Xylo-Oligosaccharides and Inulin Affect Genotoxicity and Bacterial Populations Differently in a Human Colonic Simulator Challenged with Soy Protein
Abstract
:1. Introduction
2. Experimental Section
2.1. Faecal Inoculum
2.2. Fermentation Media
2.3. Human Colonic Simulator
2.4. Growth and Maintenance of HT29 Cells
2.5. Faecal Water Genotoxicity Assay
2.6. Comet Assay
2.7. SCFA Analysis
2.8. DNA Extraction and Quantitative PCR (QPCR)
| Target | Primer | Sequence (5′–3′) | nM | Reference | Annealing | Elongation | ||
|---|---|---|---|---|---|---|---|---|
| °C | Time (s) | °C | Time (s) | |||||
| Total bacteria | 1114F 1275R | CGGCAACGAGCGCAACCC CCATTGTAGCACGTGTGTAGCC | 150 | [21] | 60 | 20 | 72 | 45 |
| Bacteroides fragilis group | Bfr-F Bfr-R | CTGAACCAGCCAAGTAGCG CCGCAAACTTTCACAACTGACTTA | 500 | [22] | 58 | 60 | 72 | 30 |
| Bifidobacterium spp. | Bif-F Bif-R | TCGCGTC(C/T)GGTGTGAAAG CCACATCCAGC(A/G)TCCAC | 600 | [23] | 58 | 20 | 72 | 30 |
| Clostridium coccoides group | g-Ccoc-F g-Ccoc-R | AAATGACGGTACCTGACTAA CTTTGAGTTTCATTCTTGCGA A | 250 | [24] | 58 | 20 | 72 | 45 |
| Clostridium leptum group | sg-Clept-F sg-Clept-R | CTTTGAGTTTCATTCTTGCGAA GCACAAGCAGTGGAGT | 250 | [24] | 58 | 20 | 72 | 45 |
| Escherichia coli | E.coli F E.coli R | CATGCCGCGTGTATGAAGAA CGGGTAACGTCAATGAGCAAA | 375 | [25] | 60 | 20 | 72 | 45 |
| Faecalibacterium praunitzii | FPR-1F FPR-2R | AGATGGCCTCGCGTCCGA CCGAAGACCTTCTTCCTCC | 500 | [26] | 62 | 20 | 72 | 40 |
| Lactobacillus group | Lacto-F Lacto-R | AGCAGTAGGGAATCTTCCA CACCGCTACACATGGAG | 500 | [27] [28] | 58 | 30 | 72 | 30 |
| SRB 1_aps 2 | APS3F APS2R | TGGCAGATCATGWTYAAYGG GGGCCGTAACCRTCYTTRAA | 400 | [29] | 58 | 30 | 72 | 60 |
| SRB_dsr 3 | DSR1F+ DSR-R | ACSCACTGGAAGCACGGCGG GTGGMRCCGTGCAKRTTGG | 400 | [30] | 65 | 15 | 72 | 30 |
2.9. Statistics
3. Results
3.1. Faecal Water Genotoxicity
3.2. Short Chain Fatty Acids
| PV (pH 5.5) | DV (pH 6.8) | PBS (control) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Pre-inulin | Inulin | Pre-XOS | XOS | Pre-inulin | Inulin | Pre-XOS | XOS | ||
| Tail Moment | 18.6 ± 1.1 | 16.0 ± 0.7 | 7.6 ± 0.4 a | 5.4 ± 0.3 b | 11.3 ± 0.5 | 12.1 ± 0.6 | 6.7 ± 0.3 b | 12.8 ± 0.6 a | 5.8 ± 0.7 |
| Tail Length | 78.9 ± 3.7 | 76.7 ± 2.8 | 44.1 ± 1.7 a | 33.3 ± 1.4 b | 54.9 ± 1.9 | 61.2 ± 2.5 | 34.5 ± 1.2 b | 62.0 ± 2.2 a | 34.3 ± 3.1 |
| SCFA | PV (pH 5.5) | DV (pH 6.8) | ||||||
|---|---|---|---|---|---|---|---|---|
| Pre-inulin | Inulin | Pre-XOS | XOS | Pre-inulin | Inulin | Pre-XOS | XOS | |
| Acetate | 43 ± 7 | 47 ± 6 | 84 ± 4 | 86 ± 7 | 95 ± 12 | 97 ± 9 | 110 ± 8 | 104 ± 3 |
| Propionate | 1 ± 0 | 1 ± 0 | 3 ± 1 | 3 ± 1 | 19 ± 1 | 22 ± 4 | 21 ± 2 | 22 ± 4 |
