Increasing Phosphatidylinositol (4,5)-Bisphosphate Biosynthesis Affects Basal Signaling and Chloroplast Metabolism in Arabidopsis thaliana
Abstract
:1. Introduction
2. Results and Discussion
2.1. Generation and Growth of HsPIPKIα Transgenic Plants
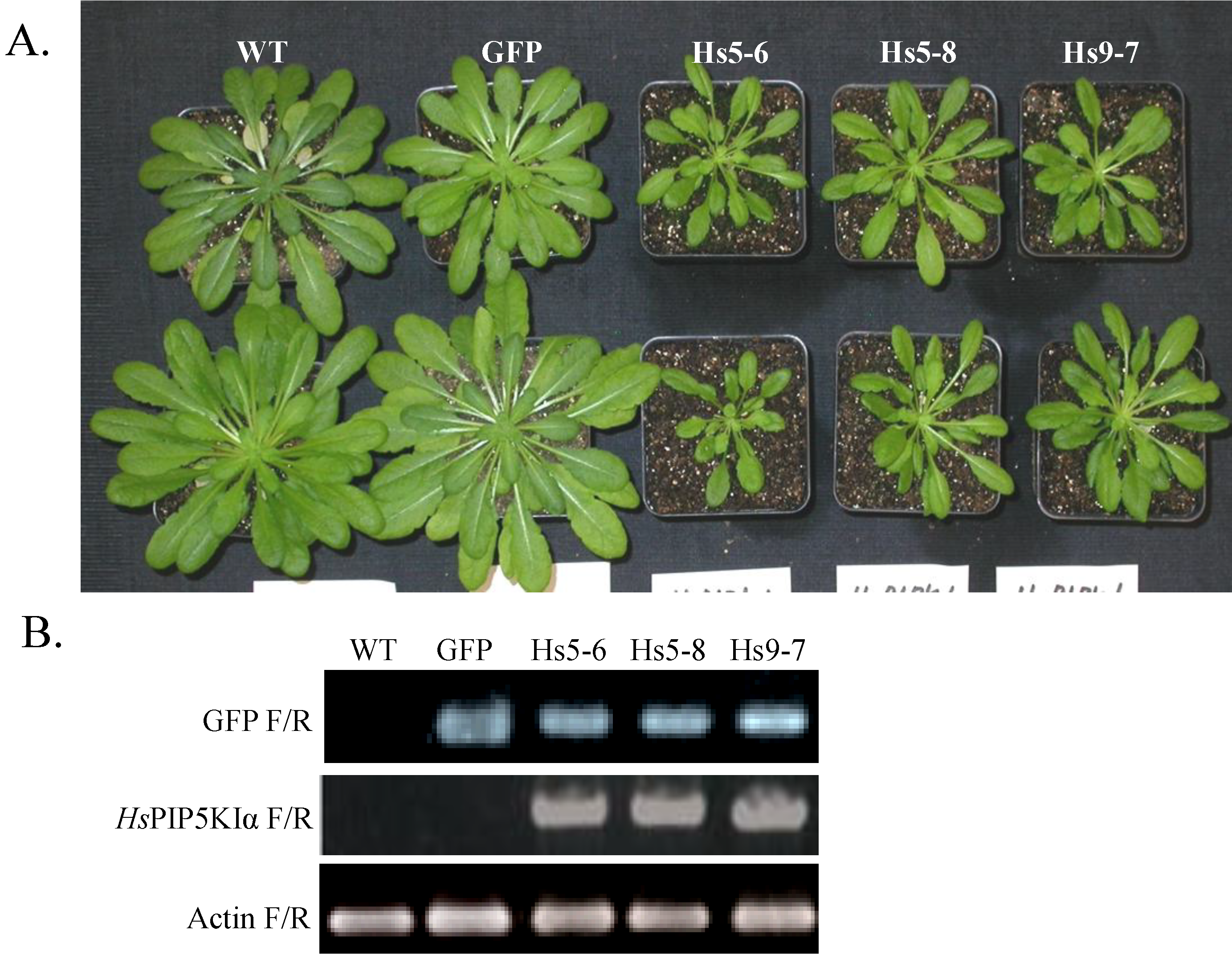


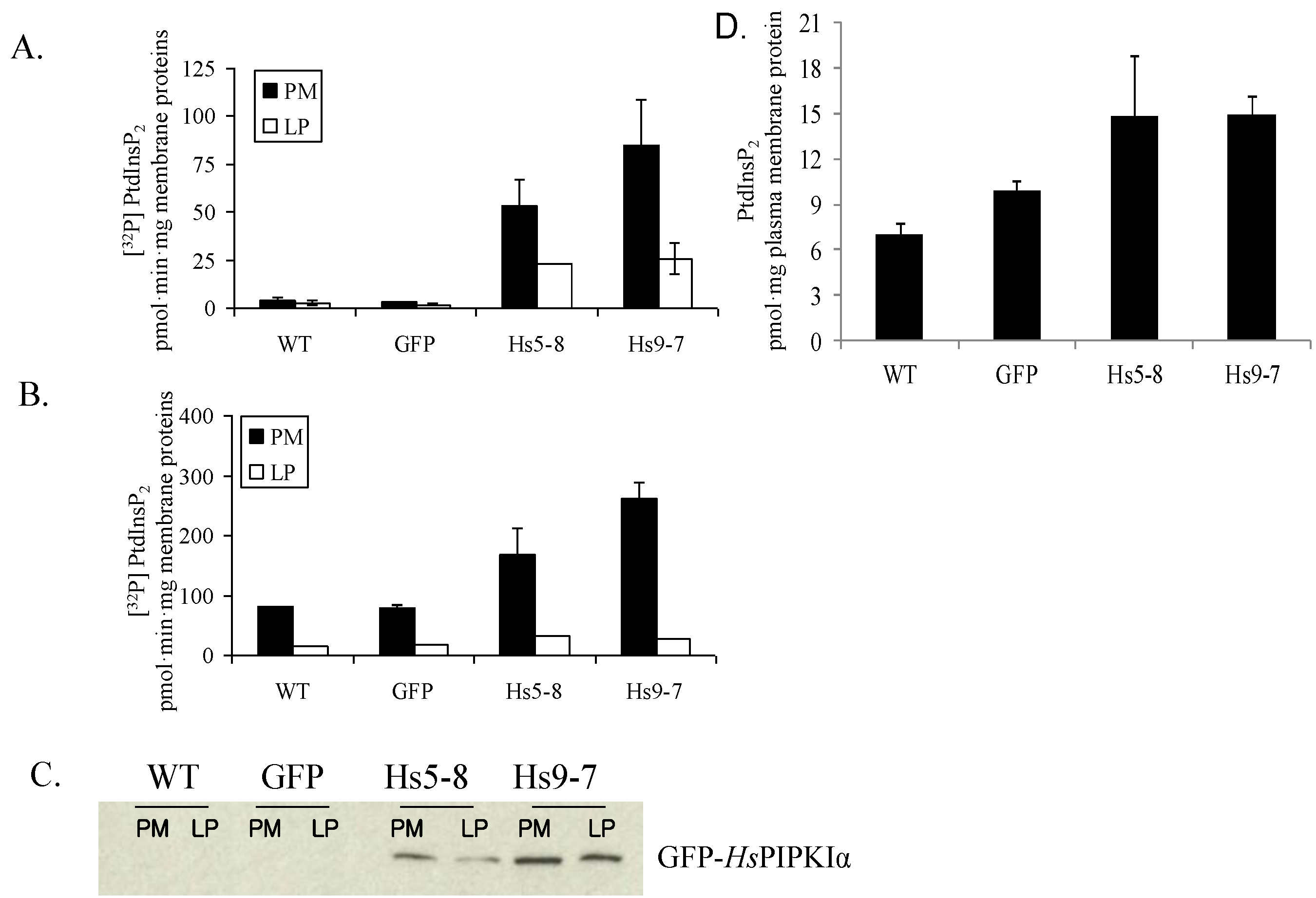
2.2. PtdInsP2 and PIPK Specific Activity Increased in Seedlings and Leaves
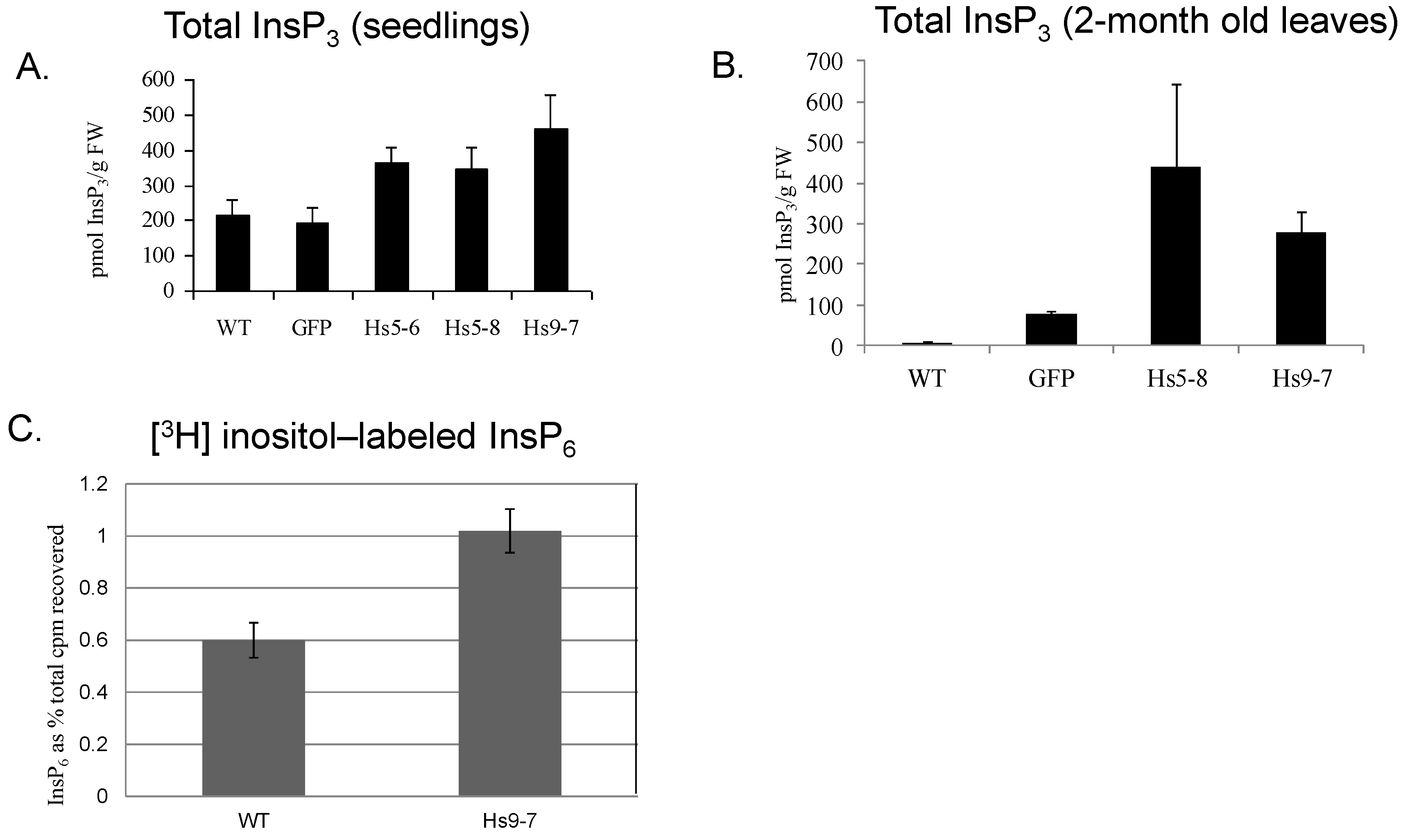
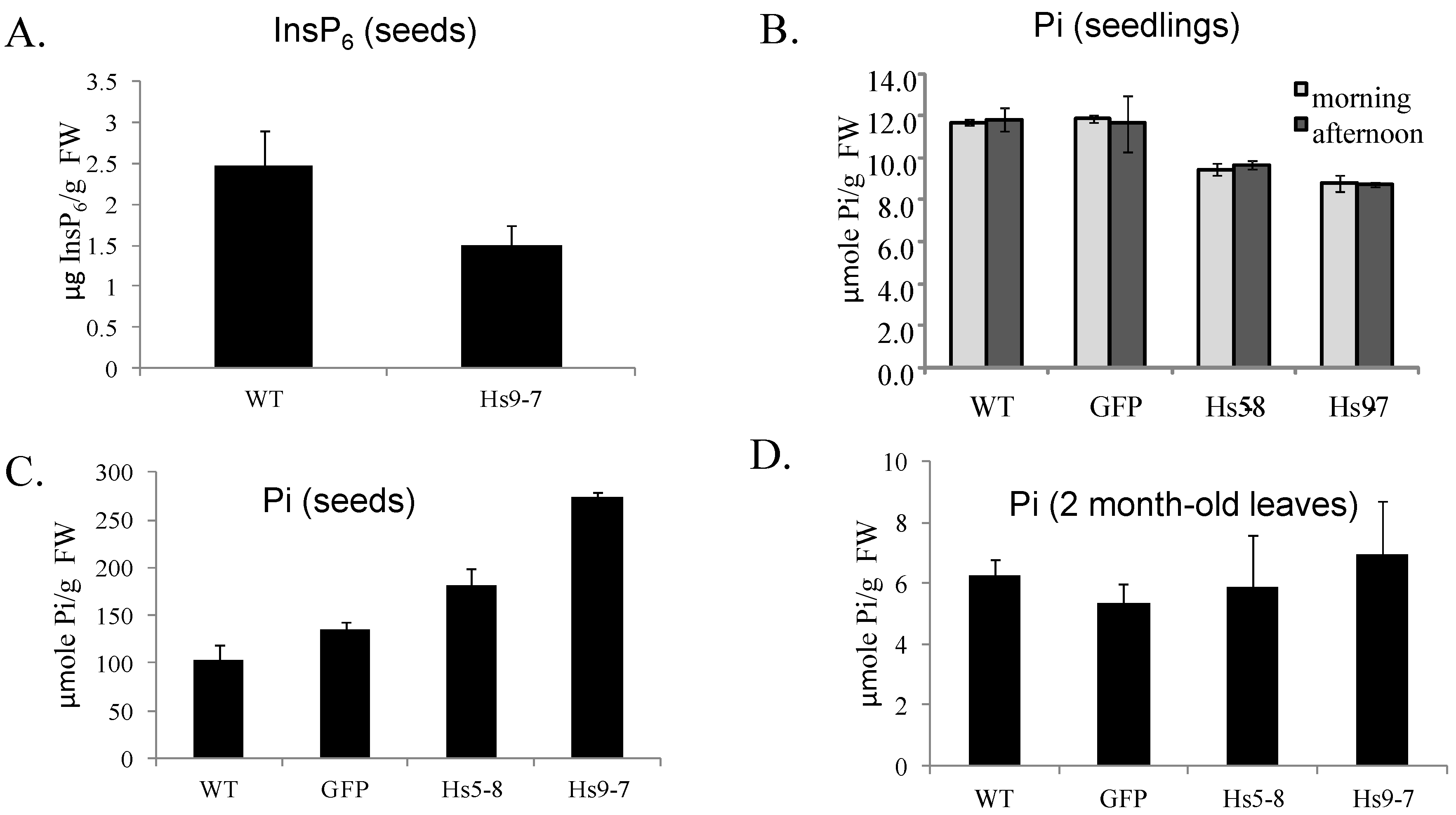
2.3. Increased Flux through the Phosphoinositide Pathway in HsPIPKIα Transgenic Plants

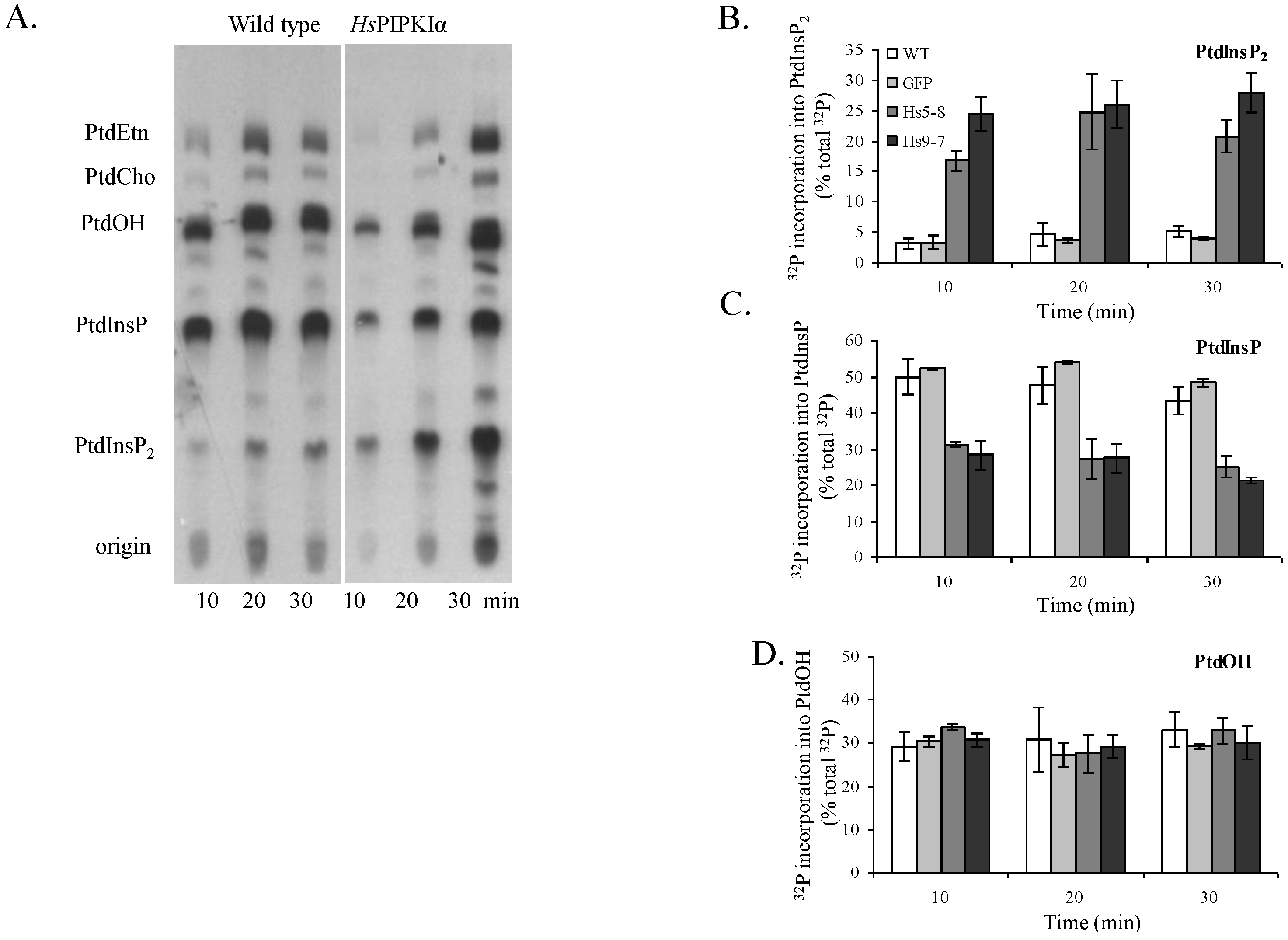
| Plant type | PtdInsP2 | PtdInsP | PtdIns | Ratio of PtdInsP/PtdInsP2 |
|---|---|---|---|---|
| WT | 0.3 ± 0.2 | 7.0 ± 0.3 | 31.9 ± 1 | 22 |
| HsPIPKIα 9-7 | 2.9 ± 0.1 | 5.9 ± 0.2 | 37.4 ± 1 | 2 |
| Plant | Concentration (mg/dry weight (g)) | ||||||
|---|---|---|---|---|---|---|---|
| P | Ca | K | Mg | S | Mn | Fe | |
| WT | 9.5 ± 0.1 | 5.8 ± 0.1 | 60.5 ± 0.6 | 2.5 ± 0.04 | 11.1 ±0.8 | 0.2 ± 0.003 | 1.0 ± 0.3 |
| GFP | 9.0 ± 0.5 | 5.1 ± 0.1 | 58.2 ± 0.6 | 2.2 ± 0.04 | 9.7 ± 0.5 | 0.2 ± 0.005 | 1.2 ± 0.2 |
| Hs5-8 | 8.6 ± 0.1 | 4.2 ± 0.1 | 61.2 ± 0.3 | 1.7 ± 0.04 | 8.8 ± 0.6 | 0.2 ± 0.006 | 0.9 ± 0.2 |
| Hs9-7 | 7.8 ± 0.4 | 4.5 ± 0.1 | 61.1 ± 1.3 | 1.8 ± 0.04 | 9.8 ± 0.9 | 0.2 ± 0.005 | 0.9 ± 0.3 |
2.4. Starch Metabolism Is Altered HsPIPKIα Plants
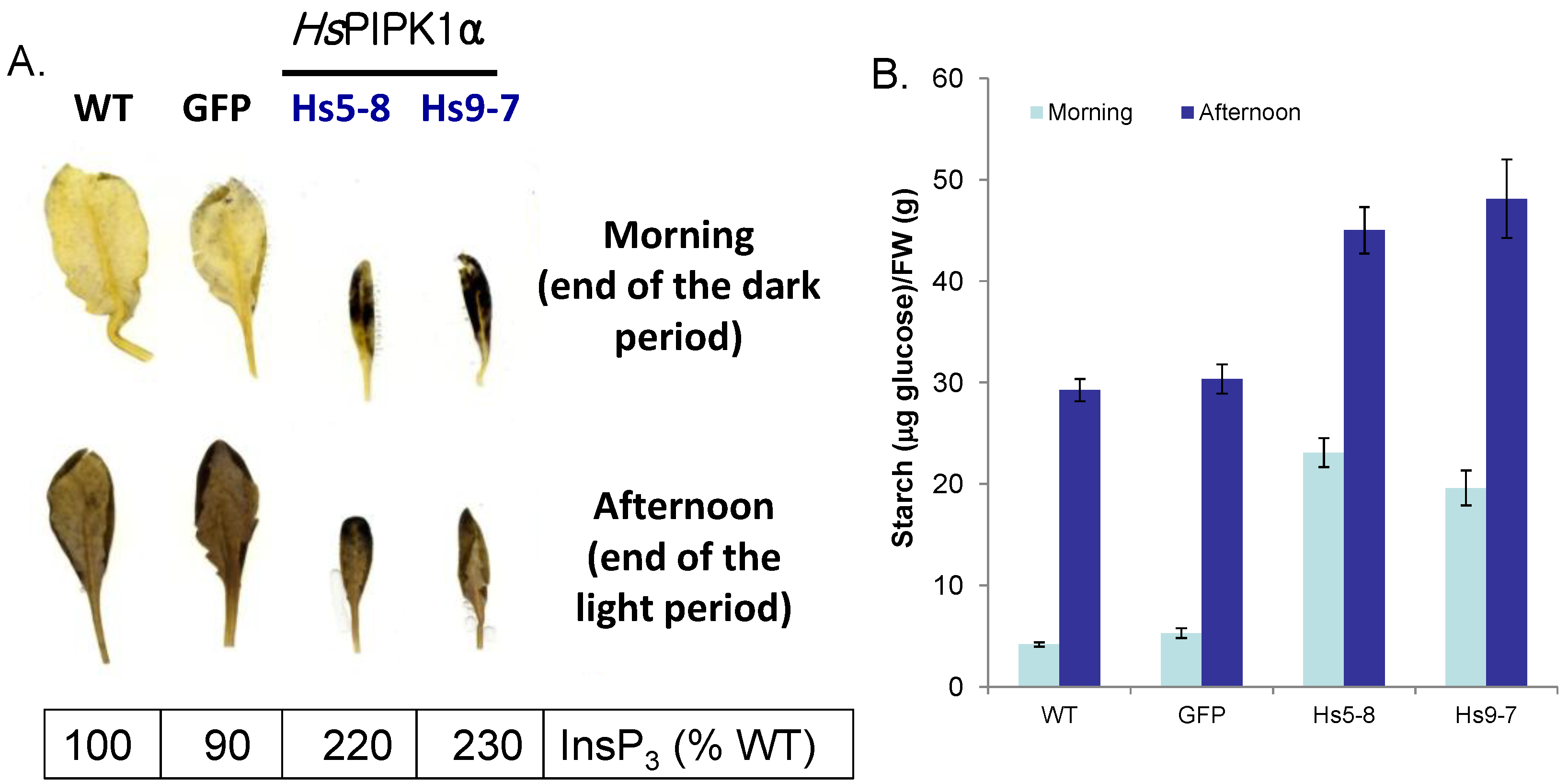
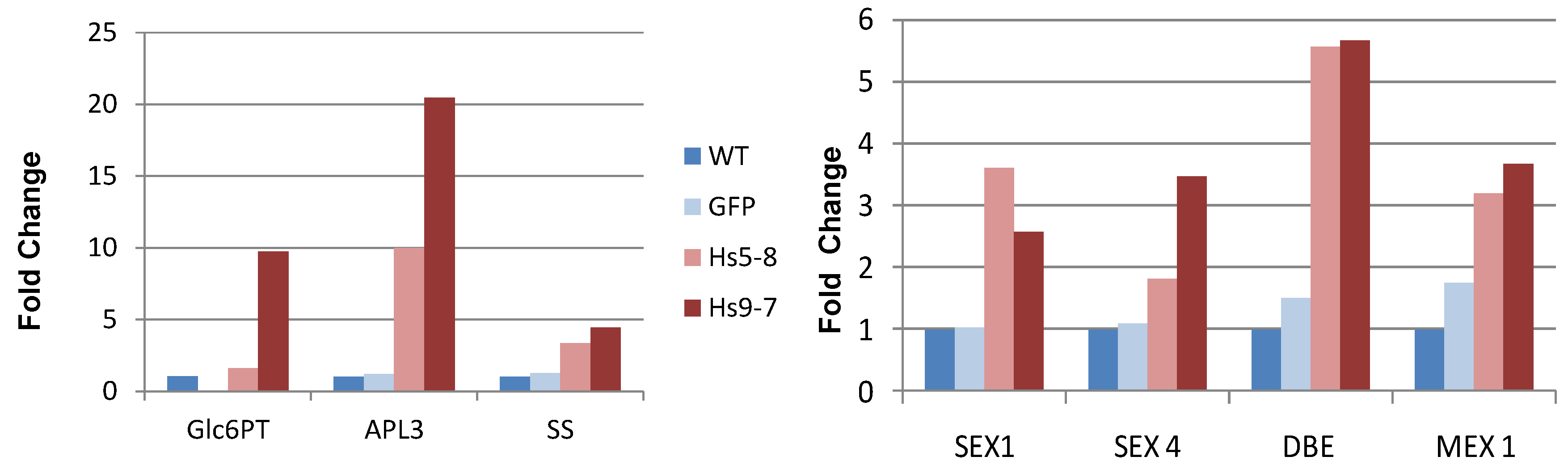
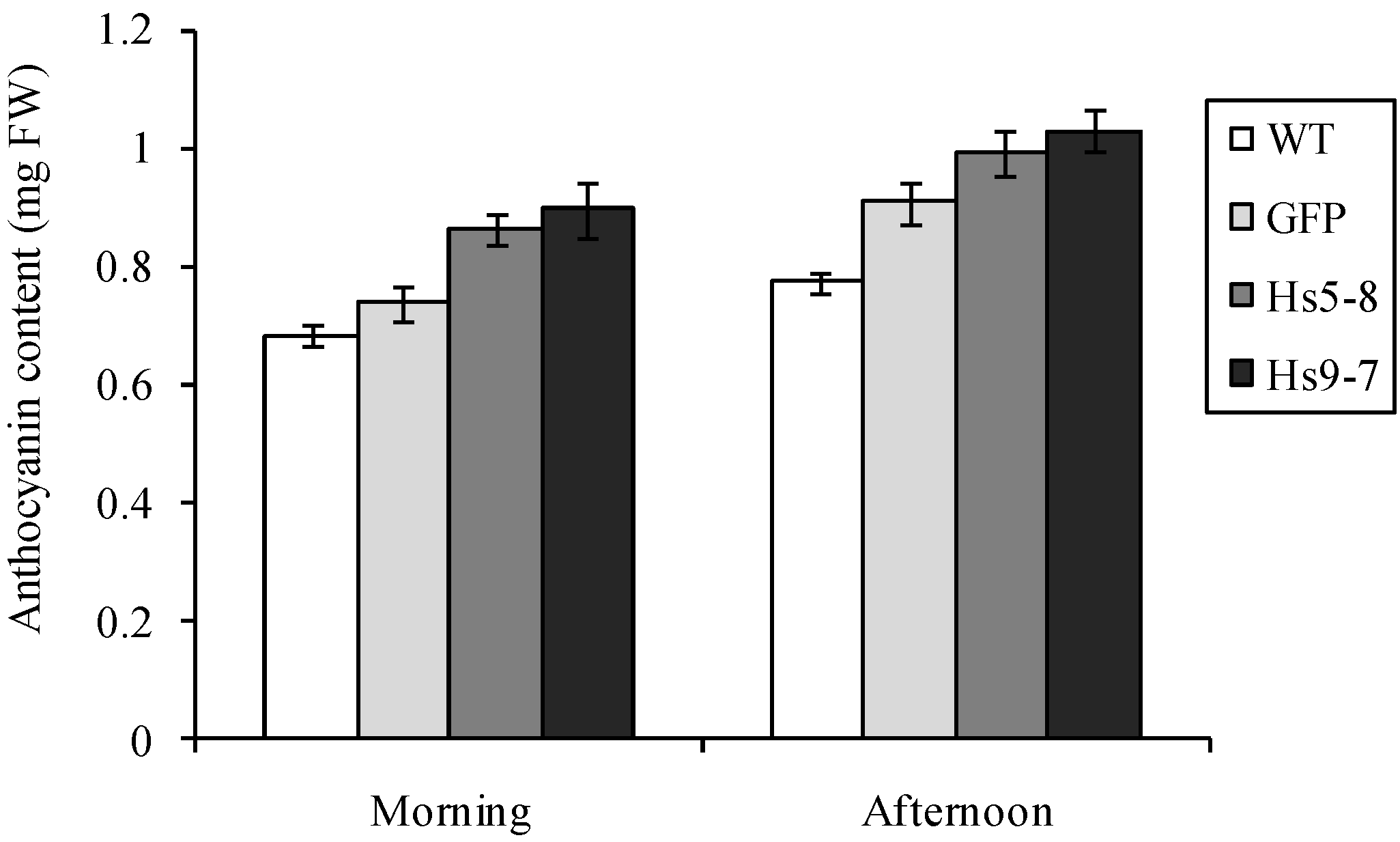
2.5. Constitutively Increasing PtdInsP2 Biosynthesis and InsP3 in Leaves did not Affect Photosynthetic Electron Transport
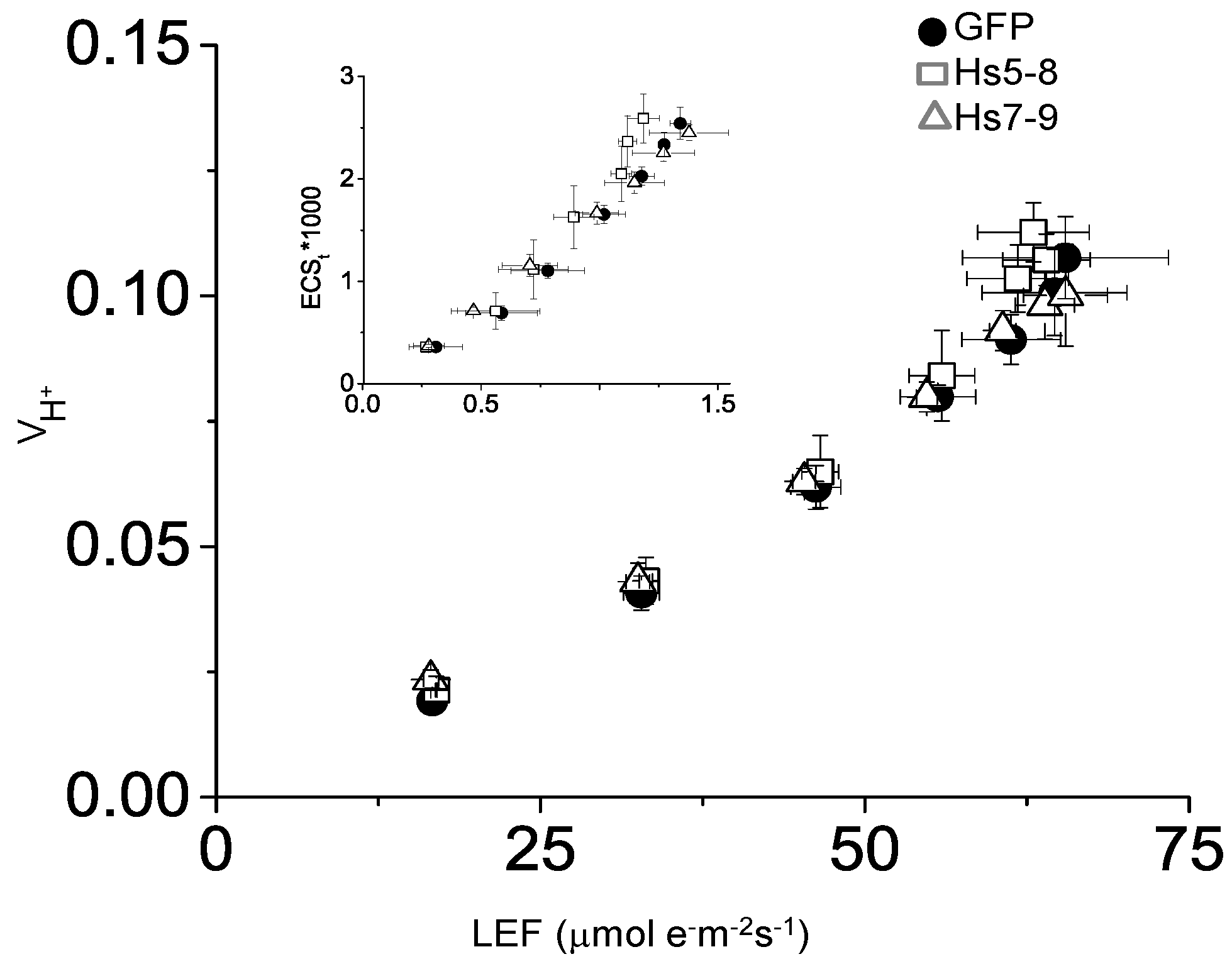
2.6. Physiological Characteristics of the HsPIPK1α Plants
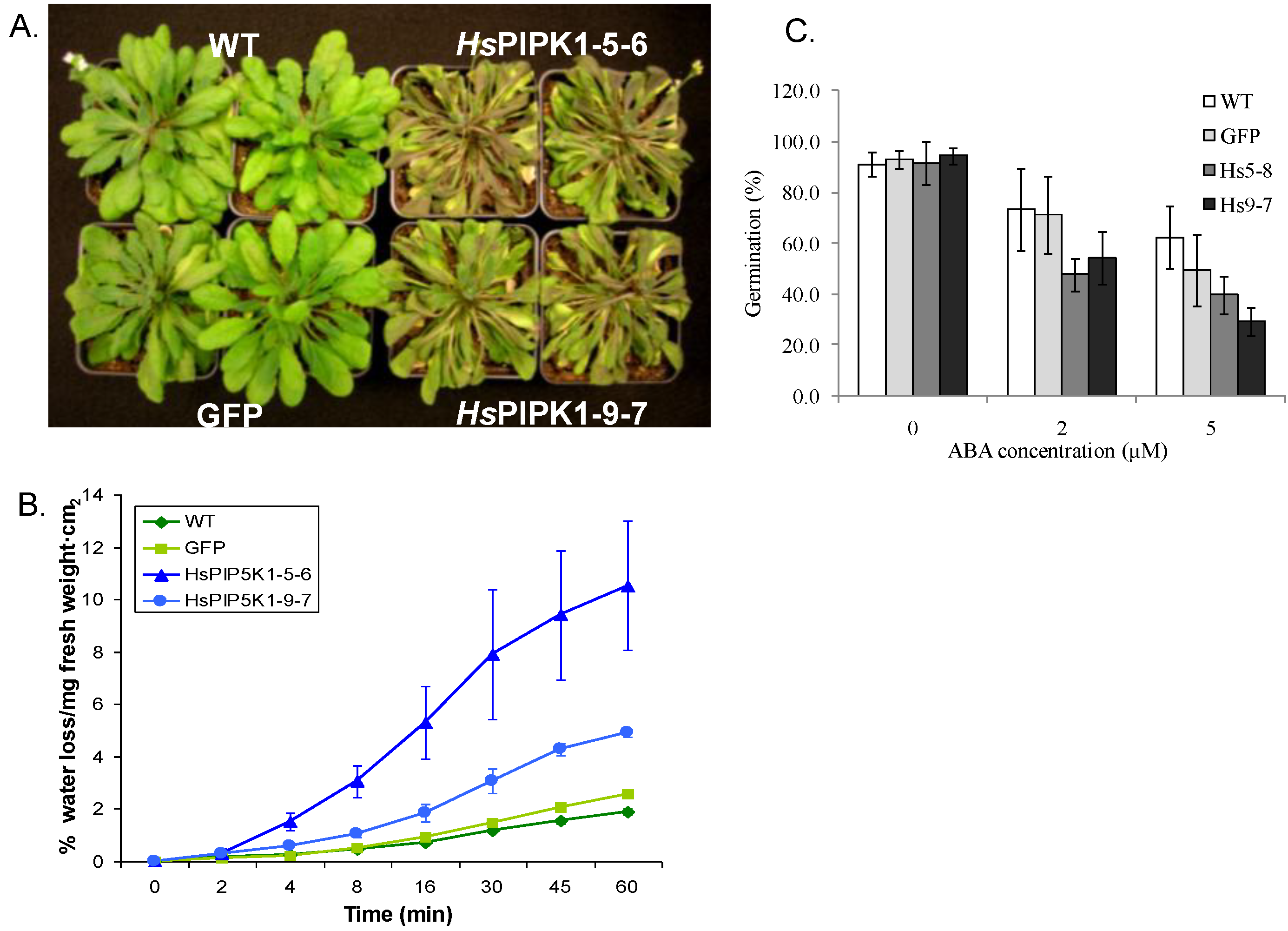
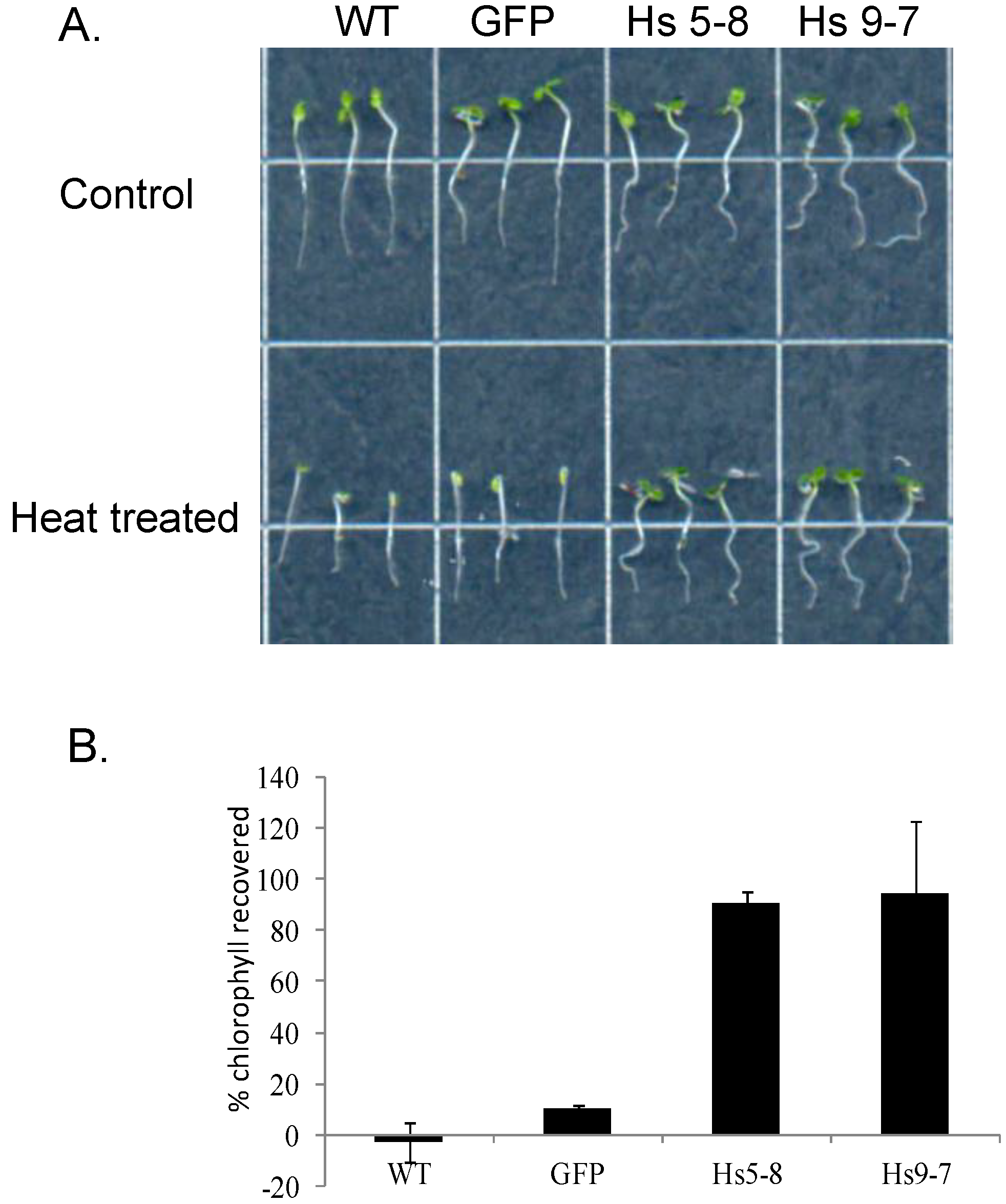
2.7. Very Few Differences in Transcript Levels Were Detected Using Microarray Analysis
| AGI Locus ID | Gene Descriptor | Microarray Fold Change | Log Ratio |
|---|---|---|---|
| Up-regulated Expression | |||
| At2g14610 | pathogenesis-related protein 1 (PR-1) | 10.97 (9-7) | 3.46 (9-7) |
| 2.40 (5-8) | 1.26 (5-8) | ||
| At3g15650 | phospholipase/carboxylesterase family protein | 5.56 (9-7) | 2.48 (9-7) |
| 4.32 (5-8) | 2.11 (5-8) | ||
| At1g73040 | jacalin lectin family protein | 5.36 (9-7) | 2.42 (9-7) |
| 4.48 (5-8) | 2.16 (5-8) | ||
| At1g69880 | thioredoxin, putative | 4.91 (9-7) | 2.30 (9-7) |
| 3.69 (5-8) | 1.88 (5-8) | ||
| At1g19960 | transmembrane receptor, putative | 4.27 (9-7) | 2.09 (9-7) |
| 2.73 (5-8) | 1.45 (5-8) | ||
| At4g23680 | major latex protein-related | 3.38 (9-7) | 1.76 (9-7) |
| 3.11 (5-8) | 1.64 (5-8) | ||
| At1g32450 | proton-dependent oligopeptide transport (POT) family protein | 3.22 (9-7) | 1.69 (9-7) |
| 2.18 (5-8) | 1.13 (5-8) | ||
| At4g15110 | cytochrome P450 97B3, putative | 2.91 (9-7) | 1.54 (9-7) |
| 3.43 (5-8) | 1.78 (5-8) | ||
| At4g32280 | auxin-responsive family protein | 2.74 (9-7) | 1.45 (9-7) |
| 2.34 (5-8) | 1.23 (5-8) | ||
| At4g12550 | protease inhibitor/seed storage/lipid transfer protein (LTP) family protein | 2.68 (9-7) | 1.42 (9-7) |
| 2.70 (5-8) | 1.43 (5-8) | ||
| At5g39110 | germin-like protein, putative | 2.48 (9-7) | 1.31 (9-7) |
| 2.48 (5-8) | 2.48 (5-8) | ||
| At5g59520 | zinc transporter (ZIP2) | 2.45 (9-7) | 1.29 (9-7) |
| 2.89 (5-8) | 1.53 (5-8) | ||
| At3g25830 | myrcene/ocimene synthase (TPS10) | 2.42 (9-7) | 1.27 (5-8) |
| 3.69 (9-7) | 1.89 (5-8) | ||
| At4g30170 | peroxidase, putative | 2.40 (9-7) | 1.26 (9-7) |
| 2.00 (5-8) | 1.00 (5-8) | ||
| At1g78340 | glutathione S-transferase, putative | 2.39 (9-7) | 1.26 99-7) |
| 2.26 (5-8) | 1.18 (5-8) | ||
| At3g28530 | gypsy-like retrotransposon family | 2.27 (9-7) | 1.18 (9-7) |
| 2.00 (5-8) | 1.00 (5-8) | ||
| At3g62040 | haloacid dehalogenase-like hydrolase family protein | 2.22 (9-7) | 1.15 (9-7) |
