA Pathways in Host Defense and Viral Pathogenesis" Viruses 5, no. 11: 2659-2678. https://doi.org/10.3390/v5112659