| Butyrate | 28 ± 4 | 37 ± 4 | 15 ± 2 | 24 ± 6 | 39 ± 5 a | 70 ± 6 b | 35 ± 2 a | 46 ± 6 b |
| Total | 72 ± 11 | 85 ± 6 | 102 ± 6 | 113 ± 13 | 153 ± 18 | 189 ± 14 | 178 ± 12 | 181 ± 14 |
3.3. Microbial Abundances
3.4. Relationships between Microbial Abundance, DNA Damage and SCFA
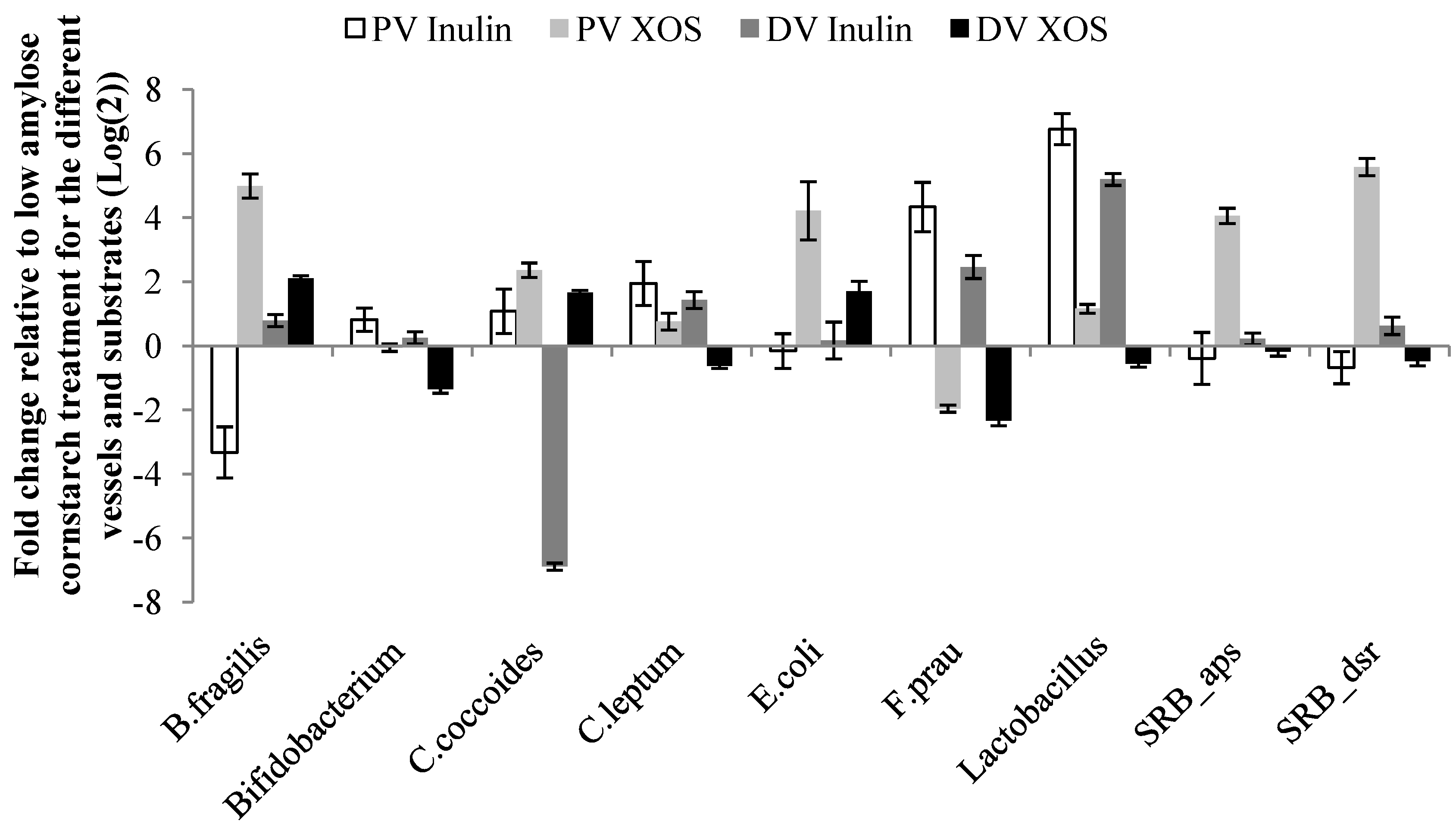
| Comparisons | B. fragilis | Bifidobacterium | C. coccoides | C. leptum | E. coli | F. prau | Lactobacillus | SRB_aps | SRB_dsr |
|---|---|---|---|---|---|---|---|---|---|
| PV inulin vs. PV XOS | ˃0.0001 | 0.02 | ns | ns | 0.0007 | ˃0.0001 | ˃0.0001 | 0.0002 | ˃0.0001 |
| PV inulin vs. DV inulin | 0.003 | ns | ˃0.0001 | ns | ns | 0.038 | 0.007 | ns | 0.046 |
| PV XOS vs. DV XOS | ˃0.0001 | ˃0.0001 | 0.008 | 0.0002 | 0.012 | ns | ˃0.0001 | ˃0.0001 | ˃0.0001 |
| DV XOS vs. DV inulin | ˃0.0001 | ˃0.0001 | ˃0.0001 | 0.0001 | 0.030 | ˃0.0001 | ˃0.0001 | ns | 0.002 |

4. Discussion
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Norat, T.; Bingham, S.; Ferrari, P.; Slimani, N.; Jenab, M.; Mazuir, M.; Overvad, K.; Olsen, A.; Tjonneland, A.; Clavel, F.; et al. Meat, fish, and colorectal cancer risk: The European prospective investigation into cancer and nutrition. J. Natl. Cancer Inst. 2005, 97, 906–916. [Google Scholar] [CrossRef]
- Howard, M.D.; Gordon, D.T.; Garleb, K.A.; Kerley, M.S. Dietary fructooligosaccharide, xylooligosaccharide and gum-arabic havevariable effects on cecal and colonic microbiota and epithelial cell proliferation in mice and rats. J. Nutr. 1995, 125, 2604–2609. [Google Scholar]
- Toden, S.; Bird, A.R.; Topping, D.L.; Conlon, M.A. Dose-dependent reduction of dietary protein-induced colonocyte DNA damage by resistant starch in rats correlates more highly with caecal butyrate than with other short chain fatty acids. Cancer Biol. Ther. 2007, 6, 253–258. [Google Scholar]
- Toden, S.; Bird, A.R.; Topping, D.L.; Conlon, M.A. Differential effects of dietary whey, casein and soya on colonic DNA damage and large bowel SCFA in rats fed diets low and high in resistant starch. Br. J. Nutr. 2007, 97, 535–543. [Google Scholar] [CrossRef]
- Toden, S.; Bird, A.R.; Topping, D.L.; Conlon, M.A. High red meat diets induce greater numbers of colonic DNA double-strand breaks than white meat in rats: Attenuation by high-amylose maize starch. Carcinogenesis 2007, 28, 2355–2362. [Google Scholar] [CrossRef]
- Swennen, K.; Courtin, C.M.; Delcour, J.A. Non-digestible oligosaccharides with prebiotic properties. Crit. Rev. Food Sci. 2006, 46, 459–471. [Google Scholar] [CrossRef]
- Gibson, G.R.; Probert, H.M.; Loo, J.V.; Rastall, R.A.; Roberfroid, M.B. Dietary modulation of the human colonic microbiota: Updating the concept of prebiotics. Nutr Res. Rev. 2004, 17, 259–275. [Google Scholar] [CrossRef]