| 3.03 (5-8) | 1.60 (5-8) | ||
| At3g08860 | alanine-glyoxylate aminotransferase | 2.17 (9-7) | 1.12 (9-7) |
| 2.50 (5-8) | 1.32 (5-8) | ||
| At2g01520 | major latex protein-related | 2.10 (9-7) | 1.07 (9-7) |
| 2.18 (5-8) | 1.12 (5-8) | ||
| At3g46130 | myb family transcription factor (MYB48) | 2.03 (9-7) | 1.02 (9-7) |
| 2.40 (5-8) | 1.26 (5-8) | ||
| At3g49160 | pyruvate kinase family protein | 2.00 (9-7) | 1.00 (9-7) |
| 3.16 (5-8) | 1.66 (5-8) | ||
| Down-regulated Expression | |||
| At3g22640 | cupin family protein | −2.08 (9-7) | −1.06 (9-7) |
| −2.48 (5-8) | −1.31 (5-8) | ||
| At5g14180 | lipase family protein | −2.40 (9-7) | −1.27 (9-7) |
| −2.27 (5-8) | −1.19 (5-8) | ||
| At2g34600 | jasmonate-zim-domain protein 7 | −2.34 (9-7) | −1.23 (9-7) |
| −2.28 (5-8) | −1.19 (5-8) | ||
| At1g66900 | α/β-hydrolase domain-containing Protein | −2.65 (9-7) | −1.41 (9-7) |
| −2.27 (5-8) | −1.18 (5-8) | ||
3. Experimental
3.1. Generation and Selection of HsPIPKIα Transgenic Plants
3.2. Plant Growth Conditions
3.3. Seed Germination Assays
3.4. RNA Extraction, RT-PCR, and qRT-PCR Analysis
3.5. Protein Isolation and Immunoblotting
3.6. PtdInsP 5-Kinase Assays
3.7. Ins(1,4,5)P3 Assays
3.8. Lipid Profiling
3.9. In Vivo Labeling Studies
3.10. Labeling Studies with [3H]myo-Inositol
3.11. Determination of Total InsP6 in Seeds
3.12. Quantification of Soluble Pi
3.13. Determination of Anthocyanin and Chlorophyll A
3.14. Staining and Quantification of Starch
3.15. Analysis of ATP and NADP(H) and NAD(H)
3.16. Sugar Analysis
3.17. ICP Analysis
3.18. In Vivo Spectroscopic Analysis
3.19. RNA Isolation for Microarray Analysis
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgements
Conflicts of Interest
References
- Balla, T. Phosphoinositides: Tiny lipids with giant impact on cell regulation. Physiol. Rev. 2013, 93, 1019–1137. [Google Scholar] [CrossRef]
- Munnik, T.; Nielsen, E. Green light for polyphosphoinositide signals in plants. Curr. Opin. Plant. Biol. 2011, 14, 498–497. [Google Scholar] [CrossRef]
- Boss, W.F.; Im, Y.J. Phosphoinositide signaling. Annu. Rev. Plant Biol. 2012, 63, 409–429. [Google Scholar] [CrossRef]
- Ischebeck, T.; Seiler, S.; Heilmann, I. At the poles across kingdoms: Phosphoinositides and polar tip growth. Protoplasma 2010, 240, 13–31. [Google Scholar] [CrossRef]
- Dowd, P.; Gilroy, S. The emerging roles of phospholipase C in plant growth and development. In Lipid Signaling in Plants, Plant Cell Monographs; Springer: Berlin/Heidelberg, Germany, 2010; Volume 16, pp. 23–37. [Google Scholar]
- Kusano, H.; Testerink, C.; Vermeer, J.E.M.; Tsuge, T.; Shimada, H.; Oka, A.; Munnik, T.; Aoyama, T. The Arabidopsis phosphatidylinositol phosphate 5-kinase PIP5K3 is a key regulator of root hair tip growth. Plant Cell 2008, 20, 367–380. [Google Scholar] [CrossRef]
- Lee, Y.; Kim, E.-S.; Choi, Y.; Hwang, I.; Staiger, C.J.; Chung, Y.-Y.; Lee, Y. The Arabidopsis phosphatidylinositol 3-kinase is important for pollen development. Plant Physiol. 2008, 147, 1886–1897. [Google Scholar] [CrossRef]
- Lee, Y.; Bak, G.; Choi, Y.; Chuang, W.-I.; Cho, H.-T.; Lee, Y. Roles of phosphatidylinositol 3-kinase in root hair growth. Plant Physiol. 2008, 147, 624–635. [Google Scholar] [CrossRef]
- Tan, X.; Calderon-Villalobos, L.I.; Sharon, M.; Zheng, C.; Robinson, C.V.; Estelle, M.; Zheng, N. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 2007, 446, 640–645. [Google Scholar] [CrossRef]
- Sheard, L.B.; Tan, X.; Mao, H.; Withers, J.; Ben-Nissan, G.; Hinds, T.R.; Kobayashi, Y.; Hsu, F.-F.; Sharon, M.; Browse, J.; et al. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature 2010, 468, 400–405. [Google Scholar] [CrossRef]
- Mosblech, A.; Thurow, C.; Gatz, C.; Feussner, I.; Heilmann, I. Jasmonic acid perception by COI1 involves inositol polyphosphates in Arabidopsis thaliana. Plant J. 2011, 65, 949–957. [Google Scholar] [CrossRef]