- Bird, A.; Conlon, M.; Christophersen, C.T.; Topping, D. Resistant starch, large bowel fermentation and a broader perspective of prebiotics and probiotics. Benef. Microbes 2010, 1, 423–431. [Google Scholar] [CrossRef]
- Roberfroid, M. Prebiotics: The concept revisited. J. Nutr. 2007, 137, 830S–837S. [Google Scholar]
- Vazquez, M.J.; Alonso, J.L.; Dominguez, H.; Parajo, J.C. Xylooligosaccharides: Manufacture and applications. Trends Food Sci. Technol. 2000, 11, 387–393. [Google Scholar] [CrossRef]
- Pool-Zobel, B.L.; Sauer, J. Overview of experimental data on reduction of colorectal cancer risk by inulin-type fructans. J. Nutr. 2007, 137, 2580S–2584S. [Google Scholar]
- Ramirez-Farias, C.; Slezak, K.; Fuller, Z.; Duncan, A.; Holtrop, G.; Louis, P. Effect of inulin on the human gut microbiota: Stimulation of Bifidobacterium adolescentis and Faecalibacterium prausnitzii. Br. J. Nutr. 2009, 101, 541–550. [Google Scholar]
- Sokol, H.; Pigneur, B.; Watterlot, L.; Lakhdari, O.; Bermudez-Humaran, L.G.; Gratadoux, J.J.; Blugeon, S.; Bridonneau, C.; Furet, J.P.; Corthier, G.; et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of crohn disease patients. Proc. Natl. Acad. Sci. USA 2008, 105, 16731–16736. [Google Scholar] [CrossRef]
- Fukuda, S.; Toh, H.; Hase, K.; Oshima, K.; Nakanishi, Y.; Yoshimura, K.; Tobe, T.; Clarke, J.; Topping, D.; Suzuki, T.; et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 2011, 469, 543–547. [Google Scholar] [CrossRef]
- Bruck, W.M.; Graverholt, G.; Gibson, G.R. Use of batch culture and a two-stage continuous culture system to study the effect of supplemental alpha-lactalbumin and glycomacropeptide on mixed populations of human gut bacteria. FEMS Microbiol. Ecol. 2002, 41, 231–237. [Google Scholar]
- McKelvey-Martin, V.J.; Green, M.H.; Schmezer, P.; Pool-Zobel, B.L.; de Meo, M.P.; Collins, A. The single cell gel electrophoresis assay (comet assay): A European review. Mutat. Res. 1993, 288, 47–63. [Google Scholar] [CrossRef]
- Pool-Zobel, B.L.; Bub, A.; Muller, H.; Wollowski, I.; Rechkemmer, G. Consumption of vegetables reduces genetic damage in humans: First results of a human intervention trial with carotenoid-rich foods. Carcinogenesis 1997, 18, 1847–1850. [Google Scholar] [CrossRef]
- McOrist, A.L.; Abell, G.C.J.; Cooke, C.; Nyland, K. Bacterial population dynamics and faecal short-chain fatty acid (SCFA) concentrations in healthy humans. Br. J. Nutr. 2008, 100, 138–146. [Google Scholar]
- Hellemans, J.; Mortier, G.; de Paepe, A.; Speleman, F.; Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef]