- Chen, X.; Lin, W.H.; Wang, Y.; Luan, S.; Xue, H.W. An inositol polyphosphate 5-phosphatase functions in PHOTOTROPIN1 signaling in Arabidopsis by altering cytosolic Ca2+. Plant Cell 2008, 20, 353–366. [Google Scholar] [CrossRef]
- Salinas-Mondragon, R.E.; Kajla, J.D.; Perera, I.Y.; Brown, C.S.; Sederoff, H.W. Role of inositol 1,4,5-triphosphate signalling in gravitropic and phototropic gene expression. Plant Cell Environ. 2010, 33, 2041–2055. [Google Scholar] [CrossRef]
- Lee, Y.; Jung, J.-Y.; Kim, Y.-W.; Kwak, J.M.; Young, J.; Hwang, J.-U.; Schroder, J.I.; Hwang, I. Phosphatidylinositol 3- and 4-phosphate are required for normal stomatal movements. Plant Cell 2002, 14, 2399–2412. [Google Scholar] [CrossRef]
- Perera, I.Y.; Hung, C.Y.; Moore, C.D.; Stevenson-Paulik, J.; Boss, W.F. Transgenic Arabidopsis plants expressing the type 1 inositol 5-phosphatase exhibit increased drought tolerance and altered abscisic acid signaling. Plant Cell 2008, 20, 2876–2893. [Google Scholar]
- Bak, G.; Lee, E.-J.; Lee, Y.; Kato, M.; Segami, S.; Sze, H.; Maeshima, M.; Hwang, J.-U.; Lee, Y. Rapid structural changes and acidification of guard cell vacuoles during stomatal closure require phosphatidylinositol 3,5-bisphosphate. Plant Cell 2013, 25, 2202–2216. [Google Scholar] [CrossRef]
- Ananieva, E.A.; Gillaspy, G.E.; Ely, A.; Burnette, R.N.; Erickson, F.L. Interaction of the WD40 domain of a myoinositol polyphosphate 5-phosphatase with SnRK1 links inositol, sugar, and stress signaling. Plant Physiol. 2008, 148, 1868–1882. [Google Scholar] [CrossRef]
- Ehrhardt, D.W.; Wais, R.; Long, S.R. Calcium spiking in plant root hairs responding to Rhizobium nodulation signals. Cell 1996, 85, 673–681. [Google Scholar] [CrossRef]
- Hong, Z.; Verma, D.P. A phosphatidylinositol 3-kinase is induced during soybean nodule organogenesis and is associated with membrane proliferation. Proc. Natl. Acad. Sci. USA 1994, 91, 9617–9621. [Google Scholar] [CrossRef]
- Dall’Armi, C.; Devereaux, K.A.; di Paolo, G. The role of lipids in the control of autophagy. Curr. Biol. 2013, 23, R33–R45. [Google Scholar]
- Tamura, N.; Oku, M.; Ito, M.; Noda, N.N.; Inagaki, F.; Sakai, Y. Atg18 phosphoregulation controls organellar dynamics by modulating its phosphoinositide-binding activity. J. Cell Biol. 2013, 202, 685–698. [Google Scholar] [CrossRef] [Green Version]
- Preuss, M.L.; Schmitz, A.J.; Thole, J.M.; Bonner, H.K.S.; Otegui, M.S.; Nielsen, E. A role for the RabA4b effector protein, PI-4Kb1, in polarized expansion of root hair cells in Arabidopsis. J. Cell Biol. 2006, 172, 991–998. [Google Scholar] [CrossRef]
- Thole, J.M.; Nielsen, E. Phosphoinositides in plants: Novel functions in membrane trafficking. Curr. Opin. Plant Biol. 2008, 11, 620–631. [Google Scholar] [CrossRef]
- Torabinejad, J.; Gillaspy, G.E. Functional genomics of inositol metabolism. In Biology of Inositols and Phosphoinositides; Majumder, A.L., Biswas, B.B., Eds.; Springer: Berlin/Heidelberg, Germany, 2006; Volume 39, pp. 47–70. [Google Scholar]
- Heilmann, I. Using genetic tools to understand plant phosphoinositide signalling. Trends Plant Sci. 2009, 14, 171–179. [Google Scholar] [CrossRef]
- Gillaspy, G. Signaling and the polyphosphoinositide phosphatases from plants. In Lipid Signaling in Plants, Plant Cell Monographs; Springer: Berlin/Heidelberg, Germany, 2010; Volume 16, pp. 117–130. [Google Scholar]
- Im, Y.; Heilmann, I.; Perera, I. The hull of fame: Lipid signaling in the plasma membrane. In The Plant Plasma Membrane, Plant Cell Monographs; Springer: Berlin/Heidelberg, Germany, 2011; Volume 19, pp. 437–455. [Google Scholar]
- Gillaspy, G.E. The cellular language of myo-inositol signaling: Tansley review. New Phytol. 2011, 192, 823–839. [Google Scholar] [CrossRef]
- Cárdenas, C.; Miller, R.A.; Smith, I.; Bui, T.; Molgó, J.; Müller, M.; Vais, H.; Cheung, K.-H.; Yang, J.; Parker, I.; et al. Essential regulation of cell bioenergetics by constitutive InsP3 receptor Ca2+ transfer to mitochondria. Cell 2010, 142, 270–283. [Google Scholar] [CrossRef]
- Kumar, S.; Dey, D.; Hasan, G. Patterns of gene expression in Drosophila InsP3 receptor mutant larvae reveal a role for InsP3 signaling in carbohydrate and energy metabolism. PLoS One 2011, 6, e24105. [Google Scholar] [CrossRef]
- Johnson, C.H.; Knight, M.R.; Kondo, T.; Masson, P.; Sedbrook, J.; Haley, A.; Trewavas, A. Circadian oscillations of cytosolic and chloroplastic free calcium in plants. Science 1995, 269, 1863–1865. [Google Scholar]
- Love, J.; Dodd, A.N.; Webb, A.A.R. Circadian and diurnal calcium oscillations encode photoperiodic information in Arabidopsis. Plant Cell 2004, 16, 956–966. [Google Scholar] [CrossRef]
- Dalchau, N.; Hubbard, K.E.; Robertson, F.C.; Hotta, C.T.; Briggs, H.M.; Stan, G.-B.; Goncalves, J.M.; Webb, A.A.R. Correct biological timing in Arabidopsis requires multiple light-signaling pathways. Proc. Natl. Acad. Sci. USA 2010, 107, 13171–13176. [Google Scholar] [CrossRef]
- Dodd, A.N.; Kudla, J.; Sanders, D. The language of calcium signaling. Annu. Rev. Plant Biol. 2010, 61, 593–620. [Google Scholar] [CrossRef]
- DeFalco, T.A.; Bender, K.W.; Snedden, W.A. Breaking the code: Ca2+ sensors in plant signalling. Biochem. J. 2010, 425, 27–40. [Google Scholar] [CrossRef]
- Johnson, C.H.; Shingles, R.; Ettinger, W. Regulation and role of calcium fluxes in the chloroplast. In The Structure and Function of Plastids; Wise, R., Hoober, J., Eds.; Springer: Berlin/Heidelberg, Germany, 2007; Volume 23, pp. 403–432. [Google Scholar]
- Sai, J.; Johnson, C.H. Dark-stimulated calcium ion fluxes in the chloroplast stroma and cytosol. Plant Cell 2002, 14, 1279–1291. [Google Scholar]
- Kreimer, G.; Melkonian, M.; Holtum, J.A.M.; Latzko, E. Stromal free calcium concentration and light-mediated activation of chloroplast fructose-1,6-bisphosphatase. Plant Physiol. 1988, 86, 423–428. [Google Scholar] [CrossRef]
- Brauer, M.; Sanders, D.; Stitt, M. Regulation of photosynthetic sucrose synthesis: A role for calcium? Planta 1990, 182, 236–243. [Google Scholar]
- Stael, S.; Wurzinger, B.; Mair, A.; Mehlmer, N.; Vothknecht, U.C.; Teige, M. Plant organellar calcium signalling: An emerging field. J. Exp. Botany 2011, 63, 1525–1542. [Google Scholar]
- Rocha, A.G.; Vothknecht, U.C. The role of calcium in chloroplasts—An intriguing and unresolved puzzle. Protoplasma 2012, 249, 957–966. [Google Scholar] [CrossRef]
- Morse, M.J.; Crain, R.C.; Satter, R.L. Light-stimulated inositol phospholipid turnover in Samanea saman leaf pulvini. Proc. Natl. Acad. Sci. USA 1987, 84, 7075–7078. [Google Scholar] [CrossRef]
- Kim, H.Y.; Cote, G.G.; Crain, R.C. Inositol 1,4,5-trisphosphate may mediate reguation of K+ channels by light and darkness in Samanea saman motor cells. Planta 1996, 198, 279–287. [Google Scholar]
- Coursol, S.; Giglioli-Guivarc’h, N.; Vidal, J.; Pierre, J.-N. An increase in phosphoinositide-specific phospholipase C activity precedes induction of C4 phosphoenolpyruvate carboxylase phosphorylation in illuminated and NH4Cl-treated protoplasts from Digitaria sanguinalis. Plant J. 2000, 23, 497–506. [Google Scholar] [CrossRef]
- Williams, M.E.; Torabinejad, J.; Cohick, E.; Parker, K.; Drake, E.J.; Thompson, J.E.; Hortter, M.; Dewald, D.B. Mutations in the Arabidopsis phosphoinositide phosphatase gene SAC9 lead to overaccumulation of PtdIns(4,5)P2 and constitutive expression of the stress-response pathway. Plant Physiol. 2005, 138, 686–700. [Google Scholar] [CrossRef]
- Gong, P.; Wu, G.; Ort, D.R. Slow dark deactivation of Arabidopsis chloroplast ATP synthase caused by a mutation in a nonplastidic SAC domain protein. Photosynth. Res. 2006, 88, 133–142. [Google Scholar] [CrossRef]