- Vandesompele, J.; de Preter, K.; Pattyn, F.; Poppe, B.; van Roy, N.; de Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3. [Google Scholar] [CrossRef]
- Denman, S.E.; McSweeney, C.S. Development of a real-time PCR assay for monitoring anaerobic fungal and cellulolytic bacterial populations within the rumen. FEMS Microbiol. Ecol. 2006, 58, 572–582. [Google Scholar] [CrossRef]
- Liu, C.; Song, Y.; McTeague, M.; Vu, A.W.; Wexler, H.; Finegold, S.M. Rapid identification of the species of the Bacteroides fragilis group by multiplex PCR assays using group- and species-specific primers. FEMS Microbiol. Lett. 2003, 222, 9–16. [Google Scholar] [CrossRef]
- Rinttilä, T.; Kassinen, A.; Malinen, E.; Krogius, L.; Palva, A. Development of an extensive set of 16S rDNA-targeted primers for quantification of pathogenic and indigenous bacteria in faecal samples by real-time PCR. Appl. Microbiol. 2004, 97, 1166–1177. [Google Scholar] [CrossRef]
- Matsuki, T.; Watanabe, K.; Fujimoto, J.; Takada, T.; Tanaka, R. Use of 16S rRNA gene-targeted group-specific primers for real-time PCR analysis of predominant bacteria in human feces. Appl. Environ. Microbiol. 2004, 70, 7220–7228. [Google Scholar] [CrossRef]
- Huijsdens, X.W.; Linskens, R.K.; Mak, M.T.; Meuwissen, S.G.M.; Vandenbroucke-Grauls, C.; Savelkoul, P.H.M. Quantification of bacteria adherent to gastrointestinal mucosa by real-time PCR. J. Clin. Microbiol. 2002, 40, 4423–4427. [Google Scholar] [CrossRef]
- Wang, R.F.; Cao, W.W.; Cerniglia, C.E. Phylogenetic analysis of Fusobacterium prausnitzii based upon the 16S rRNA gene sequence and PCR confirmation. Int. J. Syst. Bacteriol. 1996, 46, 341–343. [Google Scholar] [CrossRef]
- Walter, J.; Hertel, C.; Tannock, G.W.; Lis, C.M.; Munro, K.; Hammes, W.P. Detection of Lactobacillus, Pediococcus, Leuconostoc, and Weissella species in human feces by using group-specific PCR primers and denaturing gradient gel electrophoresis. Appl. Environ. Microbiol. 2001, 67, 2578–2585. [Google Scholar] [CrossRef]
- Heilig, H.G.H.J.; Zoetendal, E.G.; Vaughan, E.E.; Marteau, P.; Akkermans, A.D.L.; de Vos, W.M. Molecular diversity of Lactobacillus spp. and other lactic acid bacteria in the humanintestine as determined by specific amplification of 16S ribosomal DNA. Appl. Environ. Microbiol. 2002, 68, 114–123. [Google Scholar] [CrossRef]
- Christophersen, C.T.; Morrison, M.; Conlon, M.A. Abundance of sulphate-reducing bacteria in human feces is overestimated by QPCR that targets the Desulfovibrio 16S rRNA gene. Appl. Environ. Microbiol. 2011, 77, 3544–3546. [Google Scholar] [CrossRef]
- Kondo, R.; Nedwell, D.B.; Purdy, K.J.; Silva, S.Q. Detection and enumeration of sulphate-reducing bacteria in estuarine sediments by competitive PCR. Geomicrobiol. J. 2004, 21, 145–157. [Google Scholar] [CrossRef]