- Perera, I.Y.; Hung, C.Y.; Brady, S.; Muday, G.K.; Boss, W.F. A universal role for inositol 1,4,5-trisphosphate-mediated signaling in plant gravitropism. Plant Physiol. 2006, 140, 746–750. [Google Scholar] [CrossRef]
- Munnik, T.; Vermeer, J.E.M. Osmotic stress-induced phosphoinositide and inositol phosphate signalling in plants. Plant Cell Environ. 2010, 33, 655–669. [Google Scholar] [CrossRef]
- Im, Y.J.; Perera, I.Y.; Brglez, I.; Davis, A.J.; Stevenson-Paulik, J.; Phillippy, B.Q.; Johannes, E.; Allen, N.S.; Boss, W.F. Increasing plasma membrane phosphatidylinositol(4,5)bisphosphate biosynthesis increases phosphoinositide metabolism in Nicotiana tabacum. Plant Cell 2007, 19, 1603–1616. [Google Scholar] [CrossRef]
- Davis, A.J.; Perera, I.Y.; Boss, W.F. Cyclodextrins enhance recombinant phosphatidylinositol phosphate kinase activity. J. Lipid. Res. 2004, 45, 1783–1789. [Google Scholar] [CrossRef]
- Ischebeck, T.; Stenzel, I.; Heilmann, I. Type B phosphatidylinositol-4-phosphate 5-kinases mediate Arabidopsis and Nicotiana tabacum pollen tube growth by regulating apical pectin secretion. Plant Cell 2008, 20, 3312–3330. [Google Scholar] [CrossRef]
- Boss, W.F.; Sederoff, H.W.; Im, Y.J.; Moran, N.; Grunden, A.M.; Perera, I.Y. Basal signaling regulates plant growth and development. Plant Physiol. 2010, 154, 439–443. [Google Scholar]
- Stevenson-Paulik, J.; Phillippy, B.Q. Inositol polyphosphates and kinases. In Lipid Signaling in Plants; Munnik, T., Ed.; Springer Berlin Heidelberg: Berlin/Heidelberg, Germany, 2010; Volume 16, pp. 161–174. [Google Scholar]
- Zonia, L.; Munnik, T. Cracking the green paradigm: Functional coding of phosphoinositide signals in plant stress responses. Subcell. Biochem. 2006, 39, 207–237. [Google Scholar] [CrossRef]
- Brearley, C.A.; Hanke, D.E. Metabolic evidence for the order of addition of individual phosphate esters in the myo-inositol moiety of inositol hexakisphosphate in the duckweed Spirodela polyrhiza L. Biochem. J. 1996, 314, 227–233. [Google Scholar]
- Raboy, V.; Young, K.A.; Dorsch, J.A.; Cook, A. Genetics and breeding of seed phosphorus and phytic acid. J. Plant Physiol. 2001, 158, 489–497. [Google Scholar] [CrossRef]
- Dorsch, J.A.; Cook, A.; Young, K.A.; Anderson, J.M.; Bauman, A.T.; Volkmann, C.J.; Murthy, P.P.; Raboy, V. Seed phosphorus and inositol phosphate phenotype of barley low phytic acid genotypes. Phytochemistry 2003, 62, 691–706. [Google Scholar] [CrossRef]
- Hirschi, K.D. The calcium conundrum. Both versatile nutrient and specific signal. Plant Physiol. 2004, 136, 2438–2442. [Google Scholar] [CrossRef]
- Dieck, C.B.; Wood, A.; Brglez, I.; Rojas-Pierce, M.; Boss, W.F. Increasing phosphatidylinositol (4,5) bisphosphate biosynthesis affects plant nuclear lipids and nuclear functions. Plant Physiol. Biochem. 2012, 57, 32–44. [Google Scholar] [CrossRef]
- Streb, S.; Zeeman, S.C. Starch Metabolism in Arabidopsis. Arabidopsis Book 2012, 10, e0160. [Google Scholar]
- Stitt, M.; Zeeman, S.C. Starch turnover: Pathways, regulation and role in growth. Curr. Opin. Plant Biol. 2012, 15, 282–292. [Google Scholar] [CrossRef]
- Graf, A.; Schlereth, A.; Stitt, M.; Smith, A.M. Circadian control of carbohydrate availability for growth in Arabidopsis plants at night. Proc. Natl. Acad. Sci. USA 2010, 107, 9458–9463. [Google Scholar] [CrossRef]
- Lu, Y.; Jackson, P.G.; Sharkey, T.D. Daylength and circadian effects on starch degradation and maltose metabolism. Plant Physiol. 2005, 138, 2280–2291. [Google Scholar] [CrossRef]
- Kramer, D.M.; Evans, J.R. The importance of energy balance in improving photosynthetic productivity. Plant Physiol. 2011, 155, 70–78. [Google Scholar] [CrossRef]