- PASW Statistics for Windows, version 18.0; SPSS Inc.: Chicago, IL, USA, 2009.
- McArdle, B.H.; Anderson, M.J. Fitting multivariate models to community data: A comment on distance-based redundancy analysis. Ecology 2001, 82, 290–297. [Google Scholar] [CrossRef]
- Legendre, P.; Anderson, M.J. Distance-based redundancy analysis: Testing multispecies responses in multifactorial ecological experiments. Ecol. Monogr. 1999, 69, 1–24. [Google Scholar] [CrossRef]
- Petersen, A.; Bergström, A.; Andersen, J.; Hansen, M.; Lahtinen, S.; Wilcks, A.; Licht, T. Analysis of the intestinal microbiota of oligosaccharide fed mice exhibiting reduced resistance to Salmonella infection. Benef. Microbes 2010, 1, 271–281. [Google Scholar] [CrossRef]
- Scanlan, P.D.; Shanahan, F.; Marchesi, J.R. Culture-independent analysis of Desulfovibrios in the human distal colon of healthy, colorectal cancer and polypectomized individuals. FEMS Microbiol. Ecol. 2009, 69, 213–221. [Google Scholar] [CrossRef]
- Attene-Ramos, M.S.; Wagner, E.D.; Gaskins, H.R.; Plewa, M.J. Hydrogen sulfide induces direct radical-associated DNA damage. Mol. Cancer Res. 2007, 5, 455–459. [Google Scholar] [CrossRef]
- Gibson, G.R.; Cummings, J.H.; Macfarlane, G.T. Growth and activities of sulfate-reducing bacteria in the gut contents of healthy subjects and patients with ulcerative colitis. FEMS Microbiol. Lett. 1991, 86, 103–111. [Google Scholar] [CrossRef]
- Fite, A.; Macfarlane, G.T.; Cummings, J.H.; Hopkins, M.J.; Kong, S.C.; Furrie, E.; Macfarlane, S. Identification and quantitation of mucosal and faecal Desulfovibrios using real time polymerase chain reaction. Gut 2004, 53, 523–529. [Google Scholar] [CrossRef]
- Moore, J.; Babidge, W.; Millard, S.; Roediger, W. Colonic luminal hydrogen sulfide is not elevated in ulcerative colitis. Dig. Dis. Sci. 1998, 43, 162–165. [Google Scholar] [CrossRef]
- Jorgensen, J.; Mortensen, P.B. Hydrogen sulfide and colonic epithelial metabolism: Implications for ulcerative colitis. Dig. Dis. Sci. 2001, 46, 1722–1732. [Google Scholar] [CrossRef]
- Pitcher, M.C.L.; Beatty, E.R.; Cummings, J.H. The contribution of sulphate reducing bacteria and 5-aminosalicylic acid to faecal sulphide in patients with ulcerative colitis. Gut 2000, 46, 64–72. [Google Scholar] [CrossRef]
- Wallace, J.L.; Vong, L.; McKnight, W.; Dicay, M.; Martin, G.R. Endogenous and exogenous hydrogen sulfide promotes resolution of colitis in rats. Gastroenterology 2009, 137, 569–578. [Google Scholar] [CrossRef]