- Kohzuma, K.; Cruz, J.A.; Akashi, K.; Hoshiyasu, S.; Munekage, Y.N.; Yokota, A.; Kramer, D.M. The long-term responses of the photosynthetic proton circuit to drought. Plant Cell Environ. 2009, 32, 209–219. [Google Scholar] [CrossRef]
- Jia, H.; Oguchi, R.; Hope, A.B.; Barber, J.; Chow, W.S. Differential effects of severe water stress on linear and cyclic electron fluxes through photosystem I in spinach leaf discs in CO2-enriched air. Planta 2008, 228, 803–812. [Google Scholar] [CrossRef]
- Joliot, P.; Joliot, A. Quantification of cyclic and linear flows in plants. Proc. Natl. Acad. Sci. USA 2005, 102, 4913–4918. [Google Scholar] [CrossRef]
- Ma, X.; Shor, O.; Diminshtein, S.; Yu, L.; Im, Y.J.; Perera, I.; Lomax, A.; Boss, W.F.; Moran, N. Phosphatidylinositol (4,5)bisphosphate inhibits K+-efflux channel activity in NT1 tobacco cultured cells. Plant Physiol. 2008, 149, 1127–1140. [Google Scholar] [CrossRef]
- Baker, N.R.; Harbinson, J.; Kramer, D.M. Determining the limitations and regulation of photosynthetic energy transduction in leaves. Plant Cell Environ. 2007, 30, 1107–1125. [Google Scholar] [CrossRef]
- Avenson, T.J.; Cruz, J.A.; Kanazawa, A.; Kramer, D.M. Regulating the proton budget of higher plant photosynthesis. Proc. Natl. Acad. Sci. USA 2005, 102, 9709–9713. [Google Scholar] [CrossRef]
- Scheibe, R. Malate valves to balance cellular energy supply. Physiol. Plant 2004, 120, 21–26. [Google Scholar] [CrossRef]
- Tang, R.-H.; Han, S.; Zheng, H.; Cook, C.W.; Choi, C.S.; Woerner, T.E.; Jackson, R.B.; Pei, Z.-M. Coupling diurnal cytosolic Ca2+ oscillations to the CAS-IP3 pathway in Arabidopsis. Science 2007, 315, 1423–1426. [Google Scholar] [CrossRef]
- Nomura, H.; Komori, T.; Kobori, M.; Nakahira, Y.; Shiina, T. Evidence for chloroplast control of external Ca2+-induced cytosolic Ca2+ transients and stomatal closure: Chloroplast, stomatal movement and Ca2+ signal. Plant J. 2007, 53, 988–998. [Google Scholar] [CrossRef]
- Vollmer, A.H.; Youssef, N.N.; DeWald, D.B. Unique cell wall abnormalities in the putative phosphoinositide phosphatase mutant AtSAC9. Planta 2011, 234, 993–1005. [Google Scholar] [CrossRef]
- Gunesekera, B.; Torabinejad, J.; Robinson, J.; Gillaspy, G.E. Inositol polyphosphate 5-phosphatases 1 and 2 are required for regulating seedling growth. Plant Physiol. 2007, 143, 1408–1417. [Google Scholar] [CrossRef]
- Zhong, R.; Ye, Z.H. Molecular and biochemical characterization of three WD-repeat-domain-containing inositol polyphosphate 5-phosphatases in Arabidopsis thaliana. Plant Cell Physiol. 2004, 45, 1720–1728. [Google Scholar] [CrossRef]
- Wang, Y.; Chu, Y.-J.; Xue, H.-W. Inositol polyphosphate 5-phosphatase-controlled Ins(1,4,5)P3/Ca2+ is crucial for maintaining pollen dormancy and regulating early germination of pollen. Development 2012, 139, 2221–2233. [Google Scholar] [CrossRef]
- Carland, F.M.; Nelson, T. Cotyledon vascular pattern2-mediated inositol (1,4,5) triphosphate signal transduction is essential for closed venation patterns of Arabidopsis foliar organs. Plant Cell 2004, 16, 1263–1275. [Google Scholar] [CrossRef]
- Finka, A.; Cuendet, A.F.H.; Maathuis, F.J. M.; Saidi, Y.; Goloubinoff, P. Plasma membrane cyclic nucleotide gated calcium channels control land plant thermal sensing and acquired thermotolerance. Plant Cell 2012, 24, 3333–3348. [Google Scholar] [CrossRef]
- Wu, H.-C.; Hsu, S.-F.; Luo, D.-L.; Chen, S.-J.; Huang, W.-D.; Lur, H.-S.; Jinn, T.-L. Recovery of heat shock-triggered released apoplastic Ca2+ accompanied by pectin methylesterase activity is required for thermotolerance in soybean seedlings. J. Exp. Botany 2010, 61, 2843–2852. [Google Scholar] [CrossRef]
- Saidi, Y.; Finka, A.; Muriset, M.; Bromberg, Z.; Weiss, Y.G.; Maathuis, F.J.M.; Goloubinoff, P. The heat shock response in moss plants is regulated by specific calcium-permeable channels in the plasma membrane. Plant Cell 2009, 21, 2829–2843. [Google Scholar] [CrossRef]