- Campbell, J.M.; Fahey, G.C.; Wolf, B.W. Selected indigestible oligosaccharides affect large bowel mass, cecal and fecal short-chain fatty acids, pH and microflora in rats. J. Nutr. 1997, 127, 130–136. [Google Scholar]
- Hsu, C.-K.; Liao, J.-W.; Chung, Y.-C.; Hsieh, C.-P.; Chan, Y.-C. Xylooligosaccharides and fructooligosaccharides affect the intestinal microbiota and precancerous colonic lesion development in rats. J. Nutr. 2004, 134, 1523–1528. [Google Scholar]
- Havenaar, R.; Bonnin-Marol, S.; Van Dokkum, W.; Petitet, S.; Schaafsma, G. Inulin: Fermentation and microbial ecology in the intestinal tract. Food Rev. Int. 1999, 15, 109–120. [Google Scholar] [CrossRef]
- Zampa, A.; Silvi, S.; Fabiani, R.; Morozzi, G.; Orpianesi, C.; Cresci, A. Effects of different digestible carbohydrates on bile acid metabolism and SCFA production by human gut micro-flora grown in an in vitro semi-continuous culture. Anaerobe 2004, 10, 19–26. [Google Scholar] [CrossRef]
- McIntyre, A.; Gibson, P.R.; Young, G.P. Butyrate production from dietary fiber and protection against large-bowel cancer in a rat model. Gut 1993, 34, 386–391. [Google Scholar] [CrossRef]
- Le Leu, R.K.; Brown, I.L.; Hu, Y.; Esterman, A.; Young, G.P. Suppression of azoxymethane-induced colon cancer development in rats by dietary resistant starch. Cancer Biol. Ther. 2007, 6, 1621–1626. [Google Scholar] [CrossRef]
- Duncan, S.H.; Belenguer, A.; Holtrop, G.; Johnstone, A.M.; Flint, H.J.; Lobley, G.E. Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl. Environ. Microbiol. 2007, 73, 1073–1078. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Christophersen, C.T.; Petersen, A.; Licht, T.R.; Conlon, M.A. Xylo-Oligosaccharides and Inulin Affect Genotoxicity and Bacterial Populations Differently in a Human Colonic Simulator Challenged with Soy Protein. Nutrients 2013, 5, 3740-3756. https://doi.org/10.3390/nu5093740
Christophersen CT, Petersen A, Licht TR, Conlon MA. Xylo-Oligosaccharides and Inulin Affect Genotoxicity and Bacterial Populations Differently in a Human Colonic Simulator Challenged with Soy Protein. Nutrients. 2013; 5(9):3740-3756. https://doi.org/10.3390/nu5093740
Chicago/Turabian StyleChristophersen, Claus T., Anne Petersen, Tine R. Licht, and Michael A. Conlon. 2013. "Xylo-Oligosaccharides and Inulin Affect Genotoxicity and Bacterial Populations Differently in a Human Colonic Simulator Challenged with Soy Protein" Nutrients 5, no. 9: 3740-3756. https://doi.org/10.3390/nu5093740
APA StyleChristophersen, C. T., Petersen, A., Licht, T. R., & Conlon, M. A. (2013). Xylo-Oligosaccharides and Inulin Affect Genotoxicity and Bacterial Populations Differently in a Human Colonic Simulator Challenged with Soy Protein. Nutrients, 5(9), 3740-3756. https://doi.org/10.3390/nu5093740