- Mishkind, M.; Vermeer, J.E.; Darwish, E.; Munnik, T. Heat stress activates phospholipase D and triggers PIP accumulation at the plasma membrane and nucleus. Plant J. 2009, 60, 10–21. [Google Scholar] [CrossRef]
- Zhong, R.; Burk, D.H.; Nairn, C.J.; Wood-Jones, A.; Morrison, W.H.; Ye, Z.H. Mutation of SAC1, an Arabidopsis SAC domain phosphoinositide phosphatase, causes alterations in cell morphogenesis, cell wall synthesis, and actin organization. Plant Cell 2005, 17, 1449–1466. [Google Scholar] [CrossRef]
- Zhong, R.; Burk, D.H.; Morrison, W.H.; Ye, Z.H. FRAGILE FIBER3, an Arabidopsis gene encoding a type II inositol polyphosphate 5-phosphatase, is required for secondary wall synthesis and actin organization in fiber cells. Plant Cell 2004, 16, 3242–3259. [Google Scholar] [CrossRef]
- Zhong, R.; McCarthy, R.; Ye, Z.-H. Phosphoinositides and plant cell wall synthesis. In Lipid Signaling in Plants; Springer: Berlin/Heidelberg, Germany, 2010; Volume 16, pp. 175–184. [Google Scholar]
- Mikami, K.; Katagiri, T.; Iuchi, S.; Yamaguchi-Shinozaki, K.; Shinozaki, K. A gene encoding phosphatidylinositol-4-phosphate 5-kinase is induced by water stress and abscisic acid in Arabidopsis thaliana. Plant J. 1998, 15, 563–568. [Google Scholar] [CrossRef]
- Tucker, E.B.; Boss, W.F. Mastoparan-induced intracellular Ca2+ fluxes may regulate cell-to-cell communication in plants. Plant Physiol. 1996, 111, 459–467. [Google Scholar]
- Alkhalfioui, F.; Renard, M.; Frendo, P.; Keichinger, C.; Meyer, Y.; Gelhaye, E.; Hirasawa, M.; Knaff, D.B.; Ritzenthaler, C.; Montrichard, F. A novel type of thioredoxin dedicated to symbiosis in Legumes. Plant Physiol. 2008, 148, 424–435. [Google Scholar] [CrossRef]
- Mosblech, A.; Konig, S.; Stenzel, I.; Grzeganek, P.; Feussner, I.; Heilmann, I. Phosphoinositide and inositolpolyphosphate signalling in defense responses of Arabidopsis thaliana challenged by mechanical wounding. Mol. Plant 2008, 1, 249–261. [Google Scholar] [CrossRef]
- Chen, H.; Nelson, R.S.; Sherwood, J.L. Enhanced recovery of transformants of Agrobacterium tumefaciens after freeze-thaw transformation and drug selection. Biotechniques 1994, 16, 664–670. [Google Scholar]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana: Floral dip transformation of Arabidopsis. Plant. J. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Han, S.; Kim, D. AtRTPrimer: Database for Arabidopsis genome-wide homogeneous and specific RT-PCR primer-pairs. BMC Bioinform. 2006, 7, 179–187. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the DD CT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Weigel, D.; Glazebrook, J. Arabidopsis: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, CA, USA, 2002. [Google Scholar]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Perera, I.Y.; Love, J.; Heilmann, I.; Thompson, W.F.; Boss, W.F. Up-regulation of phosphoinositide metabolism in tobacco cells constitutively expressing the human type I inositol polyphosphate 5-phosphatase. Plant Physiol. 2002, 129, 1795–1806. [Google Scholar] [CrossRef]
- Kansas Lipidomics Research Center. Extraction of lipids from Arabidopsis leaf tissue. Available online: http://www.k-state.edu/lipid/lipidomics/leaf-extraction.html (accessed on 1 December 2013).
- Phillippy, B.Q.; Johnston, M.R. Determination of phytic acid in foods by ion chromatography with post-column derivatization. J. Food Sci. 1985, 50, 541–542. [Google Scholar] [CrossRef]
- Phillippy, B.Q.; Bland, J.M. Gradient ion chromatography of inositol phosphates. Anal. Biochem. 1988, 175, 162–166. [Google Scholar] [CrossRef]
- Bentsink, L.; Yuan, K.; Koornneef, M.; Vreugdenhil, D. The genetics of phytate and phosphate accumulation in seeds and leaves of Arabidopsis thaliana, using natural variation. Theor. Appl. Genet. 2003, 106, 1234–1243. [Google Scholar]
- Bartlett, G.R. Phosphorous assay in column chromatogrphy. J. Biol. Chem. 1954, 234, 466–468. [Google Scholar]
- Teng, S.; Keurentjes, J.; Bentsink, L.; Koornneef, M.; Smeekens, S. Sucrose-specific induction of anthocyanin biosynthesis in Arabidopsis requires the MYB75/PAP1 gene. Plant Physiol. 2005, 139, 1840–1852. [Google Scholar] [CrossRef]
- Smith, A.M.; Zeeman, S.C. Quantification of starch in plant tissues. Nat. Protoc. 2006, 1, 1342–1345. [Google Scholar] [CrossRef]
- Matsumura, H. Miyachi Cycling assay for nicotinamide adenine dinucleotides. Methods Enzymol. 1980, 69, 465–470. [Google Scholar]
- Hall, C.C.; Cruz, J.; Wood, M.; Zegarac, R.; Carpenter, J.; Kanazawa, A.; Kramer, D.M. Photosynthetic Measurements with the idea spec: An integrated diode emitter array spectrophotometer/fluorometer. In Photosynthesis Research for Food, Fuel and the Future, Proceedings of the 15th International Conference on Photosynthesis, Beijing, China, 22–27 August 2010; Kuang, T., Lu, C., Zhang, L., Eds.; Springer: Dordrecht, The Netherlands, 2013; pp. 184–188. [Google Scholar]
- Livingston, A.K.; Cruz, J.A.; Kohzuma, K.; Dhingra, A.; Kramer, D.M. An Arabidopsis mutant with high cyclic electron flow around photosystem I (HCEF) involving the NADPH dehydrogenase complex. Plant Cell 2010, 22, 221–233. [Google Scholar] [CrossRef]
- Livingston, A.K.; Kanazawa, A.; Cruz, J.A.; Kramer, D.M. Regulation of cyclic electron flow in C3 plants: Differential effects of limiting photosynthesis at ribulose-1,5-bisphosphate carboxylase/oxygenase and glyceraldehyde-3-phosphate dehydrogenase. Plant Cell Environ. 2010, 33, 1779–1788. [Google Scholar] [CrossRef]
- Harbinson, J.; Genty, B.; Baker, N.R. Relationship between the quantum efficiencies of photosystems I and II in pea leaves. Plant. Physiol. 1989, 90, 1029–1034. [Google Scholar] [CrossRef]
- Gentry, B.; Briantais, J.M.; Baker, N.R. The relationship between the quantum yield of photosynthetic electron transport and quenching of chlorophyll fluorescence. Biochim. Biophys. Acta 1989, 990, 87–92. [Google Scholar] [CrossRef]
- Baker, N.R. Chlorophyll fluorescence: A probe of photosynthesis in vivo. Annu. Rev. Plant Biol. 2008, 59, 89–113. [Google Scholar] [CrossRef]
- Mehlmer, N.; Parvin, N.; Hurst, C.H.; Knight, M.R.; Teige, M.; Vothknecht, U.C. A toolset of aequorin expression vectors for in planta studies of subcellular calcium concentrations in Arabidopsis thaliana. J. Exp. Bot. 2012, 63, 1751–1761. [Google Scholar] [CrossRef]
- Marti, M.C.; Stancombe, M.A.; Webb, A.A.R. Cell- and stimulus type-specific intracellular free Ca2+ signals in Arabidopsis. Plant Physiol. 2013, 163, 625–634. [Google Scholar] [CrossRef]
- Swanson, S.J.; Choi, W.-G.; Chanoca, A.; Gilroy, S. In vivo imaging of Ca2+, pH, and reactive oxygen species using fluorescent probes in plants. Annu. Rev. Plant Biol. 2011, 62, 273–297. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Im, Y.J.; Smith, C.M.; Phillippy, B.Q.; Strand, D.; Kramer, D.M.; Grunden, A.M.; Boss, W.F. Increasing Phosphatidylinositol (4,5)-Bisphosphate Biosynthesis Affects Basal Signaling and Chloroplast Metabolism in Arabidopsis thaliana. Plants 2014, 3, 27-57. https://doi.org/10.3390/plants3010027
Im YJ, Smith CM, Phillippy BQ, Strand D, Kramer DM, Grunden AM, Boss WF. Increasing Phosphatidylinositol (4,5)-Bisphosphate Biosynthesis Affects Basal Signaling and Chloroplast Metabolism in Arabidopsis thaliana. Plants. 2014; 3(1):27-57. https://doi.org/10.3390/plants3010027
Chicago/Turabian StyleIm, Yang Ju, Caroline M. Smith, Brian Q. Phillippy, Deserah Strand, David M. Kramer, Amy M. Grunden, and Wendy F. Boss. 2014. "Increasing Phosphatidylinositol (4,5)-Bisphosphate Biosynthesis Affects Basal Signaling and Chloroplast Metabolism in Arabidopsis thaliana" Plants 3, no. 1: 27-57. https://doi.org/10.3390/plants3010027




